Abstract
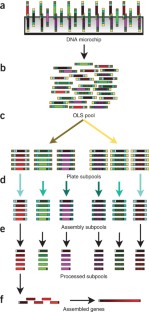
Development of cheap, high-throughput and reliable gene synthesis methods will broadly stimulate progress in biology and biotechnology1. Currently, the reliance on column-synthesized oligonucleotides as a source of DNA limits further cost reductions in gene synthesis2. Oligonucleotides from DNA microchips can reduce costs by at least an order of magnitude3,4,5, yet efforts to scale their use have been largely unsuccessful owing to the high error rates and complexity of the oligonucleotide mixtures. Here we use high-fidelity DNA microchips, selective oligonucleotide pool amplification, optimized gene assembly protocols and enzymatic error correction to develop a method for highly parallel gene synthesis. We tested our approach by assembling 47 genes, including 42 challenging therapeutic antibody sequences, encoding a total of ∼35 kilobase pairs of DNA. These assemblies were performed from a complex background containing 13,000 oligonucleotides encoding ∼2.5 megabases of DNA, which is at least 50 times larger than in previously published attempts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Carr, P.A. & Church, G.M. Genome engineering. Nat. Biotechnol. 27, 1151–1162 (2009).
Tian, J., Ma, K. & Saaem, I. Advancing high-throughput gene synthesis technology. Mol. Biosyst. 5, 714–722 (2009).
Tian, J. et al. Accurate multiplex gene synthesis from programmable DNA microchips. Nature 432, 1050–1054 (2004).
Richmond, K.E. et al. Amplification and assembly of chip-eluted DNA (AACED): a method for high-throughput gene synthesis. Nucleic Acids Res. 32, 5011–5018 (2004).
Zhou, X. et al. Microfluidic PicoArray synthesis of oligodeoxynucleotides and simultaneous assembling of multiple DNA sequences. Nucleic Acids Res. 32, 5409–5417 (2004).
Nirenberg, M.W. & Matthaei, J.H. The dependence of cell-free protein synthesis in E. coli upon naturally occurring or synthetic polyribonucleotides. Proc. Natl. Acad. Sci. USA 47, 1588–1602 (1961).
Söll, D. et al. Studies on polynucleotides, XLIX. Stimulation of the binding of aminoacyl-sRNA's to ribosomes by ribotrinucleotides and a survey of codon assignments for 20 amino acids. Proc. Natl. Acad. Sci. USA 54, 1378–1385 (1965).
Gibson, D.G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010).
Gibson, D.G. Synthesis of DNA fragments in yeast by one-step assembly of overlapping oligonucleotides. Nucleic Acids Res. 37, 6984–6990 (2009).
Li, M.Z. & Elledge, S.J. Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nat. Methods 4, 251–256 (2007).
Bang, D. & Church, G.M. Gene synthesis by circular assembly amplification. Nat. Methods 5, 37–39 (2008).
Shao, Z., Zhao, H. & Zhao, H. DNA assembler, an in vivo genetic method for rapid construction of biochemical pathways. Nucleic Acids Res. 37, e16 (2009).
Lee, C.-C., Snyder, T.M. & Quake, S.R. A microfluidic oligonucleotide synthesizer. Nucleic Acids Res. 38, 2514–2521 (2010).
Kim, C. et al. Progress in gene assembly from a MAS-driven DNA microarray. Microelectron. Eng. 83, 1613–1616 (2006).
LeProust, E.M. et al. Synthesis of high-quality libraries of long (150mer) oligonucleotides by a novel depurination controlled process. Nucleic Acids Res. 38, 2522–2540 (2010).
Patwardhan, R.P. et al. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis. Nat. Biotechnol. 27, 1173–1175 (2009).
Schlabach, M.R. et al. Synthetic design of strong promoters. Proc. Natl. Acad. Sci. USA 107, 2538–2543 (2010).
Li, J.B. et al. Multiplex padlock targeted sequencing reveals human hypermutable CpG variations. Genome Res. 19, 1606–1615 (2009).
Li, J.B. et al. Genome-wide identification of human RNA editing sites by parallel DNA capturing and sequencing. Science 324, 1210–1213 (2009).
Porreca, G.J. et al. Multiplex amplification of large sets of human exons. Nat. Methods 4, 931–936 (2007).
Xu, Q. et al. Design of 240,000 orthogonal 25mer DNA barcode probes. Proc. Natl. Acad. Sci. USA 106, 2289–2294 (2009).
Huston, J.S. et al. Medical applications of single-chain antibodies. Int. Rev. Immunol. 10, 195–217 (1993).
Carr, P.A. et al. Protein-mediated error correction for de novo DNA synthesis. Nucleic Acids Res. 32, e162 (2004).
Slater, G.S. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005).
Li, H. Maq: mapping and assembly with qualities. Welcome Trust Sanger Institute (2010). Available at: <http://maq.sourceforge.net>.
Cormack, B.P., Valdivia, R.H. & Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173, 33–38 (1996).
Acknowledgements
This work was supported by the US Office of Naval Research (N000141010144), National Human Genome Research Institute Center for Excellence in Genomics Science (P50 HG003170), Department of Energy Genomes to Life (DE-FG02-02ER63445), Defense Advanced Research Projects Agency (W911NF-08-1-0254) and the Wyss Institute for Biologically Inspired Engineering (all to G.M.C.). We thank H. Padgett for providing ErrASE and expertise during optimization and J. Boeke for advice on gene assembly protocols. We also thank S. Raman, F. Vigneault and F. Zhang for critical readings of the manuscript, G. Dantas for pZE21 (Washington University), F. Isaacs (Yale University) for pZE21G and J.S. Workman (Wyss Institute) for pSecTag2A.
Author information
Authors and Affiliations
Contributions
S.K. and N.E. wrote the paper with contributions from all authors; S.K. and G.M.C. conceived the study; S.K. wrote all algorithms and designed all sequences; S.K. and N.E. designed and performed all experiments; E.L. provided the oligonucleotides libraries; M.S. and J.F. designed the single-chained versions of commercial antibodies; J.B.L. performed the OLS high-throughput sequencing experiment and provided critical advice on the processing of subpools.
Corresponding authors
Ethics declarations
Competing interests
E.M.L. is an employee of Agilent Technologies, the commercial provider of OLS pools. G.M.C. is a co-founder of an early-stage startup company involved in gene synthesis. S.K., N.E. and G.M.C. are named inventors on a patent application on technologies described in this article. S.K. is a post-doctoral fellow whose future employment prospects depend upon refereed publications.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–3, Supplementary Figs. 1–11, Supplementary Notes 1, 2 and Supplementary Methods (PDF 4391 kb)
Rights and permissions
About this article
Cite this article
Kosuri, S., Eroshenko, N., LeProust, E. et al. Scalable gene synthesis by selective amplification of DNA pools from high-fidelity microchips. Nat Biotechnol 28, 1295–1299 (2010). https://doi.org/10.1038/nbt.1716
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1716
This article is cited by
-
DIMPLE: deep insertion, deletion, and missense mutation libraries for exploring protein variation in evolution, disease, and biology
Genome Biology (2023)
-
A multiplexed bacterial two-hybrid for rapid characterization of protein–protein interactions and iterative protein design
Nature Communications (2023)
-
DNA synthesis technologies to close the gene writing gap
Nature Reviews Chemistry (2023)
-
Magnetic DNA random access memory with nanopore readouts and exponentially-scaled combinatorial addressing
Scientific Reports (2023)
-
Purification of multiplex oligonucleotide libraries by synthesis and selection
Nature Biotechnology (2022)