Abstract
Variants at microRNA-137 (MIR137), one of the most strongly associated schizophrenia risk loci identified to date, have been associated with poorer cognitive performance. As microRNA-137 is known to regulate the expression of ~1900 other genes, including several that are independently associated with schizophrenia, we tested whether this gene set was also associated with variation in cognitive performance. Our analysis was based on an empirically derived list of genes whose expression was altered by manipulation of MIR137 expression. This list was cross-referenced with genome-wide schizophrenia association data to construct individual polygenic scores. We then tested, in a sample of 808 patients and 192 controls, whether these risk scores were associated with altered performance on cognitive functions known to be affected in schizophrenia. A subgroup of healthy participants also underwent functional imaging during memory (n=108) and face processing tasks (n=83). Increased polygenic risk within the empirically derived miR-137 regulated gene score was associated with significantly lower performance on intelligence quotient, working memory and episodic memory. These effects were observed most clearly at a polygenic threshold of P=0.05, although significant results were observed at all three thresholds analyzed. This association was found independently for the gene set as a whole, excluding the schizophrenia-associated MIR137 SNP itself. Analysis of the spatial working memory fMRI task further suggested that increased risk score (thresholded at P=10−5) was significantly associated with increased activation of the right inferior occipital gyrus. In conclusion, these data are consistent with emerging evidence that MIR137 associated risk for schizophrenia may relate to its broader downstream genetic effects.
Similar content being viewed by others
Introduction
Genome-wide association studies (GWAS) have identified significant associations between schizophrenia (SZ) and multiple single nucleotide polymorphisms (SNPs) located within or near to the microRNA-137 (MIR137) host gene on chromosome 1. In the most recent and largest SZ GWAS1 the SNP rs1702294, located intronically to MIR137,2 was identified as the second highest SZ-associated variant. This SNP is in high linkage disequilibrium (LD) with MIR137 variants previously identified in smaller GWAS3 such as rs1622579 (r2=0.99)4 and rs1198588 (r2=0.8).4, 5 In line with the proposed influence of microRNA-137 (miR-137) on proliferation, migration and maturation of neural cells,6, 7, 8 mechanisms important to the cognitive process, one of three intronic variants in high LD identified in this region (rs1625579) has been associated with a number of cognitively relevant phenotypes. These include lower performance in verbal episodic memory and vigilant attention,9 altered fronto-amygdala connectivity during a face processing task10 and decreased white matter integrity,11, 12 although a number of other studies have reported conflicting findings, with no effects of the MIR137 risk allele13, 14 on brain structure.
SZ GWAS to date suggest that the disorder is likely to be highly polygenic, involving a combination of both common risk variants of small effects, such as MIR137, and rare variants of larger effects. Modeling or quantifying this polygenic risk is therefore important, both for the broader illness phenotypes, and for specific illness dimensions related to functional outcomes, such as cognitive deficits. It is interesting to note that miR-137 is known or predicted to regulate hundreds of other genes,15 many of which are independently risk loci for SZ.3, 16 In vitro studies have suggested that miR-137 targets include BDNF, ZNF804A, TCF4, and CACNA1C,17, 18, 19, 20, 21 variants that have been shown to impact on both cognitive performance and cortical activity by our own group and others.13, 22, 23, 24, 25, 26 Because miR-137’s downstream regulatory targets include loci that are independently associated with both SZ risk and with cognitive deficits, it is possible that miR-137’s regulation of these genes may also contribute to SZ-associated deficits in cognition, in addition to the MIR137 loci’s direct effects on cognition.
The purpose of the present study was to investigate the polygenic effects on cognition of variants within genes that are downstream targets of miR-137. This was based on a previous study by Hill et al.,15 which, through miR-137 manipulation (upregulation and inhibition), assessed genome-wide transcriptional changes in a human neural progenitor cell line. This was carried out in order to elucidate pathways through which genetic disruption of miR-137 may increase SZ susceptibility. As disturbances in cognition are a cardinal feature of SZ, we investigated whether polygenic risk in this gene set explained variation in neuropsychological performance in patients and healthy participants. A key goal for this analysis was to estimate whether the variance explained by this downstream polygenic burden was independent to that explained by the single most robustly associated MIR137 SNP, rs1702294. Our hypothesis was that a higher polygenic burden within the empirically derived miR-137 regulated gene score, excluding the rs1702294 SNP, would be associated with increased cognitive deficits in a sample of healthy participants and patients with psychosis.
Materials and methods
Participants
In total, 808 cases and 192 healthy participants completed a full neuropsychological assessment battery and had full genome-wide SNP data available.1 Cases consisted of n=585 clinically stable patients with a diagnosis of SZ and schizoaffective disorder (SZA), which we refer to as ‘narrow-sense’ psychosis, and an additional n=223 patients diagnosed with bipolar disorder with psychotic features, major depressive disorder with psychotic features, delusional disorder, or psychosis not otherwise specified who combined with SZ/SZA patients to form a ‘broad-sense’ psychosis group. Patients were diagnosed by trained psychiatrists using the Structured Clinical Interview for DSM-IV Axis I Diagnosis.27 Inclusion criteria required participants to be clinically stable at time of cognitive assessment, aged between 18 and 65 years, no history of co-morbid psychiatric disorder, no substance abuse in the preceding 6 months, no prior head injury with loss of consciousness, no history of seizures, and with Irish ancestry (all four grandparents born in Ireland). Healthy participants were recruited from the general population through local media advertisements. All were aged between 18 and 65 years and had Irish-born paternal and maternal grandparents, and satisfied, on the basis of clinical interview, the criteria of having no history of major mental health problems, intellectual disability or acquired brain injury, and no substance abuse in the preceding 6 months. Exclusion criteria also included having a first-degree relative with a history of psychosis. All clinical and neuropsychological assessments were conducted in accordance with the relevant ethics committees’ approval for the six sites at which this data was collected, and all participants provided written informed consent.
Cognitive assessment
A full neuropsychological assessment designed to evaluate the cognitive deficits typically reported in SZ (general cognitive ability, memory function, attention, and social cognition) was administered to each participant. Selected subtests (Vocabulary, Similarities, Block Design, and Matrix Reasoning) of the Wechsler Adult Intelligence Scale, 3rd Edition (WAIS III)28 were used to measure general cognitive function. The Logical Memory (LM) I and II and Faces I and II subtests from the Wechsler Memory Scale, 3rd Edition (WMS III)29 were used to assess episodic and visual memory. Working memory was assessed using the Spatial Working Memory (SWM) subtest from the Cambridge Automated Neuropsychological Test Battery30 and Letter-Number Sequencing from the WMS III. Attentional control was assessed using the Continuous Performance Task (CPT) identical pairs version31 and the Sustained Attention to Response Task (SART).32 In addition to neuropsychological assessment, this study included measures of two aspects of social cognition—theory of mind (ToM) (frequently altered among patients with SZ and previously associated with MIR137(ref. 33)) and attributional style. ToM is the ability to attribute mental states—beliefs, intents, desires, pretending, knowledge and so on—to oneself and others and to understand that others have beliefs, desires, intentions and perspectives that are different from one's own, and is estimated using the Reading the Mind in the Eyes Task34 and the Hinting Task.35 Attributional style refers to the pervasive tendency to explain personally significant events in a particular manner.36 Consistency in cognitive assessment was assessed across sites by an independent researcher marking a number of patients assessed at each site.
Genotyping
Genotyping was conducted on DNA extracted from whole blood or saliva. Full GWAS data were available for all samples. A proportion of samples were genotyped with an Affymetrix 6.0 chip (Santa Clara, CA, USA; as part of the WTCCC2,37 referred to as Sample A) and the remainder on the Illumina HumanCoreExome chip (San Diego, CA, USA; referred to as Sample B). SNPs were excluded on the basis of MAF<0.1%, SNP missingness ⩽2%, and Hardy–Weinberg equilibrium P⩽10−6. Imputation was carried out on these data sets separately using 1000 Genomes Phase I integrated haplotypes (Dec 2013 release) and IMPUTE2 to give ~10 million SNPs genome-wide per sample.
Polygene score
We began by constructing the miR-137 downstream pathway based on the set of 1016 genes whose expression was identified as being altered by miR-137 manipulation in the study by Hill et al.,15 831 of these genes could be unambiguously mapped to the autosomes and this gene set was used to generate polygene scores. We then cross-referenced with unweighted P-values from the PGC2 2014 GWAS1 for variants within 20 kb of each gene in this set without LD-pruning (scores with and without pruning were highly correlated). We next defined three arbitrary threshold values based on the PGC threshold values (P<10−5, P<0.05, and P<0.5) to generate three polygene risk score values for analysis. Using these thresholds, 1020, 20 920 and 89 102 SNPs were included in the analysis, respectively. Finally, each participant was then given a weighted polygene score based on the number of risk alleles they carried within this gene list at the given gene threshold, weighted by the log of the SZ-association odds ratio from the PGC2. Genes used in this process are found in the Supplementary Information.
Statistical analysis: neuropsychological tests and polygene scores
Data were inspected for heteroscedasticity, and outliers,38 which were excluded from analysis. To estimate polygene effects on cognitive deficits associated with SZ (intelligence quotient (IQ), memory, attentional control and social cognition) linear regression analyses were performed using IBM SPSS Statistics,39 with age and gender entered as variables of no interest, and test score as the dependent variable. To take into account potential differences in results between Sample A and B due to the two differing genotyping platforms used, a linear regression was performed separately in each sample (those genotyped using Affymetrix and those with Illumina). Results from these were then meta-analyzed using the inverse variance method.40 In the inverse variance method, the weight given to each sample is chosen to be the inverse of the variance of the effect estimate (that is, one over the square of its standard error). This minimizes the imprecision of the pooled effect estimate. An estimate of sample heterogeneity and significance of such is shown in Supplementary Table 1 (I2). To maximize power to detect differences, we carried out our analysis on the full dataset of all cases and controls (n=1000). We then followed any significant results in the patient only groups (both the broad psychosis group (n=808) and narrow psychosis group (patients with SZ and schizoaffective disorder only (n=585)) to confirm the direction of effects in these groups (Supplementary Tables 2 and 3). As the meta-analyses showed little evidence of heterogeneity of results in Sample A and B despite the differing genotype platforms, the sample was analyzed as a whole to provide an estimate of effect size provide r2 and standardized β values (Supplementary Table 4). A post hoc power calculation estimated41 that n=988 has 0.88 power to detect a very conservative polygene score effect of r2=0.01, with α=0.05, although there are certain caveats to this type of calculation.42
Functional MRI
A subgroup of the healthy participants in this study (all right handed) had also undergone functional imaging during two cognitive tasks. One hundred eight participants completed a spatial working memory task and 83 had completed a facial processing task, with tasks and acquisition parameters as described by us previously.10, 33, 43, 44, 45 The task can be summarized as follows.
Spatial working memory task
This block design task was presented using Presentation software (Neurobehavioral Systems, Albany, CA, USA). Participants were asked if a white dot was in the same spatial location as a red circle (Supplementary Figure 1) during three conditions. In the baseline condition the white dot and red circle both appear at the same time. In the 1-dot condition, the white dot and red circle are separated by a 3- s delay. Finally, in the 3-dot condition, 3 white dots and a red circle are presented, separated by a 3- s delay. Task accuracy and reaction time were also measured.
Out of 108 participants, there were 3 missing this behavioral data that were excluded from fMRI analysis, and one participant was excluded due to an error with the task that caused it to run for several seconds after the scan finished. Two additional participants were excluded from analysis due to movement and 16 were excluded from all analysis due to low-quality MRI data. This left a final sample of 86 healthy participants.
The beginning of the spatial working memory task was synchronized to the first transistor-transistor logic (TTL) pulse sent from the MRI scanner at the start of the sequence. However, for a minority of participants the start of the task was not synchronized to the TTL pulse due to an experimental error; for these individuals the task was run manually at the start of the sequence. Due to the block design nature of this task, any small discrepancies in timing resulting from this error are unlikely to affect results.
Face processing task
Developed by Grosbras et al.,46 participants (n=83) watched a series of 2-s to 5-s black and white videos of faces, which started from a neutral expression, and then turned into an angry expression or neutral expression, interspersed with a control condition consisting of video clips of black and white concentric circles expanding and contracting. Blocks of video clips lasted 18- s, with 4–7 video clips presented per block. Overall, there were 19 blocks: 5 blocks of neutral face videos, 5 blocks of angry face videos and 9 control condition blocks (every second block was a control block). To make sure that individuals had paid attention to the videos, they performed a face recognition test after scanning. In this test, 5 stil-color pictures of faces were presented and participants were asked to determine whether these matched faces seen during the task. Six participants were excluded as they scored less than 4/5 correct answers or were missing data for the face recognition task. One face processing participant was excluded from analysis due to excessive movement and 5 participants were excluded due to low-quality MRI data and/or significant artefacts, resulting in a final sample of 70 participants.
Imaging pre-processing and statistical analysis
Spatial pre-processing and statistical analysis of MRI data was performed using Statistical Parametric Mapping (SPM8, revision 4290, http://www.fil.ion.ucl.ac.uk/spm/software/spm8/) and MATLAB R2011b (v7.13; http://www.mathworks.co.uk/). Functional images were realigned to the mean functional image, normalized to MNI (Montreal Neurological Institute) space with a voxel size of 3 × 3 × 3 mm3 and smoothed using a 10 mm FWHM (full-width at half maximum) isotropic Gaussian filter. After spatial pre-processing, graphical plots of the estimated time series of translations and rotations were inspected for excessive motion, which we defined as more than 3 mm translation and/or 3° rotation.
For the spatial working memory task, statistical analysis was performed using a general linear model (GLM) with two contrasts: spatial working memory (1 dot and 3 dots versus baseline), and increased spatial working memory load (3 dots>1 dot). For the face processing task, three task conditions (angry faces, neutral faces and baseline) and four contrasts consistent with our examination of neural activity associated with this task in SZ patients:33 Neutral faces versus baseline, angry faces versus baseline, all faces (angry and neutral) versus baseline, and angry faces versus neutral faces.
Participants’ contrast maps were entered into a second-level analysis to investigate effects of MIR137 network on neural activity (multiple regression analysis with the empirically derived miR-137 regulated gene score as covariate of interest). This multiple regression analysis was performed for the empirically derived MIR137 gene score at the P=10−5, P=0.05 and P=0.5 levels. Results were examined at a P<0.001 (uncorrected) level and clusters were considered statistically significant at a P<0.05 level, family-wise error (FWE) corrected for multiple comparisons across the whole brain at the cluster level. For each of these clusters, MNI coordinates of significant maxima were entered into the Anatomy toolbox in SPM8 (refs 47, 48, 49) and probable anatomical regions were identified using the AllAreas_v18_MPM atlas.
Results
Participant demographics: neuropsychological tests
Patient demographic information appears in Table 1. In all cognitive tests, healthy participants performed better than the patient groups. An ANOVA found that the miR-137 polygene scores differed significantly between diagnosis groups at each threshold. Predictably, patients showed higher polygene scores than healthy participants. A post hoc Tukey test indicated that the polygene scores for each patient group (narrow and broad sense) was significantly different from the control group (P<0.001) but not from each other.
Poorer performance on measures of IQ, memory, and attention were each found to be associated with higher miR-137 polygene scores at all P-value thresholds. When participants were combined, significant findings were found across all three polygene thresholds in domains of IQ, episodic memory, visual memory and working memory (Table 2). As hypothesized, performance on these neuropsychological measures decreased as polygene score increased. The most markedly significant domain was for declarative memory functioning. Multiple declarative memory subtests (the logical memory [immediate/delayed] and faces subtests [immediate/delayed] of the WMS III) were highly significant at each score threshold, surviving a stringent Bonferroni correction for the five cognitive domains analyzed. In contrast, the analysis of the single miR-137 risk SNP rs1702294 only showed significant influence on three domains: IQ, attention, and a ToM measure (Table 3; P-values ranging from P=0.042 to P=0.007).
As a post hoc analysis, when only patients were considered (and not controls), the same general pattern of results were observed as for the full group, with the relationship between higher polygene scores and lower test accuracy remaining significant at the P=10−5 threshold, and nominally significant at the P=0.05 threshold. Similarly, when the effects of rs1702294 were considered in patients only, nominal effects of the risk allele on poorer general cognitive ability and memory ability remained. Finally, when only SZ/SZA patients (‘narrow sense psychosis’) were analyzed, the same direction of results was also observed for both the polygene (Supplementary Tables 2 and 3) and single SNP analysis (Table 3). The effects of polygene risk on cognitive test performance were broader than that seen for the rs1702294 SNP alone (Supplementary Table 4). This was the case across each of the whole, broad psychosis and narrow psychosis samples. Whereas the risk ‘C’ allele of rs1702294 was associated with variance on only three cognitive tests, the polygenic risk was associated with multiple test scores in both the memory and general cognitive domains.
Participant demographics: fMRI
Participant demographics for the fMRI spatial working memory sample are presented in Table 4. A regression analysis was performed in SPSS (22.0.0) to examine any significant effects of MIR137 pathway risk (at P=10−5, P=0.05 and P=0.5 levels) on age, gender, years of education, SWM accuracy or SWM reaction time in our sample. Overall, there were no significant effects of MIR137 pathway risk on any of these variables (P>0.05).
Participant demographics for the overlapping fMRI face processing sample are presented in Supplementary Table 5. A regression analysis was performed in SPSS (22.0.0) to examine any significant effects of MIR137 pathway risk (at P=10−5, P=0.05 and P=0.5 levels) on age, gender, or years of education, and once again there were no significant effects of MIR137 pathway risk on any of these variables (P>0.05).
Effects of MIR137 pathway risk score on neural activity
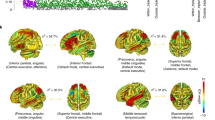
Increasing MIR137 pathway risk score (P=10−5 level) was associated with significantly increased neural activation in clusters incorporating the right middle temporal gyrus, left posterior cingulate and left thalamus with increasing spatial working memory load (3 dots versus 1 dot contrast, Supplementary Table 6). In each individual the mean parameters estimates of all voxels was calculated for each cluster showing a significant effect. These values were then entered into SPSS to check for outliers. Overall, three outliers were detected. As such, the analysis was run again, excluding these individuals. When this analysis was re-run, two clusters showed a significant effect of increasing MIR137 pathway risk score, one incorporating the right inferior occipital gyrus and right middle temporal gyrus, and another in the medial parietal region (individual cluster peaks not found on any probability maps using Anatomy toolbox). We observed that, in the high working memory load condition (3-dot versus 1-dot) increasing MIR137 polygenic risk score (P=10−5 level) was associated with increased activation across two clusters (Supplementary Table 6). The first incorporated the right inferior occipital gyrus and right middle temporal gyrus and showed peak coordinates in the right inferior occipital gyrus (MNI peak coordinates 48, −76, −2; tmax=4.80; P-value=0.011; cluster extent (voxels)=175; Figure 1). A second cluster, located in the medial parietal region, did not localize to any known probability maps using Anatomy toolbox (Cluster extent [voxel]=121, P=0.038, tmax=4.19; MNI peak coordinates 3, −34, 16), possibly because these coordinates represent a region of overlap between gray and white matter (Figure 1). No other effects were observed either for other MIR137 polygenic thresholds or other contrasts. Note that neural activity associated with the 3-dot versus 1-dot condition across all participants is presented in Supplementary Table 7 and Figure 1, showing overlap with the right inferior occipital cluster in which a MIR137 effect was observed.
Effect of increasing MIR137 pathway risk score (P=10−5 level) on neural activity during increasing spatial working memory load (3 dots versus 1 dot), 3 outliers removed (N=83; shown in red), shown alongside overall activation for this contrast in the same group (shown in blue). Cluster 1 (right inferior occipital gyrus and right middle temporal gyrus), top panels; Cluster 2 (medial parietal region) bottom panels. Clusters are significant at P<0.05, FWE-corrected at the cluster level and represent regions where neural activity was greater with increasing risk score. Each 2D axial slice is labeled with an x-coordinate (MNI space). Clusters are rendered on the ‘ch256’ brain template using MRIcroGL (http://www.mccauslandcenter.sc.edu/mricrogl/). Additional editing of figure (for example, changing the size/resolution) performed using MS Paint and/or Paint.NET v3.5.10. FWE, family-wise error; MIR137, microRNA-137; MNI, Montreal Neurological Institute.
Finally, for the face processing task, after correcting for outliers (n=1 using the same outlier detection method as for the spatial working memory data) no significant effects of the empirically derived miR-137 regulated gene score (at P=10−5, P=0.05, or P=0.5 levels) were observed for any other contrast.
Discussion
This study assessed the effects of higher polygenic risk burden in a network of genes whose expression was altered by manipulation of miR-137 expression, the gene product of a SZ risk gene. On the basis of previous studies from our group and others, we sought to establish whether this gene network would convey the same cognitive and cortical effects as have been associated with individual risk variants at this locus and, if so, whether the amount of variance explained was the same or greater. We did this based on polygene scores derived from all risk variants within this network (not including MIR137 risk SNPs themselves), calculated from three risk thresholds using the PGC-SCZ1 data (P=10−5, P=0.05 and P=0.5). Following correction for multiple testing we found that, irrespective of the threshold used, genetic variants within the defined MIR137 score were associated with lower declarative memory performance. We also found that the MIR137 pathway was nominally associated with poorer working memory and general cognitive ability. Finally, functional imaging revealed that participants showed increased cortical activation in right inferior occipital gyrus during a spatial working memory task. This was found in the absence of any association with cortical activity during a face processing task and polygenic risk. Collectively, these data indicate deleterious effects of risk variants within the empirically derived miR-137 regulated gene score was on cognitive function in patients and controls.
Studies seeking to characterize the cognitive effects of individual risk variants have been criticized previously for low power, making their findings difficult to replicate. This study, based on one of the largest single data sets available (n=1000 individuals with adequate genetic information available from a total of 1269 participants collected) attempted to overcome this difficulty through the use of a relatively large sample size and a novel approach to risk—that is, calculated on the basis of a risk gene molecular pathway. The robustness of our results are supported by the following: (1) the direction of all significant results were the same both for the full dataset and when both the narrow and broad patients groups were analyzed alone; (2) the findings were consistent across all genetic risk thresholds, even after correction for the number of cognitive functions assessed (general cognitive ability, declarative memory, working memory, attentional control, and social cognition) was made; (3) in the case of the memory subtests used, findings replicated across modalities, that is, increased polygenic risk was associated with both poorer verbal performance and poorer non-verbal performance; finally, (4) the amount of variation in cognitive performance explained by the empirically derived miR-137 regulated gene score was larger than that explained by the single MIR137 variant, rs1702294, with the polygenic pathway score explaining in the region of ~2% (Supplementary Table 1) of variance on declarative memory measures. Although still modest, this is approximately two to three times the variance explained by the individual rs1702294 SNP.
Although correction for multiple testing was carried out for the cognitive component of the study (a Bonferroni correction for five domains: IQ, working memory, visual declarative memory, attention and social cognition) we did not correct for the fact that three polygenic risk significance thresholds were used. It is still unclear what the best approach to selecting thresholds is. The original PGC analysis3 reported 10 risk thresholds. We have included three to reduce the multiple testing burden, while others have reduced this burden even further by including only one, for example, a threshold of P=0.5.50 However, for our cognitive analysis the threshold considered would appear to have been unimportant: broadly the same cognitive associations were observed for each of the three thresholds considered. This would suggest that in fact a study of only one threshold would have been sufficient to characterize the cognitive consequences of this gene set. For the neuroimaging study, an association between MIR137 polygenic risk and cortical activation during a working memory task was only significant at the most conservative threshold of P=10−5; why this might have been case is uncertain. One possibility is that this was simply a false-positive finding; the fact that only a subsample of participants were included in this analysis, albeit relatively large for fMRI studies (n=108) might support this conclusion. Other explanations include that imaging findings may be more sensitive to the signal to noise ratio inherent in high polygenic risk thresholds (that is, that more lenient thresholds harbor a greater percentage of variants not robustly associated with risk, hence making them less informative). To our knowledge, only one other polygenic fMRI study has been carried out to date, and as noted included only one threshold.50 As a consequence this relatively novel approach requires the accumulation of further studies to determine the best thresholding approach to take, as evidence for an optimum threshold (in terms of signal to noise ratio) has yet to emerge.
When examining brain regions showing increased blood oxygenation level dependent (BOLD) response with increasing spatial working memory load (contrast: 3 dots condition>1 dot condition), two clusters showed significantly increased BOLD response in individuals with higher MIR137 pathway risk score (MIR137 pathway risk score P=10−5 level). The first cluster incorporated the right inferior occipital gyrus and right middle temporal gyrus. Given that this cluster overlaps with cortical regions activated by our task (Figure 1), hyperactivation in this cluster may represent relative inefficiency in processing task stimuli in individuals with higher risk scores, although this suggestion remains speculative.
Although a second cluster, in the medial parietal region, showed significantly increased BOLD response with increasing MIR137 pathway risk score (P=10−5 level) also, no individual peak coordinates in MNI space were not found on any probability map (Figure 1). Given that visual inspection of this cluster reveals that a large proportion resides in white matter, this may be an artefactual finding and should be interpreted with caution. It is possible that as regions of the medial parietal cortex typically show task-induced decreases in BOLD response,51 the present finding of increased BOLD response in this region may relate to a relative failure to deactivate this region during the working memory task in individuals with higher risk scores. However, as noted above, the fact that these results are obtained in a sample that, although large for imaging studies, is quite modest for genetics studies, requires caution in interpretation of these results.
In addition to the issue of sample size, further study limitations include that only healthy participants were used for studying the association of polygenic risk on patterns of brain activation. Ideally the analysis will be repeated in a patient sample to confirm findings in this group. However, symptoms in SZ and cognitive deficits occur along a spectrum, so despite the lack of patient fMRI data, our current analysis does allow for investigation of the association between this polygene score and functional alterations in healthy participant endophenotypes. In addition, a recent paper evaluating the effect of MIR137 on gray matter concentration found that the effects of SNPs in MIR137 and associated pathways have greater impact on brain structure and function in SZ patients than healthy participants.52 The results of the present analysis of healthy participants may be indicative of a greater impact in a patient population. Another potential limitation here is using a gene list derived from human progenitor neural cells, in which the stability of transcriptional regulation can change over time. Nonetheless, at least some of the genetic variants identified to generate the polygenic score are present in both adult and embryonic stages, so the effect of the variant may have already had its substantial influence in neurodevelopmental stages. With regard to the effect of medication on cognition, medication dosage was not used as a covariate in our whole sample analysis, which included controls, as doing so would have limited our sample size in the analysis (n=1000) to patients with this information (n=585). In the patient groups considered separately, results from the neuropsychological results were non-significant following Bonferroni correction; including medication dosage as a covariate in this analysis did not change these results.
Taking a polygene approach to characterize genetic effects has particular significance for the MIR137 polygene score. As with other gene sets and putative biological pathways, discrepancies exist between genes that are bioinformatically predicted to be related, and those that have been empirically identified. For the MIR137 polygene scores generated here, we included only empirically derived genes. Gene expression changes in this study were found to significantly correlate with those following miR-137 manipulation in another human neural progenitor cell line in the subsequent study.21 Several lines of evidence suggest that at least part of the risk for SZ conferred by genetic variation at the MIR137 locus is via effects of miR-137 perturbation on the expression of other SZ susceptibility genes.1, 15, 20, 53 Thus, variants within this empirically derived miR-137 regulated gene score that contribute to SZ polygenic risk may be hypothesized to impact upon cognitive function that is a central feature of the disorder. This study, by charactering the cumulative polygenic effects of these downstream risk variants while excluding the effects of risk variants with the miR-137 gene itself supports this hypothesis, both at a cognitive and at a cortical level. Functional studies indicate that miR-137 is involved in proliferation, migration and maturation of neural cells,6, 7, 8 likely to be the mechanisms behind this involvement with the cognitive process. Further studies of the effects of MIR137 polygene scores may benefit from adopting a more fine-grained approach to the selection of SNPs. This could include parsing the effects on cognition of those variants in this network that are directly versus indirectly influenced by MIR137, given that both are likely to have been included in the gene set.15 This was not possible in this study as the method of gene selection could not differentiate between direct and indirect interactors of MIR137.
In conclusion, these data suggest that increased polygenic risk in a gene set consisting of downstream targets of miR-137 was associated with decreased cognitive performance, both in terms of memory function, and for general cognitive ability. Notably, the strength of association was stronger for the gene set as a whole than for the most strongly associated MIR137 SNP. These data are consistent with emerging evidence that at least some of the genetic risk for SZ conferred by MIR137 variation relates to its downstream effects on gene expression.
References
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511: 421–427.
Cunningham F, Amode MR, Barrell D, Beal K, Billis K, Brent S et al. Ensembl 2015. Nucleic Acids Res 2015; 43: D662–D669.
Schizophrenia Psychiatric Genome-Wide Association Study Consortium. Genome-wide association study identifies five new schizophrenia loci. Nat Genet 2011; 43: 969–976.
Ward LD, Kellis M . HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res 2012; 40: D930–D934.
Ripke S, O'Dushlaine C, Chambert K, Moran JL, Kahler AK, Akterin S et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet 2013; 45: 1150–1159.
Smrt RD, Szulwach KE, Pfeiffer RL, Li X, Guo W, Pathania M et al. MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase mind bomb-1. Stem Cells 2010; 28: 1060–1070.
Konopka W, Kiryk A, Novak M, Herwerth M, Parkitna JR, Wawrzyniak M et al. MicroRNA loss enhances learning and memory in mice. J Neurosci 2010; 30: 14835–14842.
Willemsen MH, Valles A, Kirkels LA, Mastebroek M, Olde Loohuis N, Kos A et al. Chromosome 1p21.3 microdeletions comprising DPYD and MIR137 are associated with intellectual disability. J Med Genet 2011; 48: 810–818.
Cummings E, Donohoe G, Hargreaves A, Moore S, Fahey C, Dinan TG et al. Mood congruent psychotic symptoms and specific cognitive deficits in carriers of the novel schizophrenia risk variant at MIR-137. Neurosci Lett 2013; 532: 33–38.
Mothersill O, Morris DW, Kelly S, Rose EJ, Fahey C, O'Brien C et al. Effects of MIR137 on fronto-amygdala functional connectivity. Neuroimage 2014; 90: 189–195.
Kuswanto CN, Sum MY, Qiu A, Sitoh YY, Liu J, Sim K . The impact of genome wide supported microRNA‐137 (MIR137) risk variants on frontal and striatal white matter integrity, neurocognitive functioning, and negative symptoms in schizophrenia. American J Med Genet B Neuropsychiatr Genet 2015; 168B: 317–326.
Lett TA, Chakavarty MM, Felsky D, Brandl EJ, Tiwari AK, Goncalves VF et al. The genome-wide supported microRNA-137 variant predicts phenotypic heterogeneity within schizophrenia. Mol Psychiatry 2013; 18: 443–450.
Cousijn H, Eissing M, Fernandez G, Fisher SE, Franke B, Zwiers M et al. No effect of schizophrenia risk genes MIR137, TCF4, and ZNF804A on macroscopic brain structure. Schizophr Res 2014; 159: 329–332.
Kelly S, Morris DW, Mothersill O, Rose EJ, Fahey C, O’Brien C et al. Genome-wide schizophrenia variant at MIR137 does not impact white matter microstructure in healthy participants. Neurosci Lett 2014; 574: 6–10.
Hill MJ, Donocik JG, Nuamah RA, Mein CA, Sainz-Fuertes R, Bray NJ . Transcriptional consequences of schizophrenia candidate miR-137 manipulation in human neural progenitor cells. Schizophr Res 2014; 153: 225–230.
Yin J, Lin J, Luo X, Chen Y, Li Z, Ma G et al. miR-137: a new player in schizophrenia. Int J Mol Sci 2014; 15: 3262–3271.
Kim AH, Parker EK, Williamson V, McMichael GO, Fanous AH, Vladimirov VI . Experimental validation of candidate schizophrenia gene ZNF804A as target for hsa-miR-137. Schizophr Res 2012; 141: 60–64.
Wright C, Turner JA, Calhoun VD, Perrone-Bizzozero N . Potential impact of miR-137 and its targets in schizophrenia. Front Genet 2013; 4: 10.3389/fgene.2013.00058.
Guan F, Zhang B, Yan T, Li L, Liu F, Li T et al. MIR137 gene and target gene CACNA1C of miR-137 contribute to schizophrenia susceptibility in Han Chinese. Schizophr Res 2014; 152: 97–104.
Kwon E, Wang W, Tsai L . Validation of schizophrenia-associated genes CSMD1, C10orf26, CACNA1C and TCF4 as miR-137 targets. Mol Psychiatry 2013; 18: 11–12.
Collins AL, Kim Y, Bloom RJ, Kelada SN, Sethupathy P, Sullivan PF . Transcriptional targets of the schizophrenia risk gene MIR137. Transl Psychiatry 2014; 4: e404.
Nicodemus K, Hargreaves A, Morris D, Anney R, Gill M, Corvin A et al. Variability in working memory performance explained by epistasis vs polygenic scores in the ZNF804A pathway. JAMA Psychiatry 2014; 71: 778–785.
Walters JT, Corvin A, Owen MJ, Williams H, Dragovic M, Quinn EM et al. Psychosis susceptibility gene ZNF804A and cognitive performance in schizophrenia. Arch Gen Psychiatry 2010; 67: 692–700.
Donohoe G, Morris DW, Robertson IH, Clarke S, McGhee KA, Schwaiger S et al. Variance in facial recognition performance associated with BDNF in schizophrenia. Am J Med Genet B Neuropsychiatr Genet 2007; 144b: 578–579.
Zhang Q, Shen Q, Xu Z, Chen M, Cheng L, Zhai J et al. The effects of CACNA1C gene polymorphism on spatial working memory in both healthy controls and patients with schizophrenia or bipolar disorder. Neuropsychopharmacology 2012; 37: 677–684.
Rose EJ, Morris DW, Fahey C, Cannon D, McDonald C, Scanlon C et al. The miR-137 schizophrenia susceptibility variant rs1625579 does not predict variability in brain volume in a sample of schizophrenic patients and healthy individuals. Am J Med Genet B Neuropsychiatr Genet 2014; 165b: 467–471.
First M, Spitzer R, Gibbon M, Williams J . Structured Clinical Interview for DSM-IV-TR Axis I Disorders, Research Version, Patient Edition (SCID-I/P):. New York State Psychiatric Institute: New York, NY, USA, 2002.
Wechsler D . WAIS-III, Wechsler Adult Intelligence Scale: Administration and Scoring Manual. Psychological Corporation: London, 1997.
Wechsler D . Wechsler Memory Scale (WMS-III). Psychological Corporation: London, 1997.
Robbins T, James M, Owen A, Sahakian B, McInnes L, Rabbitt P . Cambridge Neuropsychological Test Automated Battery (CANTAB): a factor analytic study of a large sample of normal elderly volunteers. Dement Geriatr Cogn Disord 1994; 5: 266–281.
Cornblatt BA, Risch NJ, Faris G, Friedman D, Erlenmeyer-Kimling L . The Continuous Performance Test, identical pairs version (CPT-IP): I. New findings about sustained attention in normal families. Psychiatry Res 1988; 26: 223–238.
Robertson IH, Manly T, Andrade J, Baddeley BT, Yiend J . Oops!': performance correlates of everyday attentional failures in traumatic brain injured and normal subjects. Neuropsychologia 1997; 35: 747–758.
Mothersill O, Morris DW, Kelly S, Rose EJ, Bokde A, Reilly R et al. Altered medial prefrontal activity during dynamic face processing in schizophrenia spectrum patients. Schizophr Res 2014; 157: 225–230.
Baron‐Cohen S, Wheelwright S, Hill J, Raste Y, Plumb I . The “Reading the Mind in the Eyes” test revised version: A study with normal adults, and adults with Asperger syndrome or high‐functioning autism. J Child Psychol Psychiatry 2001; 42: 241–251.
Corcoran R, Mercer G, Frith CD . Schizophrenia, symptomatology and social inference: investigating "theory of mind" in people with schizophrenia. Schizophr Res 1995; 17: 5–13.
Kinderman P, Bentall RP . A new measure of causal locus: the internal, personal and situational attributions questionnaire. Pers Individ Dif 1996; 20: 261–264.
Irish Schizophrenia Genomics Consortium, The Wellcome Trust Case Control Consortium. Genome-wide association study implicates HLA-C*01:02 as a risk factor at the major histocompatibility complex locus in schizophrenia. Biol Psychiatry 2012; 72: 620–628.
Hoaglin DC, Iglewicz B . Fine-tuning some resistant rules for outlier labeling. J Am Stat Assoc 1987; 82: 1147–1149.
IBM Corp IBM SPSS statistics for Windows, version 21.0. IBM Corp Armonk: New York, NY, USA, 2012.
Higgins JP, Green S . Cochrane Handbook for Systematic Reviews of Interventions. Vol 5. Wiley Online Library: Chichester, 2008.
Faul F, Erdfelder E, Buchner A, Lang A-G . Statistical power analyses using G* Power 3.1: Tests for correlation and regression analyses. Behav Res Methods 2009; 41: 1149–1160.
Hoenig JM, Heisey DM . The abuse of power. The American Statistician 2012; 55: 1.
Rose EJ, Greene C, Kelly S, Morris DW, Robertson IH, Fahey C et al. The NOS1 variant rs6490121 is associated with variation in prefrontal function and grey matter density in healthy individuals. Neuroimage 2012; 60: 614–622.
Rose EJ, Morris DW, Fahey C, Robertson IH, Greene C, O'Doherty J et al. The effect of the neurogranin schizophrenia risk variant rs12807809 on brain structure and function. Twin Res Hum Genet 2012; 15: 296–303.
Rose EJ, Morris DW, Hargreaves A, Fahey C, Greene C, Garavan H et al. Neural effects of the CSMD1 genome-wide associated schizophrenia risk variant rs10503253. Am J Med Genet B Neuropsychiatr Genet 2013; 162B: 530–537.
Grosbras MH, Paus T . Brain networks involved in viewing angry hands or faces. Cereb Cortex 2006; 16: 1087–1096.
Eickhoff SB, Paus T, Caspers S, Grosbras MH, Evans AC, Zilles K et al. Assignment of functional activations to probabilistic cytoarchitectonic areas revisited. Neuroimage 2007; 36: 511–521.
Eickhoff SB, Heim S, Zilles K, Amunts K . Testing anatomically specified hypotheses in functional imaging using cytoarchitectonic maps. Neuroimage 2006; 32: 570–582.
Eickhoff SB, Stephan KE, Mohlberg H, Grefkes C, Fink GR, Amunts K et al. A new SPM toolbox for combining probabilistic cytoarchitectonic maps and functional imaging data. Neuroimage 2005; 25: 1325–1335.
Whalley HC, Papmeyer M, Sprooten E, Romaniuk L, Blackwood DH, Glahn DC et al. The influence of polygenic risk for bipolar disorder on neural activation assessed using fMRI. Transl Psychiatry 2012; 2: e130.
Buckner RL, Andrews-Hanna JR, Schacter DL . The brain's default network: anatomy, function, and relevance to disease. Ann N Y Acad Sci 2008; 1124: 1–38.
Wright C, Gupta CN, Chen J, Patel V, Calhoun VD, Ehrlich S et al. Polymorphisms in MIR137HG and microRNA-137-regulated genes influence gray matter structure in schizophrenia. Transl Psychiatry 2016; 6: e724.
Wright C, Calhoun VD, Ehrlich S, Wang L, Turner JA, Bizzozero NI . Meta gene set enrichment analyses link miR-137-regulated pathways with schizophrenia risk. Front Genet 2015; 6: 147.
Acknowledgements
We thank all patients and their support staff, and all healthy volunteers for participating in the data collection on which this manuscript is based. Recruitment and genotyping was supported by Science Foundation Ireland (SFI) (grants 12.IP.1359 and 08/IN.1/B1916) and the Wellcome Trust Case Control Consortium 2 project (grants 085475 /B/08/Z and 085475/Z/08/Z) and the Wellcome Trust (grants 072894/Z/03/Z, 090532/Z/09/Z, and 075491/Z/04/B).
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
The Wellcome Trust Case Control Consortium 2 investigators Peter Donnelly, Lesley Bates, Ines Barroso, Jenefer M. Blackwell, Elvira Bramon, Matthew A. Brown, Juan P. Casas, Aiden Corvin, Panos Deloukas, Audrey Duncanson, Janusz Jankowski, Hugh S. Markus, Christopher G. Mathew, Colin N. A. Palmer, Robert Plomin, Anna Rautanen, Stephen J. Sawcer, Richard C. Trembath, Ananth C. Viswanathan, Nicholas W.Wood, Chris C. A. Spencer, Gavin Band, Céline Bellenguez, Colin Freeman, Garrett Hellenthal, Eleni Giannoulatou, Lucinda Hopkins, Matti Pirinen, Richard Pearson, Amy Strange, Zhan Su, Damjan Vukcevic, Cordelia Langford, Sarah E. Hunt, Sarah Edkins, Rhian Gwilliam, Hannah Blackburn, Suzannah J. Bumpstead, Serge Dronov, Matthew Gillman, Emma Gray, Naomi Hammond, Alagurevathi Jayakumar, Owen T.McCann, Jennifer Liddle, Simon C. Potter, Radhi Ravindrarajah, Michelle Ricketts, Matthew Waller, Paul Weston, Sara Widaa, and Pamela Whittaker.
Supplementary Information accompanies the paper on the Translational Psychiatry website
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Cosgrove, D., Harold, D., Mothersill, O. et al. MiR-137-derived polygenic risk: effects on cognitive performance in patients with schizophrenia and controls. Transl Psychiatry 7, e1012 (2017). https://doi.org/10.1038/tp.2016.286
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tp.2016.286
This article is cited by
-
Association of neurotransmitter pathway polygenic risk with specific symptom profiles in psychosis
Molecular Psychiatry (2024)
-
Synergistic, long-term effects of glutamate dehydrogenase 1 deficiency and mild stress on cognitive function and mPFC gene and miRNA expression
Translational Psychiatry (2023)
-
Polygenic analysis suggests the involvement of calcium signaling in executive function in schizophrenia patients
European Archives of Psychiatry and Clinical Neuroscience (2020)
-
A systematic review of associations between functional MRI activity and polygenic risk for schizophrenia and bipolar disorder
Brain Imaging and Behavior (2019)
-
Cognitive Characterization of Schizophrenia Risk Variants Involved in Synaptic Transmission: Evidence of CACNA1C's Role in Working Memory
Neuropsychopharmacology (2017)