Abstract
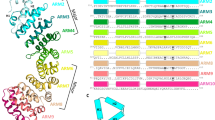
Importin α is the nuclear import receptor that recognizes classical monopartite and bipartite nuclear localization signals (NLSs). The structure of mouse importin α has been determined at 2.5 Å resolution. The structure shows a large C-terminal domain containing armadillo repeats, and a less structured N-terminal importin β-binding domain containing an internal NLS bound to the NLS-binding site. The structure explains the regulatory switch between the cytoplasmic, high-affinity form, and the nuclear, low-affinity form for NLS binding of the nuclear import receptor predicted by the current models of nuclear import. Importin β conceivably converts the low- to high-affinity form by binding to a site overlapping the autoinhibitory sequence. The structure also has implications for understanding NLS recognition, and the structures of armadillo and HEAT repeats.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Cole, C.N. & Hammel, C.M. Nucleocytoplasmic transport: driving and directing transport. Curr. Biol. 8, R368–R372 (1998).
Gorlich, D. Transport into and out of the nucleus. EMBO J. 17, 2721–2727 (1998).
Imamoto, N., Kamei, Y. & Yoneda, Y. Nuclear transport factors: function, behavior and interaction. Eur. J. Histochem. 42, 9– 20 (1998).
Izaurralde, E. & Adam, S. Transport of macromolecules between the nucleus and the cytoplasm. RNA 4, 351 –364 (1998).
Mattaj, I.W. & Englmeier, L. Nucleocytoplasmic transport: the soluble phase. Annu. Rev. Biochem. 67, 265 –306 (1998).
Melchior, F. & Gerace, L. Two-way trafficking with Ran. Trends Cell Biol. 8, 175–179 (1998).
Moroianu, J. Distinct nuclear import and export pathways mediated by members of the karyopherin beta family. Cell. Biochem. 70, 231– 239 (1998).
Ohno, M., Fornerod, M. & Mattaj, I.W. Nucleocytoplasmic transport: the last 200 nanometers. Cell 92, 327–336 (1998).
Pemberton, L.F., Blobel, G. & Rosenblum, J.S. Transport routes through the nuclear pore complex. Curr. Opin. Cell Biol. 10, 392– 399 (1998).
Weis, K. Importins and exportins: how to get in and out of the nucleus. Trends Biochem. Sci. 23, 185–189 (1998).
Wozniak, R.W., Rout, M.P. & Aitchison, J.D. Karyopherins and kissing cousins. Trends Cell Biol. 8, 184–188 ( 1998).
Dingwall, C. & Laskey, R.A. Nuclear targeting sequences — a consensus? Trends Biochem. Sci. 16, 478 –481 (1991).
Gorlich, D., Henklein, P., Laskey, R.A. & Hartmann, E. A 41 amino acid motif in importin-alpha confers binding to importin-beta and hence transit into the nucleus. EMBO J. 15, 1810–1817 (1996).
Weis, K., Ryder, U. & Lamond, A.I. The conserved amino-terminal domain of hSRP1 alpha is essential for nuclear protein import. EMBO J. 15, 1818–1825 (1996).
Moroianu, J., Blobel, G. & Radu, A. The binding site of karyopherins alpha for karyopherin beta overlaps with a nuclear localization sequence. Proc. Natl. Acad. Sci. USA 93, 6572–6576 (1996).
Peifer, M., Berg, S. & Reynolds, A.B. A repeating amino acid motif shared by proteins with diverse cellular roles. Cell 76, 789– 791 (1996).
Huber, A.H., Nelson, W.J. & Weis, W.I. Three-dimensional structure of the armadillo repeat region of beta-catenin. Cell 90, 871– 882 (1997).
Kobe, B. Leucines on a roll. Nature Struct. Biol. 3, 977–980 (1996).
Andrade, M.A. & Bork, P. HEAT repeats in the Huntington's disease protein. Nat. Genet. 11, 115– 116 (1995).
Malik, H.S., Eickbush, T.H. & Goldfarb, D.S. Evolutionary specialization of the nuclear targeting apparatus. Proc. Natl. Acad. Sci. USA 94, 13738–13742 (1997).
Conti, E., Uy, M., Leighton, L., Blobel, G. & Kuriyan, J. Crystallographic analysis of the recognition of a nuclear localization signal by the nuclear import factor karyopherins alpha. Cell 94, 193–204 ( 1998).
Kobe, B. et al. Giant protein kinases: domain interactions and structural basis of autoregulation. EMBO J. 15, 6810– 6821 (1996).
Kussel, P. & Frasch, M. Yeast Srp1, a nuclear protein related to Drosophila and mouse pendulin, is required for normal migration, division, and integrity of nuclei during mitosis. Mol. Gen. Genet. 248, 351–363 (1995).
Kobe, B. & Deisenhofer, J. A structural basis of the interactions between leucine-rich repeats and protein ligands. Nature 374, 183–186 (1995).
Barden, J.A., Sehgal, P. & Kemp, B.E. Structure of the pseudosubstrate recognition site of chicken smooth muscle myosin light chain kinase. Biochim. Biophys. Acta 1292, 106–112 ( 1996).
Kissinger, C.R. et al. Crystal structures of human calcineurin and the human FKBP12–FK506–calcineurin complex. Nature 378, 641– 644 (1995).
Khan, A.R. & James, M.N.G. Molecular mechanisms for the conversion of zymogens to active proteolytic enzymes. Protein Sci. 7, 815–836 (1998).
Kobe, B. et al. Structural basis of autoregulation of phenylalanine hydroxylase. Nature Struct. Biol. in the press (1999 ).
Huber, J. et al. Snurportin1, an m3G-cap-specific nuclear import receptor with a novel domain structure. EMBO J. 17, 4114 –4126 (1998).
Boche, I. & Fanning, E. Nucleocytoplasmic recycling of the nuclear localization signal receptors alpha subunit in vivo is dependent on a nuclear export signal, energy, and RCC1. J. Cell Biol. 139, 313–325 (1997).
Nigg, E.A. Nucleocytoplasmic transport: signals, mechanisms and regulation. Nature 386, 779–787 ( 1997).
Makkerh, J.P.S., Dingwall, C. & Laskey, R.A. Comparative mutagenesis of nuclear localization signals reveals the importance of neutral and acidic amino acids. Curr. Biol. 6, 1025–1027 ( 1996).
Jans, D.A. & Hubner, S. Regulation of protein transport to the nucleus: central role of phosphorylation. Physiol. Rev. 76, 651–685 (1996).
Hubner, S., Xiao, C.Y. & Jans, D.A. The protein kinase CK2 site (Ser111/112) enhances recognition of the simian virus 40 large T-antigen nuclear localization sequence by importin. J. Biol. Chem. 272, 17191– 17195 (1997).
Ruediger, R., Hentz, M., Fait, J., Mumby, M. & Walter, G. Molecular model of the A subunit of protein phosphatase 2A: interaction with other subunits and tumor antigens. J. Virol. 68, 123–129 ( 1994).
Tiganis, T., Flint, A.J., Adam, S.A. & Tonks, N.K. Association of the T-cell protein tyrosine phosphatase with nuclear import factor p97. J. Biol. Chem. 272, 21548–21557 (1997).
Henderson, B.R. & Percipalle, P. Interactions between HIV Rev and nuclear import and export factors: the Rev nuclear localisation signal mediates specific binding to human importin-beta. J. Mol. Biol. 274, 693–707 ( 1997).
Jones, E.Y. MHC class I and class II structures. Curr. Opin. Immunol. 9, 75–79 (1997).
Teh, T., Tiganis, T. & Kobe, B. Crystallization of importin α, the nuclear import receptor. Acta Crystallogr. D D55, 561–536 (1999).
Otwinowski, Z. & Minor, W. Processing of x-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 ( 1997).
Jones, T.A., Bergdoll, M. & Kjeldgaard, M. In Crystallographic and modeling methods in molecular design (eds Bugg, C.E. & Ealick, S.E.) 189– 195 (Springer-Verlag, New York; 1990).
DeLano, W.L. & Brünger, A.T. The direct rotation function: rotational Patterson correlation search applied to molecular replacement. Acta Crystallogr. D 51, 740– 748 (1995).
Brünger, A.T. Free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472– 475 (1992).
Lamzin, V.S. & Wilson, K.S. Automated refinement of protein models. Acta Crystallogr. D 49, 129– 147 (1993).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283– 291 (1993).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Kraulis, P. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946 –950 (1991).
Merritt, E.A. & Murphy, M.E.P. Raster3D Version 2—a program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. 15, 133–138 (1997).
Lawrence, M.C. & Colman, P.M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993).
Wallace, A.C., Laskowski, R.A. & Thornton, J.M. LIGPLOT: a program to generate schematic diagrams of protein ligand interactions. Protein Eng. 8, 127–134 (1995).
Chelsky, D., Ralph, R. & Jonak, G. Sequence requirements for synthetic peptide-mediated translocation to the nucleus. Mol. Cell. Biol. 9, 2487 –2492 (1989).
Acknowledgements
I thank T. Teh, T. Tiganis, M. Lam, B. Kemp, J. Horne and T. Gleichmann for help and discussions, and J. Kuriyan and coworkers for making the coordinates of yeast Kapα50 available on their web site before the release by the Protein Data Bank. I apologize to those whose original publications could not be cited because of space limitations. This work was supported by the Wellcome Trust and the National Health and Medical Research Council; B.K. is a Wellcome Senior Research Fellow in Medical Science in Australia.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kobe, B. Autoinhibition by an internal nuclear localization signal revealed by the crystal structure of mammalian importin α. Nat Struct Mol Biol 6, 388–397 (1999). https://doi.org/10.1038/7625
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/7625
This article is cited by
-
OsWRKY62 and OsWRKY76 Interact with Importin α1s for Negative Regulation of Defensive Responses in Rice Nucleus
Rice (2022)
-
Super-resolved 3D tracking of cargo transport through nuclear pore complexes
Nature Cell Biology (2022)
-
Structural characterization of human importin alpha 7 in its cargo-free form at 2.5 Å resolution
Scientific Reports (2022)
-
An epilepsy-associated mutation in the nuclear import receptor KPNA7 reduces nuclear localization signal binding
Scientific Reports (2020)
-
The NAD+-mediated self-inhibition mechanism of pro-neurodegenerative SARM1
Nature (2020)