Abstract
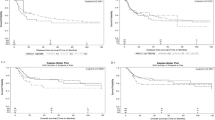
Oxaliplatin-based chemotherapy exerts its effects through generating DNA damage. Hence, genetic variants in DNA repair pathways could modulate treatment response. We used a prospective cohort of 623 colorectal cancer patients with stage II–IV disease treated with adjuvant/palliative chemotherapy to comprehensively investigate 1727 genetic variants in the DNA repair pathways as potential predictive markers for oxaliplatin treatment. Single nucleotide polymorphisms (SNP) associations with overall survival and recurrence-free survival were assessed using a Cox regression model. Pathway analysis was performed using the gamma method. Patients carrying variant alleles of rs3783819 (MNAT1) and rs1043953 (XPC) experienced a longer overall survival after treatment with oxaliplatin than patients who did not carry the variant allele, while the opposite association was found in patients who were not treated with oxaliplatin (false discovery rate-adjusted P-values for heterogeneity 0.0047 and 0.0237, respectively). The nucleotide excision repair (NER) pathway was found to be most likely associated with overall survival in patients who received oxaliplatin (P-value=0.002). Our data show that genetic variants in the NER pathway are potentially predictive of treatment response to oxaliplatin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ferlay J, Soerjomataram I, Ervik M, Dikshit R, Eser S, Mathers C et al. GLOBOCAN 2012 v1.0, Cancer Incidence and Mortality Worldwide: IARC CancerBase No. 11 [internet]. Lyon, France International Agency for Research on Cancer. Available from http://globocan.iarc.fr, accessed on 12 November 2014.
Majek O, Gondos A, Jansen L, Emrich K, Holleczek B, Katalinic A et al. Survival from colorectal cancer in Germany in the early 21st century. Br J Cancer 2012; 106: 1875–1880.
Schmiegel W, Pox C, Adler G, Fleig W, Folsch UR, Fruhmorgen P et al. [S3-Guidelines Conference ‘Colorectal Carcinoma’ 2004]. Z Gastroenterol 2004; 42: 1129–1177.
Cunningham D, Sirohi B, Pluzanska A, Utracka-Hutka B, Zaluski J, Glynne-Jones R et al. Two different first-line 5-fluorouracil regimens with or without oxaliplatin in patients with metastatic colorectal cancer. Ann Oncol 2009; 20: 244–250.
Rothenberg ML, Oza AM, Bigelow RH, Berlin JD, Marshall JL, Ramanathan RK et al. Superiority of oxaliplatin and fluorouracil-leucovorin compared with either therapy alone in patients with progressive colorectal cancer after irinotecan and fluorouracil-leucovorin: interim results of a phase III trial. J Clin Oncol 2003; 21: 2059–2069.
Goldberg RM, Sargent DJ, Morton RF, Fuchs CS, Ramanathan RK, Williamson SK et al. A randomized controlled trial of fluorouracil plus leucovorin, irinotecan, and oxaliplatin combinations in patients with previously untreated metastatic colorectal cancer. J Clin Oncol 2004; 22: 23–30.
Andre T, Boni C, Mounedji-Boudiaf L, Navarro M, Tabernero J, Hickish T et al. Oxaliplatin, fluorouracil, and leucovorin as adjuvant treatment for colon cancer. N Engl J Med 2004; 350: 2343–2351.
Raymond E, Faivre S, Chaney S, Woynarowski J, Cvitkovic E . Cellular and molecular pharmacology of oxaliplatin. Mol Cancer Ther 2002; 1: 227–235.
Arnould S, Hennebelle I, Canal P, Bugat R, Guichard S . Cellular determinants of oxaliplatin sensitivity in colon cancer cell lines. Eur J Cancer 2003; 39: 112–119.
Huang MY, Huang ML, Chen MJ, Lu CY, Chen CF, Tsai PC et al. Multiple genetic polymorphisms in the prediction of clinical outcome of metastatic colorectal cancer patients treated with first-line FOLFOX-4 chemotherapy. Pharmacogenet Genomics 2011; 21: 18–25.
Theile D, Grebhardt S, Haefeli WE, Weiss J . Involvement of drug transporters in the synergistic action of FOLFOX combination chemotherapy. Biochem Pharmacol 2009; 78: 1366–1373.
Wood RD, Mitchell M, Sgouros J, Lindahl T . Human DNA repair genes. Science 2001; 291: 1284–1289.
Sancar A, Lindsey-Boltz LA, Unsal-Kacmaz K, Linn S . Molecular mechanisms of mammalian DNA repair and the DNA damage checkpoints. Annu Rev Biochem 2004; 73: 39–85.
Wang W . Emergence of a DNA-damage response network consisting of Fanconi anaemia and BRCA proteins. Nat Rev Genet 2007; 8: 735–748.
Reardon JT, Vaisman A, Chaney SG, Sancar A . Efficient nucleotide excision repair of cisplatin, oxaliplatin, and Bis-aceto-ammine-dichloro-cyclohexylamine-platinum(IV) (JM216) platinum intrastrand DNA diadducts. Cancer Res 1999; 59: 3968–3971.
McLeod HL, Sargent DJ, Marsh S, Green EM, King CR, Fuchs CS et al. Pharmacogenetic predictors of adverse events and response to chemotherapy in metastatic colorectal cancer: results from North American Gastrointestinal Intergroup Trial N9741. J Clin Oncol 2010; 28: 3227–3233.
Ruzzo A, Graziano F, Loupakis F, Rulli E, Canestrari E, Santini D et al. Pharmacogenetic profiling in patients with advanced colorectal cancer treated with first-line FOLFOX-4 chemotherapy. J Clin Oncol 2007; 25: 1247–1254.
Stoehlmacher J, Ghaderi V, Iobal S, Groshen S, Tsao-Wei D, Park D et al. A polymorphism of the XRCC1 gene predicts for response to platinum based treatment in advanced colorectal cancer. Anticancer Res 2001; 21: 3075–3079.
Lilla C, Verla-Tebit E, Risch A, Jager B, Hoffmeister M, Brenner H et al. Effect of NAT1 and NAT2 genetic polymorphisms on colorectal cancer risk associated with exposure to tobacco smoke and meat consumption. Cancer Epidemiol Biomarkers Prev 2006; 15: 99–107.
Jansen L, Hoffmeister M, Chang-Claude J, Koch M, Brenner H, Arndt V . Age-specific administration of chemotherapy and long-term quality of life in stage II and III colorectal cancer patients: a population-based prospective cohort. Oncologist 2011; 16: 1741–1751.
Fan JB, Oliphant A, Shen R, Kermani BG, Garcia F, Gunderson KL et al. Highly parallel SNP genotyping. Cold Spring Harb Symp Quant Biol 2003; 68: 69–78.
Gabriel S, Ziaugra L, Tabbaa D . SNP genotyping using the Sequenom MassARRAY iPLEX platform. Curr Protoc Hum Genet 2009, Chapter 2: Unit 2.12.
Peters U, Hutter CM, Hsu L, Schumacher FR, Conti DV, Carlson CS et al. Meta-analysis of new genome-wide association studies of colorectal cancer risk. Hum Genet 2012; 131: 217–234.
Howie BN, Donnelly P, Marchini J . A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 2009; 5: e1000529.
Li Y, Willer CJ, Ding J, Scheet P, Abecasis GR . MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol 2010; 34: 816–834.
Schemper M, Smith TL . A note on quantifying follow-up in studies of failure time. Control Clin Trials 1996; 17: 343–346.
Grambsch PM, Therneau TM . Proportional hazards tests and diagnostics based on weighted residuals. Biometrika 1994; 81: 515–526.
Benjamini Y, Hochberg Y . Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Statist Soc B 1995; 57: 289–300.
Goenen M, Heller G . Concordance probability and discriminatory power in proportional hazards regression. Biometrika 2005; 92: 965–970.
Nagelkerke NJ . A note on a general definition of the coefficient of determination. Biometrika 1991; 78: 691–692.
Biernacka JM, Jenkins GD, Wang L, Moyer AM, Fridley BL . Use of the gamma method for self-contained gene-set analysis of SNP data. Eur J Hum Genet 2012; 20: 565–571.
Marchini J, Howie B . Genotype imputation for genome-wide association studies. Nat Rev Genet 2010; 11: 499–511.
Marinoni JC, Roy R, Vermeulen W, Miniou P, Lutz Y, Weeda G et al. Cloning and characterization of p52, the fifth subunit of the core of the transcription/DNA repair factor TFIIH. EMBO J 1997; 16: 1093–1102.
Evans E, Moggs JG, Hwang JR, Egly JM, Wood RD . Mechanism of open complex and dual incision formation by human nucleotide excision repair factors. EMBO J 1997; 16: 6559–6573.
Hoogstraten D, Nigg AL, Heath H, Mullenders LH, van DR, Hoeijmakers JH et al. Rapid switching of TFIIH between RNA polymerase I and II transcription and DNA repair in vivo. Mol Cell 2002; 10: 1163–1174.
Matsuno M, Kose H, Okabe M, Hiromi Y . TFIIH controls developmentally-regulated cell cycle progression as a holocomplex. Genes Cells 2007; 12: 1289–1300.
Chen D, Riedl T, Washbrook E, Pace PE, Coombes RC, Egly JM et al. Activation of estrogen receptor alpha by S118 phosphorylation involves a ligand-dependent interaction with TFIIH and participation of CDK7. Mol Cell 2000; 6: 127–137.
Wu X, Spitz MR, Lee JJ, Lippman SM, Ye Y, Yang H et al. Novel susceptibility loci for second primary tumors/recurrence in head and neck cancer patients: large-scale evaluation of genetic variants. Cancer Prev Res (Phila) 2009; 2: 617–624.
Qin HD, Shugart YY, Bei JX, Pan QH, Chen L, Feng QS et al. Comprehensive pathway-based association study of DNA repair gene variants and the risk of nasopharyngeal carcinoma. Cancer Res 2011; 71: 3000–3008.
Barry KH, Koutros S, Andreotti G, Sandler DP, Burdette LA, Yeager M et al. Genetic variation in nucleotide excision repair pathway genes, pesticide exposure and prostate cancer risk. Carcinogenesis 2012; 33: 331–337.
Wang SS, Gonzalez P, Yu K, Porras C, Li Q, Safaeian M et al. Common genetic variants and risk for HPV persistence and progression to cervical cancer. PLoS One 2010; 5: e8667.
Gao Y, Hayes RB, Huang WY, Caporaso NE, Burdette L, Yeager M et al. DNA repair gene polymorphisms and tobacco smoking in the risk for colorectal adenomas. Carcinogenesis 2011; 32: 882–887.
McWilliams RR, Bamlet WR, de AM, Rider DN, Cunningham JM, Petersen GM . Nucleotide excision repair pathway polymorphisms and pancreatic cancer risk: evidence for role of MMS19L. Cancer Epidemiol Biomarkers Prev 2009; 18: 1295–1302.
Buch SC, Diergaarde B, Nukui T, Day RS, Siegfried JM, Romkes M et al. Genetic variability in DNA repair and cell cycle control pathway genes and risk of smoking-related lung cancer. Mol Carcinog 2012; 51: E11–E20.
Li Y, Jin G, Wang H, Liu H, Qian J, Gu S et al. Polymorphisms of CAK genes and risk for lung cancer: a case-control study in Chinese population. Lung Cancer 2007; 58: 171–183.
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet 2013; 45: 580–585.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005; 120: 15–20.
DiGiovanna JJ, Kraemer KH . Shining a light on xeroderma pigmentosum. J Invest Dermatol 2012; 132: 785–796.
Volker M, Mone MJ, Karmakar P, van HA, Schul W, Vermeulen W et al. Sequential assembly of the nucleotide excision repair factors in vivo. Mol Cell 2001; 8: 213–224.
Yue AM, Xie ZB, Zhao HF, Guo SP, Shen YH, Wang HP . Associations of ABCB1 and XPC genetic polymorphisms with susceptibility to colorectal cancer and therapeutic prognosis in a Chinese population. Asian Pac J Cancer Prev 2013; 14: 3085–3091.
Liu D, Wu HZ, Zhang YN, Kang H, Sun MJ, Wang EH et al. DNA repair genes XPC, XPG polymorphisms: relation to the risk of colorectal carcinoma and therapeutic outcome with oxaliplatin-based adjuvant chemotherapy. Mol Carcinog 2012; 51: E83–E93.
Yin M, Yan J, Martinez-Balibrea E, Graziano F, Lenz HJ, Kim HJ et al. ERCC1 and ERCC2 polymorphisms predict clinical outcomes of oxaliplatin-based chemotherapies in gastric and colorectal cancer: a systemic review and meta-analysis. Clin Cancer Res 2011; 17: 1632–1640.
Acknowledgements
We thank all participants of the DACHS study, the interviewers, physicians and staff of the hospitals for their cooperation. Finally, the excellent technical assistance by the microarray unit of the German Cancer Research Center (especially Matthias Schick), Muhabbet Celik, Ursula Eilber, Sabine Behrens and Ute Handte-Daub was very much appreciated. We thank all those who contributed to the 1000 genomes project to make the imputation of additional SNPs possible. The DACHS study was supported by grants from the German Research Council (Deutsche Forschungsgemeinschaft, grant numbers BR 1704/6-1, BR 1704/6-3, BR 1704/6-4 and CH 390 117/1-1) and the German Federal Ministry of Education and Research (grant numbers 01ER0814 and 01ER0815). GECCO is supported by the National Cancer Institute, National Institutes of Health, US Department of Health and Human Services, grant numbers U01 CA137088; R01 CA059045.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
PowerPoint slides
Rights and permissions
About this article
Cite this article
Kap, E., Seibold, P., Richter, S. et al. Genetic variants in DNA repair genes as potential predictive markers for oxaliplatin chemotherapy in colorectal cancer. Pharmacogenomics J 15, 505–512 (2015). https://doi.org/10.1038/tpj.2015.8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2015.8
This article is cited by
-
Genetic variants in RPA1 associated with the response to oxaliplatin-based chemotherapy in colorectal cancer
Journal of Gastroenterology (2019)
-
Polymorphisms in drug-metabolizing enzymes and steady-state exemestane concentration in postmenopausal patients with breast cancer
The Pharmacogenomics Journal (2017)
-
The study of the relation of DNA repair pathway genes SNPs and the sensitivity to radiotherapy and chemotherapy of NSCLC
Scientific Reports (2016)
-
Reactive oxygen species mediate oxaliplatin-induced epithelial-mesenchymal transition and invasive potential in colon cancer
Tumor Biology (2016)