Abstract
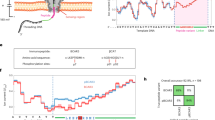
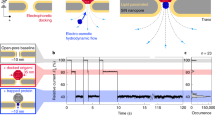
We demonstrate single-molecule, site-specific detection of protein phosphorylation with protein nanopore technology. A model protein, thioredoxin, was phosphorylated at two adjacent sites. Analysis of the ionic current amplitude and noise, as the protein unfolds and moves through an α-hemolysin pore, enables the distinction between unphosphorylated, monophosphorylated and diphosphorylated variants. Our results provide a step toward nanopore proteomics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Prabakaran, S., Lippens, G., Steen, H. & Gunawardena, J. Wiley Interdiscip. Rev. Syst. Biol. Med 4, 565–583 (2012).
Khoury, G.A., Baliban, R.C. & Floudas, C.A. Sci. Rep. 1, 90 (2011).
Choudhary, C. & Mann, M. Nat. Rev. Mol. Cell Biol. 11, 427–439 (2010).
Howorka, S. & Siwy, Z. Chem. Soc. Rev. 38, 2360–2384 (2009).
Pennisi, E. Science 336, 534–537 (2012).
Cherf, G.M. et al. Nat. Biotechnol. 30, 344–348 (2012).
Manrao, E.A. et al. Nat. Biotechnol. 30, 349–353 (2012).
Rodriguez-Larrea, D. & Bayley, H. Nat. Nanotechnol. 8, 288–295 (2013).
Nivala, J., Marks, D.B. & Akeson, M. Nat. Biotechnol. 31, 247–250 (2013).
Wallace, E.V. et al. Chem. Commun. (Camb.) 46, 8195–8197 (2010).
Tam, J.P., Yu, Q. & Miao, Z. Biopolymers 51, 311–332 (1999).
Hinner, M.J. & Johnsson, K. Curr. Opin. Biotechnol. 21, 766–776 (2010).
Howorka, S., Cheley, S. & Bayley, H. Nat. Biotechnol. 19, 636–639 (2001).
Witus, L.S. et al. J. Am. Chem. Soc. 132, 16812–16817 (2010).
Acknowledgements
We thank E. Mikhailova for the preparation of α-hemolysin pores. We thank L. Harrington for the PKA-CAT plasmid and the purification protocol for the catalytic subunit of PKA. This work has been supported by the National Institutes of Health, Oxford Nanopore Technologies and an European Research Council Advanced Grant. C.B.R. is funded by the Danish National Research Foundation (grant number DNRF78) and Aarhus University, Faculty of Science and Technology.
Author information
Authors and Affiliations
Contributions
C.B.R. performed experiments, analyzed data and wrote the paper. D.R.-L. planned the research, performed experiments, analyzed data and wrote the paper. H.B. planned the research and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
H.B. is the founder, and a director and shareholder of Oxford Nanopore Technologies, a company engaged in the development of nanopore sequencing technology. Work in the Bayley laboratory at the University of Oxford is supported in part by Oxford Nanopore Technologies.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Table 1 (PDF 1704 kb)
Rights and permissions
About this article
Cite this article
Rosen, C., Rodriguez-Larrea, D. & Bayley, H. Single-molecule site-specific detection of protein phosphorylation with a nanopore. Nat Biotechnol 32, 179–181 (2014). https://doi.org/10.1038/nbt.2799
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2799
This article is cited by
-
Peptide sequencing based on host–guest interaction-assisted nanopore sensing
Nature Methods (2024)
-
Unambiguous discrimination of all 20 proteinogenic amino acids and their modifications by nanopore
Nature Methods (2024)
-
MoS2 nanopore identifies single amino acids with sub-1 Dalton resolution
Nature Communications (2023)
-
Detection of phosphorylation post-translational modifications along single peptides with nanopores
Nature Biotechnology (2023)
-
Enzyme-less nanopore detection of post-translational modifications within long polypeptides
Nature Nanotechnology (2023)