Key Points
-
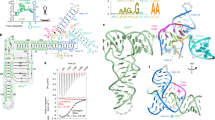
Structured regions in bacterial mRNAs can act as thermosensors, known as RNA thermometers (RNATs), that control translation efficiency of the transcript.
-
Some RNATs act like zippers that open and close, in a reversible manner, according to the ambient temperature. These RNATs control heat shock and virulence genes.
-
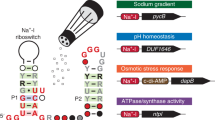
Escherichia coli uses a cascade of hierarchically organized RNATs to monitor any harmful temperature upshifts and to induce the production of protective heat shock proteins when required.
-
Known heat shock RNATs have little if any sequence conservation.
-
Recent biophysical approaches revealed details of RNAT zippers at base pair resolution. Non-Watson–Crick base pairs, internal loops and bulges, Mg2+ and the hydration shell can be crucial for RNAT stability and melting.
-
RNATs that permit translation of cold-shock and phage genes at low temperatures switch between two mutually exclusive structures. The conformation at low temperature favours translation.
-
Synthetic thermosensors inspired by natural RNATs can induce gene expression in response to a temperature upshift and bear potential as tools for large-scale industrial applications without the need for chemical inducers.
Abstract
Bacteria use complex strategies to coordinate temperature-dependent gene expression. Many genes encoding heat shock proteins and virulence factors are regulated by temperature-sensing RNA sequences, known as RNA thermometers (RNATs), in their mRNAs. For these genes, the 5′ untranslated region of the mRNA folds into a structure that blocks ribosome access at low temperatures. Increasing the temperature gradually shifts the equilibrium between the closed and open conformations towards the open structure in a zipper-like manner, thereby increasing the efficiency of translation initiation. Here, we review the known molecular principles of RNAT action and the hierarchical RNAT cascade in Escherichia coli. We also discuss RNA-based thermosensors located upstream of cold shock and other genes, translation of which preferentially occurs at low temperatures and which thus operate through a different, more switch-like mechanism. Finally, we consider the potential biotechnological applications of natural and synthetic RNATs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Konkel, M. E. & Tilly, K. Temperature-regulated expression of bacterial virulence genes. Microbes Infect. 2, 157–166 (2000).
Lim, B. & Gross, C. A. in Bacterial Stress Responses (eds Storz, G. & Hengge, R.) 93–114 (American Society of Microbiology Press, Washington DC, 2011).
Klinkert, B. & Narberhaus, F. Microbial thermosensors. Cell. Mol. Life Sci. 66, 2661–2676 (2009).
Guisbert, E., Yura, T., Rhodius, V. A. & Gross, C. A. Convergence of molecular, modeling, and systems approaches for an understanding of the Escherichia coli heat shock response. Microbiol. Mol. Biol. Rev. 72, 545–554 (2008).
Falconi, M., Colonna, B., Prosseda, G., Micheli, G. & Gualerzi, C. O. Thermoregulation of Shigella and Escherichia coli EIEC pathogenicity. A temperature-dependent structural transition of DNA modulates accessibility of virF promoter to transcriptional repressor H-NS. EMBO J. 17, 7033–7043 (1998).
Prosseda, G. et al. The virF promoter in Shigella: more than just a curved DNA stretch. Mol. Microbiol. 51, 523–537 (2004).
Servant, P., Rapoport, G. & Mazodier, P. RheA, the repressor of hsp18 in Streptomyces albus G. Microbiology 145, 2385–2391 (1999).
de Smit, M. H. & van Duin, J. Secondary structure of the ribosome binding site determines translational efficiency: a quantitative analysis. Proc. Natl Acad. Sci. USA 87, 7668–7672 (1990). A seminal study on the relationship between mRNA secondary structure and translation efficiency.
de Smit, M. H. & van Duin, J. Control of translation by mRNA secondary structure in Escherichia coli. A quantitative analysis of literature data. J. Mol. Biol. 244, 144–150 (1994).
Narberhaus, F., Waldminghaus, T. & Chowdhury, S. RNA thermometers. FEMS Microbiol. Rev. 30, 3–16 (2006).
Narberhaus, F. Translational control of bacterial heat shock and virulence genes by temperature-sensing mRNAs. RNA Biol. 7, 84–89 (2010).
Rinnenthal, J., Klinkert, B., Narberhaus, F. & Schwalbe, H. Modulation of the stability of the Salmonella fourU-type RNA thermometer. Nucleic Acids Res. 39, 8258–8270 (2011).
Rinnenthal, J., Klinkert, B., Narberhaus, F. & Schwalbe, H. Direct observation of the temperature-induced melting process of the Salmonella fourU RNA thermometer at base-pair resolution. Nucleic Acids Res. 38, 3834–3847 (2010). An NMR study revealing the molecular details of zipper-like melting for each individual base pair.
Winkler, W. C. & Breaker, R. R. Regulation of bacterial gene expression by riboswitches. Annu. Rev. Microbiol. 59, 487–517 (2005).
Henkin, T. M. Riboswitch RNAs: using RNA to sense cellular metabolism. Genes Dev. 22, 3383–3390 (2008).
Chowdhury, S., Maris, C., Allain, F. H. & Narberhaus, F. Molecular basis for temperature sensing by an RNA thermometer. EMBO J. 25, 2487–2497 (2006). The first three-dimensional structure of an RNAT to be obtained finds that several non-canonical base pairs are key to melting.
Jones, P. G. & Inouye, M. The cold-shock response — a hot topic. Mol. Microbiol. 11, 811–818 (1994).
Waldminghaus, T., Gaubig, L. C. & Narberhaus, F. Genome-wide bioinformatic prediction and experimental evaluation of potential RNA thermometers. Mol. Genet. Genom. 278, 555–564 (2007).
Shah, P. & Gilchrist, M. A. Is thermosensing property of RNA thermometers unique? PLoS ONE 5, e11308 (2010).
Neupert, J., Karcher, D. & Bock, R. Design of simple synthetic RNA thermometers for temperature-controlled gene expression in Escherichia coli. Nucleic Acids Res. 36, e124 (2008).
Waldminghaus, T., Kortmann, J., Gesing, S. & Narberhaus, F. Generation of synthetic RNA-based thermosensors. Biol. Chem. 389, 1319–1326 (2008).
Nocker, A. et al. A mRNA-based thermosensor controls expression of rhizobial heat shock genes. Nucleic Acids Res. 29, 4800–4807 (2001).
Nocker, A., Krstulovic, N. P., Perret, X. & Narberhaus, F. ROSE elements occur in disparate rhizobia and are functionally interchangeable between species. Arch. Microbiol. 176, 44–51 (2001).
Waldminghaus, T., Fippinger, A., Alfsmann, J. & Narberhaus, F. RNA thermometers are common in α- and γ-proteobacteria. Biol. Chem. 386, 1279–1286 (2005).
Waldminghaus, T., Gaubig, L. C., Klinkert, B. & Narberhaus, F. The Escherichia coli ibpA thermometer is comprised of stable and unstable structural elements. RNA Biol. 6, 455–463 (2009).
Chowdhury, S., Ragaz, C., Kreuger, E. & Narberhaus, F. Temperature-controlled structural alterations of an RNA thermometer. J. Biol. Chem. 278, 47915–47921 (2003).
Waldminghaus, T., Heidrich, N., Brantl, S. & Narberhaus, F. FourU: a novel type of RNA thermometer in Salmonella. Mol. Microbiol. 65, 413–424 (2007).
Alatossava, T., Jutte, H., Kuhn, A. & Kellenberger, E. Manipulation of intracellular magnesium content in polymyxin B nonapeptide-sensitized Escherichia coli by ionophore A23187. J. Bacteriol. 162, 413–419 (1985).
Kortmann, J., Sczodrok, S., Rinnenthal, J., Schwalbe, H. & Narberhaus, F. Translation on demand by a simple RNA-based thermosensor. Nucleic Acids Res. 39, 2855–2868 (2011).
Torok, Z. et al. Synechocystis HSP17 is an amphitropic protein that stabilizes heat-stressed membranes and binds denatured proteins for subsequent chaperone-mediated refolding. Proc. Natl Acad. Sci. USA 98, 3098–3103 (2001).
Morita, M. T. et al. Translational induction of heat shock transcription factor σ32: evidence for a built-in RNA thermosensor. Genes Dev. 13, 655–665 (1999). A comprehensive study on the complex E. coli rpoH thermometer. This study sets experimental standards for analysis of RNATs.
Storz, G. An RNA thermometer. Genes Dev. 13, 633–636 (1999).
Morita, M., Kanemori, M., Yanagi, H. & Yura, T. Heat-induced synthesis of σ32 in Escherichia coli: structural and functional dissection of rpoH mRNA secondary structure. J. Bacteriol. 181, 401–410 (1999).
Cheng, B., Zhu, C. X., Ji, C., Ahumada, A. & Tse-Dinh, Y. C. Direct interaction between Escherichia coli RNA polymerase and the zinc ribbon domains of DNA topoisomerase I. J. Biol. Chem. 278, 30705–30710 (2003).
Mah, T. F., Kuznedelov, K., Mushegian, A., Severinov, K. & Greenblatt, J. The α subunit of E. coli RNA polymerase activates RNA binding by NusA. Genes Dev. 14, 2664–2675 (2000).
Guisbert, E., Herman, C., Lu, C. Z. & Gross, C. A. A chaperone network controls the heat shock response in E. coli. Genes Dev. 18, 2812–2821 (2004).
Tomoyasu, T. et al. Escherichia coli FtsH is a membrane-bound, ATP-dependent protease which degrades the heat-shock transcription factor σ32. EMBO J. 14, 2551–2560 (1995).
Allen, S. P., Polazzi, J. O., Gierse, J. K. & Easton, A. M. Two novel heat shock genes encoding proteins produced in response to heterologous protein expression in Escherichia coli. J. Bacteriol. 174, 6938–6947 (1992).
Richmond, C. S., Glasner, J. D., Mau, R., Jin, H. & Blattner, F. R. Genome-wide expression profiling in Escherichia coli K-12. Nucleic Acids Res. 27, 3821–3835 (1999).
Gaubig, L. C., Waldminghaus, T. & Narberhaus, F. Multiple layers of control govern expression of the Escherichia coli ibpAB heat-shock operon. Microbiology 157, 66–76 (2011).
Veinger, L., Diamant, S., Buchner, J. & Goloubinoff, P. The small heat-shock protein IbpB from Escherichia coli stabilizes stress-denatured proteins for subsequent refolding by a multichaperone network. J. Biol. Chem. 273, 11032–11037 (1998).
Matuszewska, M., Kuczynska-Wisnik, D., Laskowska, E. & Liberek, K. The small heat shock protein IbpA of Escherichia coli cooperates with IbpB in stabilization of thermally aggregated proteins in a disaggregation competent state. J. Biol. Chem. 280, 12292–12298 (2005).
Bissonnette, S. A., Rivera-Rivera, I., Sauer, R. T. & Baker, T. A. The IbpA and IbpB small heat-shock proteins are substrates of the AAA+ Lon protease. Mol. Microbiol. 75, 1539–1549 (2010).
Lambris, J. D., Ricklin, D. & Geisbrecht, B. V. Complement evasion by human pathogens. Nature Rev. Microbiol. 6, 132–142 (2008).
Childers, B. M. & Klose, K. E. Regulation of virulence in Vibrio cholerae: the ToxR regulon. Future Microbiol. 2, 335–344 (2007).
Johansson, J. RNA thermosensors in bacterial pathogens. Contrib. Microbiol. 16, 150–160 (2009).
Gripenland, J. et al. RNAs: regulators of bacterial virulence. Nature Rev. Microbiol. 8, 857–866 (2011).
Papenfort, K. & Vogel, J. Regulatory RNA in bacterial pathogens. Cell Host Microbe 8, 116–127 (2010).
Skurnik, M. & Toivanen, P. LcrF is the temperature-regulated activator of the yadA gene of Yersinia enterocolitica and Yersinia pseudotuberculosis. J. Bacteriol. 174, 2047–2051 (1992).
Hoe, N. P. & Goguen, J. D. Temperature sensing in Yersinia pestis: translation of the LcrF activator protein is thermally regulated. J. Bacteriol. 175, 7901–7909 (1993).
Böhme, K. et al. Concerted actions of a thermo-labile regulator and a unique intergenic RNA thermosensor control Yersinia virulence. PLoS Pathog. 8, e1002518 (2012).
Johansson, J. et al. An RNA thermosensor controls expression of virulence genes in Listeria monocytogenes. Cell 110, 551–561 (2002). A report demonstrating that synthesis of the L. monocytogenes virulence master regulator PrfA is modulated by a temperature-sensing 5′ UTR.
Hamon, M., Bierne, H. & Cossart, P. Listeria monocytogenes: a multifaceted model. Nature Rev. Microbiol. 4, 423–434 (2006).
Freitag, N. E., Port, G. C. & Miner, M. D. Listeria monocytogenes — from saprophyte to intracellular pathogen. Nature Rev. Microbiol. 7, 623–628 (2009).
Toledo-Arana, A. et al. The Listeria transcriptional landscape from saprophytism to virulence. Nature 459, 950–956 (2009).
Loh, E. et al. A trans-acting riboswitch controls expression of the virulence regulator PrfA in Listeria monocytogenes. Cell 139, 770–779 (2009). The first report on a trans -acting riboswitch that interacts with a cis -acting RNAT.
de Smit, M. H. & van Duin, J. Translational standby sites: how ribosomes may deal with the rapid folding kinetics of mRNA. J. Mol. Biol. 331, 737–743 (2003).
Phadtare, S. Unwinding activity of cold shock proteins and RNA metabolism. RNA Biol. 8, 394–397 (2011).
Jiang, W., Hou, Y. & Inouye, M. CspA, the major cold-shock protein of Escherichia coli, is an RNA chaperone. J. Biol. Chem. 272, 196–202 (1997).
Phadtare, S. & Severinov, K. RNA remodeling and gene regulation by cold shock proteins. RNA Biol. 7, 788–795 (2010).
Bae, W., Xia, B., Inouye, M. & Severinov, K. Escherichia coli CspA-family RNA chaperones are transcription antiterminators. Proc. Natl Acad. Sci. USA 97, 7784–7789 (2000).
Yamanaka, K., Mitta, M. & Inouye, M. Mutation analysis of the 5′ untranslated region of the cold shock cspA mRNA of Escherichia coli. J. Bacteriol. 181, 6284–6291 (1999).
Giuliodori, A. M. et al. The cspA mRNA is a thermosensor that modulates translation of the cold-shock protein CspA. Mol. Cell 37, 21–33 (2010). A detailed structural and functional analysis of the cold-sensing cspA mRNA from E. coli.
Breaker, R. R. RNA switches out in the cold. Mol. Cell 37, 1–2 (2010).
Fang, L., Hou, Y. & Inouye, M. Role of the cold-box region in the 5′ untranslated region of the cspA mRNA in its transient expression at low temperature in Escherichia coli. J. Bacteriol. 180, 90–95 (1998).
Yamanaka, K. Cold shock response in Escherichia coli. J. Mol. Microbiol. Biotechnol. 1, 193–202 (1999).
Jiang, W., Fang, L. & Inouye, M. The role of the 5′-end untranslated region of the mRNA for CspA, the major cold-shock protein of Escherichia coli, in cold-shock adaptation. J. Bacteriol. 178, 4919–4925 (1996).
Cruz, J. A. & Westhof, E. The dynamic landscapes of RNA architecture. Cell 136, 604–609 (2009).
Nechooshtan, G., Elgrably-Weiss, M., Sheaffer, A., Westhof, E. & Altuvia, S. A pH-responsive riboregulator. Genes Dev. 23, 2650–2662 (2009).
Bingham, R. J., Hall, K. S. & Slonczewski, J. L. Alkaline induction of a novel gene locus, alx, in Escherichia coli. J. Bacteriol. 172, 2184–2186 (1990).
Altuvia, S., Kornitzer, D., Teff, D. & Oppenheim, A. B. Alternative mRNA structures of the cIII gene of bacteriophage λ determine the rate of its translation initiation. J. Mol. Biol. 210, 265–280 (1989). The initial description of a temperature-sensing RNA that acts like a switch and controls the lysis–lysogeny decision of phage λ.
Blount, K. F. & Breaker, R. R. Riboswitches as antibacterial drug targets. Nature Biotech. 24, 1558–1564 (2006).
Deigan, K. E. & Ferre-D'Amare, A. R. Riboswitches: discovery of drugs that target bacterial gene-regulatory RNAs. Acc. Chem. Res. 33, 1329–1338 (2011).
Horvath, I. et al. Membrane physical state controls the signaling mechanism of the heat shock response in Synechocystis PCC 6803: identification of hsp17 as a “fluidity gene”. Proc. Natl Acad. Sci. USA 95, 3513–3518 (1998).
Tsvetkova, N. M. et al. Small heat-shock proteins regulate membrane lipid polymorphism. Proc. Natl Acad. Sci. USA 99, 13504–13509 (2002).
Balogi, Z. et al. “Heat shock lipid” in cyanobacteria during heat/light-acclimation. Arch. Biochem. Biophys. 436, 346–354 (2005).
Balogi, Z. et al. A mutant small heat shock protein with increased thylakoid association provides an elevated resistance against UV-B damage in Synechocystis 6803. J. Biol. Chem. 283, 22983–22991 (2008).
Horvath, I. & Vigh, L. Cell biology: Stability in times of stress. Nature 463, 436–438 (2010).
Lee, S., Prochaska, D. J., Fang, F. & Barnum, S. R. A 16.6-kilodalton protein in the cyanobacterium Synechocystis sp. PCC 6803 plays a role in the heat shock response. Curr. Microbiol. 37, 403–407 (1998).
Roth, A. & Breaker, R. R. The structural and functional diversity of metabolite-binding riboswitches. Annu. Rev. Biochem. 78, 305–334 (2009).
Smith, A. M., Fuchs, R. T., Grundy, F. J. & Henkin, T. M. The SAM-responsive SMK box is a reversible riboswitch. Mol. Microbiol. 78, 1393–1402 (2010).
Montange, R. K. & Batey, R. T. Riboswitches: emerging themes in RNA structure and function. Annu. Rev. Biophys. 37, 117–133 (2008).
Fuchs, R. T., Grundy, F. J. & Henkin, T. M. The SMK box is a new SAM-binding RNA for translational regulation of SAM synthetase. Nature Struct. Mol. Biol. 13, 226–233 (2006).
McCabe, K. M., Lacherndo, E. J., Albino-Flores, I., Sheehan, E. & Hernandez, M. LacI(Ts)-regulated expression as an in situ intracellular biomolecular thermometer. Appl. Environ. Microbiol. 77, 2863–2868 (2011).
Win, M. N., Liang, J. C. & Smolke, C. D. Frameworks for programming biological function through RNA parts and devices. Chem. Biol. 16, 298–310 (2009).
Weigand, J. E. & Suess, B. Aptamers and riboswitches: perspectives in biotechnology. Appl. Microbiol. Biotechnol. 85, 229–236 (2009).
Blouin, S., Mulhbacher, J., Penedo, J. C. & Lafontaine, D. A. Riboswitches: ancient and promising genetic regulators. Chembiochem 10, 400–416 (2009).
Lee, J. & Kotov, N. Thermometer design at the nanoscale. Nano Today 2, 48–51 (2007).
Neupert, J. & Bock, R. Designing and using synthetic RNA thermometers for temperature-controlled gene expression in bacteria. Nature Protoc. 4, 1262–1273 (2009).
Wieland, M. & Hartig, J. S. RNA quadruplex-based modulation of gene expression. Chem. Biol. 14, 757–763 (2007).
Zuker, M. Mfold web server for nucleic acid folding and hybridizat3a cited beforeion prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Acknowledgements
RNA-related work in the Narberhaus laboratory is supported by grants from the German Research Foundation (DFG Priority Programme 1258, Sensory and Regulatory RNAs in Prokaryotes). J.K. received a fellowship from the Studienstiftung des Deutschen Volkes.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1 (table)
Overview of experimentally-characterized RNATs (PDF 190 kb)
Related links
Glossary
- Feed-forward systems
-
Modules within control networks; feed-forward systems sense external perturbations (for example, a temperature up- or downshift) before the actual impact on the network occurs.
- Ribosome-binding site
-
The part of an mRNA with which the 30S ribosome interacts to initiate translation. It contains the Shine–Dalgarno sequence and the AUG start codon.
- Riboswitches
-
Regulatory elements within an mRNA that directly affect gene expression through specific binding of a ligand which induces structural alterations.
- Aptamer
-
An oligonucleotide that specifically binds a target molecule. RNA-based aptamers form the ligand-binding domains of riboswitches.
- Shine–Dalgarno sequence
-
The primary ribosome-binding site of bacterial mRNAs, located 6–10 nucleotides upstream of the AUG start codon, in the 5′ untranslated region. It is complementary to the 3′ end of the 16S ribosomal RNA of the 30S ribosome. The Escherichia coli consensus sequence is AGGAGG.
- tRNAfMet
-
The initiating tRNA, charged with a methionine that has the amino terminus blocked by a formyl group, thus defining the start and synthesis direction of the nascent polypeptide chain.
- syn–anti base pairing
-
Non-Watson–Crick base pairing, in which one base is rotated from the typical anti position (relative to the ribose ring) to a syn position.
- Hydration shell
-
A layer of water molecules that is formed around intracellular macromolecules (for example, DNA, RNA and proteins), with a possible effect on their functionality.
- Toeprinting
-
A primer extension method that uses inhibition of reverse transcription to probe the interaction between an mRNA and the 30S ribosome.
- Pseudoknot
-
A knot-shaped tertiary RNA structure that, in its simplest form, is built by two helical structures. The loop of the first hairpin contributes to the stem of the second hairpin. Pseudoknots are a structurally diverse group with important effects in many biological processes.
Rights and permissions
About this article
Cite this article
Kortmann, J., Narberhaus, F. Bacterial RNA thermometers: molecular zippers and switches. Nat Rev Microbiol 10, 255–265 (2012). https://doi.org/10.1038/nrmicro2730
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2730
This article is cited by
-
Wie RNA-Thermometer die Sekretion von Virulenzfaktoren kontrollieren
BIOspektrum (2024)
-
Improved production of andrimid in Erwinia persicina BST187 strain by fermentation optimization
BMC Microbiology (2023)
-
Tissue-specific temperature dependence of RNA editing levels in zebrafish
BMC Biology (2023)
-
Probing the dynamic RNA structurome and its functions
Nature Reviews Genetics (2023)
-
Temperature Matters: Bacterial Response to Temperature Change
Journal of Microbiology (2023)