Abstract
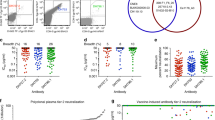
The RV144 trial demonstrated 31% vaccine efficacy at preventing human immunodeficiency virus (HIV)-1 infection1. Antibodies against the HIV-1 envelope variable loops 1 and 2 (Env V1 and V2) correlated inversely with infection risk2. We proposed that vaccine-induced immune responses against V1/V2 would have a selective effect against, or sieve, HIV-1 breakthrough viruses. A total of 936 HIV-1 genome sequences from 44 vaccine and 66 placebo recipients were examined. We show that vaccine-induced immune responses were associated with two signatures in V2 at amino acid positions 169 and 181. Vaccine efficacy against viruses matching the vaccine at position 169 was 48% (confidence interval 18% to 66%; P = 0.0036), whereas vaccine efficacy against viruses mismatching the vaccine at position 181 was 78% (confidence interval 35% to 93%; P = 0.0028). Residue 169 is in a cationic glycosylated region recognized by broadly neutralizing and RV144-derived antibodies. The predicted distance between the two signature sites (21 ± 7 Å) and their match/mismatch dichotomy indicate that multiple factors may be involved in the protection observed in RV144. Genetic signatures of RV144 vaccination in V2 complement the finding of an association between high V1/V2-binding antibodies and reduced risk of HIV-1 acquisition, and provide evidence that vaccine-induced V2 responses plausibly had a role in the partial protection conferred by the RV144 regimen.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
24 September 2012
The web PDF was replaced to correct a corrupted Ångström symbol in three places
References
Rerks-Ngarm, S. et al. Vaccination with ALVAC and AIDSVAX to prevent HIV-1 infection in Thailand. N. Engl. J. Med. 361, 2209–2220 (2009)
Haynes, B. F. et al. Immune-correlates analysis of an HIV-1 vaccine efficacy trial. N. Engl. J. Med. 366, 1275–1286 (2012)
Plotkin, S. A. & Gilbert, P. B. Nomenclature for immune correlates of protection after vaccination. Clin. Infect. Dis. 54, 1615–1617 (2012)
Rolland, M. & Gilbert, P. Evaluating immune correlates in HIV type 1 vaccine efficacy trials: what RV144 may provide. AIDS Res. Hum. Retroviruses 28, 400–404 (2012)
Gilbert, P. B., Self, S. G. & Ashby, M. A. Statistical methods for assessing differential vaccine protection against human immunodeficiency virus types. Biometrics 54, 799–814 (1998)
Gilbert, P., Self, S., Rao, M., Naficy, A. & Clemens, J. Sieve analysis: methods for assessing from vaccine trial data how vaccine efficacy varies with genotypic and phenotypic pathogen variation. J. Clin. Epidemiol. 54, 68–85 (2001)
Rolland, M. et al. Genetic impact of vaccination on breakthrough HIV-1 sequences from the STEP trial. Nature Med. 17, 366–371 (2011)
Wei, X. et al. Antibody neutralization and escape by HIV-1. Nature 422, 307–312 (2003)
Moore, P. L. et al. Limited neutralizing antibody specificities drive neutralization escape in early HIV-1 subtype C infection. PLoS Pathog. 5, e1000598 (2009)
Tomaras, G. D. et al. Polyclonal B cell responses to conserved neutralization epitopes in a subset of HIV-1-infected individuals. J. Virol. 85, 11502–11519 (2011)
Lynch, R. M. et al. The B cell response is redundant and highly focused on V1V2 during early subtype C infection in a Zambian seroconverter. J. Virol. 85, 905–915 (2011)
Robb, M. L. et al. Risk behaviour and time as covariates for efficacy of the HIV vaccine regimen ALVAC-HIV (vCP1521) and AIDSVAX B/E: a post-hoc analysis of the Thai phase 3 efficacy trial RV 144. Lancet Infect. Dis. 12, 531–537 (2012)
Gilbert, P. B., Wu, C. & Jobes, D. V. Genome scanning tests for comparing amino acid sequences between groups. Biometrics 64, 198–207 (2008)
Edlefsen, P. T. Model-based sieve analysis. Preprint at http://arxiv.org/abs/1206.6701 (2012)
Gilbert, P. B. et al. Statistical interpretation of the RV144 HIV vaccine efficacy trial in Thailand: a case study for statistical issues in efficacy trials. J. Infect. Dis. 203, 969–975 (2011)
Carlson, J. M. et al. Phylogenetic dependency networks: inferring patterns of CTL escape and codon covariation in HIV-1 Gag. PLoS Comput. Biol. 4, e1000225 (2008)
Felsenstein, J. Phylogenies and the comparative method. Am. Nat. 125, 1–15 (1985)
Pond, S. L. & Frost, S. D. Datamonkey: rapid detection of selective pressure on individual sites of codon alignments. Bioinformatics 21, 2531–2533 (2005)
Derdeyn, C. A. et al. Envelope-constrained neutralization-sensitive HIV-1 after heterosexual transmission. Science 303, 2019–2022 (2004)
Moore, P. L. et al. Potent and broad neutralization of HIV-1 subtype C by plasma antibodies targeting a quaternary epitope including residues in the V2 loop. J. Virol. 85, 3128–3141 (2011)
Doria-Rose, N. A. et al. A short segment of the HIV-1 gp120 V1/V2 region is a major determinant of resistance to V1/V2 neutralizing antibodies. J. Virol. 86, 8319–8323 (2012)
McLellan, J. S. et al. Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9. Nature 480, 336–343 (2011)
Rubinstein, N. D. et al. Computational characterization of B-cell epitopes. Mol. Immunol. 45, 3477–3489 (2008)
Gilbert, P. B., Novitsky, V. & Essex, M. Covariability of selected amino acid positions for HIV type 1 subtypes C and B. AIDS Res. Hum. Retroviruses 21, 1016–1030 (2005)
Poon, A. F., Lewis, F. I., Pond, S. L. & Frost, S. D. An evolutionary-network model reveals stratified interactions in the V3 loop of the HIV-1 envelope. PLoS Comput. Biol. 3, e231 (2007)
Prentice, R. L. et al. The analysis of failure times in the presence of competing risks. Biometrics 34, 541–554 (1978)
Lunn, M. & McNeil, D. Applying Cox regression to competing risks. Biometrics 51, 524–532 (1995)
Grambsch, P. & Therneau, T. M. Proportional hazards tests and diagnostics based on weighted residuals. Biometrika 81, 515–526 (1994)
Gilbert, P. B. Comparison of competing risks failure time methods and time-independent methods for assessing strain variations in vaccine protection. Stat. Med. 19, 3065–3086 (2000)
Storey, J. D. The positive false discovery rate: a Bayesian interpretation and the q-value. Ann. Stat. 31, 2013–2035 (2003)
Acknowledgements
We thank B. F. Haynes and F. A. Matsen for advice and comments, and I. A. Wilson and R. L. Stanfield for assistance. This study was supported in part by an Interagency Agreement Y1-AI-2642-12 between US Army Medical Research and Material Command (USAMRMC) and the National Institutes of Allergy and Infectious Diseases. This work was also supported by a cooperative agreement (W81XWH-07-2-0067) between the Henry M. Jackson Foundation for the Advancement of Military Medicine, Inc., and the US Department of Defense (DOD). Additional support was provided to P.B.G. through the NIH grant 2R37AI05465-10. The opinions herein are those of the authors and should not be construed as official or representing the views of the US Department of Defense or the Department of the Army.
Author information
Authors and Affiliations
Contributions
M.R. conducted the sequence analysis with contributions from B.B.L., W.D. and B.S.M.; P.T.E. and P.B.G. conducted the site-scanning sieve analyses. M.R., S.T., E.S.-B. and J.I.M. designed the sequencing experiments. B.B.L., L.C., P.K., S.N., J.N.S., K.W., H.Z., M.B., S.H., A. Bates, M.L., A.O’S., E.L., A. Bradfield, G.I. and V.A. performed viral sequencing. T.H., A.C.deC., C.A.M., H.A. and M.J. contributed statistical analyses. C.C., S.M. and W.R.S. developed the EPIMAP approach. J.S.M., I.G. and P.D.K. identified antibody contact residues and performed V2 structural analyses. J.M.C. performed phylogenetic dependency network analyses. R.J.O’C., M.S.deS., S.N., S.R.-N., M.L.R., N.L.M. and J.H.K. conducted the RV144 trial. M.R., P.T.E., P.B.G., J.I.M. and J.H.K. designed the studies, analysed data, prepared the manuscript (with contributions from J.M.C., P.D.K. and W.R.S.) and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information 1
This file contains Supplementary Figures 1–5, Supplementary Methods, Supplementary Tables 1–11 and Supplementary Note 1. (PDF 973 kb)
Supplementary Information 2
This Supplementary Information file contains the Code for differential VE and for the site-scanning methods. (PDF 712 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Rolland, M., Edlefsen, P., Larsen, B. et al. Increased HIV-1 vaccine efficacy against viruses with genetic signatures in Env V2. Nature 490, 417–420 (2012). https://doi.org/10.1038/nature11519
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11519
This article is cited by
-
Direct intranodal tonsil vaccination with modified vaccinia Ankara vaccine protects macaques from highly pathogenic SIVmac251
Nature Communications (2023)
-
Impact of adjuvants on the biophysical and functional characteristics of HIV vaccine-elicited antibodies in humans
npj Vaccines (2022)
-
Differential V2-directed antibody responses in non-human primates infected with SHIVs or immunized with diverse HIV vaccines
Nature Communications (2022)
-
Diverse antiviral IgG effector activities are predicted by unique biophysical antibody features
Retrovirology (2021)
-
Risk compensation after HIV-1 vaccination may accelerate viral adaptation and reduce cost-effectiveness: a modeling study
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.