Abstract
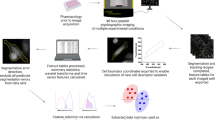
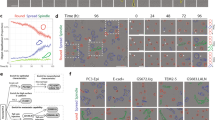
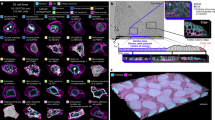
Analysis of cellular phenotypes in large imaging data sets conventionally involves supervised statistical methods, which require user-annotated training data. This paper introduces an unsupervised learning method, based on temporally constrained combinatorial clustering, for automatic prediction of cell morphology classes in time-resolved images. We applied the unsupervised method to diverse fluorescent markers and screening data and validated accurate classification of human cell phenotypes, demonstrating fully objective data labeling in image-based systems biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Conrad, C. & Gerlich, D.W. J. Cell Biol. 188, 453–461 (2010).
Goshima, G. et al. Science 316, 417–421 (2007).
Collinet, C. et al. Nature 464, 243–249 (2010).
Schmitz, M.H. et al. Nat. Cell Biol. 12, 886–893 (2010).
Neumann, B. et al. Nature 464, 721–727 (2010).
Boland, M.V. & Murphy, R.F. Bioinformatics 17, 1213–1223 (2001).
Held, M. et al. Nat. Methods 7, 747–754 (2010).
Harder, N. et al. Genome Res. 19, 2113–2124 (2009).
Wang, M. et al. Bioinformatics 24, 94–101 (2008).
Loo, L.H., Wu, L.F. & Altschuler, S.J. Nat. Methods 4, 445–453 (2007).
Jones, T.R. et al. Proc. Natl. Acad. Sci. USA 106, 1826–1831 (2009).
Conrad, C. et al. Nat. Methods 8, 246–249 (2011).
Meraldi, P., Draviam, V.M. & Sorger, P.K. Dev. Cell 7, 45–60 (2004).
Wolthuis, R. et al. Mol. Cell 30, 290–302 (2008).
Mackay, A.M., Ainsztein, A.M., Eckley, D.M. & Earnshaw, W.C. J. Cell Biol. 140, 991–1002 (1998).
Schmitz, M.H. & Gerlich, D.W. Methods Mol. Biol. 545, 113–134 (2009).
Martin-Lluesma, S., Stucke, V.M. & Nigg, E.A. Science 297, 2267–2270 (2002).
Waizenegger, I., Gimenez-Abian, J.F., Wernic, D. & Peters, J.M. Curr. Biol. 12, 1368–1378 (2002).
Neumann, B. et al. Nat. Methods 3, 385–390 (2006).
Qi, W., Tang, Z. & Yu, H. Mol. Biol. Cell 17, 3705–3716 (2006).
Thoma, C.R. et al. Nat. Cell Biol. 11, 994–1001 (2009).
Pearson, K. Philos. Mag. 2, 559–572 (1901).
Wasserman, L. All of Statistics: A Concise Course in Statistical Inference (Springer, 2003).
Jain, A.K. & Dubes, R.C. Algorithms for Clustering Data (Prentice Hall, 1988).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction (Springer, 2001).
Rabiner, L. Proc. IEEE 77, 257–286 (1989).
Chehreghani, M.H., Busetto, A.G. & Buhmann, J.M. J. Mach. Learn. Res. 22, 495–503 (2012).
Chang, C.C. & Lin, C.J. ACM Trans. Intelligent Syst. Technol. 2 (2011).
Acknowledgements
The authors thank M. Held for processing image data, D. Scheder and C. Sommer for critical comments on the manuscript and R. Stanyte for user annotations of image data. The Gerlich laboratory has received funding from the European Community's Seventh Framework Programme FP7/2007-2013 under grant agreements no. 241548 (MitoSys) and no. 258068 (Systems Microscopy), from a European Young Investigator award of the European Science Foundation, from an EMBO Young Investigator Programme fellowship to D.W.G. and from the Swiss National Science Foundation. The Buhmann laboratory has received funding from the SystemsX.ch initiative (LiverX and YeastX projects). J.P.F. was funded by an EMBO long-term fellowship.
Author information
Authors and Affiliations
Contributions
Q.Z. contributed to method design and implementation, experiments, data analysis and manuscript writing. A.G.B. contributed to method design, data analysis and manuscript writing. J.P.F. contributed to experiments and data analysis. J.M.B. contributed to method design and data analysis. D.W.G. contributed to method design, data analysis and manuscript writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 , Supplementary Tables 1–3 and Supplementary Notes 1–3 (PDF 2159 kb)
Supplementary Software
TC3 software and data (ZIP 657534 kb)
Rights and permissions
About this article
Cite this article
Zhong, Q., Busetto, A., Fededa, J. et al. Unsupervised modeling of cell morphology dynamics for time-lapse microscopy. Nat Methods 9, 711–713 (2012). https://doi.org/10.1038/nmeth.2046
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2046
This article is cited by
-
Analyzing temporal dynamics of cell deformation and intracellular movement with video feature aggregation
BioMedical Engineering OnLine (2019)
-
A curated collection of tissue microarray images and clinical outcome data of prostate cancer patients
Scientific Data (2017)
-
Microfluidic device enabled quantitative time-lapse microscopic-photography for phenotyping vegetative and reproductive phases in Fusarium virguliforme, which is pathogenic to soybean
Scientific Reports (2017)
-
Semi-automated quantification of living cells with internalized nanostructures
Journal of Nanobiotechnology (2016)
-
Image-based computational quantification and visualization of genetic alterations and tumour heterogeneity
Scientific Reports (2016)