Abstract
Different combinations of 5-fluorouracil (5-FU), oxaliplatin, irinotecan and other newly developed agents have been used to treat colorectal cancer. Despite the advent of new treatment regimens, the 5-year survival rate for metastatic colorectal cancer remains low (~10%). Knowing the drug sensitivity of a given tumor for a particular agent could significantly impact decision making and treatment planning. Biomarkers are proven to be successful in characterizing patients into different response groups. Using survival prediction analysis, we have identified three independent gene signatures, which are associated with sensitivity of colorectal cancer cells to 5-FU, oxaliplatin or irinotecan. On the basis of the three gene signatures, three score systems were developed to stratify patients from sensitive to resistance. These score systems exhibited robustness in stratify patients in two independent clinical studies. Patients with high scores in all three drugs exhibited the lowest survival.
Similar content being viewed by others
Introduction
Colorectal cancer is the third most common cancer in the world and the second leading cause of cancer-related death in the western world.1,2 Around a quarter of colorectal cancer patients are incurable at diagnosis and half of the patients who undergo potentially curative surgery will ultimately develop metastatic disease.
In many cases, chemotherapy is used in treating colorectal cancer, which aims to slow tumor growth, shrink tumor size and reduce the likelihood of metastasis development. The standard treatment for advanced colorectal cancer is based on the administration of fluoropyrimidines (5-fluorouracil (5-FU) or capecitabine) combined with oxaliplatin, the topoisomerase I (TOP1) inhibitor CPT-11 (Irinotecan) and the monoclonal antibodies cetuximab, bevacizumab or panitunumab.2,3 Although most patients with advanced colorectal cancer are initially responsive to the combined chemotherapy treatment, they later experience disease relapse due to eventual tumor recurrence and emergence of drug-resistant tumor cells.
Gaining insight of the mechanisms underlying drug resistance is important to develop more effective therapeutic approaches.2 Human cancers may be resistant to therapy at the time of drug presentation (innate drug resistance). Some cancers become resistant after an initial response (acquired drug resistance). Both innate and acquired drug resistance involve multiple mechanisms, such as altering drug metabolite potency, increasing drug efflux or decreasing drug toxicity or inhibiting cell death.4 In colorectal cancer, higher level of thymidylate synthase were found associated with tumor insensitivity to 5-FU-based therapy.5 Similarly, higher levels of TOP1 is correlated with greater sensitivity of colon tumors to camptothecin derivatives compared with normal colonic mucosa.5 Glucuronidation, involved in xenobiotic detoxification, regulates innate resistance to TOP1 inhibitors in colon cell lines and tumors.6 The resistance to oxaliplatin involves decreased drug accumulation, increased detoxification and repair, enhanced tolerance to damage, alteration in pathways involved in cell cycle kinetics and apoptosis inactivation.7 In addition, overexpression of specific drug transporters (ABCB1/P-gp, lung resistance-related protein or multidrug resistance-related protein) was shown by flow cytometry and fluorescence microscopy to occur in human colon adenocarcinoma cell lines resistant to TOP1 inhibitors. Despite the mechanisms identified, implication of these biomarkers in clinic was not confirmed. The only clinically used biomarker is K-ras. Patients harboring a K-ras mutation are excluded from being treated with epidermal growth factor receptor antibodies, as they are less likely to benefit from epidermal growth factor receptor-targeted treatment.8
Microarray technology has been widely used in biomarker discovery and clinical outcome prediction.9,10 We analyzed microarray data derived from colon cancer cells resistant to oxaliplatin, SN38 (the active metabolite of irinotecan) and 5-FU, respectively. In order to prioritize the genes regulating cancer cell response to these anti-cancer drugs, we combined microarray data of drug resistance with data of patient survival. Three gene signatures were identified for these three respective drugs. Score systems were developed for each drug based on the gene signatures. The score systems were able to stratify cancer patients into high- or poor-survival groups.
Materials and methods
Microarray data
Microarray data derived from oxaliplatin-resistant HCT116 (HCT116-Oxa), 5-FU-resistant HCT116 (HCT116-FU), SN38-resistant HCT116 (HCT116-SN) and corresponding parent cell lines were downloaded from ArrayExpress (E-MEXP-390 and E-MEXP-1171).11,12 The information of the cell lines were described previously.11,12 Microarray data with patient survival information were obtained from Gene Expression Omnibus using accessing number GSE17536 (training data set) and GSE14333 (validation data set).13,14
Gene expression analysis
Microarray files were processed using Robust Multi-array Average algorithm and imported into Partek software.15 Principle Component Analysis and sample clustering were used to inspect for the existence of batch effects, outlier and errors. Batch effects were removed using software package Partek. To determine differentially expressed genes, data were analyzed using two-way analysis of variance. The differentially expressed genes (P<0.005) were further subjected to Gene Ontology analysis. Pathway analysis was performed using Ingenuity software (www.ingenuity.com). To detect survival-related genes, univariate Cox regress analyses for the differentially expressed genes were performed using Partek.
Development and validation of a drug-resistant score system
To generate a score system for drug resistance (Table 1), we subject the candidate gene lists to BRB-arrayTools (developed by Dr Richard Simon and BRB-ArrayTools Development Team) to calculate the regression coefficient for each gene using the training data set. The drug-resistant score is the sum of the product of the expression level of a gene and its corresponding regression coefficient (drug-resistant score=sum of coefficient of Gene Gi × expression value of Gene Gi). The patients were dichotomized into groups at high or low risk using the 50th percentile (median). The coefficient derived from the training data set was directly applied to the validation data set.
Results
Differentially expressed genes derived from the 5-FU-resistant cell line
Genes (363) (P<0.005) were found to be differentially expressed between HCT116-FU and HCT116 parent cell lines. Hundred of these genes are known to regulate drug resistance in cancer, with 32 upregulated and 68 downregulated. Upregulated genes include IGF1R, NQO1, ABCC3, FOXO3, MLL, LGALS3, TOX, GNAS, IGFBP4, AKR1C2, MSTN, SULF2 and EGR2. Downregulated genes include CFLAR, FANCA, TRPM2, SLC19A1, RRM1, FOXA2, CASR, MTAP, BACE1, ELOVL6, NF2, FBXW7, DPH5 and FOLR1.
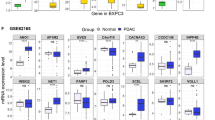
Gene Ontology analysis (Figure 1a1) indicated that these 363 genes are enriched in functions involving metabolic process (53.6%), growth (27.7%), pigmentation (12%), viral reproduction (3.8%) and cell proliferation (2.3%). Ingenuity pathway analysis indicated that the top five enriched signaling networks (Figure 1a2) were cellular assembly and organization, cellular movement, gastrointestinal disease; cell death, cellular development, hematopoiesis; cellular growth and proliferation, protein synthesis, cell cycle; cell-mediated immune response, cellular development, cellular function and maintenance; and free radical scavenging, cellular movement, gene expression. The core nodes in the signaling networks include ERK1/2, NFKB complex, Akt, PI3K complex and Ras (Supplementary Data 1, Figure 1).
(a1) The differentially expressed genes between 5-fluorouracil (5-FU)-resistant and control HCT116 cells were subjected to oncology analysis. These genes can be categorized based on their functions into metabolic process (53.6%), growth (27.7%), pigmentation (12%), viral reproduction (3.8%) and cell proliferation (2.3%). (a2) The differentially expressed genes between 5-FU-resistant and control HCT116 cells were subjected to ingenuity pathway analysis and the top five enriched signaling networks were obtained. (b1) Gene Ontology analysis indicated that these differentially expressed genes between oxaliplatin-resistant and control HCT116 cells were enriched in functions involving rhythmic process (31.7%), viral reproduction (27.2%), cellular process (26.2%), cell proliferation (7.7%) and metabolic process (6.1%). (b2) The differentially expressed genes between oxaliplatin-resistant and control HCT116 cells were subjected to ingenuity pathway analysis and the top five enriched signaling networks were obtained. (c1) Gene Ontology analysis indicated that the differentially expressed genes between SN38-resistant and control HCT116 cells were enriched in functions involving cell proliferation (39.7%), response to stimulus (16.1%), growth (15.9%), rhythmic process (11%), viral reproduction (8.6%) and cellular process (8.4%). (c2) The differentially expressed genes between SN38-resistant and control HCT116 cells were subjected to ingenuity pathway analysis and the top five enriched signaling networks were obtained.
Differentially expressed genes derived from the oxaliplatin-resistant cell line
Genes (373) (P<0.005) were found to be differentially expressed between HCT116-Oxa and HCT116 parent cell lines. Genes (104) are known to regulate drug resistance in cancer, with 37 upregulated and 67 downregulated genes. Upregulated genes are CD24, CD80, FANCD2, IL2, MMP2, HLF, CYP1B1, DGAT1, PRKCQ, STAT5B, AKT3, RICTOR, CCKBR and FLT1. Downregulated genes includes DHFR, DIABLO, EIF4E, PPP2CB, LYN, ADO, CD59, LDLR, SMAD4, TYMS, ARNT, NR3C1, MINA, MTAP, WEE1, B2M, PRKAR1A, MGST1, RAB18, HOXA9, MCAM, TUBB, ENO1, SMARCA4, BCL2L11, ELP3, CREM, MERTK, CENPV and ARG2.
Gene Ontology analysis (Figure 1b1) indicated that these 373 genes were enriched in functions involving rhythmic process (31.7%), viral reproduction (27.2%), cellular process (26.2%), cell proliferation (7.7%) and metabolic process (6.1%). Ingenuity analysis indicated that the top five enriched signaling networks (Figure 1b2) were cellular assembly and organization, cell morphology, cellular movement; cell cycle, lipid metabolism, small molecule biochemistry; cell cycle, gene expression, cell-to-cell signaling and interaction; cellular assembly and organization, DNA replication, recombination, and repair, connective tissue disorders; and RNA post-transcriptional modification, cell death, protein synthesis. The core nodes in the signaling networks include PI3K complex, ERK1/2, Akt and IL2 (Supplementary Data 1, Figure 2).
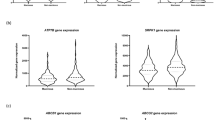
Venn diagram analysis. (a) Venn diagram of upregulated genes in 5-fluorouracil (5-FU), oxaliplatin or SN38-resistant HCT116 cells. (b) Venn diagram of downregulated genes in 5-FU, oxaliplatin or SN38-resistant HCT116 cells.
Differentially expressed genes derived from the SN38-resistant cell line
Genes (692) (P<0.005) were found to be differentially expressed between HCT116-SN and HCT116 parent cell lines. Genes (196) are known to regulate drug resistance in cancer, with 108 upregulated genes and 88 downregulated genes. Upregulated genes include HBA1, SRC, ICAM1, CD44, ANXA8, APOE, TRH, GALE, CSF1R, IGF1, NQO1, NCAM1, LOX, ERBB3, SAT1, LDLR, RET, GRP, NOS1, CDK1, IGH and CASP10. Downregulated genes include IRS1, HPRT1, GAL, VEGFA, CYP3A5, MSH3, MTR, GHR, CYP2B6, ZAK, MYB, LDLR, SOD2, DTNB, NCOR1, NR3C1, GCLC, MINA, TFPI, LPP, ADAM17, MDM4, RIPK1 and AQP3.
Gene Ontology analysis (Figure 1c1) indicated that these 496 genes were enriched in functions involving cell proliferation (39.7%), response to stimulus (16.1%), growth (15.9%), rhythmic process (11%), viral reproduction (8.6%) and cellular process (8.4%). Ingenuity analysis indicated that the top five enriched signaling networks (Figure 1c2) were: cell death, cellular compromise, antimicrobial response; cellular function and maintenance, molecular transport, cell signaling; gene expression, DNA replication, recombination, and repair, cell-to-cell signaling and interaction; cell cycle, reproductive system development and function, cell-to-cell signaling and interaction; and cellular growth and proliferation, skeletal and muscular system development and function, lipid metabolism. The core nodes in the signaling networks include NFKB complex, CACNA1A, SMARCA4, PI3K complex and MAPK (Supplementary Data 1, Figures 3, 4, 5).
Kaplan–Meier curves were drawn for conditions as follows. (a) Patients in the training data set were dichotomized into high (H_FU) and low (L_FU) drug-resistant groups based on the resistance score of 5-fluorouracil (5-FU) using the 50th percentile of the score as cutoff. (b) Patients in the training data set were similarly dichotomized into high (H_Oxa) and low (L_Oxa) drug-resistant groups based on the resistance score of oxaliplatin. (c) Patients in the training data set were similarly dichotomized into high (H_SN) and low (L_SN) drug resistance groups based on the resistance score of SN38.
Kaplan–Meier curves were drawn for conditions as follows. (a) Patients in the validation data set were similarly dichotomized into high (H_FU) and low (L_FU) drug-resistant groups based on the resistance score of 5-fluorouracil (5-FU) (b) Patients in the validation data set were similarly dichotomized into high (H_Oxa) and low (L_Oxa) drug-resistant groups based on the resistance score of oxaliplatin. (c) Patients in the validation data set were similarly dichotomized into high (H_SN) and low (L_SN) drug-resistant groups based on resistance score of SN38.
Kaplan–Meier curves were drawn for conditions as follows. (a) Patients in the training data set were grouped into four groups based on the number of high-resistance scores the patients have (3, the patients are in three high-resistance groups; 2, the patients are in two high-resistance groups; 1, the patients are in one high-resistance group; 0, the patients are in low-resistance groups). (b) Patients in the validation data set were grouped into four groups based on the number of high-resistance scores the patients have (3, the patients are in three high-resistance groups; 2, the patients are in two high-resistance groups; 1, the patients are in one high-resistance group; 0, the patients are in low-resistance groups).
Difference between the three gene lists
There are limited overlaps between differentially expressed genes derived from these three drug-resistant cell lines (Figure 2a). Only one overlapping gene (ANO8) was found between the 134 upregulated genes derived from HCT116-FU and 361 upregulated genes derived from HCT116-SN cells. Only two overlapping genes (C15orf28 and FBXW2) were found between the 229 downregulated genes derived from HCT116-FU and 331 downregulated genes derived from HCT116-SN cells. In the 143 upregulated genes derived from HCT116-Oxa cells and 361 upregulated genes derived from HCT116-SN cells, only two genes (LOC257358 and C1orf61) overlap. Between the 230 downregulated genes derived from HCT116-Oxa cells and 331 downregulated genes derived from HCT116-SN cells, there were only five genes (MINA, CLASP2, SFRS7, ZBTB20 and C2orf69). In the 134 upregulated genes derived from HCT116-FU cells and 143 upregulated genes derived from HCT116-Oxa cells, there was no overlapping gene. In the 229 downregulated genes derived from HCT116-FU cells and 230 downregulated genes derived from HCT116-Oxa cells, there was only one overlapping gene (MTAP). Among these 11 overlapping genes, the functions of 4 genes are unknown, including LOC257358, C1orf61, C15orf28 and C2orf69, whereas the other 7 genes exhibited widespread functions. ZBTB20 exhibits RNA polymerase II core promoter proximal region sequence-specific DNA-binding transcription factor activity involved in negative regulation of transcription and DNA binding.16 SFRS7 is a member of the serine/arginine-rich family of pre-mRNA-splicing factors, which constitute part of the spliceosome.17 CLASP2 exhibits microtubule plus-end binding and galactoside 2-alpha-L-fucosyltransferase activity.18 MINA is a c-Myc target gene that may have a role in cell proliferation or regulation of cell growth.19 FBXW2 regulates chaperonin-mediated protein folding and Wnt signaling pathway and pluripotency.20 MTAP is associated with metabolic pathways and cysteine and methionine metabolism, and exhibits S-methyl-5-thioadenosine phosphorylase activity and phosphorylase activity.21 ANO8 is related to stimuli-sensing channels and solute-carrier-mediated transmembrane transport, and possesses intracellular calcium-activated chloride channel activity.22
Gene ontology analysis (Figure 1a) indicated that there are different levels of enrichment in functions including metabolic process (53.6% for HCT116-FU cells and 6.1% for HCT116-Oxa cells), growth (28% for HCT116-FU cells and 15.9% for HCT116-SN cells), viral reproduction (3.8% for HCT116-FU cells, 27% for HCT116-Oxa cells and 8.6% for HCT116-SN cells), cell proliferation (2.3% for HCT116-FU cells, 7.7% for HCT116-Oxa cells and 39% for HCT116-SN cells), rhythmic process (32% for HCT116-Oxa cells and 11% for HCT116-SN cells) and cellular process (26% for HCT116-Oxa cells and 8.4% for HCT116-SN cells). Network analysis indicated that PI3K–Akt and MAPK–ERK pathways are altered in all three drug-resistant cell lines and the NFKB pathway is altered in HCT116-FU and HCT116-SN cells. However, the genes in each signaling network are totally different between cells with different drug resistance (Figures 1b and d).
Genes correlate to colorectal cancer patient survival
We analyzed the differentially expressed genes for their association with patients’ survival. In the 363 differentially expressed genes between HCT116-FU and its corresponding cell lines, 10 genes are significantly associated with patient survival (P<0.01), with 6 positively associated and 4 negatively associated. The positively correlated genes are CLIC4, CTHRC1, COL3A1, IER5L, IGFBP4 and EGR2. The negatively correlated genes are FERMT1, FN3K, ASB2 and TRPM2. In the 373 differentially expressed genes between HCT116-Oxa and its corresponding cell lines, 18 are significantly associated with patient survival (P<0.01), with 16 positively correlated and 2 negatively correlated. The positively correlated genes are FLT1, CYP1B1, KCNJ8, AKT3, STC1, DOCK4, LTBP1, CD59, RAB31, SERPINH1, MCAM, GLG1, FERMT2, RASSF8, CHST14 and NOG. The negatively correlated genes are SCFD2 and WDR77. In the 692 differentially expressed genes between HCT116-FU and its corresponding cell lines, 19 are significantly associated with patient survival (P<0.01), with 11 positively correlated and 8 negatively correlated. The positively correlated genes include RAB3B, PLAUR, APOE, VEGFA, LDLR, EPYC, RHOQ, LOX, FTH1, NUAK1 and TCEAL3. The negatively correlated genes include LRRC36, NUAK2, DTYMK, TSPAN11, AXIN2, CCDC13, C1orf151 and ANAPC5.
Development of the drug-resistant score system
To assess contribution of drug resistance-related genes to patient survival, we applied survival prediction analysis in the BRB-arrayTools. Each gene in the respective gene signature was assigned a coefficient, and the score for each sample was calculated using the coefficient of these genes. The patients were then dichotomized into a high- and low-drug-resistant group for drug resistance using the 50th percentile cutoff of the score as the threshold value. The results indicated that patient survival rates were significantly lower in the patient group with a higher drug-resistant score using the drug-resistant score system for all three drugs (5-FU, 62.5 vs 87.7%, P<2.5E-4; oxaliplatin, 56.9 vs 93.2%, P<5.3E-7 and SN38, 52.8 vs 97.3%, P<2E-10). To further validate the score systems, we used an independent data set derived from 290 patients. The drug-resistant score was similarly calculated and the patients were similarly dichotomized into a high- and low-drug-resistant group using the 50th percentile cutoff of the score as the threshold value. Results indicated that the drug-resistant score system was able to separate patients with good prognosis (high survival rates) from these with poor prognosis (low survival rates) (5-FU, 65.8 vs 87.9%, P<2E-4; oxaliplatin, 66.7 vs 87%, P<4.2E-4 and SN38, 65.8 vs 87.9%, P<1.6E-4).
As colon cancer is usually treated with a combination of drugs, we assume that the more drug the cancer cells are resistant to, the more likely the patient exhibits a lower chance of survival. We therefore combined the drug-resistant scores for 5-FU, oxaliplatin and SN38 to see whether we could further separate the patients into subgroups with different risks. The patients were categorized into four groups based on the number of drugs they predicted to be at high risk of resistance (high risk of 3, 2, 1 or 0 drug resistance). In the 177-patient data set, the patients with high risk of three drug resistances exhibit the lowest survival rates (42.7%) and the patients with no risk of drug resistance exhibit highest survival rates. In the 290-patient data set, the patients with high risk of three drug resistances exhibit the lowest survival rates (54.1%) and the patients with no risk of drug resistance exhibit the highest survival rates (95.4%).
Discussion
Knowing the drug sensitivity of a given tumor for a particular agent could significantly impact decision making and treatment planning. Starting from the gene expression pattern of drug-resistant cells, we developed three drug-resistance-based score systems to rank the patients’ response to three first-line anti-cancer drugs (5-FU, oxaliplatin and irinotecan) for colorectal cancer treatment. Patients with a high drug resistance score exhibited poor survival. Patients with a high score of all three drugs exhibited the poorest survival.
The gene expression patterns of these drug-resistant cell lines are distinct, which provide a strong basis to develop specific drug-resistant gene signatures. As genetic background/mutation in cancer cell lines may contribute to resistance to the tested drugs, we used data derived from one single cell line HCT116. No overlapping gene was found in all the three gene lists. The distinct gene expression patterns also indicated that different drug resistance mechanisms for drugs with different cytotoxic mechanisms. Indeed, cytotoxicity of 5-FU is due to misincorporation of fluoronucleotides into RNA and DNA and to the inhibition of the nucleotide synthetic enzyme thymidylate synthase.23 Oxaliplatin (1,2-diaminocyclohexane-oxalate platinum) mainly forms intrastrand adducts between two adjacent guanine residues or guanine and adenine, disrupting DNA replication and transcription.24 Irinotecan interacts with cellular Top1-DNA complexes and has S-phase-specific cytotoxicity.25
A large portion of the differentially expressed genes is related to drug resistance, indicating that the identified genes recapitulate features of drug resistance. For example, IGF1R, ABCC3 and FOXO3 are associated with drug resistance26, 27, 28 and overexpressed in 5-FU-resistant cells. Expression levels of drug-resistant-related genes such as CD24, FANCD2, CYP1B1 and STAT5B were increased in oxaliplatin-resistant cells.29, 30, 31, 32 Similarly, SRC, ICAM1, CD44, IGF1, ERBB3, RET and CDK1 correlate with drug resistance33, 34, 35, 36, 37, 38, 39 and were overexpressed in SN38-resistant cells to regulate drug resistance. Interestingly, although gene expression patterns are distinct in these drug-resistant cell lines, pathway analysis indicated that the drugs share some common core nodes of significantly changed pathways. For example, PI3K–Akt, MAPK–ERK pathways were altered in all three drug-resistant cell lines and the NFKB pathway was altered in HCT116-FU and HCT116-SN cells. The PI3K–Akt signal transduction pathway comprises of the lipid kinase, phosphatidylinositol 3-kinase (PI3K) and the serine/threonine kinase, Akt (or PKB). Activation of this pathway has a pivotal role in essential cellular functions, such as survival, proliferation, migration and differentiation that underlie the biology of human cancer.40 The Raf/MEK/ERK pathway influences chemotherapeutic drug resistance as ectopic activation of Raf induces resistance to doxorubicin and paclitaxel in breast cancer cells.41 NFKB transcription factor induces drug resistance through MDR1 expression in cancer cells.42 However, there is a lack of explicit link of gene functions to corresponding resistance to a particular drug. Caution must be taken to interpret the results.
Other molecular signatures have been reported in colon cancer. For example, a molecular signature for oncogenic BRAF is reported in human colon cancer cells.43 Colon cancer molecular subtypes were identified by expression profiling and associated with stroma, mucinous type and different clinical behavior.44 Gene expression profiling of peritoneal metastases from appendiceal and colon cancer demonstrates unique biologic signatures and predicts patient outcomes.45 Intestinal adenomagenesis involves core molecular signatures of the epithelial–mesenchymal transition.46 The new discovered drug-resistant gene signatures add new knowledge to this field and a combination of these gene signatures may reveal new mechanisms in colorectal cancer. However, the cellular model-based gene signatures may not represent all features of clinical samples. The results are preliminary and require further validation in future studies involving tumor samples from patients. Combining the drug-resistant scores derived from each single agent might not reflect reality in patients who receive these drugs together. Caution should be taken to interpret the gene-outcome correlations.
In conclusion, we identified candidate genes, gene sets and pathways related to drug resistance in colorectal cancer, which warrant further investigation. The drug-resistant score systems are able to stratify patients into good or poor survival groups, which warrant further validation.
References
Siegel R, Naishadham D, Jemal A . Cancer statistics, 2013. CA Cancer J Clin 2013; 63: 11–30.
Cartwright TH . Treatment decisions after diagnosis of metastatic colorectal cancer. Clin Colorectal Cancer 2012; 11: 155–166.
Macedo LT, da Costa Lima AB, Sasse AD . Addition of bevacizumab to first-line chemotherapy in advanced colorectal cancer: a systematic review and meta-analysis, with emphasis on chemotherapy subgroups. BMC Cancer 2012; 12: 89.
Donnenberg VS, Donnenberg AD . Multiple drug resistance in cancer revisited: the cancer stem cell hypothesis. J Clin Pharmacol 2005; 45: 872–877.
Jensen NF, Smith DH, Nygård SB, Rømer MU, Nielsen KV, Brünner N . Predictive biomarkers with potential of converting conventional chemotherapy to targeted therapy in patients with metastatic colorectal cancer. Scand J Gastroenterol 2012; 47: 340–355.
Graudens E, Boulanger V, Mollard C, Mariage-Samson R, Barlet X, Grémy G et al. Deciphering cellular states of innate tumor drug responses. Genome Biol 2006; 7: R19.
Alcindor T, Beauger N . Oxaliplatin: a review in the era of molecularly targeted therapy. Curr Oncol 2011; 18: 18–25.
Chen J, Ye Y, Sun H, Shi G . Association between KRAS codon 13 mutations and clinical response to anti-EGFR treatment in patients with metastatic colorectal cancer: results from a meta-analysis. Cancer Chemother Pharmacol 2013; 71: 265–272.
Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 2004; 351: 2817–2826.
van de Vijver MJ, He YD, van't Veer LJ, Dai H, Hart AA, Voskuil DW et al. A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 2002; 347: 1999–2009.
Boyer J, Allen WL, McLean EG, Wilson PM, McCulla A, Moore S et al. Pharmacogenomic identification of novel determinants of response to chemotherapy in colon cancer. Cancer Res 2006; 66: 2765–2777.
Allen WL, Turkington RC, Stevenson L, Carson G, Coyle VM, Hector S et al. Pharmacogenomic profiling and pathway analyses identify MAPK-dependent migration as an acute response to SN38 in p53 null and p53-mutant colorectal cancer cells. Mol Cancer Ther 2012; 11: 1724–1734.
Jorissen RN, Gibbs P, Christie M, Prakash S, Lipton L, Desai J et al. Metastasis-associated gene expression changes predict poor outcomes in patients with Dukes stage B and C colorectal cancer. Clin Cancer Res 2009; 15: 7642–7651.
Smith JJ, Deane NG, Wu F, Merchant NB, Zhang B, Jiang A et al. Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 2010; 138: 958–968.
Tong Y, Zhou J, Mizutani J, Fukuoka H, Ren SG, Gutierrez-Hartmann A et al. CEBPD suppresses prolactin expression and prolactinoma cell proliferation. Mol Endocrinol 2011; 25: 1880–1891.
Peterson ML, Ma C, Spear BT . Zhx2 and Zbtb20: novel regulators of postnatal alpha-fetoprotein repression and their potential role in gene reactivation during liver cancer. Semin Cancer Biol 2011; 21: 21–27.
van Abel D, Hölzel DR, Jain S, Lun FM, Zheng YW, Chen EZ et al. SFRS7-mediated splicing of tau exon 10 is directly regulated by STOX1A in glial cells. PLoS One 2011; 6: e21994.
Al-Bassam J, Chang F . Regulation of microtubule dynamics by TOG-domain proteins XMAP215/Dis1 and CLASP. Trends Cell Biol 2011; 21: 604–614.
Komiya K, Sueoka-Aragane N, Sato A, Hisatomi T, Sakuragi T, Mitsuoka M et al. Mina53, a novel c-Myc target gene, is frequently expressed in lung cancers and exerts oncogenic property in NIH/3T3 cells. J Cancer Res Clin Oncol 2010; 136: 465–473.
Chiang MH, Chen LF, Chen H . Ubiquitin-conjugating enzyme UBE2D2 is responsible for FBXW2 (F-box and WD repeat domain containing 2)-mediated human GCM1 (glial cell missing homolog 1) ubiquitination and degradation. Biol Reprod 2008; 79: 914–920.
Bertino JR, Waud WR, Parker WB, Lubin M et al. Targeting tumors that lack methylthioadenosine phosphorylase (MTAP) activity: current strategies. Cancer Biol Ther 2011; 11: 627–632.
Kunzelmann K, Schreiber R, Kmit A, Jantarajit W, Martins JR, Faria D et al. Expression and function of epithelial anoctamins. Exp Physiol 2012; 97: 184–192.
Longley DB, Harkin DP, Johnston PG . 5-fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer 2003; 3: 330–338.
Martinez-Cardus A, Martinez-Balibrea E, Bandrés E, Malumbres R, Ginés A, Manzano JL et al. Pharmacogenomic approach for the identification of novel determinants of acquired resistance to oxaliplatin in colorectal cancer. Mol Cancer Ther 2009; 8: 194–202.
Xu Y, Villalona-Calero MA . Irinotecan: mechanisms of tumor resistance and novel strategies for modulating its activity. Ann Oncol 2002; 13: 1841–1851.
Bose D, Zimmerman LJ, Pierobon M, Petricoin E, Tozzi F, Parikh A et al. Chemoresistant colorectal cancer cells and cancer stem cells mediate growth and survival of bystander cells. Br J Cancer 2011; 105: 1759–1767.
O'Brien C, Cavet G, Pandita A, Hu X, Haydu L, Mohan S et al. Functional genomics identifies ABCC3 as a mediator of taxane resistance in HER2-amplified breast cancer. Cancer Res 2008; 68: 5380–5389.
Hui RC, Francis RE, Guest SK, Costa JR, Gomes AR, Myatt SS et al. Doxorubicin activates FOXO3a to induce the expression of multidrug resistance gene ABCB1 (MDR1) in K562 leukemic cells. Mol Cancer Ther 2008; 7: 670–678.
Muraro MG, Mele V, Däster S, Han J, Heberer M, Cesare Spagnoli G et al. CD133+, CD166+CD44+, and CD24+CD44+ phenotypes fail to reliably identify cell populations with cancer stem cell functional features in established human colorectal cancer cell lines. Stem Cells Transl Med 2012; 1: 592–603.
Kachnic LA, Li L, Fournier L, Willers H . Fanconi anemia pathway heterogeneity revealed by cisplatin and oxaliplatin treatments. Cancer Lett 2010; 292: 73–79.
McFadyen MC, McLeod HL, Jackson FC, Melvin WT, Doehmer J, Murray GI . Cytochrome P450 CYP1B1 protein expression: a novel mechanism of anticancer drug resistance. Biochem Pharmacol 2001; 62: 207–212.
Warsch W, Kollmann K, Eckelhart E, Fajmann S, Cerny-Reiterer S, Hölbl A et al. High STAT5 levels mediate imatinib resistance and indicate disease progression in chronic myeloid leukemia. Blood 2011; 117: 3409–3420.
Chen X, Yeung TK, Wang Z . Enhanced drug resistance in cells coexpressing ErbB2 with EGF receptor or ErbB3. Biochem Biophys Res Commun 2000; 277: 757–763.
Cain JW, Hauptschein RS, Stewart JK, Bagci T, Sahagian GG, Jay DG . Identification of CD44 as a surface biomarker for drug resistance by surface proteome signature technology. Mol Cancer Res 2011; 9: 637–647.
Chong K, Subramanian A, Sharma A, Mokbel K . Measuring IGF-1, ER-alpha and EGFR expression can predict tamoxifen-resistance in ER-positive breast cancer. Anticancer Res 2011; 31: 23–32.
Skinner MA, Lackey KE, Freemerman AJ . RET activation inhibits doxorubicin-induced apoptosis in SK-N-MC cells. Anticancer Res 2008; 28: 2019–2025.
Pene-Dumitrescu T, Smithgall TE . Expression of a Src family kinase in chronic myelogenous leukemia cells induces resistance to imatinib in a kinase-dependent manner. J Biol Chem 2010; 285: 21446–21457.
Zheng Y, Yang J, Qian J, Qiu P, Hanabuchi S, Lu Y et al. PSGL-1/selectin and ICAM-1/CD18 interactions are involved in macrophage-induced drug resistance in myeloma. Leukemia 2013; 27: 702–710.
Flint MS, Kim G, Hood BL, Bateman NW, Stewart NA, Conrads TP . Stress hormones mediate drug resistance to paclitaxel in human breast cancer cells through a CDK-1-dependent pathway. Psychoneuroendocrinology 2009; 34: 1533–1541.
West KA, Castillo SS, Dennis PA . Activation of the PI3K/Akt pathway and chemotherapeutic resistance. Drug Resist Updat 2002; 5: 234–248.
McCubrey JA, Steelman LS, Chappell WH, Abrams SL, Wong EW, Chang F et al. Roles of the Raf/MEK/ERK pathway in cell growth, malignant transformation and drug resistance. Biochim Biophys Acta 2007; 1773: 1263–1284.
Bentires-Alj M, Barbu V, Fillet M, Chariot A, Relic B, Jacobs N et al. NF-kappaB transcription factor induces drug resistance through MDR1 expression in cancer cells. Oncogene 2003; 22: 90–97.
Joyce T, Oikonomou E, Kosmidou V, Makrodouli E, Bantounas I, Avlonitis S et al. A molecular signature for oncogenic BRAF in human colon cancer cells is revealed by microarray analysis. Curr Cancer Drug Targets 2012; 12: 873–898.
Perez-Villamil B, Romera-Lopez A, Hernandez-Prieto S, Lopez-Campos G, Calles A, Lopez-Asenjo JA et al. Colon cancer molecular subtypes identified by expression profiling and associated to stroma, mucinous type and different clinical behavior. BMC Cancer 2012; 12: 260.
Levine EA, Blazer DG 3rd, Kim MK, Shen P, Stewart JH 4th, Guy C . et al. Gene expression profiling of peritoneal metastases from appendiceal and colon cancer demonstrates unique biologic signatures and predicts patient outcomes. J Am Coll Surg 2012; 214: 599–606.
Chen X, Halberg RB, Burch RP, Dove WF . Intestinal adenomagenesis involves core molecular signatures of the epithelial-mesenchymal transition. J Mol Histol 2008; 39: 283–294.
Acknowledgements
This manuscript is supported by the National Cancer Institute of the National Institutes of Health under award number R00CA138914 (YT) and National Natural Science Foundation under grant number 81372216 (YT).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Zheng, Y., Zhou, J. & Tong, Y. Gene signatures of drug resistance predict patient survival in colorectal cancer. Pharmacogenomics J 15, 135–143 (2015). https://doi.org/10.1038/tpj.2014.45
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2014.45
This article is cited by
-
PPARG activation promotes the proliferation of colorectal cancer cell lines and enhances the antiproliferative effect of 5-fluorouracil
BMC Cancer (2024)
-
Bioinformatics screening of colorectal-cancer causing molecular signatures through gene expression profiles to discover therapeutic targets and candidate agents
BMC Medical Genomics (2023)
-
Identification of ferroptosis and drug resistance related hub genes to predict the prognosis in Hepatocellular Carcinoma
Scientific Reports (2023)
-
A multiparametric extraction method for Vn96-isolated plasma extracellular vesicles and cell-free DNA that enables multi-omic profiling
Scientific Reports (2021)
-
Dissecting the m6A methylation affection on afatinib resistance in non-small cell lung cancer
The Pharmacogenomics Journal (2020)