Article
|
Open Access
Featured
-
-
Article
| Open Access Epigenetically-driven anatomical diversity of synovial fibroblasts guides joint-specific fibroblast functions
Epigenetically-driven anatomical diversity of synovial fibroblasts guides joint-specific fibroblast functions
Arthritis affects different joints variably despite systemic inflammatory cues. Here the authors show anatomical differences in the transcriptome, epigenome and function of synovial fibroblasts that might affect susceptibility to site-specific joint diseases.
- Mojca Frank-Bertoncelj
- , Michelle Trenkmann
- & Caroline Ospelt
-
Article
| Open Access HOPX hypermethylation promotes metastasis via activating SNAIL transcription in nasopharyngeal carcinoma
HOPX hypermethylation promotes metastasis via activating SNAIL transcription in nasopharyngeal carcinoma
HOPX is a transcription factor epigenetically silenced in several cancers. Here the authors, by analysing methylation profiles, identify HOPX as a suppressor of metastasis in nasopharyngeal carcinoma: mechanistically HOPX inhibitsSNAILtranscription through deacetylation-mediated silencing.
- Xianyue Ren
- , Xiaojing Yang
- & Jun Ma
-
Article
| Open Access UVR2 ensures transgenerational genome stability under simulated natural UV-B in Arabidopsis t haliana
UVR2 ensures transgenerational genome stability under simulated natural UV-B in Arabidopsis t haliana
As sessile organisms, plants are exposed to recurrent solar UV-B radiation that can induce DNA damage. Here, the authors characterize mutations that occur in Arabidopsisunder light regimes simulating natural UV-B exposure and find that the UVR2 photolyase is the major component required to maintain genome stability.
- Eva-Maria Willing
- , Thomas Piofczyk
- & Ales Pecinka
-
Article
| Open Access Integrative epigenome-wide analysis demonstrates that DNA methylation may mediate genetic risk in inflammatory bowel disease
Integrative epigenome-wide analysis demonstrates that DNA methylation may mediate genetic risk in inflammatory bowel disease
Epigenetic perturbations may be an important factor in diseases where both genes and environment play a role. Here, Ventham and colleagues show that DNA methylation changes in inflammatory bowel disease are related to the underlying genotype, and are associated with cell-specific changes to gene expression.
- N. T. Ventham
- , N. A. Kennedy
- & J. Satsangi
-
Article
| Open Access Abundant DNA 6mA methylation during early embryogenesis of zebrafish and pig
Abundant DNA 6mA methylation during early embryogenesis of zebrafish and pig
DNA 6mA is a poorly understood epigenetic mark present at a low abundance in eukaryotic genomes. Here the authors observe high levels in zebrafish and pig during early embryogenesis enriched to repetitive regions of the genome and followed by attenuation during development.
- Jianzhao Liu
- , Yuanxiang Zhu
- & Chuan He
-
Article
| Open Access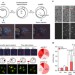 Mice produced by mitotic reprogramming of sperm injected into haploid parthenogenotes
Mice produced by mitotic reprogramming of sperm injected into haploid parthenogenotes
It is unclear what regulates gamete reprogramming competence. Here, the authors inject sperm into parthenogenetic embryos, generating viable offspring and show that mouse embryos in the mitotic cell cycle can reprogram sperm for full term development.
- Toru Suzuki
- , Maki Asami
- & Anthony C. F. Perry
-
Article
| Open Access Type II enteropathy-associated T-cell lymphoma features a unique genomic profile with highly recurrent SETD2 alterations
Type II enteropathy-associated T-cell lymphoma features a unique genomic profile with highly recurrent SETD2 alterations
Enteropathy associated T-cell lymphoma -EATL- affects the intestine and there are two different subtypes. In this study, the authors carry out exome sequencing of the type II variant and find that it is characterized by recurrent mutations in the histone methyltransferase SETD2 that are accompanied by altered H3K6 methylation.
- Annalisa Roberti
- , Maria Pamela Dobay
- & Laurence de Leval
-
Article
| Open Access Dissecting the precise role of H3K9 methylation in crosstalk with DNA maintenance methylation in mammals
Dissecting the precise role of H3K9 methylation in crosstalk with DNA maintenance methylation in mammals
There is crosstalk between the maintenance of DNA methylation and histone methylation. Here, the authors create an Uhrf1 knockin mouse model that abolishes the H3K9me2/3-binding activity of Uhrf1, and show that DNA maintenance methylation in mammals is largely independent of H3K9 methylation.
- Qian Zhao
- , Jiqin Zhang
- & Jiemin Wong
-
Article
| Open Access DNA hydroxymethylation controls cardiomyocyte gene expression in development and hypertrophy
DNA hydroxymethylation controls cardiomyocyte gene expression in development and hypertrophy
5-hydroxymethylation of cysteine (5-hmC) plays a role in epigenetic regulation. Here the authors analyse the hydroxymethylome in embryonic, neonatal, adult and hypertrophic mouse cardiomyocytes and show that the dynamic modulation of hydroxymethylated DNA is important for cardiomyocyte gene expression programming in heart development and failure.
- Carolina M. Greco
- , Paolo Kunderfranco
- & Gianluigi Condorelli
-
Article
| Open Access Extra-coding RNAs regulate neuronal DNA methylation dynamics
Extra-coding RNAs regulate neuronal DNA methylation dynamics
DNA methylation in the brain is a dynamic process, but gene-specific regulation of this process is poorly understood. Here, Day and colleagues show that extra-coding RNAs interact with DNA methyltransferases and regulate neuronal DNA methylation to control gene expression in locus-specific manner in neurons.
- Katherine E. Savell
- , Nancy V. N. Gallus
- & Jeremy J. Day
-
Article
| Open Access MTHFD1 controls DNA methylation in Arabidopsis
MTHFD1 controls DNA methylation in Arabidopsis
DNA methylation contributes to transcriptional silencing. Here, Groth et al.show that mutant plants defective in MTHFD1, an enzyme involved in folate metabolism, have a DNA hypomethylation phenotype highlighting the link between one-carbon metabolism and DNA methylation, which is mediated by SAM as a common methyl donor.
- Martin Groth
- , Guillaume Moissiard
- & Steven E. Jacobsen
-
Article
| Open Access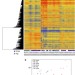 Joint-specific DNA methylation and transcriptome signatures in rheumatoid arthritis identify distinct pathogenic processes
Joint-specific DNA methylation and transcriptome signatures in rheumatoid arthritis identify distinct pathogenic processes
Rheumatoid arthritis is an inflammatory disease that selectively affects different joints. Here the authors show that gene expression and DNA methylation patterns of fibroblast-like synoviocytes differ between hip and knee joints in patients with RA, thus providing epigenetic and transcriptomic evidence for this anatomic selectivity of inflammation.
- Rizi Ai
- , Deepa Hammaker
- & Wei Wang
-
Article
| Open Access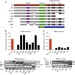 DNMT3B isoforms without catalytic activity stimulate gene body methylation as accessory proteins in somatic cells
DNMT3B isoforms without catalytic activity stimulate gene body methylation as accessory proteins in somatic cells
De novoDNA methylation is carried out by DNA methyltransferase DNMT3A/B, although DNMT3B isoforms without active catalytic domains are widely expressed. Here, the authors show that DNMT3B isoforms stimulate gene body methylation and re-methylation independently of the isoforms' catalytic activity.
- Christopher E. Duymich
- , Jessica Charlet
- & Gangning Liang
-
Article
| Open Access Genetic and environmental influences interact with age and sex in shaping the human methylome
Genetic and environmental influences interact with age and sex in shaping the human methylome
Differential impact of genetic and environmental influences on DNA methylation may result in sex- and age-related physiological variation and disease susceptibility. By analysing DNA methylome of 2,603 individuals from twin families, here, the authors establish a catalogue of between-individual variation in DNA methylation.
- Jenny van Dongen
- , Michel G. Nivard
- & Dorret I. Boomsma
-
Article
| Open Access Hemi-methylated DNA opens a closed conformation of UHRF1 to facilitate its histone recognition
Hemi-methylated DNA opens a closed conformation of UHRF1 to facilitate its histone recognition
UHRF1 is involved in the maintenance of DNA methylation, but the regulatory mechanism of this epigenetic regulator is unclear. Here, the authors show that it has a closed conformation and are able to make conclusions about the mechanism of recognition of epigenetic marks.
- Jian Fang
- , Jingdong Cheng
- & Yanhui Xu
-
Article
| Open Access Sequence features accurately predict genome-wide MeCP2 binding in vivo
Sequence features accurately predict genome-wide MeCP2 binding in vivo
MeCP2 is critical for proper brain development, and mutations in the gene encoding MeCP2 are responsible for several neurological disorders. Here, the authors show that the previously reported genome-wide preference of MeCP2 to methylated CpGs is in part due to MeCP2's affinity to GC-rich chromatin.
- H. Tomas Rube
- , Wooje Lee
- & Qizhi Gong
-
Article
| Open Access Direct evidence for sequence-dependent attraction between double-stranded DNA controlled by methylation
Direct evidence for sequence-dependent attraction between double-stranded DNA controlled by methylation
Theoretical studies suggest that homologous DNA duplexes can preferentially associate with one another in the absence of proteins. Here, the authors show that GC-rich DNA with methylated cytosine and AT-rich DNA duplexes associate more strongly than GC-rich duplexes regardless of the sequence homology.
- Jejoong Yoo
- , Hajin Kim
- & Taekjip Ha
-
Article
| Open Access Genome-wide DNA methylation levels and altered cortisol stress reactivity following childhood trauma in humans
Genome-wide DNA methylation levels and altered cortisol stress reactivity following childhood trauma in humans
Exposure to childhood trauma is a major risk factor for the development of almost all psychiatric disorders. By epigenome-wide studies, here, Houtepen et al. show that DNA methylation at a locus in the Kit ligand gene (KITLG) mediates the relationship between childhood trauma and cortisol stress reactivity.
- Lotte C. Houtepen
- , Christiaan H. Vinkers
- & Marco P. M. Boks
-
Article
| Open Access Epigenetic regulation of diacylglycerol kinase alpha promotes radiation-induced fibrosis
Epigenetic regulation of diacylglycerol kinase alpha promotes radiation-induced fibrosis
Radiotherapy can induce fibrosis in cancer patients, limiting its use in clinical settings. Here, the authors identify a differentially methylated enhancer of the lipid kinase DGKA in fibroblasts from breast cancer patients developing fibrosis after radiotherapy and they show that DGKA inhibition affects lipid homeostasis and reduces pro-fibrotic fibroblast activation.
- Christoph Weigel
- , Marlon R. Veldwijk
- & Odilia Popanda
-
Article
| Open Access Biochemical reconstitution of TET1–TDG–BER-dependent active DNA demethylation reveals a highly coordinated mechanism
Biochemical reconstitution of TET1–TDG–BER-dependent active DNA demethylation reveals a highly coordinated mechanism
Cytosine methylation is a dynamic DNA modification with the involvement of the base excision repair pathway suspected to be involved in demethylation. Here the authors show that TET1 and TDG interact to target modified bases and coordinate BER to avoid double strand breaks.
- Alain R. Weber
- , Claudia Krawczyk
- & Primo Schär
-
Article
| Open Access Mutation allele burden remains unchanged in chronic myelomonocytic leukaemia responding to hypomethylating agents
Mutation allele burden remains unchanged in chronic myelomonocytic leukaemia responding to hypomethylating agents
Chronic myelomonocytic leukaemia is treated with agents that modify DNA methylation but whether they have direct cytotoxic effects is unclear. Here, the authors show that cells from treated patients show marked methylation changes without altered somatic mutation burden, suggesting that cytotoxicity is not a major factor in therapeutic efficacy.
- Jane Merlevede
- , Nathalie Droin
- & Eric Solary
-
Article
| Open Access Maternal plasma folate impacts differential DNA methylation in an epigenome-wide meta-analysis of newborns
Maternal plasma folate impacts differential DNA methylation in an epigenome-wide meta-analysis of newborns
Folic acid is routinely recommended for women trying to conceive to ensure proper fetal development. Here, the authors perform a large epigenomics study to examine which fetal epigenetic changes are associated with varied maternal plasma folate levels.
- Bonnie R. Joubert
- , Herman T. den Dekker
- & Stephanie J. London
-
Article
| Open Access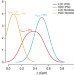 Non-CG DNA methylation is a biomarker for assessing endodermal differentiation capacity in pluripotent stem cells
Non-CG DNA methylation is a biomarker for assessing endodermal differentiation capacity in pluripotent stem cells
The methylation of non-CpG residues is a poorly understood marker of pluripotent cells, gradually lost as cells differentiate. Here the authors show non-CG methylation can be used as a marker of differentiation potential.
- Lee M. Butcher
- , Mitsuteru Ito
- & Stephan Beck
-
Article
| Open Access DNA methylation outliers in normal breast tissue identify field defects that are enriched in cancer
DNA methylation outliers in normal breast tissue identify field defects that are enriched in cancer
Altered epigenetics is a feature of cancer but whether these changes occur early in tumour development is unclear. Here, the authors analyse methylation events in breast cancer and adjacent normal pairs, and show that methylation changes in the normal tissue are also found in the tumour, suggesting that some of these events occur early in cancer.
- Andrew E Teschendorff
- , Yang Gao
- & Martin Widschwendter
-
Article
| Open Access NSD1 mutations generate a genome-wide DNA methylation signature
NSD1 mutations generate a genome-wide DNA methylation signature
Sotos syndrome is an growth syndrome characterized by advanced growth in childhood, characteristic facial appearance and intellectual disability. Here the authors identify a genome-wide DNA methylation signature that accurately diagnoses Sotos Syndrome and distinguishes it from similar conditions.
- S. Choufani
- , C. Cytrynbaum
- & R. Weksberg
-
Article
| Open Access H19 lncRNA alters DNA methylation genome wide by regulating S-adenosylhomocysteine hydrolase
H19 lncRNA alters DNA methylation genome wide by regulating S-adenosylhomocysteine hydrolase
DNA methylation is an important regulatory process and is essential for correct development and physiology. Here the authors show the long non-coding RNA H19regulates methylation by binding and inhibiting S-adenosylhomocysteine hydrolase.
- Jichun Zhou
- , Lihua Yang
- & Yingqun Huang
-
Article
| Open Access Hypomethylation of smoking-related genes is associated with future lung cancer in four prospective cohorts
Hypomethylation of smoking-related genes is associated with future lung cancer in four prospective cohorts
Smoking tobacco is known to alter DNA methylation. Here, the authors show that hypomethylation of smoke-related genes is associated with future increase in lung cancer risk.
- Francesca Fasanelli
- , Laura Baglietto
- & Paolo Vineis
-
Article
| Open Access An LSC epigenetic signature is largely mutation independent and implicates the HOXA cluster in AML pathogenesis
An LSC epigenetic signature is largely mutation independent and implicates the HOXA cluster in AML pathogenesis
Leukaemic stem cells are prevalent in acute myeloid leukemia. Here, Jung and colleagues derive a signature of 71 methylated genes that characterise these stem cells and find multiple HOXAgenes within the signature.
- Namyoung Jung
- , Bo Dai
- & Andrew P. Feinberg
-
Article
| Open Access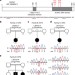 Mutations in CDCA7 and HELLS cause immunodeficiency–centromeric instability–facial anomalies syndrome
Mutations in CDCA7 and HELLS cause immunodeficiency–centromeric instability–facial anomalies syndrome
Immunodeficiency-centromeric instability-facial anomalies syndrome is a life threatening autosomal recessive disorder caused by mutations in DNMT3B and ZBTB24. Here Thijssen et al. identify mutations in CDCA7 and HELLSin previously unexplained cases.
- Peter E. Thijssen
- , Yuya Ito
- & Hiroyuki Sasaki
-
Article
| Open Access DNA methylation of oestrogen-regulated enhancers defines endocrine sensitivity in breast cancer
DNA methylation of oestrogen-regulated enhancers defines endocrine sensitivity in breast cancer
The molecular factors influencing patient response to endocrine therapy are poorly understood. Here Stone et al.characterize the DNA methylome of endocrine response and show that methylation of oestrogen receptor-associated enhancers underpins endocrine sensitivity in human breast cancer.
- Andrew Stone
- , Elena Zotenko
- & Susan J. Clark
-
Article
| Open Access Obesity-induced DNA hypermethylation of the adiponectin gene mediates insulin resistance
Obesity-induced DNA hypermethylation of the adiponectin gene mediates insulin resistance
The hormone adiponectin is produced by fat cells and has positive metabolic effects. Here, Kim et al.show that DNA methyltransferase 1 (DNMT1) represses adiponectin expression through hypermethylation of its promoter, and that inflammatory cytokines enhance DNMT1 activity in obese mice and humans.
- A. Young Kim
- , Yoon Jeong Park
- & Jae Bum Kim
-
Article
| Open Access Integrated genetic and epigenetic analysis defines novel molecular subgroups in rhabdomyosarcoma
Integrated genetic and epigenetic analysis defines novel molecular subgroups in rhabdomyosarcoma
Rhabdomyosarcoma is a common childhood soft-tissue cancer. Here Seki and Nishimura analyse the exome, transcriptome, copy number and DNA methylome of 60 sarcomas and identify distinct methylation subgroups associated with genetic and clinical features.
- Masafumi Seki
- , Riki Nishimura
- & Junko Takita
-
Article
| Open Access Single molecule-level detection and long read-based phasing of epigenetic variations in bacterial methylomes
Single molecule-level detection and long read-based phasing of epigenetic variations in bacterial methylomes
Bacterial DNA methylation is involved in many processes, from host defense to antibiotic resistance, however current methods for examining methylated genomes lack single-cell resolution. Here Beaulaurier et al. present Single Molecule Modification Analysis of Long Reads, a new tool for de novodetection of epigenetic heterogeneity.
- John Beaulaurier
- , Xue-Song Zhang
- & Gang Fang
-
Article
| Open Access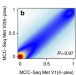 Characterization of functional methylomes by next-generation capture sequencing identifies novel disease-associated variants
Characterization of functional methylomes by next-generation capture sequencing identifies novel disease-associated variants
Currently, genome-wide methylation studies are limited to using targeted arrays or enrichment to assess large sample sizes. Here, Allum et al. demonstrate MethylC-Capture Sequencing, a cost-effective method for investigating genetic and epigenetic variation.
- Fiona Allum
- , Xiaojian Shao
- & Elin Grundberg
-
Article |
 Dissecting the role of aberrant DNA methylation in human leukaemia
Dissecting the role of aberrant DNA methylation in human leukaemia
Chronic myeloid leukaemia is characterized by the genetic translocation t(9;22) encoding for BCR-ABL oncogene; however, the molecular mechanisms of disease progression are poorly understood. Here Amabile et al. show that aberrant methylation is promoted by BCR-ABL, driving the evolution of the disease.
- Giovanni Amabile
- , Annalisa Di Ruscio
- & Daniel G. Tenen
-
Article
| Open Access Molecular mechanism for USP7-mediated DNMT1 stabilization by acetylation
Molecular mechanism for USP7-mediated DNMT1 stabilization by acetylation
DNMT1 is a methyl-transferase involved in maintaining tissue-specific patterns of DNA methylation. Here the authors solve the structure of a DNMT1-USP7 complex and demonstrate the mechanism by which DNMT1 stability is regulated through acetylation by preventing association with the deubiquitinase USP7.
- Jingdong Cheng
- , Huirong Yang
- & Yanhui Xu
-
Article |
 Epigenetic variation in the Egfr gene generates quantitative variation in a complex trait in ants
Epigenetic variation in the Egfr gene generates quantitative variation in a complex trait in ants
Variation in complex traits is generated by the interaction of genetic and environmental factors. Here, the authors show that genome-wide DNA methylation indirectly regulates quantitative methylation of the Egfrgene to generate continuous size variation of larvae workers in the carpenter ant.
- Sebastian Alvarado
- , Rajendhran Rajakumar
- & Moshe Szyf
-
Article |
 Genome-wide association study identifies peanut allergy-specific loci and evidence of epigenetic mediation in US children
Genome-wide association study identifies peanut allergy-specific loci and evidence of epigenetic mediation in US children
Food allergy is a growing clinical and public health burden. Here, the authors carry out a genome-wide association study in samples with well-defined allergies to a variety of foods, and identify the 6p21.32 region that significantly increases risk of developing peanut allergy.
- Xiumei Hong
- , Ke Hao
- & Xiaobin Wang
-
Article |
 Developmental enhancers revealed by extensive DNA methylome maps of zebrafish early embryos
Developmental enhancers revealed by extensive DNA methylome maps of zebrafish early embryos
DNA methylation undergoes dynamic changes during development and cell differentiation. Here, by comparing DNA methylomes from different stages of embryonic development of the zebrafish, the authors suggest that developmental enhancers are a major target of DNA methylation changes during embryogenesis.
- Hyung Joo Lee
- , Rebecca F. Lowdon
- & Ting Wang
-
Article
| Open Access Intermediate DNA methylation is a conserved signature of genome regulation
Intermediate DNA methylation is a conserved signature of genome regulation
Many loci in the mammalian genome are intermediately methylated. Here, by comprehensively identifying these loci and quantifying their relationship with gene activity, the authors show that intermediate methylation is an evolutionarily conserved epigenomic signature of gene regulation.
- GiNell Elliott
- , Chibo Hong
- & Joseph F. Costello
-
Article
| Open Access Epigenetic regulation of Atrophin1 by lysine-specific demethylase 1 is required for cortical progenitor maintenance
Epigenetic regulation of Atrophin1 by lysine-specific demethylase 1 is required for cortical progenitor maintenance
Histone modification is critical for gene expression regulation during development. Here, the authors show that the demethylase LSD1 and its target gene ATN1are responsible for maintenance of neural progenitor cells during mouse cortical development.
- Feng Zhang
- , Dan Xu
- & Zhiheng Xu
-
Article
| Open Access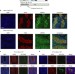 MORC1 represses transposable elements in the mouse male germline
MORC1 represses transposable elements in the mouse male germline
The Microrchidia (Morc) family of GHKL ATPases are important repressors of transposons and other DNA-methylated and silent genes in A. thaliana. Here, the authors show that MORC1 is responsible for repression and methylation of specific classes of transposons in the mouse male germline.
- William A. Pastor
- , Hume Stroud
- & Steven E. Jacobsen
-
Article
| Open Access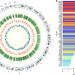 An integrated epigenomic analysis for type 2 diabetes susceptibility loci in monozygotic twins
An integrated epigenomic analysis for type 2 diabetes susceptibility loci in monozygotic twins
Type 2 diabetes (T2D) is a highly heterogeneous disease with a strong genetic component. Here the authors examine genome-wide methylation patterns in T2D-discordant, T2D-concordant and healthy concordant monozygotic twin pairs, and identify DNA methylation signals that may represent new biomarkers or drug targets for T2D.
- Wei Yuan
- , Yudong Xia
- & Tim D. Spector
-
Article
| Open Access An epigenomic roadmap to induced pluripotency reveals DNA methylation as a reprogramming modulator
An epigenomic roadmap to induced pluripotency reveals DNA methylation as a reprogramming modulator
Somatic cell reprogramming can induce distinct pluripotent states. Here the authors perform time-resolved whole-genome bisulfite sequencing during the reprogramming of mouse embryonic fibroblasts and report dynamic global DNA methylation changes.
- Dong-Sung Lee
- , Jong-Yeon Shin
- & Jeong-Sun Seo
-
Article
| Open Access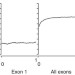 DNA methylation signatures link prenatal famine exposure to growth and metabolism
DNA methylation signatures link prenatal famine exposure to growth and metabolism
The long-term effect of prenatal nutrition on gene regulation is largely unknown. Here the authors identify differentially methylated regions in whole blood from individuals exposed to famine early after conception, and show that these epigenetic changes may have adverse metabolic consequences later in life.
- Elmar W. Tobi
- , Jelle J. Goeman
- & Bastiaan T. Heijmans
-
Article |
 Age-related variations in the methylome associated with gene expression in human monocytes and T cells
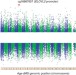
Age-related variations in the methylome associated with gene expression in human monocytes and T cells
The functional relevance of age-related variation in DNA methylation is unclear. Here, Reynolds et al. analyze how patterns of genome-wide gene expression and DNA methylation data vary with age in circulating monocytes and T cells, and report age-associated methylation signals that are correlated with cis-gene expression and vascular aging.
- Lindsay M. Reynolds
- , Jackson R. Taylor
- & Yongmei Liu
-
Article
| Open Access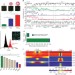 Dynamic DNA methylation orchestrates cardiomyocyte development, maturation and disease
Dynamic DNA methylation orchestrates cardiomyocyte development, maturation and disease
DNA methylation is essential for proper gene expression, development and genome stability. Here the authors present whole-genome DNA methylation analyses of purified mouse cardiomyocytes from newborn, adult and failing hearts and find highly dynamic patterns between the three phenotypes of cardiomyocytes.
- Ralf Gilsbach
- , Sebastian Preissl
- & Lutz Hein
-
Article |
 The meta-epigenomic structure of purified human stem cell populations is defined at cis-regulatory sequences
The meta-epigenomic structure of purified human stem cell populations is defined at cis-regulatory sequences
There is epigenetic variability in the same cell type among healthy individuals, but the mechanism or significance of this variability is not clear. Here, the authors purify CD34+ cells from different individuals and use meta-epigenomic approaches to analyse and explain the epigenetic variability observed.
- N. Ari Wijetunga
- , Fabien Delahaye
- & John M. Greally
-
Article
| Open Access Matrix softness regulates plasticity of tumour-repopulating cells via H3K9 demethylation and Sox2 expression
Matrix softness regulates plasticity of tumour-repopulating cells via H3K9 demethylation and Sox2 expression
Soft 3D gels can promote the growth of tumour-repopulating cells, a self-renewing subpopulation of cancer cells critical in cancer progression. Here, the authors investigate the mechanism behind this phenomenon and show that the histone 3 lysine residue 9 methylation and Sox2 are controlling this process.
- Youhua Tan
- , Arash Tajik
- & Ning Wang

 Tet2 loss leads to hypermutagenicity in haematopoietic stem/progenitor cells
Tet2 loss leads to hypermutagenicity in haematopoietic stem/progenitor cells