Abstract
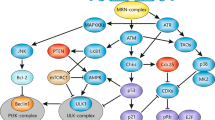
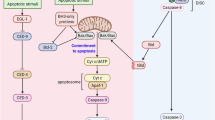
DNA damage frequently triggers death by apoptosis. The irreversible decision to die can be facilitated or forestalled through integration of a wide variety of stimuli from within and around the cell. Here we address some fundamental questions that arise from this model. Why should DNA damage initiate apoptosis in the first place? In damaged cells, what are the alternatives to death and why should they be selected in some circumstances but not others? What signals register DNA damage and how do they impinge on the effector pathways of apoptosis? Is there a suborganellar apoptosome complex effecting the integration of death signals within the nucleus, just as there is in the cytoplasm? And what are the consequences of failure to initiate apoptosis in response to DNA damage?
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Nagata, S. & Golstein, P. The Fas death factor. Science 267, 1449–1456 ( 1995).
Kroemer, G. & Reed, J. C. Mitochondrial control of cell death . Nature Med. 6, 513–519 (2000).
Korsmeyer, S. J. BCL-2 gene family and the regulation of programmed cell death. Cancer Res. 59, 1693–1700 (1999).
Friedberg, E., Walker, G. & Siede, W. DNA Repair and Mutagenesis 108– 133 (ASM Press, Washington DC, 1995).
MacCallum, D. E. et al. The p53 response to ionising radiation in adult and developing murine tissues. Oncogene 13, 2575– 2587 (1996).
Bach, S. P., Renehan, A. G. & Potten, C. S. Stem cells: the intestinal stem cell as a paradigm . Carcinogenesis 21, 469– 476 (2000).
Lowe, S. W., Schmitt, E. M., Smith, S. W., Osborne, B. A. & Jacks, T. p53 is required for radiation-induced apoptosis in mouse thymocytes. Nature 362, 847–849 (1993).
Clarke, A. R. et al. Thymocyte apoptosis induced by p53-dependent and independent pathways. Nature 362, 849– 852 (1993).
Matsuyama, S., Nouraini, S. & Reed, J. C. Yeast as a tool for apoptosis research. Curr. Opin. Microbiol. 2, 618–623 (1999).
Aravind, L., Dixit, V. M. & Koonin, E. V. The domains of death: evolution of the apoptosis machinery. Trends Biochem. Sci. 24, 47– 53 (1999).
Oshima, H., Oshima, M., Kobayashi, M., Tsutsumi, M. & Taketo, M. M. Morphological and molecular processes of polyp formation in Apc(delta716) knockout mice. Cancer Res. 57, 1644–1649 ( 1997).
Pieper, A. A., Verma, A., Zhang, J. & Snyder, S. H. Poly (ADP-ribose) polymerase, nitric oxide and cell death. Trends Pharmacol. Sci. 20, 171–181 ( 1999).
White, K. et al. Genetic control of programmed cell death in Drosophila. Science 264, 677–683 ( 1994).
Toft, N. J. et al. Msh2 status modulates both apoptosis and mutation frequency in the murine small intestine. Proc. Natl Acad. Sci. USA 96, 3911–3915 (1999).
Griffiths, S. D. et al. Absence of p53 permits propagation of mutant cells following genotoxic damage. Oncogene 14, 523– 531 (1997).
Hartwell, L. H. & Weinert, T. A. Checkpoints: controls that ensure the order of cell cycle events. Science 246, 629–634 (1989).
Hanahan, D. & Weinberg, R. The hallmarks of cancer. Cell 100, 57–70 ( 2000).
Van Sloun, P. P. et al. The role of nucleotide excision repair in protecting embryonic stem cells from genotoxic effects of UV-induced DNA damage. Nucleic Acids Res. 27, 3276–3282 (1999).
Martin, S., Laroche, T., Suka, N., Grunstein, M. & Gasser, S. Relocalization of telomeric Ku and SIR proteins in response to DNA strand breaks in yeast. Cell 97, 621–633 (1999).
Galy, V. et al. Nuclear pore complexes in the organization of silent telomeric chromatin. Nature 403, 108– 112 (2000).
Hsu, H. L., Gilley, D., Blackburn, E. H. & Chen, D. J. Ku is associated with the telomere in mammals. Proc. Natl Acad. Sci. USA 96, 12454–1248 (1999).
Gasser, S. A sense of the end. Science 288, 1377– 1379 (2000).
Lieber, M. Pathological and physiological double-strand breaks. Roles in cancer, aging and the immune system. Am. J. Pathol. 153, 1323–1332 (1998).
Artandi, S. E. & DePinho, R. A. A critical role for telomeres in suppressing and facilitating carcinogenesis. Curr. Opin. Genet. Dev. 10, 39–46 (2000).
Rotman, G. & Shiloh, Y. ATM: from gene to function. Hum. Mol. Genet. 7, 1555–1563 (1998).
Smith, G. C. M. & Jackson, S. P. The DNA-dependent protein kinase. Genes Dev. 13, 916– 934 (1999).
Hoekstra, M. F. Responses to DNA damage and regulation of cell cycle checkpoints by the ATM protein kinase family. Curr. Opin. Genet. Dev. 7, 170–175 (1997).
Smith, G. C. et al. Purification and DNA binding properties of the ataxia-telangiectasia gene product ATM. Proc. Natl Acad. Sci. USA 96, 11134–11139 (1999).
Goldbeter, A. & Koshland, D. E. Jr An amplified sensitivity arising from covalent modification in biological systems. Proc. Natl Acad. Sci. USA 78, 6840– 6844 (1981).
Aguda, B. Instabilities in phosphorylation–dephosphorylation cascades and cell cycle checkpoints. Oncogene 18, 2846– 2851 (1998).
Huang, L. C., Clarkin, K. C. & Wahl, G. M. Sensitivity and selectivity of the DNA damage sensor responsible for activating p53-dependent G1 arrest. Proc. Natl Acad. Sci. USA 93, 4827–4832 (1996).
Dasika, G. et al. DNA damage-induced cell cycle checkpoints and DNA strand break repair in development and tumorigenesis. Oncogene 18 , 7883–7899 (1999).
Bunz, F. et al. Requirement for p53 and p21 to sustain G2 arrest after DNA damage . Science 282, 1497–1501 (1998).
Pandita, T. K. et al. Ionizing radiation activates the ATM kinase throughout the cell cycle. Oncogene 19, 1386– 1391 (2000).
Lohrum, M. & Vousden, K. Regulation and function of the p53-related proteins: same family, different rules. Trends Cell Biol. 10, 197–202 (2000).
Chan, T., Hermeking, H., Lengauer, C., Kinzler, K. & Vogelstein, B. 14-3-3σ is required to prevent mitotic catastrophe after DNA damage. Nature 401, 616–620 (1999).
Lim, D. S. et al. ATM phosphorylates p95/nbs1 in an S-phase checkpoint pathway . Nature 404, 613–617 (2000).
Sionov, R. V. & Haupt, Y. The cellular response to p53: the decision between life and death. Oncogene 18, 6145–6157 (1999).
Meek, D. Mechanisms of switching on p53: a role for covalent modification? Oncogene 18, 7666–7675 ( 1999).
Clarke, A. R., Gledhill, S., Hooper, M. L., Bird, C. C. & Wyllie, A. H. p53 dependence of early apoptotic and proliferative responses within the mouse intestinal epithelium following gamma-irradiation. Oncogene 9, 1767– 1773 (1994).
Merritt, A. J. et al. The role of p53 in spontaneous and radiation-induced apoptosis in the gastrointestinal tract of normal and p53-deficient mice. Cancer Res. 54, 614–617 (1994).
Wu, G. S. et al. Induction of the TRAIL receptor KILLER/DR5 in p53-dependent apoptosis but not growth arrest. Oncogene 18, 6411 –6418 (1999).
Brodsky, M. H. et al. Drosophila p53 binds a damage response element at the reaper locus. Cell 101, 103– 113 (2000).
O'Connor, L., Harris, A. W. & Strasser, A. CD95 (Fas/APO-1) and p53 signal apoptosis independently in diverse cell types. Cancer Res. 60, 1217 –1220 (2000).
Munsch, D. et al. Human and mouse Fas (APO-1/CD95) death receptor genes each contain a p53-responsive element that is activated by p53 mutants unable to induce apoptosis. J. Biol. Chem. 275, 3867 –3872 (2000).
Zhao, R. et al. Analysis of p53-regulated gene expression patterns using oligonucleotide arrays. Genes Dev. 14, 981– 993 (2000).
Attardi, L. D. et al. PERP, an apoptosis-associated target of p53, is a novel member of the PMP-22/gas3 family. Genes Dev. 14, 704–718 (2000).
Kannan, K., Amariglio, N., Rechavi, G. & Givol, D. Profile of gene expression regulated by induced p53: connection to the TGF-β family. FEBS Lett. 470, 77– 82 (2000).
Marchenko, N. D., Zaika, A. & Moll, U. M. Death signal-induced localization of p53 protein to mitochondria. A potential role in apoptotic signaling. J. Biol. Chem. 275, 16202–16212 (2000).
Morris, V. B., Brammall, J., Noble, J. & Reddel, R. p53 localizes to the centrosomes and spindles of mitotic cells in the embryonic chick epiblast, human cell lines, and a human primary culture: an immunofluorescence study . Exp. Cell Res. 256, 122– 130 (2000).
Loughran, O. & La Thangue, N. B. Apoptotic and growth-promoting activity of E2F modulated by MDM2. Mol. Cell. Biol. 20, 2186–2197 (2000).
Blattner, C., Sparks, A. & Lane, D. Transcription factor E2F-1 is upregulated in response to DNA damage in a manner analogous to that of p53. Mol. Cell. Biol. 19, 3704–3713 ( 1999).
Lissy, N. A., Davis, P. K., Irwin, M., Kaelin, W. G. & Dowdy, S. F. A common E2F-1 and p73 pathway mediates cell death induced by TCR activation. Nature 407, 642 –645 (2000).
Irwin, M. et al. Role for the p53 homologue p73 in E2F-1-induced apoptosis. Nature 407, 645–648 ( 2000).
Wu, X. & Levine, A. J. p53 and E2f-1 co-operate to mediate apoptosis. Proc. Natl Acad. Sci. USA 91, 3602–3606 (1994).
Evan, G. I. et al. Induction of apoptosis in fibroblasts by c-myc protein. Cell 69, 119–128 ( 1992).
Lowe, S. W., Jacks, T., Housman, D. E. & Ruley, H. E. Abrogation of oncogene-associated apoptosis allows transformation of p53-deficient cells. Proc. Natl Acad. Sci. USA 91, 2026 –2030 (1994).
Van Etten, R. A. Cycling, stressed-out and nervous: cellular functions of c-Abl. Trends Cell Biol. 9, 179–186 (1999).
Kharbanda, S., Yuan, Z. M., Weichselbaum, R. & Kufe, D. Determination of cell fate by c-Abl activation in the response to DNA damage . Oncogene 17, 3309–3318 (1998).
Shaul, Y. c-Abl: activation and nuclear targets. Cell Death Differ. 7, 10–16 (2000).
Nickerson, J. Nuclear dreams: the malignant alteration of nuclear architecture. J. Cell. Biochem. 70, 172–180 (1998).
Wang, Y. et al. BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures. Genes Dev. 14, 927–939 ( 2000).
Wang, Z. G. et al. PML is essential for multiple apoptotic pathways. Nature Genet. 20, 266–272 (1998).
Zhong, S. et al. A role for PML and the nuclear body in genomic stability. Oncogene 18, 7941–7947 ( 1999).
Hodges, M., Tissot, C., Howe, K., Grimwade, D. & Freemont, P. S. Structure, organization, and dynamics of promyelocytic leukemia protein nuclear bodies. Am. J. Hum. Genet. 63, 297–304 (1998).
Maul, G. G., Negorev, D., Bell, P. & Ishov, A. M. Review: properties and assembly mechanisms of ND10, PML bodies, or PODs. J. Struct. Biol. 129, 278–287 ( 2000).
Lombard, D. B. & Guarente, L. Nijmegen breakage syndrome disease protein and MRE11 at PML nuclear bodies and meiotic telomeres . Cancer Res. 60, 2331– 2334 (2000).
Zhong, S. et al. Promyelocytic leukemia protein (PML) and Daxx participate in a novel nuclear pathway for apoptosis. J. Exp. Med. 191, 631–640 (2000).
Bochar, D. et al. BRCA1 is associated with a human SWI/SNF-related complex: linking chromatin remodeling to breast cancer. Cell 102, 257–265 (2000).
Linares-Cruz, G. et al. p21WAF-1 reorganizes the nucleus in tumor suppression. Proc. Natl Acad. Sci. USA 95, 1131– 1135 (1998).
Gray, J. & Collins, C. Genome changes and gene expression in human solid tumors. Carcinogenesis 21, 443–452 (2000).
Schrock, E. et al. Multicolor spectral karyotyping of human chromosomes. Science 273, 494–497 ( 1996).
Anderson, W. Gene therapy scores against cancer. Nature Med. 6, 862–863 (2000).
Pearson, M. et al. PML regulates p53 acetylation and premature senescence induced by oncogenic Ras. Nature 406, 207– 210 (2000).
Karran, P. DNA double strand breaks in mammalian cells. Curr. Opin. Geneti. Dev. 10, 144–150 ( 2000).
Liang, F., Han, M., Romanienko, P. J. & Jasin, M. Homology-directed repair is a major double-strand break repair pathway in mammalian cells. Proc. Natl Acad. Sci. USA 95, 5172–5177 (1998).
Kolodner, R. & Marsischky, G. Eukaryotic DNA mismatch repair . Curr. Opin. Genet. Dev. 9, 89– 96 (1999).
Krokan, H. E., Nilsen, H., Skorpen, F., Otterlei, M. & Slupphaug, G. Base excision repair of DNA in mammalian cells. FEBS Lett. 476, 73–77 ( 2000).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rich, T., Allen, R. & Wyllie, A. Defying death after DNA damage. Nature 407, 777–783 (2000). https://doi.org/10.1038/35037717
Issue Date:
DOI: https://doi.org/10.1038/35037717
This article is cited by
-
Cellular prion protein offers neuroprotection in astrocytes submitted to amyloid β oligomer toxicity
Molecular and Cellular Biochemistry (2023)
-
Sperm DNA integrity does play a crucial role for embryo development after ICSI, notably when good-quality oocytes from young donors are used
Biological Research (2022)
-
Peptidylarginine deiminase 2 promotes T helper 17-like T cell activation and activated T cell-autonomous death (ACAD) through an endoplasmic reticulum stress and autophagy coupling mechanism
Cellular & Molecular Biology Letters (2022)
-
The power of an idea: Andrew Wyllie
Cell Death & Differentiation (2022)
-
DNA double-strand breaks of human peripheral blood lymphocyte induced by CT examination of oral and maxillofacial region
Clinical Oral Investigations (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.