Abstract
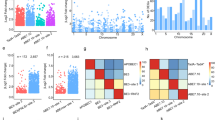
Alternative splicing as a mean to control gene expression and diversify function is suspected to considerably influence drug response and clearance. We report the quantitative expression profiles of the human UGT genes including alternatively spliced variants not previously annotated established by deep RNA-sequencing in tissues of pharmacological importance. We reveal a comprehensive quantification of the alternative UGT transcriptome that differ across tissues and among individuals. Alternative transcripts that comprise novel in-frame sequences associated or not with truncations of the 5′- and/or 3′- termini, significantly contribute to the total expression levels of each UGT1 and UGT2 gene averaging 21% in normal tissues, with expression of UGT2 variants surpassing those of UGT1. Quantitative data expose preferential tissue expression patterns and remodeling in favor of alternative variants upon tumorigenesis. These complex alternative splicing programs have the strong potential to contribute to interindividual variability in drug metabolism in addition to diversify the UGT proteome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Bellemare J, Rouleau M, Harvey M, Guillemette C . Modulation of the human glucuronosyltransferase UGT1A pathway by splice isoform polypeptides is mediated through protein-protein interactions. J Biol Chem 2010; 285: 3600–3607.
Chhibber A, French CE, Yee SW, Gamazon ER, Theusch E, Qin X et al. Transcriptomic variation of pharmacogenes in multiple human tissues and lymphoblastoid cell lines. Pharmacogenomics J 2016; 17: 137–145.
Menard V, Collin P, Margaillan G, Guillemette C . Modulation of the UGT2B7 enzyme activity by C-terminally truncated proteins derived from alternative splicing. Drug Metab Dispos 2013; 41: 2197–2205.
Guillemette C, Levesque E, Rouleau M . Pharmacogenomics of human uridine diphospho-glucuronosyltransferases and clinical implications. Clin Pharmacol Ther 2014; 96: 324–339.
Stingl JC, Bartels H, Viviani R, Lehmann ML, Brockmoller J . Relevance of UDP-glucuronosyltransferase polymorphisms for drug dosing: a quantitative systematic review. Pharmacol Ther 2014; 141: 92–116.
Meech R, Rogers A, Zhuang L, Lewis BC, Miners JO, Mackenzie PI . Identification of residues that confer sugar selectivity to UDP-glycosyltransferase 3A (UGT3A) enzymes. J Biol Chem 2012; 287: 24122–24130.
Meech R, Mubarokah N, Shivasami A, Rogers A, Nair PC, Hu DG et al. A novel function for UDP glycosyltransferase 8: galactosidation of bile acids. Mol Pharmacol 2015; 87: 442–450.
Tourancheau A, Margaillan G, Rouleau M, Gilbert I, Villeneuve L, Levesque E et al. Unravelling the transcriptomic landscape of the major phase II UDP-glucuronosyltransferase drug metabolizing pathway using targeted RNA sequencing. Pharmacogenomics J 2016; 16: 60–70.
Langmead B, Salzberg SL . Fast gapped-read alignment with Bowtie 2. Nat Methods 2012; 9: 357–359.
Bolger AM, Lohse M, Usadel B . Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014; 30: 2114–2120.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL . TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 2013; 14: R36.
Goff L, Trapnell C, Kelley D . cummeRbund: analysis, exploration, manipulation, and visualization of Cufflinks high-throughput sequencing data. R Package Version 282 2013.
R Development Core Team, R.C R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria, 2008.
den Dunnen JT, Dalgleish R, Maglott DR, Hart RK, Greenblatt MS, McGowan-Jordan J et al. HGVS recommendations for the description of sequence variants: 2016 update. Hum Mutat 2016; 37: 564–569.
MacKenzie PI, Rogers A, Elliot DJ, Chau N, Hulin JA, Miners JO et al. The novel UDP glycosyltransferase 3A2: cloning, catalytic properties, and tissue distribution. Mol Pharmacol 2011; 79: 472–478.
Mackenzie PI, Rogers A, Treloar J, Jorgensen BR, Miners JO, Meech R . Identification of UDP glycosyltransferase 3A1 as a UDP N-acetylglucosaminyltransferase. J Biol Chem 2008; 283: 36205–36210.
McCarroll SA, Hadnott TN, Perry GH, Sabeti PC, Zody MC, Barrett JC et al. Common deletion polymorphisms in the human genome. Nat Genet 2006; 38: 86–92.
Bellemare J, Rouleau M, Harvey M, Popa I, Pelletier G, Tetu B et al. Immunohistochemical expression of conjugating UGT1A-derived isoforms in normal and tumoral drug-metabolizing tissues in humans. J Pathol 2011; 223: 425–435.
Court MH, Zhang X, Ding X, Yee KK, Hesse LM, Finel M . Quantitative distribution of mRNAs encoding the 19 human UDP-glucuronosyltransferase enzymes in 26 adult and 3 fetal tissues. Xenobiotica 2012; 42: 266–277.
Margaillan G, Rouleau M, Fallon JK, Caron P, Villeneuve L, Turcotte V et al. Quantitative profiling of human renal UDP-glucuronosyltransferases and glucuronidation activity: a comparison of normal and tumoral kidney tissues. Drug Metab Dispos 2015; 43: 611–619.
Margaillan G, Rouleau M, Klein K, Fallon JK, Caron P, Villeneuve L et al. Multiplexed targeted quantitative proteomics predicts hepatic glucuronidation potential. Drug Metab Dispos 2015; 43: 1331–1335.
Ohno S, Nakajin S . Determination of mRNA expression of human UDP-glucuronosyltransferases and application for localization in various human tissues by real-time reverse transcriptase-polymerase chain reaction. Drug Metab Dispos 2009; 37: 32–40.
Hart T, Komori HK, LaMere S, Podshivalova K, Salomon DR . Finding the active genes in deep RNA-seq gene expression studies. BMC Genomics 2013; 14: 778.
Menard V, Levesque E, Chen S, Eap O, Joy MS, Ekstrom L et al. Expression of UGT2B7 is driven by two mutually exclusive promoters and alternative splicing in human tissues: changes from prenatal life to adulthood and in kidney cancer. Pharmacogenet Genomics 2013; 23: 684–696.
Rouleau M, Roberge J, Bellemare J, Guillemette C . Dual roles for splice variants of the glucuronidation pathway as regulators of cellular metabolism. Mol Pharmacol 2014; 85: 29–36.
Rouleau M, Tourancheau A, Girard-Bock C, Villeneuve L, Vaucher J, Duperre AM et al. Divergent expression and metabolic functions of human glucuronosyltransferases through alternative splicing. Cell Rep 2016; 17: 114–124.
Nagy E, Maquat LE . A rule for termination-codon position within intron-containing genes: when nonsense affects RNA abundance. Trends Biochem Sci 1998; 23: 198–199.
Bushey RT, Dluzen DF, Lazarus P . Importance of UDP-glucuronosyltransferases 2A2 and 2A3 in tobacco carcinogen metabolism. Drug Metab Dispos 2013; 41: 170–179.
Bushey RT, Lazarus P . Identification and functional characterization of a novel UDP-glucuronosyltransferase 2A1 splice variant: potential importance in tobacco-related cancer susceptibility. J Pharmacol Exp Ther 2012; 343: 712–724.
Audet-Delage Y, Rouleau M, Rouleau M, Roberge J, Miard S, Picard F et al. Crosstalk between alternatively spliced UGT1A isoforms and colon cancer cell metabolism. Mol Pharmacol 2016; 91: 167–177.
Shkreta L, Bell B, Revil T, Venables JP, Prinos P, Elela SA et al. Cancer-associated perturbations in alternative pre-messenger RNA splicing. Cancer Treat Res 2013; 158: 41–94.
Levesque E, Menard V, Laverdiere I, Bellemare J, Barbier O, Girard H et al. Extensive splicing of transcripts encoding the bile acid-conjugating enzyme UGT2B4 modulates glucuronidation. Pharmacogenet Genomics 2010; 20: 195–210.
Menard V, Eap O, Roberge J, Harvey M, Levesque E, Guillemette C . Transcriptional diversity at the UGT2B7 locus is dictated by extensive pre-mRNA splicing mechanisms that give rise to multiple mRNA splice variants. Pharmacogenet Genomics 2011; 21: 631–641.
Acknowledgements
This work was supported by the Canadian Institutes of Health Research (CIHR) (MOP-42392, to CG); and the Canada Research Chair in Pharmacogenomics (Tier I) to CG. We thank the Genomics Analysis Platform of the Institut de Biologie Integrative et des Systèmes (IBIS; Laval University, Québec, QC, Canada; specifically Brian Boyle) for their help with RNA-Seq experiments and Anne-Marie Duperré for help with preparation of figures. We also acknowledge the excellent artwork by France Couture.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Rights and permissions
About this article
Cite this article
Tourancheau, A., Rouleau, M., Guauque-Olarte, S. et al. Quantitative profiling of the UGT transcriptome in human drug-metabolizing tissues. Pharmacogenomics J 18, 251–261 (2018). https://doi.org/10.1038/tpj.2017.5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2017.5
This article is cited by
-
Non-canonical transcriptional regulation of the poor prognostic factor UGT2B17 in chronic lymphocytic leukemic and normal B cells
BMC Cancer (2024)
-
Emerging roles for UDP-glucuronosyltransferases in drug resistance and cancer progression
British Journal of Cancer (2020)
-
The Fusarium metabolite culmorin suppresses the in vitro glucuronidation of deoxynivalenol
Archives of Toxicology (2019)