Abstract
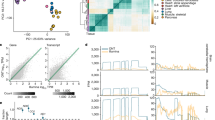
Gene expression is an important phenotype that informs about genetic and environmental effects on cellular state. Many studies have previously identified genetic variants for gene expression phenotypes using custom and commercially available microarrays1,2,3,4,5. Second generation sequencing technologies are now providing unprecedented access to the fine structure of the transcriptome6,7,8,9,10,11,12,13,14. We have sequenced the mRNA fraction of the transcriptome in 60 extended HapMap individuals of European descent and have combined these data with genetic variants from the HapMap3 project15. We have quantified exon abundance based on read depth and have also developed methods to quantify whole transcript abundance. We have found that approximately 10 million reads of sequencing can provide access to the same dynamic range as arrays with better quantification of alternative and highly abundant transcripts. Correlation with SNPs (small nucleotide polymorphisms) leads to a larger discovery of eQTLs (expression quantitative trait loci) than with arrays. We also detect a substantial number of variants that influence the structure of mature transcripts indicating variants responsible for alternative splicing. Finally, measures of allele-specific expression allowed the identification of rare eQTLs and allelic differences in transcript structure. This analysis shows that high throughput sequencing technologies reveal new properties of genetic effects on the transcriptome and allow the exploration of genetic effects in cellular processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Data deposits
All RNA-Seq data in raw and normalized form is available in ArrayExpress under accession numbers E-MTAB-197 and E-MTAB-198 and at http://jungle.unige.ch/rnaseq_CEU60/.
References
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428 (2008)
Göring, H. H. et al. Discovery of expression QTLs using large-scale transcriptional profiling in human lymphocytes. Nature Genet. 39, 1208–1216 (2007)
Moffatt, M. F. et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature 448, 470–473 (2007)
Morley, M. et al. Genetic analysis of genome-wide variation in human gene expression. Nature 430, 743–747 (2004)
Stranger, B. E. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853 (2007)
Wilhelm, B. T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453, 1239–1243 (2008)
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nature Rev. Genet. 10, 57–63 (2009)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature Methods 5, 621–628 (2008)
Sultan, M. et al. A global view of gene activity and alternative splicing by deep sequencing of the human transcriptome. Science 321, 956–960 (2008)
’t Hoen, P. A. C. et al. Deep sequencing-based expression analysis shows major advances in robustness, resolution and inter-lab portability over five microarray platforms. Nucleic Acids Res. 36, e141 (2008)
Maher, C. A. et al. Transcriptome sequencing to detect gene fusions in cancer. Nature 458, 97–101 (2009)
Wang, E. T. et al. Alternative isoform regulation in human tissue transcriptomes. Nature 456, 470–476 (2008)
Pan, Q., Shai, O., Lee, L. J., Frey, B. J. & Blencowe, B. J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nature Genet. 40, 1413–1415 (2008)
Cloonan, N. et al. Stem cell transcriptome profiling via massive-scale mRNA sequencing. Nature Methods 5, 613–619 (2008)
Frazer, K. A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007)
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008)
Hubbard, T. J. et al. Ensembl 2009. Nucleic Acids Res. 37, D690–D697 (2009)
Zheng, S. & Chen, L. A hierarchical Bayesian model for comparing transcriptomes at the individual transcript isoform level. Nucleic Acids Res. 37, e75 (2009)
Hiller, D., Jiang, H., Xu, W. & Wong, W. H. Identifiability of isoform deconvolution from junction arrays and RNA-Seq. Bioinformatics (2009)
Marioni, J. C., Mason, C. E., Mane, S. M., Stephens, M. & Gilad, Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 18, 1509–1517 (2008)
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nature Genet. 39, 906–913 (2007)
Stranger, B. E. et al. Population genomics of human gene expression. Nature Genet. 39, 1217–1224 (2007)
Stranger, B. E. et al. Genome-wide associations of gene expression variation in humans. PLoS Genet. 1, e78 (2005)
Pickrell, J. K. et al. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature XXX, XXX–XXX (2010)
Veyrieras, J. B. et al. High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet. 4, e1000214 (2008)
Pastinen, T. & Hudson, T. J. Cis-acting regulatory variation in the human genome. Science 306, 647–650 (2004)
Verlaan, D. J. et al. Targeted screening of cis-regulatory variation in human haplotypes. Genome Res. 19, 118–127 (2009)
Zhang, K. et al. Digital RNA allelotyping reveals tissue-specific and allele-specific gene expression in human. Nature Methods 6, 613–618 (2009)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Sabatti, C. & Risch, N. Homozygosity and linkage disequilibrium. Genetics 160, 1707–1719 (2002)
Marioni, J. C., Mason, C. E., Mane, S. M., Stephens, M. & Gilad, Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 18, 1509–1517 (2008)
Sammeth, M. Alternative splicing events are bubbles in splicing graphs. J. Comput. Biol. 16, 1117–1140 (2009)
Oshlack, A. & Wakefield, M. J. Transcript length bias in RNA-seq data confounds systems biology. Biol. Direct 4, 14 (2009)
Ahuja, R. K., Magnanti, T. L. & Orlin, J. B. Network Flows: Theory, Algorithms and Applications (Prentice Hall, 1993)
Cormen, T. H., Leiserson, C. E., Rivest R. L & Stein, C. in Introduction to Algorithms, 2nd ed., Ch. 29 770–821 (MIT Press and McGraw-Hill, 2001)
Acknowledgements
The authors would like to acknowledge H. Li, S. White, J. O’Brien, S. Searle, M. Quail, S. V. V. Deevi and the Sequencing Core facility at the Wellcome Trust Sanger Institute. We would also like to thank C. Beazley, A. Nica, L. Jostins, K. Morley, J. Barrett and V. Anttila. Funding was provided by the Wellcome Trust, the Louis-Jeantet foundation and the Swiss National Science Foundation NCCR (‘Frontiers in Genetics’) to E.T.D. and Spanish Ministry of Science and Consolider Ingenio 2010 to R.G.
Author Contributions S.B.M. and E.T.D. conceived and designed the study. S.B.M. performed most of the analysis. S.B.M. and E.T.D. wrote the manuscript. M.S. and R.G. contributed analysis, text and comments to the manuscript. M.G.-A. and R.P.L. helped with the analysis. C.I. and J.N. performed experimental work.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-23 with legends, and Supplementary Table 1. (PDF 2296 kb)
Rights and permissions
About this article
Cite this article
Montgomery, S., Sammeth, M., Gutierrez-Arcelus, M. et al. Transcriptome genetics using second generation sequencing in a Caucasian population. Nature 464, 773–777 (2010). https://doi.org/10.1038/nature08903
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08903
This article is cited by
-
Statistical Learning of Large-Scale Genetic Data: How to Run a Genome-Wide Association Study of Gene-Expression Data Using the 1000 Genomes Project Data
Statistics in Biosciences (2024)
-
5-Methylcytosine profiles in mouse transcriptomes suggest the randomness of m5C formation catalyzed by RNA methyltransferase
BMC Research Notes (2022)
-
Splicing QTL analysis focusing on coding sequences reveals mechanisms for disease susceptibility loci
Nature Communications (2022)
-
Experimental method for haplotype phasing across the entire length of chromosome 21 in trisomy 21 cells using a chromosome elimination technique
Journal of Human Genetics (2022)
-
Platelet transcriptome profiles provide potential therapeutic targets for elderly acute myelocytic leukemia patients
Journal of Translational Medicine (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.