Abstract
The ontogeny of haematopoietic niches in vertebrates is essentially unknown. Here we show that the stromal cells of the caudal haematopoietic tissue (CHT), the first niche where definitive haematopoietic stem/progenitor cells (HSPCs) home in zebrafish development, derive from the caudal somites through an epithelial–mesenchymal transition (EMT). The resulting stromal cell progenitors accompany the formation of the caudal vein sinusoids, the other main component of the CHT niche, and mature into reticular cells lining and interconnecting sinusoids. We characterize a zebrafish mutant defective in definitive haematopoiesis due to a deficiency in the nascent polypeptide-associated complex alpha subunit (NACA). We demonstrate that the defect resides not in HSPCs but in the CHT niche. NACA-deficient stromal cell progenitors initially develop normally together with the sinusoids, and HSPCs home to the resulting niche, but stromal cell maturation is compromised, leading to a niche that is unable to support HSPC maintenance, expansion and differentiation.
Similar content being viewed by others
Introduction
In vertebrate ontogeny, emerging definitive haematopoietic stem/progenitor cells (HSPCs) migrate and colonize successive specialized haematopoietic niches, which in turn regulate proliferation and differentiation of HSPCs, and release of their progeny into the blood. In the most studied haematopoietic niche, the bone marrow of mammals, the cellular constituents so far identified as important for haematopoietic niche function have been endothelial cells of venous sinusoids and arterioles, osteoblasts and associated cells at the endosteal surface, and various stromal cell subsets often closely associated with the blood vessels. Among these are CXCL12-abundant reticular cells1, Nestin+ perivascular cells2, and within the latter subset, Leptin receptor+ cells with reticular morphology around venous sinusoids3, and NG2+ cells with pericyte morphology around arterioles4. The relationship and degree of overlap between these various stromal cell subsets is still to be understood, as well as their relative localization and interaction with HSPCs and with other stromal elements in vivo.
In zebrafish development, as in mammals, definitive HSPCs initially emerge by an endothelial-to-haematopoietic transition from the ventral wall (‘floor’) of the dorsal aorta (VDA)5, and then migrate through the blood to colonize the first niche where they will expand and differentiate into multiple blood lineages. In fish, this initial niche is the caudal haematopoietic tissue (CHT6,7), while in mammals it is the fetal liver. The fetal liver haematopoietic niche is still poorly characterized8. We previously reported that the CHT of zebrafish is organized around a transient network of venous sinusoids, the caudal vein (CV) plexus, and that its ultrastructural organization is quite similar to the peri-sinusoidal haematopoietic microenvironment in the haematopoietic niche of adults—the bone marrow in mammals, or the kidney marrow in fish—with typical fibroblastic reticular cells (FRCs) in close contact with the abluminal side of venous sinusoids and extending a network between the latter, within which haematopoietic cells expand and differentiate6. In the CHT, these FRCs appear to be the main stromal cell type making up the niche together with the endothelium of the CV plexus/sinusoids. Thanks to the transparency of the zebrafish larva, the CHT is uniquely accessible to non invasive in vivo observation, offering a unique opportunity to explore in situ the development of this haematopoietic niche at a cellular level.
In the present study, we show that the stromal FRCs of the CHT niche derive from the ventral border of the caudal somites, through an epithelial–mesenchymal transition (EMT). We find that a deficiency in the α-subunit of the nascent polypeptide-associated complex (NACA, αNAC) prevents the maturation of the EMT-derived mesenchymal progenitors of the FRCs, and that this results in a CHT niche that can still trap circulating HSPCs, but is no longer able to retain them and support their expansion and differentiation. The NACA-deficient mutant thus highlights the essential role of these stromal FRCs in the making of a functional haematopoietic niche.
Results
The oloca mutant is defective in definitive haematopoiesis
We identified the zebrafish mutant oloca as a recessive mutant with fewer neutrophils by 3.5–4 days post fertilization (d.p.f.), using Sudan Black (SB) staining in a F3 mutant screen following ENU mutagenesis. SB staining of the oloca mutant showed fewer neutrophils in the head (Fig. 1b), and no neutrophils in the CHT (Fig. 1d), which normally produces numerous neutrophils by this stage9 (Fig. 1a,c). The few SB+cells found in the oloca tail were not localized within the CHT but at the body surface (Fig. 1d, arrows). Oloca mutants develop apparently normally up to 2.5–3 d.p.f. Morphological defects become obvious after 3.5 d.p.f.: a small head with immature lower jaw, cardiac oedema, disorganized iridophores in eyes, reduced circulation from 90 h post fertilization (h.p.f.) onward, and finally the larvae die by ∼4.5 d.p.f.
(a–d) Sudan black (SB) staining of sibling and oloca mutant at 3.5 d.p.f. in the rostral (a,b) and caudal (CHT; c,d) area. Arrows in (d) indicate SB+ cells located outside of the CHT. Brown spots are pigment cells. (e–z) Whole-mount in situ hybridization using indicated probes in oloca and sibling larvae. (e–h) cmyb at 72 and 90 h.p.f. (i–p) lysozyme c, c/ebp 1, nephrosin and csf1ra at 90 h.p.f. (q,r) βE1-globin at 90 h.p.f.. White arrowheads and black arrows indicate definitive haematopoietic clusters and circulating primitive erythrocytes, respectively. (s,t) ikaros at 90 h.p.f.; arrowheads indicate definitive haematopoietic clusters. (u–z) cmyb, ikaros and lck at 90 h.p.f.; arrows and arrowheads indicate thymus and kidney, respectively. Non-specific signal is seen in the notochord in panels (i,k,o,q,s). Scale bars, 100 μm.
As the SB phenotype suggested a lack of definitive granulopoiesis in the CHT of oloca larvae, we analysed the status of definitive haematopoiesis in the CHT by whole mount in situ hybridization (WISH). At 72 h.p.f., much fewer cmyb+ HSPCs were found in the oloca CHT compared with siblings (Fig. 1e,f), and none at all from 90 h.p.f. onwards (Fig. 1g,h). Accordingly, by 90 h.p.f., granulopoiesis, probed by lysC, cebp1 and nephrosin expression (Fig. 1i–n), erythropoiesis, probed by ße1-globin expression (Fig. 1q,r), and lymphopoiesis probed by ikaros expression (Fig. 1s,t) were all strongly reduced. The final definitive haematopoietic tissues, the thymus and kidney, which normally become seeded by HSPCs from the CHT by 3 and 5 d.p.f. respectively6, similarly lacked cmyb expression in oloca embryos (Fig. 1u,v), and the lymphoid markers ikaros and lck were missing in the mutant thymus (Fig. 1w–z). In contrast with the other haematopoietic lineages, csf1ra expression, which marks the macrophage lineage, was similar to siblings or even slightly increased in the CHT of oloca mutants (Fig. 1o,p). This is consistent with our observations that primitive macrophages, born in the yolk sac, colonize the nascent CHT through blood circulation before definitive haematopoiesis takes place there6.
Oloca is a mutant in the naca gene
To identify the mutation responsible for the oloca phenotype, we used simple sequence length polymorphism mapping. The mutation was mapped between flanking markers z7590 and w27205 (Fig. 2a) on chromosome 23. This interval is ∼450 kb and contains around 20 genes. One of them, naca, had been previously identified in an insertional mutagenesis screen as no knack10. no knack is an essential gene and the terminal phenotype was similar to oloca. In a cross with oloca, no knack failed to complement, strongly suggesting that oloca was a naca mutant. Sequencing of the zebrafish naca gene, in mutant and sibling embryos, revealed a premature stop codon in the mutant allele due to a C→T transition, resulting in a Q>stop alteration at codon 69 (Fig. 2b,c). The naca gene encodes the α subunit of the heterodimeric nascent polypeptide-associated complex (αNAC/NACA), which binds to nascent polypeptides on the ribosome11. NACA appears to be part of the chaperon system that ensures the proper folding of nascent proteins, notably those to be translocated in the endoplasmic reticulum, ER12,13. The naca gene also encodes a muscle-specific, much larger isoform, skNAC, through differential RNA splicing14,15. NACA and skNAC have also been described as transcriptional co-activators in osteoblasts and myotubes, respectively14,16,17.
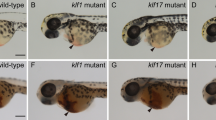
(a) Mapping of the oloca mutation using co-segregation of known simple sequence length polymorphism markers; oloca was located between w27205 and z7590 on linkage group (LG) 23. (b,c) The naca gene contains 8 exons and a C-to-T substitution was found in exon 4 in the mutant (asterisk). Splice-blocking MOs MO1 and MO2 were designed at the positions indicated. (d–g) wild-type embryos were injected with 4 ng of control or naca MO1 and analysed by WISH using a cmyb probe and by SB staining at 3.5 d.p.f. (h–k) Co-injection of naca MO1 with 125 pg of wild-type or mutant naca RNA and subsequent analysis of the caudal area at 3 d.p.f. as in (d–g). (l–q) WISH for naca expression in the caudal area from 1 to 5 d.p.f. White asterisks show the stronger expression of naca in the ventral half of somites at 25 and 30 h.p.f. in the siblings (l,n), while no signal is detected in the mutant embryos (m). Arrowheads in (n) indicate the first, mesenchymal-shaped cells expressing naca in the nascent CHT at 30 h.p.f. At 42 h.p.f., naca expression has greatly diminished in the somites and is mostly observed in the CHT in a small cell population (arrows and inset), that becomes much larger by 3 and 5 d.p.f. (p,q). ca, caudal artery; cv, definitive caudal vein (ventral-most branch of the CV plexus). Scale bars, 100 μm (d,h,l) and 50 μm (n,q).
To provide independent evidence that the naca mutation causes the oloca phenotype, we tested whether morpholinos (MOs) targeted against naca RNA would phenocopy the oloca mutant, and whether wild-type naca RNA injection would rescue the oloca phenotype. Two naca splice-blocking MOs, MO1 and MO2, phenocopied the oloca mutant. In the CHT of naca MO1-injected embryos, very few cmyb-positive HSPCs and almost no SB stained neutrophil were observed at 3.5 d.p.f. (Fig. 2d–g). Co-injection of 125 or 250 pg of naca wild-type RNA with 4 ng of naca MO1 resulted in rescue of cmyb+ HSPCs and neutrophil differentiation (SB staining) in the naca morphants (Fig. 2h,j). In contrast, co-injection of the same amounts of naca mutant RNA with naca MO1 did not rescue the morphant phenotype to any extent (Fig. 2i,k).
WISH analysis through development revealed ubiquitous expression of naca mRNA at 25–30 h.p.f., with a stronger signal in the ventral half of somites (Fig. 2l, asterisks) and in mesenchymal cells ventral to the somites in the caudal area (Fig. 2n, arrowheads). By 3 and 5 d.p.f. (by which stage haematopoiesis peaks in the CHT), naca has become specifically expressed in the CHT (Fig. 2p,q, arrows), a trend that appears to initiate by 42 h.p.f. (Fig. 2o, arrows). In oloca mutant embryos, WISH detected no naca transcripts, suggesting nonsense mediated mRNA decay (Fig. 2m).
HSPCs initially home to the mutant CHT
We used naca MO1 to assess the early effects of NACA deficiency on haematopoiesis, at stages at which mutant embryos cannot yet be distinguished morphologically. naca MO1-injected embryos showed normal primitive haematopoiesis, including granulopoiesis of yolk sac origin (Supplementary Fig. 1, arrows), and erythrocyte circulation. By 36 h.p.f., cmyb-positive definitive HSPCs were found along the aorta ventral wall (VDA) and nascent CHT of naca morphant embryos as in wild type (Fig. 3a,b). To follow their fate in live embryos, we turned to transgenic Tg[CD41:gfp] embryos, as definitive HSPCs are CD41:GFPlow from as soon as they emerge from the VDA18 (from here onwards, gfp will refer to the transgene, and, for example, CD41:GFP to the fluorescence of the GFP protein expressed from the CD41:gfp transgene). Consistently, naca MO1-injected Tg[CD41:gfp] embryos showed normal emergence of CD41:GFPlow HSPCs from the VDA at 54 h.p.f. (Fig. 3c,d,k) and their settling in the CHT (Fig. 3e,f,k). Thus, definitive HSPCs were normally specified and produced from the morphant DA floor (VDA), and initially colonized the CHT as in wild type.
(a–k) Tg[CD41:gfp] embryos were injected with control or naca MOs. (a,b) Expression of cmyb at 36 h.p.f. marking HSPC emergence in control (a) and naca morphant (b) embryos. (c–j) in vivo combined VE-DIC/fluorescence microscopy of control (c,e,g,i) and naca MO-injected embryos (d,f,h,j). (c,d) Arrows indicate emergence of CD41:GFPlow HSPCs along the VDA at 54 h.p.f. (e,f) Colonization of the CHT by CD41:GFPlow HSPCs is seen in both control and morphant embryos at 54 h.p.f. The thickness of CV plexus is almost the same between control and morphant embryos (double arrows). CV plexus segments devoid of any haematopoietic cells were observed in the morphant CHT only (f, asterisks). (g,h) At 84 h.p.f., the CHT has become much thinner in the morphant (h, double arrow) and the number of CD41-GFPlow HSPCs and CD41-GFPhigh prothrombocytes is much lower. (i,j) At 84 h.p.f., no CD41-GFPlow lymphoid progenitors were detected in the morphant thymus (j, dotted circle), unlike in control larvae (i). Number of CD41:GFPlow cells in the VDA (at 54 h.p.f.) and CHT (54 and 84 h.p.f.) of control or naca MO-injected (k), and oloca [CD41:gfp] sibling and mutant (l) embryos (mean±s.d., n=2 separate experiments). Statistical significance determined using a Student’s t-test. ****P<0.0001. n.s., not significant. cv, caudal vein; ca, caudal artery; pcv, posterior cardinal vein; da, dorsal aorta; UGO, uro-genital opening. Scale bars, 100 μm (a,b) and 50 μm (c–j).
In contrast, 30 h later, by 84 h.p.f., the morphant CHT contained much less CD41:GFPlow HSPCs than control embryos (Fig. 3g,h,k). This decline in HSPC number in the morphant CHT was accompanied by a strong reduction in the dorso-ventral width of the CHT. In addition, whereas CD41:GFPlow lymphoid progenitors normally start colonizing the thymus rudiment from 54 h.p.f. onwards18, the naca morphant thymus rudiment showed no such colonization even at 84 h.p.f. (Fig. 3i,j), consistent with our previous WISH data with the oloca mutant (Fig. 1u–z). Thus NACA deficiency did not affect HSPC emergence from the aorta, nor their initial homing in the CHT, but their subsequent maintenance, expansion and differentiation in the CHT. The same outcomes were observed with oloca:CD41gfp mutant embryos (Fig. 3l).
The mutant CHT niche is unable to support HSPC development
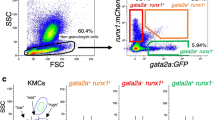
To analyse whether the defect of definitive haematopoiesis in the mutant CHT is cell-autonomous or non-cell-autonomous, we transplanted wild-type CMTPX-labelled whole kidney marrow (WKM) cells from adult Tg[mpx:gfp] fish, or GFP+ neutrophils sorted from the WKM preparation, into the sinus venosus of non-transgenic oloca mutant and sibling embryos at 48 h.p.f., and we followed up their fate in the live larvae. 1.5–2.5 h post transplantation (h.p.t.), many CMTPX-stained WKM or mpx:GFP+ transplanted cells had already homed to the CHT plexus and to the pronephric kidney area in both sibling and mutant embryos (Fig. 4a and Supplementary Fig. 2a–d). By 7 h.p.t., virtually all transplanted cells had settled in the haematopoietic tissues, in similar amounts in mutant and sibling embryos. Then in the resulting sibling larvae, the adult CMPTX+ WKM cells remained in the CHT and kidney area, and by 30 h.p.t. (78 h.p.f.), they had already strongly expanded there (Fig. 4b,c). In sharp contrast, in the mutant recipient larvae, CMTPX+ cells were no longer observed in the CHT and very few transplanted cells were detected in the kidney area by this stage (Fig. 4b,d). The same outcome occurred with the transplanted mpx:GFP+ adult WKM-derived neutrophils that homed to the CHT. In the sibling larvae, by 36 h.p.t. they were still in the CHT and kidney area (Supplementary Fig. 2e,g, Supplementary Movie 1) where they had expanded, and some were seen undergoing mitosis (as commonly observed also with endogenous larval mpx:GFP+ neutrophils). In contrast, in the mutant larvae, the transplanted mpx:GFP+ cells, which had similarly homed to the CHT up to 7 h.p.t., started leaving it from 12 h.p.t. onwards, entering circulation as well as migrating into surrounding superficial tissues, such that by 36 h.p.t., very few mpx:GFP+ cells were remaining in the CHT and kidney area (Supplementary Fig. 2f,h, Supplementary Movie 1). TUNEL staining on the recipient mutant larvae confirmed that the transplanted mpx:GFP+ cells that initially homed to the CHT did not disappear by apoptosis.
(a) Fluorescence microscopy of sibling and oloca mutant transplanted at 48 h.p.f. with CMTPX-stained WKM cells obtained from adult Tg[mpx:gfp] fish. Numerous mpx:GFP+ (green) and/or CMTPX+ (red) WKM cells were observed in the CHT niche of both oloca mutant (olc) and sibling embryos at 2 h post transplantation (h.p.t.=50 h.p.f.). (b) Almost no remaining transplanted cells were detected in the CHT of olc embryo at 30 h.p.t. whereas CMTPX+ WKM cells had expanded in the sibling CHT (arrows). (c,d) In the kidney area, stable colonization of transplanted WKM cells was detected in the sibling larva whereas very few transplanted cells were observed in the olc embryo at 30 h.p.t. (arrowheads). (e–i) Confocal microscopy analysis at 60 h.p.f. of parabiotic embryo pairs composed of a control MO-injected Tg[lysC:DsRed] embryo and a naca MO-injected Tg[mpx:gfp] embryo. A parabiotic pair is shown in (e), and magnified images of the head (f,h) and CHT (g,i) of both parabiotic partners are shown. (j) For six such parabiotic pairs, the number of GFP+ and DsRed+ cells was counted in the CHT of both partners (‘cont’ and ‘naca’, mean±s.d.; n=3 separate experiments). Scale bars, 200 μm (a,b,e), 150 μm (c,d) and 100 μm (f–i). Statistical significance determined using a Student’s t-test. ****P<0.0001.
HSPCs are not intrinsically compromised in the oloca mutant
We next addressed whether the haematopoietic defect in the oloca mutant was solely due to its deficient CHT niche, or whether it also involved a cell-autonomous defect of mutant HSPCs. To address this point, we took advantage of a recent method for generating parabiotic embryo pairs19, applying it to Tg[mpx:gfp] and Tg[lysC:DsRed] embryos, that give rise to green and red fluorescent neutrophils, respectively. When we generated parabiotic wild-type Tg[mpx:gfp]/Tg[lysC:DsRed] embryo pairs, by 60 h.p.f., the CHTs of both embryos displayed both GFP+ and DsRed+ neutrophils, indicating that the definitive HSPCs born from the aortas of both embryos had similarly homed to both CHTs and given rise to (resp. GFP+ and DsRed+) neutrophils (Supplementary Fig. 2i–m). We then performed the same parabiosis experiment but using naca MO-injected Tg[mpx:gfp] and control MO-injected Tg[lysC:DsRed] partner embryos (Fig. 4e). By 60 h.p.f., mpx:GFP+ and lysC:DsRed+ cells were similarly observed in the rostral region of both control and naca MO-injected embryos (Fig. 4f,h), consistent with the unimpaired primitive granulopoiesis in naca mutants. In contrast, numerous mpx:GFP+ and lysC:DsRed+ cells were observed in the CHT of control MO-injected lysC:DsRed embryos whereas only few cells of either origin were found in the naca morphant CHT (Fig. 4g,i,j). These results confirm that naca deficiency leads to a deficient CHT niche and show that, in contrast, the homing capacity and granulocytic differentiation ability of NACA-deficient definitive HSPCs is intact.
CHT stromal cells originate from the caudal somites
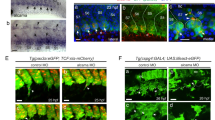
In our initial description of the CHT niche6, our electron microscopy data suggested it to consist in two main cell types, the endothelial cells of the CV plexus sinusoids, and mesenchymal cells in between the sinusoids that appeared to evolve into what we called FRCs, owing to their ultrastructural resemblance to FRCs in mammalian bone marrow and adult fish kidney marrow. Therefore we screened various existing transgenic reporters in search of one that would highlight the FRCs and their progenitors. We found that a pair of Wnt signalling reporter lines, Tg[TCF:nls-mCherry] and Tg[TCF:gfp]20, respectively expressing nuclear mCherry and whole-cell GFP, appeared to do so. As previously described20, by 20 h.p.f. (24-somites), these two reporters are highly expressed in the caudal somites. Time-lapse confocal imaging of Tg[TCF:gfp] embryos from 21 h.p.f. revealed that TCF:GFP+ cells belonging to the ventral side of the caudal somites started to migrate ventrally, and settled in the future CHT by 25–28 h.p.f. (Supplementary Fig. 3a and Supplementary Movie 2). The same cells and migration pattern were revealed using Tg[TCF:nls-mCherry] embryos (Supplementary Movie 3a). We thus combined the time-lapse confocal imaging of the TCF:nls-mCherry+ nuclei of these emigrating cells with vital staining with the interstitial fluorescent marker Bodipy ceramide, which has been used to delineate somite and cell contours within the somitic epithelium21,22 (Supplementary Fig. 3b). The four-dimensional tracking of individual nuclei of the emigrating cells confirmed that they were initially all intra-somitic, and located within the entire medio-lateral extent of the ventral border of the somites (Supplementary Fig. 3c–e and Supplementary Movie 4). We studied this emigration process in more detail by combining fluorescence with video-enhanced Differential Interference Contrast (VE-DIC) microscopy6. From 19 h.p.f., epithelial cells from the ventral part of the caudal somites underwent an EMT. They started to show active blebbing through the somite basal lamina (Fig. 5a and Supplementary Movie 5), then more extended protrusions in ventral direction (Fig. 5b), and emigrated as strings of mesenchymal cells into the more ventral caudal space, with frequent mitoses before and during their migration (Fig. 5c–e, Supplementary Movie 6). In that space, at the same time, the primordial CV started sprouting in ventral direction to give rise to the CV plexus by 30 h.p.f. To observe the interaction between the two processes, we crossed Tg[TCF:nls-mCherry] with Tg[fli1:gfp] fish. At 25 h.p.f., the angiogenic sprouting of endothelial cells from the primordial CV was very dynamic, and TCF:nls-mcherry+ cells were found interspersed with the endothelial sprouts (Supplementary Movie 3b), interacting with them and with each other through filopodial extensions (Fig. 5f,g). By 35 h.p.f., the CV plexus was complete and functional6. Between 35 and 48 h.p.f., the cytomorphology and distribution of nls-mCherry+ mesenchymal cells in the CHT evolved. By 35 h.p.f. they displayed a large nucleus, scant cytoplasm and long filopodia by which they connected to each other and to endothelial cells (Fig. 5i,j). By 48 h.p.f., their nuclei had become smaller, with a central, well-defined nucleolus, and while they were still making a dynamic network of interconnecting filopodial extensions connecting them to each other and other cells (Fig. 5k and Supplementary Movie 7), many of them now had their cell body in close apposition to the abluminal side of endothelial cells of the CV plexus (Fig. 5h,l). This configuration is identical to that of the FRCs that we previously described by electron microscopy as the main stromal cell type making up the CHT niche together with the endothelial cells of the CV plexus6. We further evidenced the somitic origin of these cells by transiently expressing GFP from a skeletal muscle α-actin promoter in Tg[TCF:nls-mCherry] embryos. By 22 h.p.f., [GFP+; nls-mCherry+] double positive cells were observed throughout the somites (Supplementary Fig. 4a–c). Then double-positive cells were observed by 24 h.p.f. emigrating from the somites (Supplementary Fig. 4d–f), and by 36 h.p.f. among the developing stromal cells both between and in close apposition to the CV plexus endothelia (Supplementary Fig. 4g–m) as previously observed (Fig. 5h,l).
in vivo observation of [TCF:nls-mCherry; Fli1:gfp] double transgenic embryos. (a–e) VE-DIC microscopy reveals the epithelial–mesenchymal transition giving rise to stromal progenitors. (a) Active cell blebbing (arrows and Supplementary Movie 5) through the somite basal lamina (arrowheads) was often observed at the ventral side of caudal somite epithelial cells by 21 h.p.f. (b) An hour later (22 h.p.f.), the transient blebs were replaced by more stable protrusions (arrows), then the cells started to emigrate from the somite as mesenchymal cells (c,d); mitosis was often observed around the onset of this emigration (d, arrowheads), and the emigrating cells tended to form strings (e, arrows) and interact through filopodia. (f,g) By the same stage (25 h.p.f.), VE-DIC and fluorescence overlay shows physical interactions between these nls-mCherry+ stromal cell progenitors (asterisks) and the Fli1:GFP+ endothelial cells of the developing CV plexus (f, arrows); the inset in (g) shows such an interaction at higher magnification. (h) One day later, among the nls-mCherry+ stromal cells (asterisks), some have now settled in adventitial position (arrows) along the endothelial cells forming the main caudal vein. (i–l) VE-DIC imaging of the cytomorphological maturation of stromal progenitors in relation to the CV plexus. (i,j) At 36 h.p.f., young stromal progenitors display a large nucleus (arrows) and interact with endothelial cells (arrowheads) through their filopods (asterisks). (k,l) At 48 h.p.f., two types of TCF:nls-mCherry+ stromal cells are observed; interconnecting (k) and adventitial (l) stromal cells. Nucleoli were often clearly visible in those stromal cells at this stage (arrows). Arrowheads and asterisks indicate endothelial cells and filopods of stromal cells, respectively. som, somite; sp, stromal progenitor; end, endothelial cell; cv, caudal vein. Scale bars, 10 μm (a–h) and 5 μm (i–l).
Finally, in vivo observation of Tg[sox10:gfp] which highlights neural crest cells (NCCs)23, revealed no NCC contribution to the CHT other than pigment cells (Supplementary Fig. 5a,c). Moreover, sox10 mutants lacking most NCC derivatives revealed no sox10:GFP+ cells in the CHT, yet no morphological abnormality of the CHT, and the presence of typical stromal cells as in the wild type (Supplementary Fig. 5a–h), indicating that NCCs do not contribute to stromal cell development during CHT niche formation. We therefore conclude that the FRCs/stromal cells initially arise from the caudal somites through a typical EMT, and that their progenitors accompany the formation of the CV plexus, leading altogether to the final haematopoietic niche. From now on, we will designate these cells as stromal cells or stromal reticular cells (SRCs).
Naca is required for stromal cell maturation
We next compared the development of caudal endothelial (Fli1:GFP+) and stromal (TCF:nls-mCherry+) cells between control and naca morphant embryos. Time-lapse confocal analysis on [Fli1:gfp; TCF:nls-mCherry] double transgenic embryos indicated that the initial sprouting of Fli1:GFP+ CV endothelial cells and parallel emigration of TCF:nls-mCherry+ stromal cell progenitors from the somites occurred normally in morphant embryos compared with controls (Fig. 6a,b).
(a–f) Live confocal fluorescence imaging of the CHT area in control and naca MO-injected [TCF:nls-mCherry; Fli1:gfp] double transgenic embryos at 24, 48 and 60 h.p.f. Double arrows indicate the thickness of the developing CV plexus. (a,b) Arrows indicate some of the stromal cell progenitors accompanying the protrusion of Fli1:GFP+ endothelial cells sprouted from the primordial CV. (c–f) Asterisks indicate intervascular spaces within the CV plexus; note that these spaces in the morphant (d) are larger and flatter at 48 h.p.f. than in control embryo (c), and that they contain fewer nls-mCherry+ stromal cells at 48 h.p.f., and virtually none at 60 h.p.f. (f). (g) Number of TCF:nls-mCherry+ cells in the CHT of control and naca MO-injected embryos at 48 h.p.f. Six control and 10 naca MO-injected embryos were analysed. (h,i) Comparison of the thickness (h) and number of intervascular spaces (i) of the CV plexus between control and naca morphant embryos at 48 h.p.f. Nine control and 14 naca MO-injected embryos were analysed. Error bars represent the s.d. Statistical significance determined using a Student’s t-test. **P<0.01; ***P<0.001; ****P<0.0001. som, somite; ca, caudal artery; cp, caudal vein plexus; isv, inter-somitic vessel. Scale bars, 50 μm.
By 48 h.p.f., about two times less TCF:nls-mCherry+ stromal cells were detected in the naca morphant CHT relative to controls (Fig. 6c,d,g). However this did not necessarily reflect a true decrease in stromal cell number at this stage, because the mean mCherry fluorescence per detected cell was also diminished over 2-fold in both the somites and CHT stromal cells (Supplementary Fig. 6a–d) but not in the fin mesenchymal cells (Supplementary Fig. 6e) in morphants relative to controls. Close inspection by combined VE-DIC and fluorescence microscopy did confirm the presence, only in the morphants, of stromal cells visible by VE-DIC that had virtually no more detectable mCherry signal (Supplementary Fig. 6b, arrowheads). By 60–72 h.p.f., both fluorescence and VE-DIC microscopy evidenced a virtually complete lack of stromal cells in the naca morphant (Fig. 6e,f, Supplementary Fig. 6f,g).
In parallel, the CV plexus was slightly thinner in the naca morphant at 48 h.p.f. and displayed fewer and often larger intervascular spaces in sagittal optical section (Fig. 6c,d,h,i). Between 48 and 60 h.p.f., these spaces flattened and the difference in CV plexus thickness between control and morphant larvae kept increasing (Fig. 6e,f, Supplementary Movie 8), as observed above in the Tg[CD41:gfp] background (Fig. 3e–h). In addition, in the thinner naca morphant CHT, blood kept circulating in the various branches of the CV plexus, unlike in controls in which blood eventually flows only through the ventral-most branch, the definitive CV, as the more dorsal branches become packed with haematopoietic cells6. Both features thus are likely consequent to the lack of haematopoietic cells accumulation in the naca morphant CHT.
NACA is cell-autonomously required for stromal cell survival
Our data suggest naca deficiency affects stromal cells survival during CHT niche maturation. To further investigate whether the requirement for naca in stromal cell development is cell-autonomous, gastrula transplantation was performed to generate chimeric embryos, in which naca function is partially recovered or affected within the stromal cell population. At early gastrula stage (40% epiboly), cells located at the outer margin of the blastoderm (where mesoderm fates arise) of wild-type (control MO-injected) Tg[H2A-gfp] (ref. 24) embryos (which express nuclear GFP ubiquitously) were transplanted into naca MO-injected Tg[TCF:nls-mCherry] at the same stage and place (Fig. 7a–d). Reciprocally, cells from naca MO-injected Tg[H2A-gfp] embryos were transplanted in the same way into control MO-injected Tg[TCF:nls-mCherry] embryos (Fig. 7e–h). Recipient embryos were individually followed up at 42 and 60 h.p.f. At 42 h.p.f., SRCs originating from the wild-type H2A-GFP+ donor could be observed in the naca morphant recipient in 7/120 successful transplants (Fig. 7a,b). These transplanted H2A-GFP+ stromal cells had slightly expanded by 60 h.p.f., while the naca-deficient endogenous TCF:nls-mCherry+ stromal cells had mostly disappeared (Fig. 7c,d,i). In the reciprocal experiment, cells transplanted from naca MO-injected H2A-GFP+ donors into wt recipient Tg[TCF:nls-mCherry] embryos also provided stromal cells associated with the caudal plexus at 42 h.p.f., in 6/120 successful transplants (Fig. 7e,f); however their number had greatly decreased by 60 h.p.f., unlike the TCF:nls-mCherry+ stromal cells from the wild-type recipient which had slightly expanded (Fig. 7g,h,j). These results demonstrate that naca is cell-autonomously required for SRC survival.
(a–d) Cells at the ventral margin of control MO-injected H2A-GFP+ embryos at 40% epiboly stage were transplanted into the same territory of naca MO-injected TCF:nls-mCherry embryos; a transplanted embryo is shown at 42 h.p.f. (a,b) and 60 h.p.f. (c,d); arrows and arrowheads indicate transplanted wild-type H2A-GFP+ and endogenous naca-deficient TCF:nls-mCherry+ stromal cells, respectively. (e–h) Reciprocal experiment, transplanting cells from naca MO-injected H2A-GFP+ embryos into control MO-injected TCF:nls-mCherry embryos; transplanted embryo at 42 h.p.f. (e,f) and 60 h.p.f. (g,h); arrows and arrowheads indicate transplanted naca-deficient H2A-GFP+ and wild-type TCF:nls-mCherry+ stromal cells, respectively. Asterisks indicate intervascular spaces within the developing CHT plexus typically harbouring stromal cells at these stages. (i,j) Quantification of CHT stromal cells of donor and recipient origin at 42 and 60 h.p.f. in the embryos that contained donor-derived CHT stromal cells at 42 h.p.f. Statistical significance determined using paired t-test. *P<0.05, ***P<0.001. ca, caudal artery; cv, caudal vein. Scale bars, 50 μm.
NACA deficiency triggers the UPR and stromal cell apoptosis
Previous reports have shown that NACA depletion can cause ER stress and ER stress-induced apoptosis through activation of the unfolded protein response (UPR) pathway in mammalian cultured cells25. We therefore analysed the expression of seven UPR-related genes in sibling and mutant larvae at 3.5 d.p.f. Four genes (atf4, bip, chop and xbp-1) coding for UPR effectors were significantly up-regulated in the mutant (Fig. 8a). In contrast, three genes (atf6, ire1 and perk) encoding ER stress sensors showed little or no significant difference in expression between oloca mutant and sibling larvae (Fig. 8b). To clarify whether NACA-deficient cells may die by apoptosis, we performed whole mount TUNEL staining in oloca mutants in different transgenic backgrounds. By 3 d.p.f., TUNEL+ signals were observed in the CHT of mutant but not sibling larvae. Using tyramide-amplified immunodetection, we could detect double positive, TCF:nls-mCherry+/TUNEL+ apoptotic bodies, evidencing apoptotic death of TCF:nls-mCherry+ mutant stromal cells (Fig. 8c–e). In contrast we found virtually no TUNEL+ apoptotic bodies among gata1:DsRed+ erythroid cells, mpx:GFP+ neutrophils, CD41:GFPlow HSPCs and ikaros:GFP+ lymphoid progenitors in the mutant CHT (Supplementary Fig. 7a–h).
(a,b) qPCR reveals significant increase in transcript levels of atf4, bip, chop and xbp-1 (a) but not in atf6, ire1 and perk (b) in oloca mutants relative to siblings (n=3 biological replicates, average of 3 independent experiments). (c–e) TUNEL staining was performed on oloca mutants and siblings at 3 d.p.f. in the TCF:nls-mCherry transgenic background. Arrows and arrowhead indicate TCF:nls-mCherry/TUNEL double positive cells and TUNEL single positive cells, respectively. (f) Number of TUNEL+ cells in the CHT of control (DMSO)- or 4PBA-treated oloca mutant and sibling embryos at 3.5 d.p.f. (n=2 separate treatments). (g–j) WISH for cmyb expression in the CHT of control (DMSO) or 4PBA-treated oloca mutant and sibling embryos at 3.5 d.p.f. (k) Number of TUNEL+ cells in the CHT of Bcl-XL mRNA injected or control embryos at 3.5 d.p.f. (n=2 separate injections). (l) in vivo counting of TCF:nls-mCherry+ stromal cells in the CHT of Bcl-XL mRNA injected or control embryos at 60 h.p.f. (m–o'). (m) in vivo counting of CD41-GFPlow HSPCs in the CHT of Bcl-XL mRNA injected or control embryos at 3 d.p.f. (q–s). n=2 separate injections for (l) and (m). Error bars represent the s.d. and statistical significance determined using a Student’s t-test. *P<0.05; ***P<0.001; and ****P<0.0001. n.s., not significant. Scale bars, 25 μm (c), 50 μm (n) and 100 μm (h,q).
We then sought to block the ER stress/UPR pathway during CHT niche formation to determine whether activation of the UPR pathway causes stromal cell death and consequent defects in niche formation. Chemical chaperones such as 4-phenylbutyric acid (4-PBA) enhance protein folding and thus act as a suppressor of ER stress. When we treated oloca sibling and mutant embryos with 4-PBA, we found first that the fold enhancement of UPR genes (atf4, atf6, bip and chop) expression in mutant relative to sibling embryos was significantly decreased (Supplementary Fig. 7i); second, the number of TUNEL+ cells was also decreased in the CHT of oloca mutants (Fig. 8f). Third, 4-PBA treatment significantly rescued the number of HSPCs marked by cmyb in situ hybridization at 3.5 d.p.f. (Fig. 8g–j).
Finally, injection of mRNA encoding the anti-apoptotic protein Bcl-XL in oloca mutant embryos totally suppressed the occurrence of TUNEL+ cells in the mutant CHT (Fig. 8k). It only partly rescued the number of TCF:nls-mCherry+ stromal cells at 60 h.p.f. (Fig. 8l,n–p), possibly reflecting the faster extinction of the transgene in the mutant, and it fully rescued the number of CD41-GFP+ HSPCs at 3 d.p.f., up to the wild-type level (Fig. 8m,q–s).
Discussion
We previously described that in zebrafish development, the CHT—the initial niche where definitive HSPCs home, expand and differentiate—consists of two main cell types: the venous endothelial cells of the CV plexus, and a cytomorphologically homogeneous stromal cell type which we named the FRCs, given their similarity in cytology and spatial organization to the FRCs of adult fish kidney marrow and mammalian bone marrow6. The origin of these FRCs was so far unknown. Here we have discovered that they originate from the ventral border of the caudal somites, through a typical EMT. Having now better described the development of these cells in vivo, we prefer to designate them as SRCs, as this name represents their two salient characteristics. Like NCCs after their EMT from the neural tube, the SRCs emigrate from the caudal somites as strings of cells that often divide on the way. They will constitute an excellent, readily accessible model for future in vivo studies of EMT. Although very little is known about the corresponding haematopoietic niche in mammals, in the fetal liver, Chagraoui et al.26 reported that in vitro haematopoiesis-supportive stromal cells isolated from fetal liver consisted of ‘EMT cells’, as defined by co-immunostaining with a series of epithelial and mesenchymal markers.
Recently, Nguyen et al.27 demonstrated that trunk somites give rise to cells that incorporate in the DA and facilitate the hemogenic specification of other endothelial cells in the DA floor. Thus, our present work and this study together reveal an amazingly pervasive, and so far entirely unsuspected, cellular contribution of somite-derived cells in nurturing HSC development and then function, in first the specification of HSCs within the aorta27, and later their expansion and differentiation in the first niche where they home (this work).
In the CHT, the ventral-wards emigration of SRC progenitors from the somite border closely accompanies the formation of the CV plexus in the same space, such that by 32–35 h.p.f., the very onset of HSPC budding from the aorta floor, the CV plexus is complete, with blood circulating through all its branches, and SRC progenitors interconnecting the CV branches. The niche is thus ready just in time to trap the first circulating aorta-derived HSPCs. Over the next 24 h, the SRC progenitors reorganize and mature into SRCs partly lining the abluminal face of the venous sinusoids.
In mammals, neural crests contribute to bone marrow niche formation via nestin+ mesenchymal stem cells28. In contrast, we observed no contribution of NCCs to the CHT other than pigment cells, and found normal SRCs and CHT haematopoiesis in a sox10 null mutant which lacks most neural crest derivatives.
Here we could observe the mesenchymal SRC progenitors in vivo due to their labelling through a Wnt signalling reporter transgene. This does not necessarily imply that these cells keep receiving and transducing Wnt signals. Indeed, the TCF:GFP or TCF:nls-mCherry reporter is highly expressed in the somites and consequently in the SRC progenitors as they emerge from them, but then the fluorescent signal progressively decays in these cells, such that by 3 d.p.f., they are no longer detectable in vivo by this means, suggesting that the reporter protein was mostly inherited from the somitic origin.
The second main result of our study is that a deficiency in NACA drastically compromises the sustainability and differentiation of HSPCs in the CHT, due to a defect not in the HSPCs, but in the CHT niche, making the oloca mutant the first of this kind in the zebrafish literature so far. In the oloca mutant, HSPCs, whether autologous or transplanted from wild-type adult kidney marrow home to the CHT niche just as in a wt larva, but then are unable to stay, expand and differentiate there. We traced this defect of the CHT niche to a defective maturation of the SRCs. Initially, SRC progenitors emerge normally from the somites, and accompany the formation of a functional CV plexus, which traps circulating HSPCs as well as in wild type. But then, over the next 2 days, the SRC progenitors progressively disappear instead of maturing into SRCs, and the resulting CHT is unable to support HSPC maintenance. We found that NACA is required cell autonomously for the maturation and survival of SRCs.
Why does NACA deficiency cause the disappearance of SRC progenitors in particular? In mammals, NACA deficiency can cause ER stress and consequently the UPR, which, depending on the context, can trigger apoptosis25,29. We found that NACA deficiency does trigger the UPR in zebrafish larvae, and that treatment of naca mutants with the chemical chaperon 4-PBA significantly suppressed SRC apoptosis, and restored CHT haematopoiesis. Interestingly, EMT-derived mesenchymal cells in general were recently shown to be exquisitely sensitive to ER stress-induced apoptosis30. This feature was shown to be linked to their high level of extracellular matrix secretory activity, which involves steady ER development. Here in the CHT, the mesenchymal SRC progenitors are most likely responsible for secreting the large amount of extracellular matrix that surrounds the CV plexus by 48 h.p.f.6, which would render them more sensitive to the ER stress caused by NACA deficiency.
Finally, the transplantation of WKM cells from adult fish into 2 d.p.f. larvae revealed an unexpected feature. The WKM cells readily homed not only to the CHT, but also to the kidney area. This was surprising because the developmental colonization of kidney by larval HSPCs only starts by 4 d.p.f.6,31. This unexpected observation suggests that developmentally, the kidney niche is potentially able to trap circulating HSPCs already by 2 d.p.f., but that larval HSPCs are not yet able to home to it. Now, whereas wild-type WKM cells similarly homed to the wt and oloca mutant kidney short after transplantation, 2 days later these cells had disappeared from the mutant kidney area, while they had expanded in the wild type. This observation suggests that NACA deficiency may affect not only the CHT niche, but also the kidney marrow niche, in a similar way. The early (pronephric) kidney niche has not been much studied yet, but similar SRCs are found in the CHT and adult kidney marrow niches6. Therefore we suggest that NACA deficiency may affect CHT and kidney marrow SRCs/FRCs in the same way, with similar consequences. The same may well apply to mesenchymal cells of mammalian haematopoietic niches such as the bone marrow.
Methods
Zebrafish
Transgenic and mutant stocks of zebrafish were raised and staged according to Westerfield32. AB wild-type fish and transgenic lines Tg[fli1:gfp]33, Tg[mpx:gfp]34, Tg[lysC:DsRed]35, Tg[lmo2:DsRed]36, Tg[gata1:DsRed]37, Tg[CD41:gfp]38, Tg[ikaros:gfp]39, Tg[TCF:gfp], Tg[TCF:nls-mCherry]20, Tg[H2A-gfp]24 and sox10 clsm618 mutant40 in Tg[sox10:egfp]23 background were used.
Generation and identification of the oloca mutant
The oloca mutant (allele t30953) was identified in a large-scale N-ethyl-N-nitrosourea (ENU) mutagenesis screen (Tübingen 2005 Screen Consortium) by whole-mount Sudan Black staining of neutrophils9 in 4 d.p.f. larvae. Meiotic mapping of olocat30953 was performed using standard simple sequence length polymorphisms. The naca gene was cloned from cDNA made from mutant and wild-type siblings (primers nacaF1 and nacaR1). DNA was sequenced on both strands (primers nacaF2 and nacaR2). Genomic sequencing was performed by PCR amplification of naca exon 3 from mutant and wild-type larvae (primers nacaF3, nacaR3, nacaF4 and nacaR4). Primers used are listed in Supplementary Table 1. The naca wild-type and mutant cDNAs were cloned into pCS2+ for mRNA injections (primers nacaF5 and nacaR5).
In situ hybridization
Whole-mount RNA in situ hybridization was performed according to Thisse (https://wiki.zfin.org/display/prot/Thisse+Lab+-+In+Situ+Hybridization+Protocol+-+2010+update) on embryos fixed at 26, 35, 48 and 72 h.p.f., 4 and 5 d.p.f. Riboprobes were synthesized for cmyb41, gata1 (ref. 42), ßE1-globin43, c/ebp1 (ref. 44), ikaros45, lck46, lysozyme C47, nephrosin (GenBank accession number. AY330223), csf1ra48. The naca wild-type cDNA cloned into pCS2+ vector was used for naca probe synthesis.
Morpholino and mRNA injections
One to two-cell-stage embryos were microinjected with 4 ng of a splice blocking antisense morpholino (MO) synthesized against naca; naca MO1, 5′-TTAAAAGCAGTACTAACCCGTCTCA-3′, Genetools). Efficacy of MO knockdown was assessed by typical morphological abnormalities of oloca mutant and following SB staining. Parallel studies using another splice blocking MO (naca MO2, 5′-AAAAGAAAGGCATCTTACCTGTGCC-3′) produced similar results, so all data presented in this study were derived from using naca MO1. For control embryos, the same amount of a standard control morpholino (5′-CCTCTTACCTCAGTTACAATTTATA-3′) was used. Wild-type and mutant mRNA used in the rescue assays were generated using the SP6 mMessage mMachine kit (Ambion) and co-injected with naca MO1 at a concentration of 125 ng μl−1.
Wide-field microscopy
VE-DIC and fluorescence microscopy were performed as described previously6,9 through the 60x/1.00 NA water-immersion objective of a Nikon 90i microscope. Time-lapse recording of VE-DIC images was performed using BTVpro software (Bensoftware, London).
Bodipy ceramide staining
Vital staining using Bodipy ceramide (Molecular probes) was performed as described previously22. Briefly, embryos at 19.5 h.p.f. were incubated in the staining solution (100 μM Bodipy ceramide, 2% dimethylsulphoxide (DMSO) in 10 mM HEPES (pH7.4)/E3 medium) for 30 min. at 28 °C. Embryos were rinsed in Volvic water and subjected to confocal microscopy.
Confocal microscopy, image processing and quantification
Confocal fluorescence microscopy was performed on Leica SPE and SP8 setups as described previously18. For quantification of the mean nls-mCherry fluorescence intensity per cell type in double Tg [TCF:nls-mCherry; Fli1:gfp] embryos, a volume of interest (VOI) was defined in 10 cells of each population of interest (stromal, somite muscle and fin mesenchymal cells) per embryo, and for each VOI the mean fluorescence/voxel was calculated using Imaris software (Bitplane). Eight control and eight naca MO-injected embryos were thus processed. The mean of 10 cells from each cell population was subjected to statistical analysis using unpaired two-tailed Student’s t-test (GraphPad Prism software). For nuclear tracking of Bodipy ceramide-stained TCF:nls-mCherry embryo, Imaris software was used for semiautomated four-dimensional tracking of nuclei over time; the tracks were manually checked under stereoscopic viewing, using Nvidia 3D-Vision glasses coupled to a double-frequency (120 Hz) video display.
Whole kidney marrow transplantation
Six adult Tg[mpx:gfp] transgenic fish were deeply anesthetized and all parts of the kidney (head, trunk and tail) were isolated in ice-cold PBS. WKM cells were prepared as described previously49. The cell suspension (5–8 × 105 ml−1 in PBS) of WKM was labelled with 5 μM of CellTracker Red CMTPX (Invitrogen) in PBS at 32 °C for 15 min, then washed several times in the zebrafish culture media previously reported by Stachura et al.50, then incubated at 32 °C for another 40 min to release excess CMTPX. Stained cells were prepared in the zebrafish culture media at a concentration of 5 × 107 ml−1 for the transplantation. Cell sorting for mpx-GFP+ neutrophils was based on forward and side scatter and GFP fluorescence as described previously37,51, using a FACSAria III flow cytometer. Sorted mpx-GFP+ cells were spun down and resuspended in DPBS (Invitrogen) at a concentration of 3 × 107 ml−1 for transplantation. CMTPX-labelled WKM cells or sorted mpx-GFP+ cells were transplanted into 2 d.p.f. larvae (mutant or sibling) using the same setup as in Sarris et al.52. The cells were transplanted into the duct of Cuvier so as to enter circulation. Larvae with similar numbers of transplanted cells were kept overnight at 28 °C in E3 medium (5 mM NaCl, 0.15 mM KCl, 0.33 mM CaCl2, 0.33 mM MgSO4, methylene blue and 1-phenyl-2-thiourea, PTU). Two to thirty hours post transplantation larvae were mounted in 1% low melting point agarose in E3 medium supplemented with 0.16 mg ml−1 tricaine (Sigma) and used for imaging.
Generation of parabiotic zebrafish embryos
Four ng of naca MO or control MO were injected in 1-cell stage Tg[mpx:gfp] or Tg[lysC:DsRed] embryos, respectively. Injected morphants were raised until 1k-cell (late blastula) stage, then the two blastoderms were fused superficially as described19 and let to develop into partially fused larvae. Non-injected Tg[mpx:gfp] and Tg[lysC:DsRed] embryos were fused for the control experiment.
Transient expression of α-actin:GFP
The plasmid pαact-GFPI2 containing skeletal muscle α-actin (acta1a) promoter driving GFP was co-injected with I-SceI meganuclease into 1-cell stage Tg[TCF:nls-mCherry] embryos (2.5–5 pg per embryo) as previously described53.
Gastrula transplantation
naca or control MO-injected embryos were dechorionated in a glass petri dish filled with high-calcium Ringer solution19 at late blastula stage. Two donor (naca or control MO-injected H2A-gfp) embryos and 4–5 host (control or naca MO-injected TCF:nls-mCherry) embryos were gently loaded using a glass pipet on a thin layer of 3% methylcellulose spread on a 30 mm petri dish filled with Ringer solution with antibiotics (penicillin/ streptomycin; 1% v/v). The transplant capillary was cut at 25 μm inner diameter and filled with Ringer solution before use. At 40% epiboly stage (5 h.p.f.), ∼50–100 cells from the donor embryos were aspirated, targeting the ventral margin of the blastoderm that will give rise to somitic mesoderm, using the CellTram micromanipulator (Eppendorf). Around 50 cells were transplanted into the exact same location of each recipient embryo. The transplanted embryos were kept in the same solution overnight at 28 °C, then the medium was replaced with Volvic/PTU the next day. The number of GFP and nls-mCherry positive stromal cells in the CHT was counted at 42 and 60 h.p.f. for in each successfully transplanted recipient embryo. Statistical analysis was performed using paired t-test (GraphPad Prism software).
4-PBA treatment
oloca mutant and sibling embryos were treated with 50 μM 4-phenylbutyrate (4-PBA; Sigma) in 10 mM HEPES (pH 7.4)/E3 medium from 24 h.p.f. The medium was renewed every day during the treatment period. Control embryos were incubated with vehicle (DMSO) alone at the equivalent concentration (0.01%). The 4-PBA- or DMSO-treated embryos were separated into sibling and mutant groups then fixed at 3.5 d.p.f. for in situ hybridization or TUNEL staining. Total RNAs for quantitative PCR (qPCR) analysis were extracted from groups of 15 oloca sibling or mutant embryos at 3.5 d.p.f.
Bcl-XL mRNA injection
For synthesis of Bcl-XL mRNA, the plasmid pCS2+flag-zBclXL54 was linearized with Not I then 1 μg of linearized plasmid was subjected to in vitro transcription using SP6 mMessage mMachine kit (Ambion). oloca mutant and sibling embryos in CD41:gfp or TCF:nls-mCherry transgenic background were injected with 200 pg of Bcl-XL mRNA per embryo at the one-cell stage. The injected embryos and non-injected controls were raised till 60 h.p.f. (oloca:TCF:nls-mCherry) or 3 d.p.f. (oloca:CD41:gfp), then subjected to confocal microscopy for cell counting. For the detection of oloca:TCF:nls-mCherry mutant embryos, confocal images of the CHT from 27 randomly chosen embryos were acquired at 60 h.p.f., and the embryos were then raised separately until 3 d.p.f. to ascertain their mutant or sibling phenotype.
qPCR and TUNEL staining
oloca sibling and mutant larvae were collected separately at 3–3.5 d.p.f. Total RNA was extracted using TRIzol reagent (Sigma) from each pool of 30 sibling and mutant embryos and 3 μg of total RNA were reverse transcribed to cDNA using M-MLV reverse transcriptase (Promega). Gene-specific primers were designed to detect an amplicon of each target gene of ∼150 bp. Each PCR reaction was performed with 10 ng cDNA using Power SYBR Green Master Mix (Life Technologies) and 250 nM of each primer. Primer sets used in this study are listed in Supplementary Table 2. qPCR was carried out for three biological replicates with measurements taken from three technical replicates using a ABI7300 real-time PCR machine (Life Technologies). The relative expression of each gene was determined after being normalized to zebrafish elongation factor 1a (ef1a). Data in bars represent mean±s.d. Statistical analyses were performed using unpaired two-tailed Student’s t-test (GraphPad Prism software) and P values <0.001 were considered as significant.
For the detection of apoptosis, oloca sibling and mutant embryos in various transgenic backgrounds were fixed at 3 d.p.f. in 4% methanol-free formaldehyde overnight then kept in methanol at −20 °C for at least 2 h. TUNEL staining was performed using ApopTag Red or Fluorescein In Situ Apoptosis Detection Kit (Chemicon) combined with peroxidase (POD) coupled anti-DIG antibody (Roche #11207733910, 1:200) followed by tyramide-based amplification9. To visualize the transgenic reporter proteins, rabbit anti-GFP (MBL #598, 1:300) or rabbit anti-DsRed/mCherry (Clontech #63296, 1:300) antibodies were co-incubated with the POD-coupled anti-DIG antibody, then revealed by anti-rabbit-AlexaFluor 488 (Life Technologies; #A11070, 1:200) and anti-rabbit-Cy3 (Jackson Lab; #711166152, 1:300) coupled antibodies, except for the [TCF:nls-mCherry] larvae, which were subjected to tyramide-based detection of POD-coupled anti-rabbit antibodies (Life Technologies #G21234, 1:300).
Additional information
How to cite this article: Murayama, E. et al. NACA deficiency reveals the crucial role of somite-derived stromal cells in haematopoietic niche formation. Nat. Commun. 6:8375 doi: 10.1038/ncomms9375 (2015).
Accession codes
Change history
29 September 2015
The HTML version of this paper was updated shortly after publication, following a technical error that resulted in the Supplementary Movie legends being omitted. This has now been corrected in the HTML; the PDF version of the paper was correct from the time of publication.
References
Sugiyama, T., Kohara, H., Noda, M. & Nagasawa, T. Maintenance of the hematopoietic stem cell pool by CXCL12-CXCR4 chemokine signaling in bone marrow stromal cell niches. Immunity 25, 977–988 (2006).
Mendez-Ferrer, S. et al. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature 466, 829–834 (2010).
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Kunisaki, Y. et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature 502, 637–643 (2013).
Kissa, K. & Herbomel, P. Blood stem cells emerge from aortic endothelium by a novel type of cell transition. Nature 464, 112–115 (2010).
Murayama, E. et al. Tracing hematopoietic precursor migration to successive hematopoietic organs during zebrafish development. Immunity 25, 963–975 (2006).
Huttenhuis, H. B. et al. Ontogeny of the common carp (Cyprinus carpio L.) innate immune system. Dev. Comp. Immunol. 30, 557–574 (2006).
Charbord, P. et al. Comparative study of stromal cell lines derived from embryonic, fetal, and postnatal mouse blood-forming tissues. Exp. Hematol. 30, 1202–1210 (2002).
Le Guyader, D. et al. Origins and unconventional behavior of neutrophils in developing zebrafish. Blood 111, 132–141 (2008).
Amsterdam, A. et al. A large-scale insertional mutagenesis screen in zebrafish. Genes Dev. 13, 2713–2724 (1999).
Yotov, W. V. & St-Arnaud, R. Mapping of the human gene for the alpha-NAC/1.9.2 (NACA/1.9.2) transcriptional coactivator to chromosome 12q23-24.1. Mamm. Genome 7, 163–164 (1996).
Lauring, B., Sakai, H., Kreibich, G. & Wiedmann, M. Nascent polypeptide-associated complex protein prevents mistargeting of nascent chains to the endoplasmic reticulum. Proc. Natl Acad. Sci. USA 92, 5411–5415 (1995).
Kirstein-Miles, J., Scior, A., Deuerling, E. & Morimoto, R. I. The nascent polypeptide-associated complex is a key regulator of proteostasis. EMBO J. 32, 1451–1468 (2013).
Yotov, W. V. & St-Arnaud, R. Differential splicing-in of a proline-rich exon converts alphaNAC into a muscle-specific transcription factor. Genes Dev. 10, 1763–1772 (1996).
Li, H., Randall, W. R. & Du, S. J. skNAC (skeletal Naca), a muscle-specific isoform of Naca (nascent polypeptide-associated complex alpha), is required for myofibril organization. FASEB J. 23, 1988–2000 (2009).
Moreau, A., Yotov, W. V., Glorieux, F. H. & St-Arnaud, R. Bone-specific expression of the alpha chain of the nascent polypeptide-associated complex, a coactivator potentiating c-Jun-mediated transcription. Mol. Cell. Biol. 18, 1312–1321 (1998).
Yotov, W. V., Moreau, A. & St-Arnaud, R. The alpha chain of the nascent polypeptide-associated complex functions as a transcriptional coactivator. Mol. Cell. Biol. 18, 1303–1311 (1998).
Kissa, K. et al. Live imaging of emerging hematopoietic stem cells and early thymus colonization. Blood 111, 1147–1156 (2008).
Demy, D. L. et al. Generating parabiotic zebrafish embryos for cell migration and homing studies. Nat. Methods 10, 256–258 (2013).
Moro, E. et al. In vivo Wnt signaling tracing through a transgenic biosensor fish reveals novel activity domains. Dev. Biol. 366, 327–340 (2012).
Cortes, F. et al. Cadherin-mediated differential cell adhesion controls slow muscle cell migration in the developing zebrafish myotome. Dev. Cell 5, 865–876 (2003).
Cooper, M. S., D'Amico, L. A. & Henry, C. A. Confocal microscopic analysis of morphogenetic movements. Methods Cell Biol. 59, 179–204 (1999).
Carney, T. J. et al. A direct role for Sox10 in specification of neural crest-derived sensory neurons. Development 133, 4619–4630 (2006).
Pauls, S., Geldmacher-Voss, B. & Campos-Ortega, J. A. A zebrafish histone variant H2A.F/Z and a transgenic H2A.F/Z:GFP fusion protein for in vivo studies of embryonic development. Dev. Genes Evol. 211, 603–610 (2001).
Hotokezaka, Y. et al. alphaNAC depletion as an initiator of ER stress-induced apoptosis in hypoxia. Cell Death Differ. 16, 1505–1514 (2009).
Chagraoui, J., Lepage-Noll, A., Anjo, A., Uzan, G. & Charbord, P. Fetal liver stroma consists of cells in epithelial-to-mesenchymal transition. Blood 101, 2973–2982 (2003).
Nguyen, P. D. et al. Haematopoietic stem cell induction by somite-derived endothelial cells controlled by meox1. Nature 512, 314–318 (2014).
Isern, J. et al. The neural crest is a source of mesenchymal stem cells with specialized hematopoietic stem cell niche function. Elife 3, e03696 (2014).
Arsenovic, P. T., Maldonado, A. T., Colleluori, V. D. & Bloss, T. A. Depletion of the C. elegans NAC engages the unfolded protein response, resulting in increased chaperone expression and apoptosis. PloS ONE 7, e44038 (2012).
Feng, Y. X. et al. Epithelial-to-mesenchymal transition activates PERK-eIF2alpha and sensitizes cells to endoplasmic reticulum stress. Cancer Discov. 4, 702–715 (2014).
Willett, C. E., Cortes, A., Zuasti, A. & Zapata, A. G. Early hematopoiesis and developing lymphoid organs in the zebrafish. Dev. Dyn. 214, 323–336 (1999).
Westerfield, M. The Zebrafish Book: A Guide for yhr Laboratory use of Zebrafish (Danio rerio) University of Oregon Press (1995).
Lawson, N. D. & Weinstein, B. M. In vivo imaging of embryonic vascular development using transgenic zebrafish. Dev. Biol. 248, 307–318 (2002).
Renshaw, S. A. et al. A transgenic zebrafish model of neutrophilic inflammation. Blood 108, 3976–3978 (2006).
Hall, C., Flores, M. V., Storm, T., Crosier, K. & Crosier, P. The zebrafish lysozyme C promoter drives myeloid-specific expression in transgenic fish. BMC Dev. Biol. 7, 42 (2007).
Zhu, H. et al. Regulation of the lmo2 promoter during hematopoietic and vascular development in zebrafish. Dev. Biol. 281, 256–269 (2005).
Traver, D. et al. Transplantation and in vivo imaging of multilineage engraftment in zebrafish bloodless mutants. Nat. Immunol. 4, 1238–1246 (2003).
Lin, H. F. et al. Analysis of thrombocyte development in CD41-GFP transgenic zebrafish. Blood 106, 3803–3810 (2005).
Bajoghli, B. et al. Evolution of genetic networks underlying the emergence of thymopoiesis in vertebrates. Cell 138, 186–197 (2009).
Dutton, K. A. et al. Zebrafish colourless encodes sox10 and specifies non-ectomesenchymal neural crest fates. Development 128, 4113–4125 (2001).
Thompson, M. A. et al. The cloche and spadetail genes differentially affect hematopoiesis and vasculogenesis. Dev. Biol. 197, 248–269 (1998).
Detrich, H. W. 3rd et al. Intraembryonic hematopoietic cell migration during vertebrate development. Proc. Natl Acad. Sci. USA 92, 10713–10717 (1995).
Quinkertz, A. & Campos-Ortega, J. A. A new beta-globin gene from the zebrafish, betaE1, and its pattern of transcription during embryogenesis. Dev. Genes Evol. 209, 126–131 (1999).
Lyons, S. E. et al. Molecular cloning, genetic mapping, and expression analysis of four zebrafish c/ebp genes. Gene 281, 43–51 (2001).
Willett, C. E., Kawasaki, H., Amemiya, C. T., Lin, S. & Steiner, L. A. Ikaros expression as a marker for lymphoid progenitors during zebrafish development. Dev. Dyn. 222, 694–698 (2001).
Langenau, D. M. et al. In vivo tracking of T cell development, ablation, and engraftment in transgenic zebrafish. Proc. Natl Acad. Sci. USA 101, 7369–7374 (2004).
Liu, F. & Wen, Z. Cloning and expression pattern of the lysozyme C gene in zebrafish. Mech. Dev. 113, 69–72 (2002).
Parichy, D. M., Ransom, D. G., Paw, B., Zon, L. I. & Johnson, S. L. An orthologue of the kit-related gene fms is required for development of neural crest-derived xanthophores and a subpopulation of adult melanocytes in the zebrafish, Danio rerio. Development 127, 3031–3044 (2000).
LeBlanc, J., Bowman, T. V. & Zon, L. Transplantation of whole kidney marrow in adult zebrafish. J. Vis. Exp. 159, doi:10.3791/159 (2007).
Stachura, D. L. et al. Zebrafish kidney stromal cell lines support multilineage hematopoiesis. Blood 114, 279–289 (2009).
Stachura, D. L. et al. Clonal analysis of hematopoietic progenitor cells in the zebrafish. Blood 118, 1274–1282 (2011).
Sarris, M. et al. Inflammatory chemokines direct and restrict leukocyte migration within live tissues as glycan-bound gradients. Curr. Biol. 22, 2375–2382 (2012).
Soroldoni, D., Hogan, B. M. & Oates, A. C. Simple and efficient transgenesis with meganuclease constructs in zebrafish. Methods Mol. Biol. 546, 117–130 (2009).
Prudent, J. et al. Bcl-wav and the mitochondrial calcium uniporter drive gastrula morphogenesis in zebrafish. Nat. Commun. 4, 2330 (2013).
Acknowledgements
The oloca mutant was identified through a large-scale genetic screen carried out at Max-Planck Institute in Tübingen, within the EU-funded ‘ZF-Models’ Integrated Project. We thank E. Moro and F. Argenton (University of Padova) for providing TCF:nls-mCherry and TCF:eGFP transgenic lines, and R. Kelsh (University of Bath) for providing the sox10m618 mutant in Tg[sox10:gfp] background. We thank D. Soroldoni, A. Oates (University College London) and G. Gillet (University of Lyon) for providing the pαact-GFPI2 plasmid and pCS2+flag-zBcl-XL plasmid, respectively. We thank Mylène Lancino for her help to analyse three-dimensional data, and Ana Cumano, Anne Schmidt and Estelle Hirsinger for their critical reading of our manuscript. This work was supported by grants to P.H. from the European Commission through the FP6 ‘ZF-Models’ Integrated Project and E-RARE ‘Neutro-Net’ project, from the Fondation pour la Recherche Médicale (FRM 2012 team, #DEQ20120323714), and from the Laboratoire d'Excellence Revive (Investissement d'Avenir; ANR-10-LABX-73).
Author information
Authors and Affiliations
Contributions
E.M. designed and performed experiments. M.R. and M.S. performed mutation mapping/cloning and WKM cell transplantation experiments, respectively. D.LG., C.V. and W.H. performed experiments. N.T. assisted with helpful discussion. E.M. and P.H. analysed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-7 and Supplementary Tables 1-2 (PDF 1753 kb)
Supplementary Movie 1
Time-lapse imaging of transplanted adult mpx:GFP+ neutrophils in sibling and oloca mutant larva. oloca sibling and mutant embryos were transplanted at 48 hpf with sorted mpx:GFP+ neutrophils obtained from the WKM of adult Tg[mpx:gfp] fish. Larvae were imaged every 5 min. from 72 hpf for 15 hrs. Rostral is to the upper left. Note that the transplanted mpx:GFP+ cells are stably settled in the sibling CHT whereas those visible in the mutant wander all over the caudal area. (MOV 15840 kb)
Supplementary Movie 2
TCF:GFP+ ventral somite cells emigrate into the presumptive CHT. 4D Time-lapse confocal imaging of caudal somites in a Tg[TCF:gfp] embryo imaged every 5 min between 21 and 24 hpf (24- to 30-somites). The trajectories of cells emigrating from the ventral border of the somites were automatically tracked by Imaris software. See also Supplementary Fig. 3a. (MOV 5511 kb)
Supplementary Movie 3
Simultaneous migration of stromal progenitors and CV endothelial cells. Time-lapse confocal sequence of developing stromal progenitors (red) and endothelial cells (green) in a [TCF:nls-mCherry; Fli1:gfp] double transgenic embryo imaged every 8 min. between 22 and 24.5 hpf. Arrows point at the nuclei of migrating TCF:nls-mCherry+ stromal progenitors. som, somite; UGO, uro-genital opening. (MOV 1047 kb)
Supplementary Movie 4
Stromal progenitors emerge from the ventral border of somites. 4D nuclear tracking analysis in a Tg[TCF:nls-mCherry] embryo vitally stained with Bodipy ceramide. The first two seconds of the movie, to be viewed frame by frame, illustrate by going through the confocal planes how each of the tracked TCF:nls-mCherry+ nuclei can be ascribed to a particular Bodipy ceramide delineated somite cell at T=0 (22 hpf); 3D rotations follow to show the 3D position of the 9 selected nuclei within the somite; then the tracking of their emigration towards the future CHT is shown. The embryo was imaged for 5 hrs every 6 min. See also Supplementary Fig. 3c-e. In the lowermost and uppermost part of the movie, the Bodipy ceramide intracellular staining partially leaking into the red channel corresponds to the perinuclear endosomal staining of cells at the body surface (periderm) that endocytosed Bodipy ceramide from the bath. (MOV 17457 kb)
Supplementary Movie 5
Active blebbing at the ventral border of caudal somites. Time-lapse VE-DIC imaging of the ventral border of caudal somites showing active blebbing through the somite basal lamina (arrows). The embryo was imaged every 10 sec. for 1 hr from 18.5 hpf (19-somites). som, somite; pe, primitive proerythroblast; pm, primitive myeloid cell. (MOV 20228 kb)
Supplementary Movie 6
High-resolution VE-DIC imaging of ventral somite cells mitoses and emigration into the presumptive CHT. The embryo was imaged every 20 sec. for 4 hrs from 25/26-som stage. Red and cyan asterisks show the nuclear position of two emigrating cells, whose pseudopodia are pointed by red and cyan arrows, respectively; then a green asterisk indicates another somitic cell that follows the emigration of its neighbor (red asterisk). Mitotic cells and their daughter cells are indicated by black arrows and asterisks, respectively; yellow and blue asterisks indicate two sister cells that start emigrating just after the mitosis, with a blue arrow showing the leading edge of the blue asterisk labelled cell. som, somite; endo, endothelial cells. (MOV 28231 kb)
Supplementary Movie 7
Dynamic network of stromal reticular cells in the CHT from 48 hpf. In this double transgenic Tg[TCF:GFP; TCF:nls-mCherry] embryo, the SRCs are labelled with GFP (green), and their nucleus additionally with nls-mCherry (red), in a variable green-to-red ratio. Arrowheads point at some of the dynamic filopodia that interconnect these cells to each other; arrows point at some SRCs that undergo mitosis. The SRCs aligned horizontally at the bottom of the image line the endothelial wall of the definitive CV. The embryo was imaged every 8 min. for 10 hrs from 48 hpf. som, somites. (MOV 13913 kb)
Supplementary Movie 8
Enhanced contraction of the CV plexus past 48 hpf in naca MO injected embryos. Time-lapse imaging of Tg[Fli1:gfp] embryos injected with control (left) or naca (right) MO. Rostral is to the upper left. Embryos were imaged every 10 min. from 48 hpf for 15 hrs. Asterisks indicate the larger intervascular spaces observed in naca morphants at 48 hpf, which start shrinking by 5 hrs of recording (arrows). (MOV 7749 kb)
Rights and permissions
About this article
Cite this article
Murayama, E., Sarris, M., Redd, M. et al. NACA deficiency reveals the crucial role of somite-derived stromal cells in haematopoietic niche formation. Nat Commun 6, 8375 (2015). https://doi.org/10.1038/ncomms9375
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms9375
This article is cited by
-
Exploring hematopoiesis in zebrafish using forward genetic screening
Experimental & Molecular Medicine (2024)
-
Alcam-a and Pdgfr-α are essential for the development of sclerotome-derived stromal cells that support hematopoiesis
Nature Communications (2023)
-
BTF3 sustains cancer stem-like phenotype of prostate cancer via stabilization of BMI1
Journal of Experimental & Clinical Cancer Research (2019)
-
VCAM-1+ macrophages guide the homing of HSPCs to a vascular niche
Nature (2018)
-
Systemic inoculation of Escherichia coli causes emergency myelopoiesis in zebrafish larval caudal hematopoietic tissue
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.