Abstract
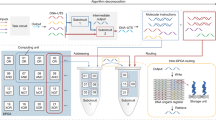
We describe a structure-specific cleavage–based READOUT strategy for surface-based DNA computing. The strategy was demonstrated in the solution of a 4-variable/3-satisfiability (SAT) problem. The READOUT step identifies the DNA molecules present at the end of the computational process. The specificity of the sequence detection used here derives from the sequence specificity of DNA hybridization coupled with the structure specificity of the enzymatic cleavage. The process is linear, yielding a higher uniformity of detection of the DNA computing products compared to that obtained with PCR amplification. The structure-specific cleavage–based readout is simple, accurate, and compatible with multiple-word DNA computing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Adleman, L.M. Molecular computation of solutions to combinatorial problems. Science 266, 1021–1024 (1994).
Lipton, R.J. DNA solution of hard computational problems. Science 268, 542–545 (1995).
Guarnieri, F., Fliss, M. & Bancroft, C. Making DNA add. Science 273, 220–223 (1996).
Ouyang, Q., Kaplan, P.D., Liu, S. & Libchaber, A. DNA solution of the maximal clique problem. Science 278, 446–449 (1997).
Winfree, E., Liu, F., Wenzler, L.A. & Seeman, N.C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Morimoto, N., Arita M. & Suyama, A. Solid phase DNA solution to the Hamiltonian Path problem. Proc. DIMACS: DNA Based Computers (III), 83–92 (1997).
Yoshida, H. & Suyama, A. Solution to 3SAT by breadth first search . Preliminary Proc. DIMACS: DNA Based Computers (V), 9–20 (1999).
Faulhammer, D., Cukras, A.R., Lipton, R.J. & Landweber, L.F. Molecular computation: RNA solutions to chess problems. Proc. Natl. Acad. Sci. USA 97, 1385–1389 (2000).
Mao, C.D., LaBean, T.H., Reif, J.H. & Seeman, N.C. Logical computation using algorithmic self-assembly of DNA triple-crossover molecules. Nature 407, 493–496 (2000).
Liu, Q. et al. DNA computing on surfaces. Nature 403, 175–179 (2000).
Saiki, R.K. et al. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239, 487–491 (1988).
Erlich, G.D. PCR based diagnostics in infectious disease. (eds Erlich, G.D. & Greenberg, S.J.) 3–18 (Blackwell Scientific, Oxford; 1994).
Kwok, S. & Higuchi, R. Avoiding false positives with PCR. Nature 339, 237–238 (1989).
Farrell, R.E. DNA amplification. Immunol. Invest. 26, 3–7 (1997).
Vaneechoutte, M. & Van Eldere, J. The possibilities and limitations of nucleic acid amplification technology in diagnostic microbiology. J. Med. Microbiol. 46, 188–194 (1997).
Barnard, R., Futo, V., Pecheniuk, N., Slattery, M. & Walsh, T. PCR bias toward the wild-type k-ras and p53 sequences: implications for PCR detection of mutations and cancer diagnosis. Biotechniques 25, 684–691 (1998).
Lyamichev, V. et al. Polymorphism identification and quantitative detection of genomic DNA by invasive cleavage of oligonucleotide probes. Nat. Biotechnol. 17, 292–296 (1999).
Jeff, H.G. et al. Sensitive detection of DNA polymorphisms by the serial invasive signal amplification reaction. Proc. Natl. Acad. Sci. USA 97, 8272–8277 (2000).
Frutos, A.G. et al. Demonstration of a word design strategy for DNA computing on surfaces. Nucleic Acids Res. 25, 4748–4757 (1997).
Garey, M.R. & Johnson, D.S. Computers and intractability: a guide to the theory of NP-completeness. (Freeman, New York, NY; 1979).
Matayoshi, E.D., Wang, G.T., Krafft, G.A. & Erickson, J. Novel fluorogenic substrates for assaying retroviral proteases by resonance energy transfer. Science 247, 954–958 (1990).
Wang, G.T., Matayoshi, E., Huffaker, H.J. & Krafft, G.A. Design and synthesis of new fluorogenic HIV protease substrates based on resonance energy-transfer. Tetrahedron Lett. 31, 6493–6496 (1990).
Allawi, H.T. & SantaLucia, J. Thermodynamics and NMR of internal GT mismatches in DNA. Biochemistry 36, 10581–10594 (1997).
Lyamichev, V.I. et al. Experimental and theoretical analysis of the invasive signal amplification reaction. Biochemistry 39, 9523–9532 (2000).
Kaiser, M.W. et al. A comparison of eubacterial and archaeal structure-specific 5′-exonuclease. J. Biol. Chem. 274, 21387–21394 (1999).
Jordan, C.E., Frutos, A.G., Thiel, A.J. & Corn, R.M. Surface plasmon resonance imaging measurements of DNA hybridization adsorption and streptavidin/DNA multilayer formation at chemically modified gold surfaces. Anal. Chem. 69, 4939–4947 (1997).
Griffin, T.J., Hall, J.G., Prudent, J.R. & Smith, L.M. Direct genetic analysis by matrix-assisted laser desorption/ionization mass spectrometry. Proc. Natl. Acad. Sci. USA 96, 6301–6306 (1999).
Ross, P., Hall, L., Smirnov, I. & Haff, L. High level multiplex genotyping by MALDI-TOF mass spectrometry. Nat. Biotechnol. 16, 1347–1351 (1998).
Mein, C.A. et al. Evaluation of single nucleotide polymorphism typing with invader on PCR amplicons and its automation. Genome Res. 10, 330–343 (2000).
Smith, L.M. Fluorescence detection in automated DNA-sequence analysis. Nature 321, 674–679 (1986).
Eisen, M.B. & Brown, P.O. DNA arrays for analysis of gene expression. Methods Enzymol. 303, 179–205 (1999).
Brown, P.O. & Botstein, D. Exploring the new world of the genome with DNA microarrays. Nat. Genet. 21 (Suppl. 1), 33–37 (1999).
Acknowledgements
The authors would like to thank Professor Robert Corn, Dr. Victor I. Lyamichev, and Dr. Bruce P. Neri for useful discussions. This work was supported by the National Science Foundation, no. 98-SC-NSF-1003/subcontract through Duke University, GenTel, Inc., and Third Wave Technologies, Inc. L.M.S. has a financial interest in both GenTel, Inc. and Third Wave Technologies, Inc.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, L., Hall, J., Lu, M. et al. A DNA computing readout operation based on structure-specific cleavage. Nat Biotechnol 19, 1053–1059 (2001). https://doi.org/10.1038/nbt1101-1053
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt1101-1053
This article is cited by
-
DNA multi-bit non-volatile memory and bit-shifting operations using addressable electrode arrays and electric field-induced hybridization
Nature Communications (2018)
-
Experimental analysis of the basic idea on the transcription-based diagnostic automata controlled by programmed molecules
Natural Computing (2008)
-
Linearly programmed DNA-based molecular computer operated on magnetic particle surface in test-tube
Chinese Science Bulletin (2004)
-
Genetic algorithm in DNA computing: A solution to the maximal clique problem
Chinese Science Bulletin (2004)