Featured
-
-
Article
| Open Access Determinants of transcription factor regulatory range
Determinants of transcription factor regulatory range
Characterization of the distance over which TF binding influences gene expression is important for inferring target genes. Here the authors study this relationship using thousands of genomic data sets, finding two classes of TFs with distinct ranges of regulatory influence modulated by chromatin states of topologically associated domains.
- Chen-Hao Chen
- , Rongbin Zheng
- & X. Shirley Liu
-
Article
| Open Access Integrating molecular markers into metabolic models improves genomic selection for Arabidopsis growth
Integrating molecular markers into metabolic models improves genomic selection for Arabidopsis growth
An increase in genomic selection (GS) accuracy can accelerate genetic gain by shortening the breeding cycles. Here, the authors introduce a network-based GS method that uses metabolic models and improves the prediction accuracy of Arabidopsis growth within and across environments.
- Hao Tong
- , Anika Küken
- & Zoran Nikoloski
-
Article
| Open Access Integration of multiple biological contexts reveals principles of synthetic lethality that affect reproducibility
Integration of multiple biological contexts reveals principles of synthetic lethality that affect reproducibility
Defining robust, penetrant synthetic lethal targets for cancer therapeutics is a challenge. Here, the authors demonstrate how pathway, genetic and cellular context dictates dependence on cancer targets.
- Angel A. Ku
- , Hsien-Ming Hu
- & Sourav Bandyopadhyay
-
Article
| Open Access Remodeling of active endothelial enhancers is associated with aberrant gene-regulatory networks in pulmonary arterial hypertension
Remodeling of active endothelial enhancers is associated with aberrant gene-regulatory networks in pulmonary arterial hypertension
Pulmonary arterial hypertension (PAH) is a heterogeneous disease, causing severe breathing problems and cardiac morbidity. Here, the authors study chromatin marks in pulmonary arterial endothelial cells from PAH patients and controls and find changes in transcription factor and enhancer activity that suggest an aberrant response to signalling in PAH.
- Armando Reyes-Palomares
- , Mingxia Gu
- & Judith B. Zaugg
-
Article
| Open Access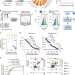 Pan-cancer analysis reveals cooperativity of both strands of microRNA that regulate tumorigenesis and patient survival
Pan-cancer analysis reveals cooperativity of both strands of microRNA that regulate tumorigenesis and patient survival
5p and 3p miRNA strands have different mRNA-targeting sequences and may both functionally impact gene expression in cancer. Here, the authors undertake a pan-cancer analysis that indicates 5p/3p miRNA strands function together to regulate tumorigenic processes.
- Ramkrishna Mitra
- , Clare M. Adams
- & Christine M. Eischen
-
Article
| Open Access Integrative pathway enrichment analysis of multivariate omics data
Integrative pathway enrichment analysis of multivariate omics data
Multi-omics datasets pose major challenges to data interpretation and hypothesis generation owing to their high-dimensional molecular profiles. Here, the authors develop ActivePathways method, which uses data fusion techniques for integrative pathway analysis of multi-omics data and candidate gene discovery.
- Marta Paczkowska
- , Jonathan Barenboim
- & Christian von Mering
-
Article
| Open Access The ETFL formulation allows multi-omics integration in thermodynamics-compliant metabolism and expression models
The ETFL formulation allows multi-omics integration in thermodynamics-compliant metabolism and expression models
Accounting for the effects of genetic expression in genome-scale metabolic models is challenging. Here, the authors introduce a model formulation that efficiently simulates thermodynamic-compliant fluxes, enzyme and mRNA concentration levels, allowing omics integration and broad analysis of in silico cellular physiology.
- Pierre Salvy
- & Vassily Hatzimanikatis
-
Article
| Open Access Agreement between two large pan-cancer CRISPR-Cas9 gene dependency data sets
Agreement between two large pan-cancer CRISPR-Cas9 gene dependency data sets
Integrating independent large-scale pharmacogenomic screens can enable unprecedented characterization of genetic vulnerabilities in cancers. Here, the authors show that the two largest independent CRISPR-Cas9 gene-dependency screens are concordant, paving the way for joint analysis of the data sets.
- Joshua M. Dempster
- , Clare Pacini
- & Francesco Iorio
-
Article
| Open Access Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers
Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers
Mass-spectrometry-based profiling can be used to stratify tumours into molecular subtypes. Here, by classifying over 500 tumours, the authors show that this approach reveals proteomic subgroups which cut across tumour types.
- Fengju Chen
- , Darshan S. Chandrashekar
- & Chad J. Creighton
-
Article
| Open Access A Bayesian machine learning approach for drug target identification using diverse data types
A Bayesian machine learning approach for drug target identification using diverse data types
Drug target identification is a crucial step in drug development. Here, the authors introduce a Bayesian machine learning framework that integrates multiple data types to predict the targets of small molecules, enabling identification of a new set of microtubule inhibitors and the target of the anti-cancer molecule ONC201.
- Neel S. Madhukar
- , Prashant K. Khade
- & Olivier Elemento
-
Article
| Open Access A network-based approach to identify deregulated pathways and drug effects in metabolic syndrome
A network-based approach to identify deregulated pathways and drug effects in metabolic syndrome
Metabolic syndrome is characterized by complex phenotypes that increases the risk of cardiovascular disease and type 2 diabetes. Here the authors’ integrative network analysis suggests BTK inhibitor ibrutinib to be a promising treatment through its obesity-associated inflammation lowering effect.
- Karla Misselbeck
- , Silvia Parolo
- & Corrado Priami
-
Article
| Open Access Macrophage-associated wound healing contributes to African green monkey SIV pathogenesis control
Macrophage-associated wound healing contributes to African green monkey SIV pathogenesis control
Here, the authors compare gene expression signatures in rectal tissues of African green monkeys (AGMs) and rhesus macaques (RMs) acutely infected with simian immunodeficiency virus and find that AGMs rapidly activate and maintain evolutionarily conserved regenerative wound healing mechanisms.
- Fredrik Barrenas
- , Kevin Raehtz
- & Michael Gale Jr
-
Article
| Open Access Enhanced and unified anatomical labeling for a common mouse brain atlas
Enhanced and unified anatomical labeling for a common mouse brain atlas
Anatomical brain atlases elucidate the anatomical and functional organisation across species but different atlases have conflicting anatomical border and 3D coordinates. The authors integrated two atlases into a unified and highly segmented anatomical labelling system of the mouse brain.
- Uree Chon
- , Daniel J. Vanselow
- & Yongsoo Kim
-
Article
| Open Access Full-length transcriptome reconstruction reveals a large diversity of RNA and protein isoforms in rat hippocampus
Full-length transcriptome reconstruction reveals a large diversity of RNA and protein isoforms in rat hippocampus
It is challenging to characterize diverse transcript isoforms by short-read sequencing. Here the authors report full-length transcriptomes in rat hippocampus by hybrid-sequencing, predict isoform-specific translational status, and reconstruct open reading frames validated by mass spectrometry.
- Xi Wang
- , Xintian You
- & Wei Chen
-
Article
| Open Access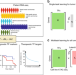 Chromatin-informed inference of transcriptional programs in gynecologic and basal breast cancers
Chromatin-informed inference of transcriptional programs in gynecologic and basal breast cancers
Epigenomic data on chromatin accessibility and transcription factor occupancy can reveal enhancer landscapes in cancer. Here, the authors develop a computational strategy called PSIONIC (patient-specific inference of networks informed by chromatin) to model the impact of enhancers on transcriptional programs in gynecologic and basal breast cancers.
- Hatice U. Osmanbeyoglu
- , Fumiko Shimizu
- & Christina S. Leslie
-
Article
| Open Access Integrative transcriptome imputation reveals tissue-specific and shared biological mechanisms mediating susceptibility to complex traits
Integrative transcriptome imputation reveals tissue-specific and shared biological mechanisms mediating susceptibility to complex traits
PrediXcan is a widely used gene expression imputation method that links genetic variants to gene expression. Here, the authors develop EpiXcan which leverages epigenetic annotations to inform transcriptomic imputation and further use the obtained gene-trait associations for computational drug repurposing.
- Wen Zhang
- , Georgios Voloudakis
- & Panos Roussos
-
Article
| Open Access A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing
A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing
The advent of single-cell RNA sequencing has revealed significant transcriptional heterogeneity in cancer, but its relationship to genomic heterogeneity remains unclear. Focusing on acute myeloid leukemia samples, the authors describe a general approach for linking mutation-containing cells to their transcriptional phenotypes using single-cell RNA sequencing data.
- Allegra A. Petti
- , Stephen R. Williams
- & Timothy J. Ley
-
Article
| Open Access Proteogenomic landscape of squamous cell lung cancer
Proteogenomic landscape of squamous cell lung cancer
Squamous cell lung cancer has dismal prognosis due to the dearth of effective treatments. Here, the authors perform an integrated proteogenomic analysis of the disease, revealing three proteomics-based subtypes and suggesting potential therapeutic opportunities.
- Paul A. Stewart
- , Eric A. Welsh
- & Eric B. Haura
-
Article
| Open Access Quantifying the impact of public omics data
Quantifying the impact of public omics data
Increasing amount of public omics data are important and valuable resources for the research community. Here, the authors develop a set of metrics to quantify the attention and impact of biomedical datasets and integrate them into the framework of Omics Discovery Index (OmicsDI).
- Yasset Perez-Riverol
- , Andrey Zorin
- & Henning Hermjakob
-
Article
| Open Access Integrating biomedical research and electronic health records to create knowledge-based biologically meaningful machine-readable embeddings
Integrating biomedical research and electronic health records to create knowledge-based biologically meaningful machine-readable embeddings
The Scalable Precision Medicine Oriented Knowledge Engine (SPOKE) is a heterogeneous knowledge network that integrates information from 29 public databases. Here, Nelson et al. extend SPOKE to embed clinical data from electronic health records to create medically meaningful barcodes for each medical variable.
- Charlotte A. Nelson
- , Atul J. Butte
- & Sergio E. Baranzini
-
Article
| Open Access Improving the diagnostic yield of exome- sequencing by predicting gene–phenotype associations using large-scale gene expression analysis
Improving the diagnostic yield of exome- sequencing by predicting gene–phenotype associations using large-scale gene expression analysis
A genetic diagnosis remains unattainable for many individuals with a rare disease because of incomplete knowledge about the genetic basis of many diseases. Here, the authors present the web-based tool GADO (GeneNetwork Assisted Diagnostic Optimization) that uses public RNA-seq data for prioritization of candidate genes.
- Patrick Deelen
- , Sipko van Dam
- & Lude Franke
-
Article
| Open Access Optimal control nodes in disease-perturbed networks as targets for combination therapy
Optimal control nodes in disease-perturbed networks as targets for combination therapy
Synergistic interactions may arise between regulators in complex molecular networks. Here, the authors develop OptiCon, a computational method for de novo identification of synergistic key regulators and investigate their potential roles as candidate targets for combination therapy.
- Yuxuan Hu
- , Chia-hui Chen
- & Kai Tan
-
Article
| Open Access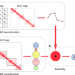 Integrating Hi-C and FISH data for modeling of the 3D organization of chromosomes
Integrating Hi-C and FISH data for modeling of the 3D organization of chromosomes
Methodological advances have increased our understanding of chromatin structure through chromosome conformation capture techniques and high resolution imaging, but integration of these datasets is challenging. Here the authors propose GEM-FISH, a method for reconstructing the 3D models of chromosomes through systematically integrating both Hi-C and FISH data with the prior biophysical knowledge of a polymer model.
- Ahmed Abbas
- , Xuan He
- & Jianyang Zeng
-
Article
| Open Access Metascape provides a biologist-oriented resource for the analysis of systems-level datasets
Metascape provides a biologist-oriented resource for the analysis of systems-level datasets
With the increasing obtainability of multi-OMICs data comes the need for easy to use data analysis tools. Here, the authors introduce Metascape, a biologist-oriented portal that provides a gene list annotation, enrichment and interactome resource and enables integrated analysis of multi-OMICs datasets.
- Yingyao Zhou
- , Bin Zhou
- & Sumit K. Chanda
-
Article
| Open Access Integrated systems approach defines the antiviral pathways conferring protection by the RV144 HIV vaccine
Integrated systems approach defines the antiviral pathways conferring protection by the RV144 HIV vaccine
The RV144 vaccine trial showed reduced risk of HIV-1 acquisition, but mechanisms underlying protection are poorly understood. Here, Fourati et al. assess the transcriptomic profile of blood collected from 223 vaccinees and 40 placebo recipients and identify IRF7 as a mediator of protection.
- Slim Fourati
- , Susan Pereira Ribeiro
- & Rafick-Pierre Sékaly
-
Article
| Open Access Towards a data-integrated cell
Towards a data-integrated cell
Integration of omics data remains a challenge. Here, the authors introduce iCell, a framework to integrate tissue-specific protein–protein interaction, co-expression and genetic interaction data, enabling identification of the most rewired genes in cancer, unidentifiable in individual data layers.
- Noël Malod-Dognin
- , Julia Petschnigg
- & Nataša Pržulj
-
Article
| Open Access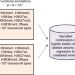 A semi-supervised approach for predicting cell-type specific functional consequences of non-coding variation using MPRAs
A semi-supervised approach for predicting cell-type specific functional consequences of non-coding variation using MPRAs
Predicting the functional consequences of non-coding genetic variants is a challenge. Here, He et al. present GenoNet, a semi-supervised method that combines information from experimentally confirmed regulatory variants with cell type- and tissue specific annotation for function prediction.
- Zihuai He
- , Linxi Liu
- & Iuliana Ionita-Laza
-
Article
| Open Access Integrative epigenetic taxonomy of primary prostate cancer
Integrative epigenetic taxonomy of primary prostate cancer
The Androgen Receptor (AR) is the main driver of prostate cancer and functions in conjunction with chromatin modifications. Here, the authors comprehensively profile 100 primary prostate carcinomas by sequencing RNA transcripts in combination with ChIP-sequencing for AR and the active histone marks H3K27ac, H3K4me3 and repressive mark H3K27me3.
- Suzan Stelloo
- , Ekaterina Nevedomskaya
- & Wilbert Zwart
-
Article
| Open Access Improved estimation of cancer dependencies from large-scale RNAi screens using model-based normalization and data integration
Improved estimation of cancer dependencies from large-scale RNAi screens using model-based normalization and data integration
Integrated analyses of multiple large-scale screenings can be complicated by batch effects and technical artefacts. McFarland et al. introduce DEMETER2, a hierarchical model coupled with model-based normalization, which allows the assessment of differential dependencies across genes and cell lines.
- James M. McFarland
- , Zandra V. Ho
- & Aviad Tsherniak
-
Article
| Open Access Network integration of multi-tumour omics data suggests novel targeting strategies
Network integration of multi-tumour omics data suggests novel targeting strategies
Tumours of different tissues can show similarities in genomic alterations. Here, the authors combine tumour transcriptome and protein interaction data in a network-based analysis of 11 tumours types, and identify clusters of tumours with specific signatures for multi-tumour drug targeting and survival prognosis.
- Ítalo Faria do Valle
- , Giulia Menichetti
- & Daniel Remondini
-
Article
| Open Access Expression-based drug screening of neural progenitor cells from individuals with schizophrenia
Expression-based drug screening of neural progenitor cells from individuals with schizophrenia
Unbiased large scale screening of small molecules for drug discovery in psychiatric disease is technically challenging and financially costly. Here, Readhead and colleagues integrate in silico and in vitro approaches to design and conduct transcriptomic drug screening in schizophrenia patient-derived neural cells, in order to survey novel pathologies and points of intervention.
- Benjamin Readhead
- , Brigham J. Hartley
- & Kristen J. Brennand
-
Article
| Open Access Temporal genetic association and temporal genetic causality methods for dissecting complex networks
Temporal genetic association and temporal genetic causality methods for dissecting complex networks
Temporal omics data have the potential to dissect complex biological networks. Here the authors develop methods for detecting temporal genetic loci (teQTLs) of quantitative traits monitored over time and inferring causal relationships between traits linked to the locus.
- Luan Lin
- , Quan Chen
- & Jun Zhu
-
Article
| Open Access The gut microbiota promotes hepatic fatty acid desaturation and elongation in mice
The gut microbiota promotes hepatic fatty acid desaturation and elongation in mice
The role of the gut microbiota in hepatic lipid metabolism is controversial and incompletely understood. Here the authors perform multi-omics analyses of altered lipid metabolic processes in germ-free and specific pathogen-free mice, revealing how the gut microbiota affects hepatic fatty acid desaturation and elongation.
- Alida Kindt
- , Gerhard Liebisch
- & Josef Ecker
-
Article
| Open Access Distinct epigenomic patterns are associated with haploinsufficiency and predict risk genes of developmental disorders
Distinct epigenomic patterns are associated with haploinsufficiency and predict risk genes of developmental disorders
Predicting haploinsufficient genes helps to understand the genetic risk underlying developmental disorders. Here, the authors develop a Random Forest-based method that uses epigenomic data to predict haploinsufficiency, Episcore, which is complementary to methods based on mutation intolerance scores.
- Xinwei Han
- , Siying Chen
- & Yufeng Shen
-
Article
| Open Access Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics
Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics
Phenotypic variation and diseases are influenced by factors such as genetic variants and gene expression. Here, Barbeira et al. develop S-PrediXcan to compute PrediXcan results using summary data, and investigate the effects of gene expression variation on human phenotypes in 44 GTEx tissues and >100 phenotypes.
- Alvaro N. Barbeira
- , Scott P. Dickinson
- & Hae Kyung Im
-
Article
| Open Access Massive mining of publicly available RNA-seq data from human and mouse
Massive mining of publicly available RNA-seq data from human and mouse
Publicly available RNA-seq data is provided mostly in raw form, resulting in a barrier for integrative analyses. Here, Lachmann et al. develop a high-throughput processing infrastructure and search database (ARCHS4) that provides processed RNA-seq data for 187,946 publicly available mouse and human samples to support exploration and reuse.
- Alexander Lachmann
- , Denis Torre
- & Avi Ma’ayan
-
Article
| Open Access Integrative proteomics in prostate cancer uncovers robustness against genomic and transcriptomic aberrations during disease progression
Integrative proteomics in prostate cancer uncovers robustness against genomic and transcriptomic aberrations during disease progression
Understanding of molecular events in cancer requires proteome-level characterisation. Here, proteome profiling of patient samples representing primary and progressed prostate cancer enables the authors to identify pathway alterations that are not reflected at the genomic and transcriptomic levels.
- Leena Latonen
- , Ebrahim Afyounian
- & Tapio Visakorpi
-
Article
| Open Access Integrative analysis of omics summary data reveals putative mechanisms underlying complex traits
Integrative analysis of omics summary data reveals putative mechanisms underlying complex traits
The identification of the causal gene at a GWAS locus remains to be a challenging task. Here, using the SMR & HEIDI method to integrate GWAS, eQTL and mQTL data, Wu et al. map DNA methylation sites to the transcriptome and thereby prioritize functionally relevant genes for 12 human complex traits.
- Yang Wu
- , Jian Zeng
- & Jian Yang
-
Article
| Open Access Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor
Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor
Single cell RNA-sequencing analysis poses challenges in replication due to technical biases and analytic variability among bioinformatics pipelines. Here, Crow et al develop MetaNeighbor for measuring cell-type replication across datasets, and use it to identify marker genes for neuron subtypes with evidence of replication.
- Megan Crow
- , Anirban Paul
- & Jesse Gillis
-
Article
| Open Access Inference of differentiation time for single cell transcriptomes using cell population reference data
Inference of differentiation time for single cell transcriptomes using cell population reference data
Single cell transcriptome data can be used to determine developmental lineage trajectories. Here the authors map single cell transcriptomes onto a differentiation trajectory defined by cell population transcriptomes and show that cell cycle regulators have a role in differentiation timing.
- Na Sun
- , Xiaoming Yu
- & Jing-Dong J. Han
-
Article
| Open Access Functional mapping and annotation of genetic associations with FUMA
Functional mapping and annotation of genetic associations with FUMA
Prioritizing genetic variants is a major challenge in genome-wide association studies. Here, the authors develop FUMA, a web-based bioinformatics tool that uses a combination of positional, eQTL and chromatin interaction mapping to prioritize likely causal variants and genes.
- Kyoko Watanabe
- , Erdogan Taskesen
- & Danielle Posthuma
-
Article
| Open Access A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information
Network-based data integration for drug–target prediction is a promising avenue for drug repositioning, but performance is wanting. Here, the authors introduce DTINet, whose performance is enhanced in the face of noisy, incomplete and high-dimensional biological data by learning low-dimensional vector representations.
- Yunan Luo
- , Xinbin Zhao
- & Jianyang Zeng
-
Article
| Open Access Topologically associating domains are ancient features that coincide with Metazoan clusters of extreme noncoding conservation
Topologically associating domains are ancient features that coincide with Metazoan clusters of extreme noncoding conservation
Metazoan genomes contain many clusters of conserved noncoding elements. Here, the authors provide evidence that these clusters coincide with distinct topologically associating domains in humans and Drosophila, revealing a conserved regulatory genomic architecture.
- Nathan Harmston
- , Elizabeth Ing-Simmons
- & Boris Lenhard
-
Article
| Open Access Bayesian association scan reveals loci associated with human lifespan and linked biomarkers
Bayesian association scan reveals loci associated with human lifespan and linked biomarkers
Along with various environmental factors, a complex genetic architecture influences human lifespan. Here, McDaid and colleagues reveal novel loci associated with human lifespan and linked biomarkers by Bayesian association scan.
- Aaron F. McDaid
- , Peter K. Joshi
- & Zoltán Kutalik
-
Article
| Open Access Pancancer modelling predicts the context-specific impact of somatic mutations on transcriptional programs
Pancancer modelling predicts the context-specific impact of somatic mutations on transcriptional programs
Cancer genomic data sets contain a wealth of data that can be used to predict prognosis and further understand disease. Here, the authors integrate multiple genomics data types to identify transcriptional dysregulation in response to somatic mutations.
- Hatice U. Osmanbeyoglu
- , Eneda Toska
- & Christina S. Leslie
-
Article
| Open Access Tissue-specific and convergent metabolic transformation of cancer correlates with metastatic potential and patient survival
Tissue-specific and convergent metabolic transformation of cancer correlates with metastatic potential and patient survival
Cancer cells reprogramme their metabolism with unclear clinical implications. Here, the authors analyse the expression of metabolic genes across 20 types of solid cancers and find that clinical aggressiveness, poor survival and metastasis are associated with the deregulation of mitochondrial metabolism.
- Edoardo Gaude
- & Christian Frezza
-
Article
| Open Access Multi-omics integration accurately predicts cellular state in unexplored conditions for Escherichia coli
Multi-omics integration accurately predicts cellular state in unexplored conditions for Escherichia coli
Multi-omics data integration is a great challenge. Here, the authors compile a database of E. coliproteomics, transcriptomics, metabolomics and fluxomics data to train models of recurrent neural network and constrained regression, enabling prediction of bacterial responses to perturbations.
- Minseung Kim
- , Navneet Rai
- & Ilias Tagkopoulos
-
Article
| Open Access Extraction and analysis of signatures from the Gene Expression Omnibus by the crowd
Extraction and analysis of signatures from the Gene Expression Omnibus by the crowd
A wealth of gene expression data is publicly available, yet is little use without additional human curation. Ma’ayan and colleagues report a crowdsourcing project involving over 70 participants to annotate and analyse thousands of human disease-related gene expression datasets.
- Zichen Wang
- , Caroline D. Monteiro
- & Avi Ma’ayan
-
Article
| Open Access High-dimensional genomic data bias correction and data integration using MANCIE
High-dimensional genomic data bias correction and data integration using MANCIE
Analyses of data from high-throughput genomic technologies are challenging given large data dimensionality. Here, Liu and colleagues describe a method called MANCIE (Matrix Analysis and Normalization by Concordant Information Enhancement) that can conduct genomic data normalization and bias correction to detect biologically relevant information.
- Chongzhi Zang
- , Tao Wang
- & X. Shirley Liu

 Construction of a web-based nanomaterial database by big data curation and modeling friendly nanostructure annotations
Construction of a web-based nanomaterial database by big data curation and modeling friendly nanostructure annotations