Abstract
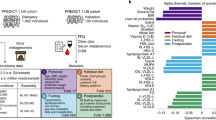
Type 2 diabetes (T2D) is a result of complex gene–environment interactions, and several risk factors have been identified, including age, family history, diet, sedentary lifestyle and obesity. Statistical models that combine known risk factors for T2D can partly identify individuals at high risk of developing the disease. However, these studies have so far indicated that human genetics contributes little to the models, whereas socio-demographic and environmental factors have greater influence1. Recent evidence suggests the importance of the gut microbiota as an environmental factor, and an altered gut microbiota has been linked to metabolic diseases including obesity2,3, diabetes4 and cardiovascular disease5. Here we use shotgun sequencing to characterize the faecal metagenome of 145 European women with normal, impaired or diabetic glucose control. We observe compositional and functional alterations in the metagenomes of women with T2D, and develop a mathematical model based on metagenomic profiles that identified T2D with high accuracy. We applied this model to women with impaired glucose tolerance, and show that it can identify women who have a diabetes-like metabolism. Furthermore, glucose control and medication were unlikely to have major confounding effects. We also applied our model to a recently described Chinese cohort4 and show that the discriminant metagenomic markers for T2D differ between the European and Chinese cohorts. Therefore, metagenomic predictive tools for T2D should be specific for the age and geographical location of the populations studied.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Noble, D., Mathur, R., Dent, T., Meads, C. & Greenhalgh, T. Risk models and scores for type 2 diabetes: systematic review. BMJ 343, d7163 (2011)
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006)
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009)
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012)
Karlsson, F. H. et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nature Commun. 3, 1245 (2012)
Mueller, S. et al. Differences in fecal microbiota in different European study populations in relation to age, gender, and country: a cross-sectional study. Appl. Environ. Microbiol. 72, 1027–1033 (2006)
Biagi, E. et al. Through ageing, and beyond: gut microbiota and inflammatory status in seniors and centenarians. PLoS ONE 5, e10667 (2010)
Claesson, M. J. et al. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc. Natl Acad. Sci. USA 108, 4586–4591 (2011)
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012)
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl Acad. Sci. USA 107, 14691–14696 (2010)
Brohall, G., Behre, C. J., Hulthe, J., Wikstrand, J. & Fagerberg, B. Prevalence of diabetes and impaired glucose tolerance in 64-year-old Swedish women: experiences of using repeated oral glucose tolerance tests. Diabetes Care 29, 363–367 (2006)
Fagerberg, B., Kellis, D., Bergstrom, G. & Behre, C. J. Adiponectin in relation to insulin sensitivity and insulin secretion in the development of type 2 diabetes: a prospective study in 64-year-old women. J. Intern. Med. 269, 636–643 (2011)
Salonen, A. et al. Comparative analysis of fecal DNA extraction methods with phylogenetic microarray: effective recovery of bacterial and archaeal DNA using mechanical cell lysis. J. Microbiol. Methods 81, 127–134 (2010)
Gaetti-Jardim, E., Jr, Marcelino, S. L., Feitosa, A. C., Romito, G. A. & Avila-Campos, M. J. Quantitative detection of periodontopathic bacteria in atherosclerotic plaques from coronary arteries. J. Med. Microbiol. 58, 1568–1575 (2009)
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010)
Wang, Y., Rimm, E. B., Stampfer, M. J., Willett, W. C. & Hu, F. B. Comparison of abdominal adiposity and overall obesity in predicting risk of type 2 diabetes among men. Am. J. Clin. Nutr. 81, 555–563 (2005)
Louis, P., Young, P., Holtrop, G. & Flint, H. J. Diversity of human colonic butyrate-producing bacteria revealed by analysis of the butyryl-CoA:acetate CoA-transferase gene. Environ. Microbiol. 12, 304–314 (2010)
Vrieze, A. et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in subjects with metabolic syndrome. Gastroenterology 143, 913–916 (2012)
Furet, J. P. et al. Differential adaptation of human gut microbiota to bariatric surgery-induced weight loss: links with metabolic and low-grade inflammation markers. Diabetes 59, 3049–3057 (2010)
Lundberg, V., Stegmayr, B., Asplund, K., Eliasson, M. & Huhtasaari, F. Diabetes as a risk factor for myocardial infarction: population and gender perspectives. J. Intern. Med. 241, 485–492 (1997)
Vendrame, F. & Gottlieb, P. A. Prediabetes: prediction and prevention trials. Endocrinol. Metab. Clin. North Am. 33, 75–92 (2004)
Oliveira, A. P., Patil, K. R. & Nielsen, J. Architecture of transcriptional regulatory circuits is knitted over the topology of bio-molecular interaction networks. BMC Syst. Biol. 2, 17 (2008)
Patil, K. R. & Nielsen, J. Uncovering transcriptional regulation of metabolism by using metabolic network topology. Proc. Natl Acad. Sci. USA 102, 2685–2689 (2005)
Finegold, S. M. et al. Clostridium clostridioforme: a mixture of three clinically important species. Eur. J. Clin. Microbiol. Infect. Dis. 24, 319–324 (2005)
Larsen, N. et al. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS ONE 5, e9085 (2010)
Karjalainen, K. M., Knuuttila, M. L. & Kaar, M. L. Salivary factors in children and adolescents with insulin-dependent diabetes mellitus. Pediatr. Dent. 18, 306–311 (1996)
Alberti, K. G. & Zimmet, P. Z. Definition, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis and classification of diabetes mellitus provisional report of a WHO consultation. Diabet. Med. 15, 539–553 (1998)
Bingley, P. J., Bonifacio, E. & Mueller, P. W. Diabetes Antibody Standardization Program: first assay proficiency evaluation. Diabetes 52, 1128–1136 (2003)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008)
Zhu, W., Lomsadze, A. & Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 38, e132 (2010)
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006)
Hess, M. et al. Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331, 463–467 (2011)
Dongen, S. v. Graph Clustering by Flow Simulation. PhD thesis, Univ. Utrecht. (2000)
R. Development Core Team. R: A Language and Environment for Statistical Computinghttp://www.R-project.org (R Foundation for Statistical Computing, 2012)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate — a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. B 57, 289–300 (1995)
Knights, D., Costello, E. K. & Knight, R. Supervised classification of human microbiota. FEMS Microbiol. Rev. 35, 343–359 (2011)
Acknowledgements
We would like to acknowledge R. Perkins for editing the manuscript and M.-L. Ekholm, B. Jannemark, G. Östergren-Lundén, H. Kling Bäckhed, C. Schmidt and U. Prahl for technical assistance. This study was supported by the Swedish Research Council, the NovoNordisk foundation, Torsten Söderberg’s foundation, Ragnar Söderberg’s foundation, Swedish Diabetes foundation, Swedish Heart Lung Foundation, IngaBritt och Arne Lundbergs foundation, Chalmers foundation, Bioinformatics Infrastructure for Life Sciences (BILS), Knut and Alice Wallenberg foundation, the Swedish Foundation for Strategic Research, AstraZeneca R&D Mölndal, Sweden and the regional agreement on medical training and clinical research (ALF) between Region Västra Götaland and Sahlgrenska University Hospital. The computations were performed on Chalmers C3SE computing resources.
Author information
Authors and Affiliations
Contributions
C.J.B., B.F. and F.B. conceived and designed the project. F.H.K., V.T., C.J.B. and G.B. performed the experiments. F.H.K., I.N. and J.N. analysed the sequence data. All authors contributed to writing and editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.N. and F.B. are shareholders in MetaboGen AB. All other authors declare no competing financial interests.
Supplementary information
Supplementary Information
This fie contains Supplementary Results, Supplementary Figures 1-17, Supplementary Tables 1-2 and additional references. (PDF 2892 kb)
Supplementary Data
This file contains Supplementary Tables 3-21. (XLSX 668 kb)
Rights and permissions
About this article
Cite this article
Karlsson, F., Tremaroli, V., Nookaew, I. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498, 99–103 (2013). https://doi.org/10.1038/nature12198
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12198
This article is cited by
-
Gut microbiota in relationship to diabetes mellitus and its late complications with a focus on diabetic foot syndrome: A review
Folia Microbiologica (2024)
-
Gut microbiota and type 2 diabetes mellitus: a focus on the gut-brain axis
Endocrine (2024)
-
Short-chain fatty acids as a link between diet and cardiometabolic risk: a narrative review
Lipids in Health and Disease (2023)
-
Effects of Candidatus Liberibacter asiaticus infection on metagenome of Diaphorina citri gut endosymbiont
Scientific Data (2023)
-
Alterations of gut microbiota in biopsy-proven diabetic nephropathy and a long history of diabetes without kidney damage
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.