Abstract
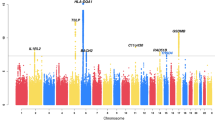
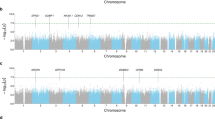
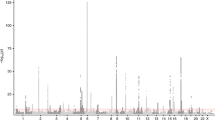
Reversibility of airway obstruction in response to β2-agonists is highly variable among asthmatics, which is partially attributed to genetic factors. In a genome-wide association study of acute bronchodilator response (BDR) to inhaled albuterol, 534 290 single-nucleotide polymorphisms (SNPs) were tested in 403 white trios from the Childhood Asthma Management Program using five statistical models to determine the most robust genetic associations. The primary replication phase included 1397 polymorphisms in three asthma trials (pooled n=764). The second replication phase tested 13 SNPs in three additional asthma populations (n=241, n=215 and n=592). An intergenic SNP on chromosome 10, rs11252394, proximal to several excellent biological candidates, significantly replicated (P=1.98 × 10−7) in the primary replication trials. An intronic SNP (rs6988229) in the collagen (COL22A1) locus also provided strong replication signals (P=8.51 × 10−6). This study applied a robust approach for testing the genetic basis of BDR and identified novel loci associated with this drug response in asthmatics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Masoli M, Fabian D, Holt S, Beasley R . The global burden of asthma: executive summary of the GINA Dissemination Committee report. Allergy 2004; 59: 469–478.
Sears MR, Lotvall J . Past, present and future–beta2-adrenoceptor agonists in asthma management. Respir Med 2005; 99: 152–170.
Drazen JM, Silverman EK, Lee TH . Heterogeneity of therapeutic responses in asthma. Br Med Bull 2000; 56: 1054–1070.
Barnett SB, Nurmagambetov TA . Costs of asthma in the United States: 2002–2007. J Allergy Clin Immunol 2011; 127: 145–152.
Litonjua AA, Lasky-Su J, Schneiter K, Tantisira KG, Lazarus R, Klanderman B et al. ARG1 is a novel bronchodilator response gene: screening and replication in four asthma cohorts. Am J Respir Crit Care Med 2008; 178: 688–694.
Martinez FD, Graves PE, Baldini M, Solomon S, Erickson R . Association between genetic polymorphisms of the beta2-adrenoceptor and response to albuterol in children with and without a history of wheezing. J Clin Invest 1997; 100: 3184–3188.
Green SA, Turki J, Innis M, Liggett SB . Amino-terminal polymorphisms of the human beta 2-adrenergic receptor impart distinct agonist-promoted regulatory properties. Biochemistry 1994; 33: 9414–9419.
Lima JJ, Thomason DB, Mohamed MH, Eberle LV, Self TH, Johnson JA . Impact of genetic polymorphisms of the beta2-adrenergic receptor on albuterol bronchodilator pharmacodynamics. Clin Pharmacol Ther 1999; 65: 519–525.
Martin AC, Zhang G, Rueter K, Khoo SK, Bizzintino J, Hayden CM et al. Beta2-adrenoceptor polymorphisms predict response to beta2-agonists in children with acute asthma. J Asthma 2008; 45: 383–388.
Poon AH, Tantisira KG, Litonjua AA, Lazarus R, Xu J, Lasky-Su J et al. Association of corticotropin-releasing hormone receptor-2 genetic variants with acute bronchodilator response in asthma. Pharmacogenet Genomics 2008; 18: 373–382.
Tantisira KG, Small KM, Litonjua AA, Weiss ST, Liggett SB . Molecular properties and pharmacogenetics of a polymorphism of adenylyl cyclase type 9 in asthma: interaction between beta-agonist and corticosteroid pathways. Hum Mol Genet 2005; 14: 1671–1677.
Himes BE, Jiang X, Hu R, Wu AC, Lasky-Su J, Klanderman B et al. Genome-wide association analysis in asthma subjects identifies SPATS2L as a novel bronchodilator response gene. PLoS Genet 2012; 8: e1002824.
The Childhood Asthma Management Program Research Group. Long-term effects of budesonide or nedocromil in children with asthma.. N Engl J Med 2000; 343: 1054–1063.
The Childhood Asthma Management Program Research Group. The Childhood Asthma Management Program (CAMP): design, rationale, and methods. Control Clin Trials 1999; 20: 91–120.
Himes BE, Hunninghake GM, Baurley JW, Rafaels NM, Sleiman P, Strachan DP et al. Genome-wide association analysis identifies PDE4D as an asthma-susceptibility gene. Am J Hum Genet 2009; 84: 581–593.
Baron RM, Palmer LJ, Tantisira K, Gabriel S, Sonna LA, Le L et al. DNA sequence variants in epithelium-specific ETS-2 and ETS-3 are not associated with asthma. Am J Respir Crit Care Med 2002; 166: 927–932.
Silverman ES, Palmer LJ, Subramaniam V, Hallock A, Mathew S, Vallone J et al. Transforming growth factor-beta1 promoter polymorphism C-509T is associated with asthma. Am J Respir Crit Care Med 2004; 169: 214–219.
Peters SP, Anthonisen N, Castro M, Holbrook JT, Irvin CG, Smith LJ et al. Randomized comparison of strategies for reducing treatment in mild persistent asthma. N Engl J Med 2007; 356: 2027–2039.
American Lung Association Asthma Clinical Research Centers. Clinical trial of low-dose theophylline and montelukast in patients with poorly controlled asthma. Am J Respir Crit Care Med 2007; 175: 235–242.
Israel E, Chinchilli VM, Ford JG, Boushey HA, Cherniack R, Craig TJ et al. Use of regularly scheduled albuterol treatment in asthma: genotype-stratified, randomised, placebo-controlled cross-over trial. Lancet 2004; 364: 1505–1512.
International HapMap Consortium. A haplotype map of the human genome. Nature 2005; 437: 1299–1320.
Willer CJ, Sanna S, Jackson AU, Scuteri A, Bonnycastle LL, Clarke R et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat Genet 2008; 40: 161–169.
Hunninghake GM, Soto-Quiros ME, Avila L, Su J, Murphy A, Demeo DL et al. Polymorphisms in IL13, total IgE, eosinophilia, and asthma exacerbations in childhood. J Allergy Clin Immunol 2007; 120: 84–90.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Lange C, DeMeo D, Silverman EK, Weiss ST, Laird NM . PBAT: tools for family-based association studies. Am J Hum Genet 2004; 74: 367–369.
Liptak T . On the combination of independent tests. Magyar Tud Akad Mat Kutato Int Kozl 1958; 3: 171–197, 171-197.
Du P, Kibbe WA, Lin SM . lumi: a pipeline for processing Illumina microarray. Bioinformatics 2008; 24: 1547–1548.
Tantisira KG, Fuhlbrigge AL, Tonascia J, Van Natta M, Zeiger RS, Strunk RC et al. Bronchodilation and bronchoconstriction: predictors of future lung function in childhood asthma. J Allergy Clin Immunol 2006; 117: 1264–1271.
Hawkins GA, Meyers DA, Bleecker ER, Pack AI . Identification of coding polymorphisms in human circadian rhythm genes PER1, PER2, PER3, CLOCK, ARNTL, CRY1, CRY2 and TIMELESS in a multi-ethnic screening panel. DNA Seq 2008; 19: 44–49.
Chaudhary D, Kasaian M . PKCtheta: a potential therapeutic target for T-cell-mediated diseases. Curr Opin Investig Drugs 2006; 7: 432–437.
Mosyak L, Xu Z, Joseph-McCarthy D, Brooijmans N, Somers W, Chaudhary D . Structure-based optimization of PKCtheta inhibitors. Biochem Soc Trans 2007; 35 (Pt 5): 1027–1031.
Cole DC, Asselin M, Brennan A, Czerwinski R, Ellingboe JW, Fitz L et al. Identification, characterization and initial hit-to-lead optimization of a series of 4-arylamino-3-pyridinecarbonitrile as protein kinase C theta (PKCtheta) inhibitors. J Med Chem 2008; 51: 5958–5963.
Ruckert R, Brandt K, Braun A, Hoymann HG, Herz U, Budagian V et al. Blocking IL-15 prevents the induction of allergen-specific T cells and allergic inflammation in vivo. J Immunol 2005; 174: 5507–5515.
Doganci A, Karwot R, Maxeiner JH, Scholtes P, Schmitt E, Neurath MF et al. IL-2 receptor beta-chain signaling controls immunosuppressive CD4+ T cells in the draining lymph nodes and lung during allergic airway inflammation in vivo. J Immunol 2008; 181: 1917–1926.
Mgbemena V, Segovia J, Chang T, Bose S . Kruppel-like factor 6 regulates transforming growth factor-beta gene expression during human respiratory syncytial virus infection. Virol J 2011; 8: 409.
McGraw DW, Elwing JM, Fogel KM, Wang WC, Glinka CB, Mihlbachler KA et al. Crosstalk between Gi and Gq/Gs pathways in airway smooth muscle regulates bronchial contractility and relaxation. J Clin Invest 2007; 117: 1391–1398.
Einstein R, Jordan H, Zhou W, Brenner M, Moses EG, Liggett SB . Alternative splicing of the G protein-coupled receptor superfamily in human airway smooth muscle diversifies the complement of receptors. Proc Natl Acad Sci USA 2008; 105: 5230–5235.
Pando MP, Morse D, Cermakian N, Sassone-Corsi P . Phenotypic rescue of a peripheral clock genetic defect via SCN hierarchical dominance. Cell 2002; 110: 107–117.
DeBruyne JP, Weaver DR, Reppert SM . Peripheral circadian oscillators require CLOCK. Curr Biol 2007; 17: R538–R539.
Travnickova-Bendova Z, Cermakian N, Reppert SM, Sassone-Corsi P . Bimodal regulation of mPeriod promoters by CREB-dependent signaling and CLOCK/BMAL1 activity. Proc Natl Acad Sci USA 2002; 99: 7728–7733.
Takata M, Burioka N, Ohdo S, Fukuoka Y, Miyata M, Endo M et al. Beta2-adrenoceptor agonists induce the mammalian clock gene, hPer1, mRNA in cultured human bronchial epithelium cells in vitro. Chronobiol Int 2005; 22: 777–783.
Petrie GR, Chookang JY, Hassan WU, Morrison JF, O’Reilly JF, Pearson SB et al. Bambuterol: effective in nocturnal asthma. Respir Med 1993; 87: 581–585.
Bergholtz B . Nocturnal asthma. Causes and treatment. Tidsskr Nor Laegeforen 1989; 109: 1796–1797.
Hetzel MR, Clark TJ . Comparison of normal and asthmatic circadian rhythms in peak expiratory flow rate. Thorax 1980; 35: 732–738.
Wong YH, Lee TY, Liang HK, Huang CM, Wang TY, Yang YH et al. KinasePhos 2.0: a web server for identifying protein kinase-specific phosphorylation sites based on sequences and coupling patterns. Nucleic Acids Res 2007; 35 (Web Server issue): W588–W594.
Ng PC, Henikoff S . SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 2003; 31: 3812–3814.
Sunyaev S, Ramensky V, Koch I, Lathe W, Kondrashov AS, Bork P . Prediction of deleterious human alleles. Hum Mol Genet 2001; 10: 591–597.
Acknowledgements
This work was supported by U01 HL65899 and P01 HL083069 from the National Heart, Lung and Blood Institute (NHLBI). We thank all families for their enthusiastic participation in the CAMP Genetics Ancillary Study and the CAMP investigators and research teams, who were supported by the NHLBI N01 HR16049. Additional support for this research came from NHLBI grants N01 HR16044, HR16045, HR16046, HR16047, HR16048, HR16049, HR16050, HR16051 and HR16052. All data collection from the CAMP Genetic Ancillary Study was conducted at the Channing Laboratory of the Brigham and Women’s Hospital under appropriate CAMP policies and human subject’s protections. The CAMP Genetics Ancillary Study is supported by U01 HL075419, U01 HL65899, P01 HL083069, R01 HL086601 and T32 HL07427 from the NIH/NHLBI. Collection of microarray data from immortalized lymphoblastoid cell lines of CAMP subjects was supported by K23 HG003983 and R01 HL092197 from the NIH/NHGRI. We acknowledge the American Lung Association (ALA) and the ALA’s Asthma Clinical Research Centers investigators and research teams for use of LOCCS and LODO data, with additional funding from HL071394 and HL074755 from the NHLBI, and Nemours Children's’ Clinic. GlaxoSmithKline supported the conduct of the LOCCS Trial by an unrestricted grant to the ALA. We acknowledge Sepracor for the use of the Asthma Trial data. The Single-Nucleotide Polymorphism Health Association Asthma Resource Project (SHARP) was funded by grants from the NHLBI U01 HL51510, U01 HL51834, U01 HL51831, U01 HL51845, U01 HL51843, M01 RR00079 and M01 RR03186, and was carried out by researchers from the Asthma Clinical Research Network (ACRN), CAMP and Childhood Asthma Research and Education (CARE) Network. Details are available in the Online Repository and on the dbGaP (database of Genotypes and Phenotypes) website: www.ncbi.nlm.nih.gov/sites/entrez?Db=gap. The GACRS was supported by HL04370 and HL66289 from the NIH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Rights and permissions
About this article
Cite this article
Duan, Q., Lasky-Su, J., Himes, B. et al. A genome-wide association study of bronchodilator response in asthmatics. Pharmacogenomics J 14, 41–47 (2014). https://doi.org/10.1038/tpj.2013.5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2013.5
Keywords
This article is cited by
-
A genome-wide analysis of the response to inhaled β2-agonists in chronic obstructive pulmonary disease
The Pharmacogenomics Journal (2016)
-
Asthma Pharmacogenomics: 2015 Update
Current Allergy and Asthma Reports (2015)