Featured
-
-
Article
| Open Access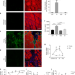 Age-related declines in α-Klotho drive progenitor cell mitochondrial dysfunction and impaired muscle regeneration
Age-related declines in α-Klotho drive progenitor cell mitochondrial dysfunction and impaired muscle regeneration
While young muscle faithfully regenerates damaged myofibers, aged muscle is impaired. Here the authors show the “anti-aging” protein α-Klotho is upregulated in young muscle after damage via promoter demethylation and this regulation is lost in aging, resulting in mitochondrial damage and an impaired healing response.
- A. Sahu
- , H. Mamiya
- & F. Ambrosio
-
Article
| Open Access Epigenetic profiling for the molecular classification of metastatic brain tumors
Epigenetic profiling for the molecular classification of metastatic brain tumors
The treatment of brain metastases is often limited by the ability to diagnose their origins. Here the authors generate DNA methylomes from the three most frequent types of brain metastases, identify epigenetic signatures specific to each type of metastasis and construct a DNA methylation-based classifier (BrainMETH) to advance brain metastasis diagnosis.
- Javier I. J. Orozco
- , Theo A. Knijnenburg
- & Diego M. Marzese
-
Article
| Open Access Arabidopsis AGDP1 links H3K9me2 to DNA methylation in heterochromatin
Arabidopsis AGDP1 links H3K9me2 to DNA methylation in heterochromatin
DNA methylation and H3K9 dimethylation are two linked epigenetic marks of silenced chromatin in plants that depend on the activity of CMT3/2 and SUVH4/5/6. Here the authors identify AGDP1 as an H3K9me2-binding protein required for heterochromatic non-CG DNA methylation, H3K9 dimethylation, and transcriptional silencing.
- Cuijun Zhang
- , Xuan Du
- & Jian-Kang Zhu
-
Article
| Open Access Methylation of all BRCA1 copies predicts response to the PARP inhibitor rucaparib in ovarian carcinoma
Methylation of all BRCA1 copies predicts response to the PARP inhibitor rucaparib in ovarian carcinoma
Around 10% of high-grade serous ovarian carcinomas (HGSOC) harbor BRCA1 promoter methylation, but it is uncertain how it predicts response to PARP inhibition. Here, the authors show that homozygous BRCA1 methylation predicts response to rucaparib while heterozygous methylation of BRCA1 predicts resistance in HGSOC.
- Olga Kondrashova
- , Monique Topp
- & Clare L. Scott
-
Article
| Open Access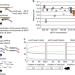 Robust single-cell DNA methylome profiling with snmC-seq2
Robust single-cell DNA methylome profiling with snmC-seq2
Single-cell DNA methylome profiling allows the study of epigenomic heterogeneity in tissues but has been impeded by library quality. Here the authors demonstrate snmC-seq2 which improves mapping, throughput and library complexity.
- Chongyuan Luo
- , Angeline Rivkin
- & Joseph R. Ecker
-
Article
| Open Access Autosomal genetic variation is associated with DNA methylation in regions variably escaping X-chromosome inactivation
Autosomal genetic variation is associated with DNA methylation in regions variably escaping X-chromosome inactivation
DNA methylation is critically involved in X chromosome inactivation (XCI) and dosage compensation, yet some X-chromosomal genes escape XCI. Here, Lujik et al. identify three autosomal genetic loci that associate with differential DNA methylation near genes that variably escape XCI in females.
- René Luijk
- , Haoyu Wu
- & Bastiaan T. Heijmans
-
Article
| Open Access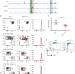 Stage-specific epigenetic regulation of CD4 expression by coordinated enhancer elements during T cell development
Stage-specific epigenetic regulation of CD4 expression by coordinated enhancer elements during T cell development
The expression of CD4, a critical co-receptor providing T cell help in adaptive immunity, is finely tuned during development. Here the authors show that two enhancer elements, E4p and the newly-defined E4m, coordinate the expression and heritable demethylation of Cd4 in thymocytes but are dispensable for its sustained expression in peripheral T cells.
- Priya D. Issuree
- , Kenneth Day
- & Dan R. Littman
-
Article
| Open Access High-fidelity CRISPR/Cas9- based gene-specific hydroxymethylation rescues gene expression and attenuates renal fibrosis
High-fidelity CRISPR/Cas9- based gene-specific hydroxymethylation rescues gene expression and attenuates renal fibrosis
Suppression of gene expression due to aberrant promoter methylation contributes to organ fibrosis. Here, the authors couple a deactivated Cas9 to the TET3 catalytic domain to induce expression of four antifibrotic genes, and show that lentiviral-mediated delivery is effective in reducing kidney fibrosis in mouse models.
- Xingbo Xu
- , Xiaoying Tan
- & Michael Zeisberg
-
Article
| Open Access The effect of maternal care on gene expression and DNA methylation in a subsocial bee
The effect of maternal care on gene expression and DNA methylation in a subsocial bee
Development may be plastic and influenced by parental care. Here, the authors show that experimental reduction of maternal care in the small carpenter bee leads to extensive changes in gene expression and splicing, minor changes in methylation, and greater offspring aggression and social avoidance.
- Samuel V. Arsenault
- , Brendan G. Hunt
- & Sandra M. Rehan
-
Article
| Open Access Epigenetic dysregulation of naive CD4+ T-cell activation genes in childhood food allergy
Epigenetic dysregulation of naive CD4+ T-cell activation genes in childhood food allergy
Immunoglobulin E (IgE)-mediated food allergy is a major issue that affects 2–10% of infants. Here the authors study the epigenetic regulation of the naive CD4+ T cell activation response among children with IgE-mediated food allergy finding epigenetic dysregulation in the early stages of signal transduction through the T cell receptor complex.
- David Martino
- , Melanie Neeland
- & Richard Saffery
-
Article
| Open Access Epigenome-wide DNA methylation profiling in Progressive Supranuclear Palsy reveals major changes at DLX1
Epigenome-wide DNA methylation profiling in Progressive Supranuclear Palsy reveals major changes at DLX1
Progressive supranuclear palsy (PSP) is a neurodegenerative disease characterized by aggregation of Tau, encoded by MAPT. Here, the authors perform an EWAS for PSP in prefrontal lobe tissue and find hypermethylation of DLX1 and its antisense transcript DLX1AS to associate with MAPT expression.
- Axel Weber
- , Sigrid C. Schwarz
- & Ulrich Müller
-
Article
| Open Access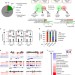 Uhrf1 regulates active transcriptional marks at bivalent domains in pluripotent stem cells through Setd1a
Uhrf1 regulates active transcriptional marks at bivalent domains in pluripotent stem cells through Setd1a
Uhrf1 is a known regulator of heterochromatin and DNA methylation in embryonic stem cells (ESCs). Here, the authors demonstrate that Uhrf1 acts together with the Set1/COMPASS complex regulator of active transcription to promote H3K4 methylation at bivalent loci and Uhrf1 loss results in disruption of differentiation.
- Kun-Yong Kim
- , Yoshiaki Tanaka
- & In-Hyun Park
-
Article
| Open Access DNA methylation as a mediator of HLA-DRB1*15:01 and a protective variant in multiple sclerosis
DNA methylation as a mediator of HLA-DRB1*15:01 and a protective variant in multiple sclerosis
The human leukocyte antigen (HLA) haplotype DRB1*15:01 is the major risk factor for multiple sclerosis (MS). Here the authors find that DNA methylation at HLA-DRB1 gene mediates the effect of DRB1*15:01 and of a protective HLA variant on HLA-DRB1 expression and the risk of MS.
- Lara Kular
- , Yun Liu
- & Maja Jagodic
-
Article
| Open Access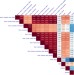 Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood
Identifying gene targets for brain-related traits using transcriptomic and methylomic data from blood
To comprehend the genetic regulatory mechanisms underlying brain-related traits in humans, Qi et al. estimate the correlation of expression and DNA methylation QTL effects in cis between blood and brain and show that using blood eQTL/mQTL data of large sample size can increase power in gene discovery for brain-related traits and diseases.
- Ting Qi
- , Yang Wu
- & Jian Yang
-
Article
| Open Access Tet1 and Tet2 maintain mesenchymal stem cell homeostasis via demethylation of the P2rX7 promoter
Tet1 and Tet2 maintain mesenchymal stem cell homeostasis via demethylation of the P2rX7 promoter
Tet-mediated DNA oxidation converts 5-methylcytosine (5-mC) to 5-hydroxymethylcytosine (5-hmC), which is essential to regulate different biological processes. Here the authors show that Tet1 and Tet2 regulate mesenchymal stem cell and bone homeostasis through demethylation of P2rX7 promoter.
- Ruili Yang
- , Tingting Yu
- & Songtao Shi
-
Article
| Open Access Identification of rare de novo epigenetic variations in congenital disorders
Identification of rare de novo epigenetic variations in congenital disorders
A proportion of neurodevelopmental disorder and congenital anomaly cases remain without a genetic diagnosis. Here, the authors study aberrations of DNA methylation in such cases and find that epivariations might provide an explanation for some of these undiagnosed patients.
- Mafalda Barbosa
- , Ricky S. Joshi
- & Andrew J. Sharp
-
Article
| Open Access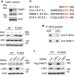 Methylated DNMT1 and E2F1 are targeted for proteolysis by L3MBTL3 and CRL4DCAF5 ubiquitin ligase
Methylated DNMT1 and E2F1 are targeted for proteolysis by L3MBTL3 and CRL4DCAF5 ubiquitin ligase
Lysine methylation is increasingly being implicated in the modification of non-histone proteins. Here the authors find that the methylation of DNMT1 and E2F1 are recognized by the protein L3MBTL3 and the ubiquitin E3 ligase CRL4DCAF5, which cooperatively target these methylated proteins for ubiquitin-dependent proteolysis.
- Feng Leng
- , Jiekai Yu
- & Hui Zhang
-
Article
| Open Access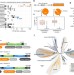 Recurrent acquisition of cytosine methyltransferases into eukaryotic retrotransposons
Recurrent acquisition of cytosine methyltransferases into eukaryotic retrotransposons
Cytosine methyltransferases (DNMTs) often silence transposons in eukaryotic genomes. Here the authors describe the recurrent acquisition of DNMTs by transposons from two distantly-related eukaryotes and suggest that methylation of CG dinucleotides by transposon DNMTs could modify the host epigenome in dinoflagellates.
- Alex de Mendoza
- , Amandine Bonnet
- & Ryan Lister
-
Article
| Open Access Heritable DNA methylation marks associated with susceptibility to breast cancer
Heritable DNA methylation marks associated with susceptibility to breast cancer
DNA methylation is associated with breast cancer risk. Here the authors measure DNA methylation in the blood of individuals from 25 Australian families with multiple cases of breast cancer but not known mutations associated with breast cancer risk to identify possible heritable methylation markers.
- Jihoon E. Joo
- , James G. Dowty
- & Yoland Antill
-
Article
| Open Access Co-occurring expression and methylation QTLs allow detection of common causal variants and shared biological mechanisms
Co-occurring expression and methylation QTLs allow detection of common causal variants and shared biological mechanisms
Most expression QTLs (eQTLs) co-occur with a DNA methylation QTL (meQTL), suggesting a common causal variant. Here the authors analyse DNA and RNA from blood and identify eQTL-meQTL pairs likely to share a causal variant, finding that expression and methylation are often genetically co-regulated.
- Brandon L. Pierce
- , Lin Tong
- & Habibul Ahsan
-
Article
| Open Access scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells
scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells
Relationships between DNA methylation and transcription, and methylation and DNA accessibility can be probed but interrogating all three in the same single cells has not been possible. Here, the authors report the first single-cell method for parallel chromatin accessibility, DNA methylation and transcriptome profiling.
- Stephen J. Clark
- , Ricard Argelaguet
- & Wolf Reik
-
Article
| Open Access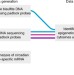 Cytosine modifications exhibit circadian oscillations that are involved in epigenetic diversity and aging
Cytosine modifications exhibit circadian oscillations that are involved in epigenetic diversity and aging
While epigenetic factors have been implicated in the circadian rhythm, the detection of circadian cytosine modifications has remained elusive. Here the authors identify a large number of epigenetically variable cytosines that show circadian oscillations in their modification status in mice.
- Gabriel Oh
- , Sasha Ebrahimi
- & Art Petronis
-
Article
| Open Access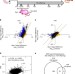 Epigenetic modulation of Fgf21 in the perinatal mouse liver ameliorates diet-induced obesity in adulthood
Epigenetic modulation of Fgf21 in the perinatal mouse liver ameliorates diet-induced obesity in adulthood
FGF21 exerts beneficial metabolic effects on multiple tissues. Here the authors show that the Fgf21 gene is demethylated during the postnatal suckling period, creating an epigenetic memory that determines the responsiveness of the Fgf21 gene to inducers such as PPARα activators or fasting in adulthood.
- Xunmei Yuan
- , Kazutaka Tsujimoto
- & Yoshihiro Ogawa
-
Article
| Open Access Methylation profiling identifies two subclasses of squamous cell carcinoma related to distinct cells of origin
Methylation profiling identifies two subclasses of squamous cell carcinoma related to distinct cells of origin
Cutaneous squamous cell carcinoma (cSCC) is a skin cancer that normally progresses from UV-induced actinic keratosis (AK). Here, the authors investigate the epigenomics of cSCC and highlight two distinct subclasses of AK and cSCC originating from distinct keratinocyte differentiation stages.
- Manuel Rodríguez-Paredes
- , Felix Bormann
- & Frank Lyko
-
Article
| Open Access A naturally occurring epiallele associates with leaf senescence and local climate adaptation in Arabidopsis accessions
A naturally occurring epiallele associates with leaf senescence and local climate adaptation in Arabidopsis accessions
Epigenetic variation underlies various aspects of phenotypic diversity of plants. Here, He et al show a naturally occurring epiallele controls Arabidopsis leaf senescence by regulating the expression of PHEOPHYTIN PHEOPHORBIDE HYDROLASE (PPH), and is associated with local climate adaptation.
- Li He
- , Wenwu Wu
- & Jian-Kang Zhu
-
Article
| Open Access Distinct epigenetic programs regulate cardiac myocyte development and disease in the human heart in vivo
Distinct epigenetic programs regulate cardiac myocyte development and disease in the human heart in vivo
How the cardiac myocyte epigenome is rearranged during development, postnatal maturation and disease is not well understood. Here, the authors investigate the human cardiac myocyte epigenome during development and chronic heart failure and identify distinct epigenetic programs regulating these processes.
- Ralf Gilsbach
- , Martin Schwaderer
- & Lutz Hein
-
Article
| Open Access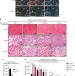 Lsd1 regulates skeletal muscle regeneration and directs the fate of satellite cells
Lsd1 regulates skeletal muscle regeneration and directs the fate of satellite cells
Satellite cells can differentiate both into myocytes and brown adipocytes. Here, the authors show that the histone demethylase Lsd1 prevents adipogenic differentiation of satellite cells by repressing expression of Glis1, and that its ablation changes satellite cell fate towards brown adipocytes and delays muscle regeneration in mice.
- Milica Tosic
- , Anita Allen
- & Roland Schüle
-
Article
| Open Access A PRDX1 mutant allele causes a MMACHC secondary epimutation in cblC patients
A PRDX1 mutant allele causes a MMACHC secondary epimutation in cblC patients
Inborn errors of vitamin B12 metabolism of the cblC class are caused by mutations in the MMACHC gene. Here, Guéant et al. report epi-cblC, a class of cblC in which patients are compound heterozygous for a genetic mutation and a secondary epimutation at the MMACHC locus.
- Jean-Louis Guéant
- , Céline Chéry
- & David S. Rosenblatt
-
Article
| Open Access Diabetes impairs wound healing by Dnmt1-dependent dysregulation of hematopoietic stem cells differentiation towards macrophages
Diabetes impairs wound healing by Dnmt1-dependent dysregulation of hematopoietic stem cells differentiation towards macrophages
Type 2 diabetes is associated with impaired wound healing, which can lead to limb loss. Here, the authors show that in Type 2 diabetic mouse models, Dnmt1 is upregulated in hematopoietic stem cells, leading to impaired differentiation towards macrophages, reduced macrophage infiltration in the wound and skewed M1/M2 polarization.
- Jinglian Yan
- , Guodong Tie
- & Louis M. Messina
-
Article
| Open Access RAS-pathway mutation patterns define epigenetic subclasses in juvenile myelomonocytic leukemia
RAS-pathway mutation patterns define epigenetic subclasses in juvenile myelomonocytic leukemia
Juvenile myelomonocytic leukemia (JMML) is an aggressive disease with limited options for treatment. Here, the authors analyse the DNA methylome and mutational profile of JMML to define three subgroups with unique molecular and clinical characteristics.
- Daniel B. Lipka
- , Tania Witte
- & Christoph Plass
-
Article
| Open Access Stable transgenerational epigenetic inheritance requires a DNA methylation-sensing circuit
Stable transgenerational epigenetic inheritance requires a DNA methylation-sensing circuit
DNA methylation patterns are inherited over many generations in plants. Here, Williams and Gehring show that the 5-methylcytosine DNA glycosylase ROS1 functions as part of a methylation-sensitive circuit that ensures long-term epigenetic fidelity in Arabidopsis.
- Ben P. Williams
- & Mary Gehring
-
Article
| Open Access Ancestral perinatal obesogen exposure results in a transgenerational thrifty phenotype in mice
Ancestral perinatal obesogen exposure results in a transgenerational thrifty phenotype in mice
Early life exposure to endocrine disrupting chemicals has been linked to increased adiposity during adulthood. Here Chamorro-García et al. show that ancestral exposure to the obesogen tributyltin causes obesity in untreated F4 generation male descendants by inducing heritable changes in genome architecture that promote a thrifty phenotype.
- Raquel Chamorro-Garcia
- , Carlos Diaz-Castillo
- & Bruce Blumberg
-
Article
| Open Access DNA methylation signatures follow preformed chromatin compartments in cardiac myocytes
DNA methylation signatures follow preformed chromatin compartments in cardiac myocytes
Chromatin is organized in higher order A and B compartments, reflecting active and inactive chromatin. Here, the authors provide evidence that in cardiac myocytes DNA methylation is established in preformed chromatin compartments and may be dispensable for higher order chromatin organization.
- Stephan Nothjunge
- , Thomas G. Nührenberg
- & Ralf Gilsbach
-
Article
| Open Access Hit-and-run epigenetic editing prevents senescence entry in primary breast cells from healthy donors
Hit-and-run epigenetic editing prevents senescence entry in primary breast cells from healthy donors
“Although aberrant promoter DNA hypermethylation is a hallmark of cancer, it is not clear whether it is sufficient to drive transformation. Here, the authors use CRISPR-dCas9 to perform hit-and-run epigenetic editing, which prevents senescence entry in primary breast cells from healthy donors.”
- Emily A. Saunderson
- , Peter Stepper
- & Gabriella Ficz
-
Article
| Open Access Genome-wide genetic and epigenetic analyses of pancreatic acinar cell carcinomas reveal aberrations in genome stability
Genome-wide genetic and epigenetic analyses of pancreatic acinar cell carcinomas reveal aberrations in genome stability
Pancreatic acinar cell carcinoma (ACC) is an aggressive exocrine tumor with largely unknown biology. Here, the authors perform genome- and epigenome-wide analyses from normal and ACC pancreatic tissue that identify aberrations in genome stability and cell cycle control.
- Cornelia Jäkel
- , Frank Bergmann
- & Peter Schmezer
-
Article
| Open Access Epigenome-wide association studies identify DNA methylation associated with kidney function
Epigenome-wide association studies identify DNA methylation associated with kidney function
Genome-wide association studies of kidney function show enrichment of associated genetic variants in regulatory regions. Here, the authors perform epigenome-wide association studies of kidney function and disease, identifying 19 CpG sites significantly associated with these.
- Audrey Y. Chu
- , Adrienne Tin
- & Anna Köttgen
-
Article
| Open Access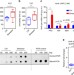 DNA N6-methyladenine is dynamically regulated in the mouse brain following environmental stress
DNA N6-methyladenine is dynamically regulated in the mouse brain following environmental stress
N6-methyladenine is a covalent epigenetic modification of the genome. Here, Yao and colleagues show that N6-methyladenine level in the mouse brain is dynamic following environmental stress, and the subsequent differential gene expression is correlated with LINE transposon expression.
- Bing Yao
- , Ying Cheng
- & Peng Jin
-
Article
| Open Access DNA methylation and transcriptional trajectories during human development and reprogramming of isogenic pluripotent stem cells
DNA methylation and transcriptional trajectories during human development and reprogramming of isogenic pluripotent stem cells
While DNA methylation and gene expression data are widely available for animal models, comprehensive data from human development is rarer. Here, the authors generated transcriptional and DNA methylation data from 21 organs during human development and 6 isogenic induced pluripotent stem cell lines.
- Matthias S. Roost
- , Roderick C. Slieker
- & Susana M. Chuva de Sousa Lopes
-
Article
| Open Access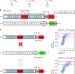 Deciphering TAL effectors for 5-methylcytosine and 5-hydroxymethylcytosine recognition
Deciphering TAL effectors for 5-methylcytosine and 5-hydroxymethylcytosine recognition
Transcription activator-like effector proteins recognise specific DNA sequences via tandem repeats. Here the authors demonstrate TALEs can recognise the methylated bases 5mC and 5hmC, enabling them to detect epigenetic modifications.
- Yuan Zhang
- , Lulu Liu
- & Chengqi Yi
-
Article
| Open Access Caloric restriction delays age-related methylation drift
Caloric restriction delays age-related methylation drift
Caloric restriction has been shown to increase lifespan in mammals. Here, the authors provide evidence that age-related methylation drift correlates with lifespan and that caloric restriction in mice and rhesus monkeys results in attenuation of age-related methylation drift.
- Shinji Maegawa
- , Yue Lu
- & Jean-Pierre J. Issa
-
Article
| Open Access Blood monocyte transcriptome and epigenome analyses reveal loci associated with human atherosclerosis
Blood monocyte transcriptome and epigenome analyses reveal loci associated with human atherosclerosis
The molecular mechanisms mediating the impact of environmental factors in atherosclerosis are unclear. Here, the authors examine CD14+ blood monocyte’s transcriptome and epigenome signatures to find differential methylation and expression of ARID5B to be associated with human atherosclerosis.
- Yongmei Liu
- , Lindsay M. Reynolds
- & James H. Stein
-
Article
| Open Access Targeted DNA methylation in vivo using an engineered dCas9-MQ1 fusion protein
Targeted DNA methylation in vivo using an engineered dCas9-MQ1 fusion protein
Understanding how DNA methylation regulates gene expression requires the capacity to deploy it to regions of interest. The authors generate a highly rapid and locus-specific CpG methylation tool by fusing dCas9 to MQ1 DNA methyltransferase and show efficacy at multiple sitesin vitro and in vivo.
- Yong Lei
- , Xiaotian Zhang
- & Margaret A. Goodell
-
Article
| Open Access Ten-eleven translocation 2 interacts with forkhead box O3 and regulates adult neurogenesis
Ten-eleven translocation 2 interacts with forkhead box O3 and regulates adult neurogenesis
Epigenetic modifications, such as DNA methylation, play an important role in adult neurogenesis. Here the authors show that Tet2, which converts 5mC to 5hmC, interacts with the transcription factor Foxo3a and regulates critical genes related to the proliferation and differentiation of adult neural stem cells.
- Xuekun Li
- , Bing Yao
- & Peng Jin
-
Article
| Open Access Drug-seeking motivation level in male rats determines offspring susceptibility or resistance to cocaine-seeking behaviour
Drug-seeking motivation level in male rats determines offspring susceptibility or resistance to cocaine-seeking behaviour
Drug addiction is partially heritable but the non-genetic inheritance mechanisms are not well understood. The authors show that motivation of male rats in response to cocaine self-administration elicit susceptibility and/or decreased resistance to developing addiction like behaviour in offspring.
- Qiumin Le
- , Biao Yan
- & Lan Ma
-
Article
| Open Access Design of synthetic epigenetic circuits featuring memory effects and reversible switching based on DNA methylation
Design of synthetic epigenetic circuits featuring memory effects and reversible switching based on DNA methylation
Recording systems would allow synthetic organisms to store a ‘memory’ of a past event for future reference. Here the authors design an epigenetic memory system inE. colithat methylates DNA in response to exogenous and endogenous signals.
- Johannes A. H. Maier
- , Raphael Möhrle
- & Albert Jeltsch
-
Article
| Open Access Genetic architecture of epigenetic and neuronal ageing rates in human brain regions
Genetic architecture of epigenetic and neuronal ageing rates in human brain regions
Studies on the ‘epigenetic clock’, a recently identified ageing biomarker, suggest that pathology might be linked to tissue-specific accelerated ageing. Here, the authors investigate ageing in the human brain and identify genetic loci associated with accelerated ageing in different brain regions.
- Ake T. Lu
- , Eilis Hannon
- & Steve Horvath
-
Article
| Open Access Intron retention is regulated by altered MeCP2-mediated splicing factor recruitment
Intron retention is regulated by altered MeCP2-mediated splicing factor recruitment
Intron retention is a conserved mechanism that controls gene expression but its regulation is poorly understood. Here, the authors provide evidence that DNA methylation regulates intron retention and find reduced MeCP2 occupancy and splicing factor recruitment near affected splice junctions.
- Justin J. -L. Wong
- , Dadi Gao
- & John E. J. Rasko
-
Article
| Open Access Tet2 loss leads to hypermutagenicity in haematopoietic stem/progenitor cells
Tet2 loss leads to hypermutagenicity in haematopoietic stem/progenitor cells
TET2 catalyses DNA demethylation and is mutated in various blood cancers; in particularTet2null mice develop haematological neoplasms. Here the authors show that this effect could be due to the increased frequency of mutation associated with TET2 loss in haematopoietic stem/progenitor cells.
- Feng Pan
- , Thomas S. Wingo
- & Mingjiang Xu
-
Article
| Open Access Epigenetically-driven anatomical diversity of synovial fibroblasts guides joint-specific fibroblast functions
Epigenetically-driven anatomical diversity of synovial fibroblasts guides joint-specific fibroblast functions
Arthritis affects different joints variably despite systemic inflammatory cues. Here the authors show anatomical differences in the transcriptome, epigenome and function of synovial fibroblasts that might affect susceptibility to site-specific joint diseases.
- Mojca Frank-Bertoncelj
- , Michelle Trenkmann
- & Caroline Ospelt

 Comprehensive human cell-type methylation atlas reveals origins of circulating cell-free DNA in health and disease
Comprehensive human cell-type methylation atlas reveals origins of circulating cell-free DNA in health and disease