Abstract
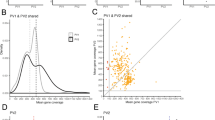
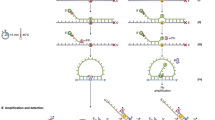
Single-nucleotide polymorphisms (SNPs) can be assayed using DNA isolated from archival formalin-fixed, paraffin-embedded (FFPE) samples, making retrospective pharmacogenetic studies possible. In this study, we describe methods that significantly increase the number of SNP determinations possible using FFPE samples. Quantifying the amount of DNA amenable to PCR (amplification-quality DNA, AQ-DNA) allows a significant reduction in the amount of sample required for Taqman-based SNP assays. Optimizing AQ-DNA input increases PCR amplification efficiency and SNP determination accuracy. DNA was extracted from 39 FFPE tumor sections and matched tumor and stromal cores, which were of the type used to generate tissue microarrays. Sections and tumor cores yielded sufficient AQ-DNA for more than 1000 SNP determinations. Seven SNPs were assessed following individual assay optimization for minimal AQ-DNA. Genotypes from tumor cores for single SNPs were 92.3–100% concordant with those obtained from sections. Using these methods, the number of SNP genotypes that can be determined from single FFPE samples is greatly increased expanding the genetic association studies possible from limited archival specimens. The use of tumor cores is of particular importance as the harvesting of tumor cores has minimal impact on the utility of the donor blocks for other purposes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Marsh S, Mallon MA, Goodfellow P, McLeod HL . Concordance of pharmacogenetic markers in germline and colorectal tumor DNA. Pharmacogenomics 2005; 6: 873–877.
Rae JM, Cordero KE, Scheys JO, Lippman ME, Flockhart DA, Johnson MD . Genotyping for polymorphic drug metabolizing enzymes from paraffin-embedded and immunohistochemically stained tumor samples. Pharmacogenetics 2003; 13: 501–507.
Schneider BP, Skaar TC, Sledge GW, Badve S, Li L, Flockhart DA . Analysis of angiogenesis genes from paraffin-embedded breast tumor and lymph nodes. Breast Cancer Res Treat 2006; 96: 209–215.
Weiss JR, Baer MR, Ambrosone CB, Blanco JG, Hutson A, Ford LA et al. Concordance of pharmacogenetic polymorphisms in tumor and germ line DNA in adult patients with acute myeloid leukemia. Cancer Epidemiol Biomarkers Prev 2007; 16: 1038–1041.
Farragher SM, Tanney A, Kennedy RD, Paul Harkin D . RNA expression analysis from formalin fixed paraffin embedded tissues. Histochem Cell Biol 2008; 130: 435–445.
Greer CE, Peterson SL, Kiviat NB, Manos MM . PCR amplification from paraffin-embedded tissues. Effects of fixative and fixation time. Am J Clin Pathol 1991; 95: 117–124.
Iwamoto KS, Mizuno T, Ito T, Akiyama M, Takeichi N, Mabuchi K et al. Feasibility of using decades-old archival tissues in molecular oncology/epidemiology. Am J Pathol 1996; 149: 399–406.
Romero RL, Juston AC, Ballantyne J, Henry BE . The applicability of formalin-fixed and formalin fixed paraffin embedded tissues in forensic DNA analysis. J Forensic Sci 1997; 42: 708–714.
Hui L, DelMonte T, Ranade K . Genotyping using the TaqMan assay. Curr Protoc Hum Genet 2008; 56: 2.10.1–2.10.8 (Chapter 2: Unit 2.10).
van Beers EH, Joosse SA, Ligtenberg MJ, Fles R, Hogervorst FB, Verhoef S et al. A multiplex PCR predictor for aCGH success of FFPE samples. Br J Cancer 2006; 94: 333–337.
Jin Y, Desta Z, Stearns V, Ward B, Ho H, Lee KH et al. CYP2D6 genotype, antidepressant use, and tamoxifen metabolism during adjuvant breast cancer treatment. J Natl Cancer Inst 2005; 97: 30–39.
Goetz MP, Rae JM, Suman VJ, Safgren SL, Ames MM, Visscher DW et al. Pharmacogenetics of tamoxifen biotransformation is associated with clinical outcomes of efficacy and hot flashes. J Clin Oncol 2005; 23: 9312–9318.
Xie B, Freudenheim JL, Cummings SS, Singh B, He H, McCann SE et al. Accurate genotyping from paraffin-embedded normal tissue adjacent to breast cancer. Carcinogenesis 2006; 27: 307–310.
Mort R, Mo L, McEwan C, Melton DW . Lack of involvement of nucleotide excision repair gene polymorphisms in colorectal cancer. Br J Cancer 2003; 89: 333–337.
Jaremko M, Justenhoven C, Abraham BK, Schroth W, Fritz P, Brod S et al. MALDI-TOF MS and TaqMan assisted SNP genotyping of DNA isolated from formalin-fixed and paraffin-embedded tissues (FFPET). Hum Mutat 2005; 25: 232–238.
Wang Y, Carlton VE, Karlin-Neumann G, Sapolsky R, Zhang L, Moorhead M et al. High quality copy number and genotype data from FFPE samples using Molecular Inversion Probe (MIP) microarrays. BMC Med Genomics 2009; 2: 8.
Johnson NA, Hamoudi RA, Ichimura K, Liu L, Pearson DM, Collins VP et al. Application of array CGH on archival formalin-fixed paraffin-embedded tissues including small numbers of microdissected cells. Lab Invest 2006; 86: 968–978.
Koch I, Slotta-Huspenina J, Hollweck R, Anastasov N, Hofler H, Quintanilla-Martinez L et al. Real-time quantitative RT-PCR shows variable, assay-dependent sensitivity to formalin fixation: implications for direct comparison of transcript levels in paraffin-embedded tissues. Diagn Mol Pathol 2006; 15: 149–156.
Andreassen CN, Sorensen FB, Overgaard J, Alsner J . Optimisation and validation of methods to assess single nucleotide polymorphisms (SNPs) in archival histological material. Radiother Oncol 2004; 72: 351–356.
Lehmann U, Kreipe H . Real-time PCR analysis of DNA and RNA extracted from formalin-fixed and paraffin-embedded biopsies. Methods 2001; 25: 409–418.
Siwoski A, Ishkanian A, Garnis C, Zhang L, Rosin M, Lam WL . An efficient method for the assessment of DNA quality of archival microdissected specimens. Mod Pathol 2002; 15: 889–892.
Farrand K, Jovanovic L, Delahunt B, McIver B, Hay ID, Eberhardt NL et al. Loss of heterozygosity studies revisited: prior quantification of the amplifiable DNA content of archival samples improves efficiency and reliability. J Mol Diagn 2002; 4: 150–158.
Gilbert MT, Sanchez JJ, Haselkorn T, Jewell LD, Lucas SB, Van Marck E et al. Multiplex PCR with minisequencing as an effective high-throughput SNP typing method for formalin-fixed tissue. Electrophoresis 2007; 28: 2361–2367.
Godfrey TE, Kim SH, Chavira M, Ruff DW, Warren RS, Gray JW et al. Quantitative mRNA expression analysis from formalin-fixed, paraffin-embedded tissues using 5′ nuclease quantitative reverse transcription-polymerase chain reaction. J Mol Diagn 2000; 2: 84–91.
Abrahamsen HN, Steiniche T, Nexo E, Hamilton-Dutoit SJ, Sorensen BS . Towards quantitative mRNA analysis in paraffin-embedded tissues using real-time reverse transcriptase-polymerase chain reaction: a methodological study on lymph nodes from melanoma patients. J Mol Diagn 2003; 5: 34–41.
Rae JM, Sikora MJ, Henry NL, Li L, Kim S, Oesterreich S et al. Cytochrome P450 2D6 activity predicts discontinuation of tamoxifen therapy in breast cancer patients. Pharmacogenomics J 2009; 9: 258–264.
Gaedigk A, Simon SD, Pearce RE, Bradford LD, Kennedy MJ, Leeder JS . The CYP2D6 activity score: translating genotype information into a qualitative measure of phenotype. Clin Pharmacol Ther 2008; 83: 234–242.
Acknowledgements
We thank Dr Sharoni Jacobs for assistance in developing the quality control multiplex PCR. MD and JS are grateful for funding from Breakthrough Breast Cancer and the Royal Marsden NIHR Biomedical Research Centre. This work was supported in part by the Breast Cancer Research Foundation Grant N003173 and by U-01 GM61373 and T-32 GM007767 from the National Institute of General Medical Sciences, Bethesda, MD, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the The Pharmacogenomics Journal website
Rights and permissions
About this article
Cite this article
Sikora, M., Thibert, J., Salter, J. et al. High-efficiency genotype analysis from formalin-fixed, paraffin-embedded tumor tissues. Pharmacogenomics J 11, 348–358 (2011). https://doi.org/10.1038/tpj.2010.50
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tpj.2010.50
Keywords
This article is cited by
-
DNA and RNA analysis of blood and muscle from bodies with variable postmortem intervals
Forensic Science, Medicine, and Pathology (2014)