Abstract
The hypothetical ‘AXAS’ gene network model that profiles functional patterns of heterogeneous DNA variants overrepresented in autism spectrum disorder (ASD), X-linked intellectual disability, attention deficit and hyperactivity disorder and schizophrenia was used in this current study to analyze whole exome sequencing data from an Australian ASD cohort. An optimized DNA variant filtering pipeline was used to identify loss-of-function DNA variations. Inherited variants from parents with a broader autism phenotype and de novo variants were found to be significantly associated with ASD. Gene ontology analysis revealed that putative rare causal variants cluster in key neurobiological processes and are overrepresented in functions involving neuronal development, signal transduction and synapse development including the neurexin trans-synaptic complex. We also show how a complex gene network model can be used to fine map combinations of inherited and de novo variations in families with ASD that converge in the L1CAM pathway. Our results provide an important step forward in the molecular characterization of ASD with potential for developing a tool to analyze the pathogenesis of individual affected families.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD) is a highly heritable neurodevelopmental disorder that is clinically characterized by impaired social interaction and communication deficits, as well as restricted and repetitive behavior.1 ASD is generally first apparent in childhood and affects up to ~1–2% of the population.2,3 The emergence of massively parallel DNA sequencing has identified many rare single-nucleotide variants (SNVs) and small insertion/deletion (indels) variations associated with ASD. Recently, whole exome sequencing (WES) has verified the contribution of de novo variants (DNVs) in ASD, including an increased rate of DNVs associated with aged paternity.4, 5, 6, 7 WES studies have similarly clarified the role of rare inherited variants (IVs), such as rare gene deletion (inherited loss-of-function homozygous, compound heterozygous or X-chromosome variants in males),8 recessive homozygous,9 bi-allelic variants10 and coinheritance of variants from multiplex families.11 More recently, whole genome sequencing has identified rare genetic variants associated with noncoding DNA and also variants affecting the differential splicing of genes.12,13 These discoveries have led to the insight that hundreds of rare genetic variants can influence numerous biological functions associated with ASD.14 Many of these variations occur in genes that are comorbid for ASD and other symptom complexes (for example, specific language impairments, intellectual disabilities, epilepsies, schizophrenia) and monogenic syndromes (for example, Rett syndrome, fragile X syndrome, tuberous sclerosis, Timothy syndromes).3
Notwithstanding these conceptual advances, large gaps remain in our understanding of the genetic basis of ASD. Genetic heterogeneity adds to the challenge of understanding the relative contributions of numerous rare genetic variants to the severity of ASD.15, 16, 17 The influence of differing combinations of rare ASD-associated variants on the occurrence of ASD is poorly understood.18 In spite of these problems, attempts have been made to reconcile the functional relevance of case-specific rare variants. However, these studies are few in number and reductionist approaches have struggled to accommodate potentially hundreds of DNA variants associated with ASD. Single gene/variant functional studies fail to account for the likely combinations of weak alleles contributing to the cumulative liability threshold for ASD. Similarly, the genetic tool-box of model species is not accessible for analysis of the relative contribution of differing combinations of rare heterozygous human DNA variants. In the face of this scientific challenge, the field of molecular psychiatric genetics is trying to build a paradigm to grasp the ‘bigger picture’ concerning the molecular and biological basis of ASD. One such way forward is the use of systems approaches.19, 20, 21
We recently reported a hypothetical gene network model that was used to profile the functional patterns of heterogeneous DNA variants overrepresented in ASD, X-linked intellectual disorder, attention deficit and hyperactivity disorder and schizophrenia.1 In the current study, we have applied the AXAS model to analyze WES data from an Australian ASD cohort. We show how combinations of loss-of-function variants cluster in functional processes and putatively contribute to the causal profile of individuals with an ASD. We also show how DNA variants inherited from parents with a broader autism phenotype (BAP)22 and DNVs have a significant association with ASD.
Materials and methods
Recruitment and behavioral assessment
Participants were drawn from the Western Australian Autism Biological Registry, at the Telethon Institute for Child Health Research in Perth, Western Australia. Participants were recruited via advertisement and children with a DSM-IV-based23 clinical diagnosis of autistic disorder, Asperger’s syndrome or pervasive developmental disorder-not otherwise specified were included in the study. Diagnosis of ASD was obtained by consensus following a multidisciplinary assessment by a team comprising a pediatrician, clinical psychologist and speech pathologist.24
Forty families from the biological registry, who had either one child or multiple children (trio and sibling families) diagnosed with an ASD, were randomly selected for the current study. The total sample comprised 48 cases and 80 parent controls (Supplementary Figure 1). All probands were administered the autism diagnostic observation schedule-generic,25 and were found to meet the criteria for ASD. Parents were asked to complete the autism-spectrum quotient, a self-report questionnaire that provides a quantitative measure of autistic-like traits in the general population26 and a score of 22 or above on the autism-spectrum quotient was indicative of the BAP27—subclinical expression of behaviors characteristic of ASD.
The final participant sample comprised 35 simplex families and 5 multiplex families: 37 families were of European origin and 3 families were of Asian origin (Supplementary Figure 1; Supplementary Table 2). Among the 48 probands, there were 43 children with a diagnosis of autistic disorder, 2 with Asperger’s disorder and 3 with pervasive developmental disorder-not otherwise specified. Among the 80 parents (controls), there were 23 parents (6 mothers and 17 fathers) classified as BAP, 55 parents (34 mothers and 21 fathers) classified as non-BAP (Supplementary Table 2) and 2 fathers who did not complete the BAP assessment.
Exome capture sequencing
Blood was obtained via venipuncture from all 142 participants and genomic DNA was extracted from whole blood. We used 1 μg of genomic DNA to construct whole genome libraries. Multiplexed genomic libraries of four different individuals were subjected to whole exome capture using NimbleGen SeqCap version 3. Exome libraries were sequenced with paired-end 100 bp reads on the Illumina HiSeq2000 (San Diego, CA, USA) using Illumina SBS V3 chemistry. This approach captures 62 Mb of the genomic sequence comprising exonic variants (variants in protein-coding portion), ncRNA variants (variants in noncoding RNAs), untranslated region (UTR) or intronic variants (variants in UTRs and intron regions) and intergenic variants (variants between mRNA transcripts of two genes).
Variant identification
Raw sequencing data were mapped by Burrows–Wheeler Aligner28 on the hg19 reference genome from 1000 Genome Project (human_g1k_v37) followed by removal of PCR duplicates using Picard software (http://picard.sourceforge.net). Realignment of the mapped reads was performed using the Genome Analysis Toolkit (version 2.6–4-g3e5ff60).29 For variant calling, we used the Genome Analysis Toolkit UnifiedGenotyper to call raw genotypes from sequenced regions that contain single-nucleotide polymorphisms (SNPs) and small indels. To achieve a high-quality call set, we used multisample calling and Variant Quality Score Recalibrator with training data sets (HapMap3, 1k genome and dbSNP).30,31 We discarded genotypes that were likely to be false positives or of poor quality from the list of variants with the following criteria: (1) heterozygous genotypes in the X-chromosome in males, (2) genotypes in the Y chromosome in females, (3) genotypes covered by fewer than 20 reads, (4) genotypes with a Phred-scaled base quality score lower than 30 and (5) genotypes with indels that are frequently observed across multiple samples.
We assessed SNPs and indels using ANNOVAR software.32 We annotated variants using the hg19 reference genome and prioritized these by their minor allele frequency using the 1000 Genomes Project (the April 2012 release for European and Asian populations) and the NHLBI-6500 Exomes (this version includes sex chromosomes and indels for all ethnicity groups). We determined the functional significance of variants using scale-invariant feature transform (SIFT) and PolyPhen2 (the LJB version 2) for missense mutations33 and used the SIFT Indel tool (http://sift.bii.a-star.edu.sg/www/SIFT_indels2.html) for frameshift indels.34 Finally, we excluded genomic regions that are likely to contribute false-positive signals in variant calling of exome sequencing.35
De novo and inherited variant selection
We used the phase-by-transmission, a Genome Analysis Toolkit module, to identify DNVs and IVs from given familial information. The most likely genotype was estimated by computing the posterior probability for all possible genotypes in each family from the raw genotype likelihoods and expected prior probability of transmission (P<0.001). For DNVs, we chose de novo calls with ⩾20 Transmission Probability (a Phred-scaled probability of erroneous call ⩽1%) and ⩾20 read depth. We further reduced de novo calls in cases where the depth in number of sequence reads of alternative allele is greater than 10% in the maternal or paternal genotype. We then randomly selected 10 DNVs and successfully validated their genotype by targeted Sanger sequencing using custom primers (Supplementary Figure 2). For IVs, we only considered genotypes that were successfully phased in the parents.
MicroRNA gene and target site variants
Target site prediction from the 3′-UTR of gene transcripts found in UCSC ‘knownGene’ annotation on hg19 reference genome were conducted for all microRNAs (miRNAs) found in miRBase Release 19.36 Predicted sites were then mapped back onto the genome, and compared with the refSeq gene annotation. Two miRNA target site predictions tools miRanda (v3.3a)37 and RNAhybrid (v2.1.1)38 were used to confirm target sites with a minimum free energy less than −18 and parameters optimized for human 3′-UTRs (options: −b 2000, −e −18, −s 3utr_human). MiRanda was run with default parameters. The predicted miRNA target sites were then filtered using a free energy of −25 or less for RNAhybrid and −18 for miRanda. The sites used in this analysis are those predicted by miRanda, with a 75% reciprocal overlap with RNAhybrid predictions. Bedtools was used to find SNVs that were located in predicted miRNA target regions.39
Protein–protein interaction network
We utilized the protein–protein interaction (PPI) network based on the AXAS model,1 for variant-classification and systems biology analyses. We retrieved all possible interactions between primary candidate genes and their first-order interactors from the whole PPI network. The PPI network visualization and functional analysis was performed as previously reported1 using Cytoscape.40 We examined nonredundant DNA variants found in our sequence analysis using a binomial distribution with a standardized Z-score to test whether sets of genes are overrepresented in the AXAS-PPI network of four different mental health disorders; ASD, X-linked intellectual disorder, attention deficit and hyperactivity disorder and schizophrenia.1
Functional ontology analysis
We used ClueGo version 1.8 (a Cytoscape plug-in),41 to perform functional enrichment analysis based on annotations of three functional ontology databases: gene ontology (GO) (http://geneontology.org/), Kyoto encyclopedia of genes and genomes (KEGG) pathways (http://www.genome.jp/kegg/) and Reactome (http://www.reactome.org/). The statistical significance of functional terms was calculated with the Fisher’s Exact Test and adjusted using Bonferroni step-down correction. We only used GO terms that are annotated based on experimental evidence in the biological process ontology, and excluded all KEGG disease pathways.
Results
Exome sequencing
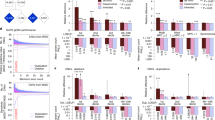
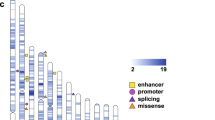
Exome sequencing was performed on ASD families (Supplementary Figure 3), with a 57.9 × average sequence read depth and 71.5% capture efficiency across the target regions (Supplementary Table 2). Approximately 50 Mb of the exome was covered by ⩾20 × reads from which we confidently identified DNA variants. There was an average of 38 686 SNPs and 2420 indels per individual exome. Of these, 13 919 SNPs and 155 indels were located in protein-coding sequences, and 24 767 SNPs and 2265 indels found in noncoding sequences (Figure 1a). Most of the SNPs were identified in exonic (13 919; 36.0%) and intronic regions (14 087; 36.4%), whereas indels were mostly found in intronic regions (1382; 57.1%) and 3′-UTRs (453; 18.7%). Among variants in noncoding sequences, there were 1773 SNPs and 103 indels in regions that encode a miRNA or long noncoding RNA (Figure 1b). There were a total of 9650 SNVs that occur within predicted miRNA target sites of 3'-UTRs (from 2 624 853 predicted sites in all).
Genome distribution of DNA variants. Circle plots show the type and localization of filtered DNA variants detected by exome sequence analysis. The location of variants was annotated using the Reference Sequence hg19 database with ANNOVAR software. (a) SNPs were equally distributed in both intronic and exonic regions, whereas most indels were observed in intronic and 3’-UTR regions of gene transcripts. (b) DNA variants were also found to occur in noncoding RNA sequences. SNP, single-nucleotide polymorphism; UTR, untranslated region.
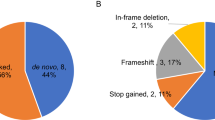
Regarding Mendelian inheritance, most of the variants were inherited from parents but 44 variants newly occurred (de novo) in 28 ASD children. Among 44 DNVs, there were 29 associated with protein-coding sequences, 9 were intronic, 4 were intergenic and 2 occur in 3’-UTR regions of genes. Aside from variants associated with noncoding sequences, we selected variants that result in changes to protein-coding sequences (missense, nonsense and frameshift variants) for analysis. Although coding variants can more obviously be associated with loss of function, we are mindful that noncoding variants detected using exome sequencing offer important insights into the aberrant expression of coding genes and noncoding RNAs (Figure 1b).
Putative causal variants
Many previous exome studies have used public databases for prioritizing putative causal variants from raw variants based on information of allele frequency or a functional prediction score. For example, databases of the 1000 Genomes Project and NHLBI-6500si Exomes were used for measuring minor allele frequency (MAF) of variants, and those of SIFT and PolyPhen2 were used for predicting a functional impact of an amino acid substitution caused by variants. Typically, exome studies select putative causal variants that are rare in a population (those variants with <5% MAF or unknown in the 1000 Genomes Project and NHLBI-6500si Exomes) or alter a protein function (those variants classified as ‘damaging’ in SIFT and PolyPhen2 predictions).11,42, 43, 44, 45
We tested how the filtering process affects the prioritization of causal variants (Supplementary Figure 4) using various thresholds and exclusion parameters with the AXAS-PPI model which assumes that there is a set of genes significantly associated with ASD. We retrieved genes for which variants occurred in cases or controls (parents) to test whether these genes are overrepresented in the ASD-PPI network model.1 We found that by lowering the MAF threshold we were able to more stringently filter DNA variants on the basis of allele frequency. The differences in association of variants with ASD between cases and controls became smaller when using a <2% MAF setting (with the 1000 Genomes Project data set; Supplementary Figure 4a) and <1% MAF (with the NHLBI-6500si Exomes data set; Supplementary Figure 4b), respectively. When examining the SIFT variant filtering process, the association of variants from cases was higher than from controls, within the range of thresholds for the prediction of the damaging classification (0⩽ SIFT score ⩽0.05; Supplementary Figure 4c). The association of variants from parent controls decreased with a SIFT score of 0.1, the threshold associated with a damaging classification. When using PolyPhen2, the association of variants from cases became higher when increasing the cutoff within the range of thresholds for the prediction of the damaging classification (0.447⩽ PolyPhen2 score ⩽1; Supplementary Figure 4d), whereas association of variants from controls did not change much within this range. Therefore, rare variants with low allele frequency and significant functional impact as determined by the AXAS model are most likely to be associated with ASD.
On the basis of these observations, we defined causal variants as those meeting the following criteria: (1) variants annotated as a missense/nonsense and frameshift indel, (2) variants that have minor allele frequency (MAF) <0.01 or are not reported in the 1000 Genomes Project and NHLBI-6500si Exomes, (3) missense/nonsense variants classified as damaging by both SIFT (0⩽ prediction score ⩽0.05) and PolyPhen2 (0.447⩽ prediction score ⩽1) and (4) frameshift variants classified as damaging by SIFT Indel tool. Next, we conducted the process of variant filtering and reduced the number of variants by subsequent combinations of variant databases (Figure 2a). On the basis of this stringent variant filtering pipeline, combining four variant databases with appropriate thresholds, we were able to successfully predict rare DNA variants in ASD cases. These putative causal variants were shown to be significantly associated with ASD but not with other neurodevelopmental or neuropsychiatric disorders (Figure 2b).
Overview of variant filtering pipeline. (a) A detailed diagram of the computational pipeline and databases used to filter DNA variants found in WES data. Shown are the average number of variants per individual. (b) Comparison of databases showing the association of filtered variants with ASD. The level of association was measured using a binomial distribution with a standardized Z-score as determined by the AXAS model.1 ADHD, attention deficit and hyperactivity disorder; ASD, autism spectrum disorder; MAF, minor allele frequency; PPI, protein–protein interaction; SZ, schizophrenia; WES, whole exome sequencing; XLID, X-linked intellectual disorder.
Variant classification
Our exome analysis identified DNA variations in 1754 and 2607 candidate genes in cases and parent controls, respectively (Supplementary Table 4). As described above, we used the AXAS-PPI network as a biological assumption set to test the association of these genes with ASD. We found that variations in candidate genes of ASD children were significantly associated with the ASD-PPI network (P=0.04) (Figure 3a). Notably, variants of candidate genes found in parent controls were not significantly associated with ASD.
Variant classification using the AXAS model. Radar plots showing binomial distributions with standardized Z-scores used to examine the association of putative causal DNA variants with the AXAS-PPI model.1 Z-scores ⩾1.65 show significant association with a neurodevelopmental disorder. Inherited variants from BAP parents (g) and de novo variants (c) show the greatest association with ASD; see also Supplementary Table S1. ADHD, attention deficit and hyperactivity disorder; ASD, autism spectrum disorder; BAP, broader autism phenotype; SZ, schizophrenia; XLID, X-linked intellectual disorder.
Although putative causal variants in control parents did not reach statistical significance, these variants were tightly associated with ASD and a higher Z-score for ASD compared with other disorders (Figure 3a). Therefore, we assessed prospective causal variants by examining their association with BAP and non-BAP parents. We found that 1053 and 2174 candidate genes containing putative causal variants occurred in 23 parents with BAP and 55 parents without BAP, respectively (Supplementary Table 4). The causal variants associated with candidate genes in parents with BAP were shown to be significantly associated with ASD (P=0.02), whereas those in parents without BAP were not (Figures 3d and 3f). Importantly, this suggests that parents with weaker or milder behavioral deficits confer significant ASD risk as determined by our variant-classification system.
Mendelian inheritance of ASD risk
We further examined whether Mendelian inheritance confers a risk for ASD by comparing causal variants inherited from parents with BAP (IVs-BAP) and those from parents without BAP (IVs non-BAP). We found that a total of 764 and 1271 candidate genes that contained causal variants occur in these groups, respectively (Supplementary Table 4). Variant classification showed that IVs-BAP are significantly associated with the ASD-PPI network (P=0.002), whereas IVs from non-BAP parents are not (Figures 3e and 3g). This finding was consistently observed when causal variants were separated using the weak ASD phenotype status of the parents to measure the basis of ASD inheritance (Supplementary Table 1). Therefore ASD-associated variants found in parents with BAP are more likely to confer ASD risk to their children.
Our results also revealed that candidate genes containing DNVs from ASD cases have significant association with the ASD-PPI network (P=0.01). This is consistent with recent findings that DNVs are important genetic factors that contribute to sporadic ASD cases.46 We also examined the possible relationship of paternal age with frequency of DNVs. We found that there was a significant positive correlation between DNVs and the age of the father, confirming recent reports4, 5, 6, 7 of an increased risk of ASD with paternal age (Supplementary Figure 5). There is, however, a possibility that some filtered DNVs may arise somatically in immune cell lineages found in blood. Future targeted re-sequencing of DNA from a second (nonimmune) cell type would comprehensively verify parental origin of DNVs.
Convergent pathway in ASD cases
Although there were many putative ASD variants identified in probands, it still remains unclear how these variants contribute to biological/behavioral phenotypes associated with ASD. Examining these variants in control parents with BAP does, however, provide an insight as to how these are likely to combine in the next generation and contribute to the presentation of an ASD. We postulated that causal variants of different origins, inherited and de novo, converge in molecular pathways and process so as to contribute to a critical threshold of liability that results in an ASD. To test this hypothesis, we examined inherited and de novo causal variants from selected ASD cases, and constructed the PPI network of these families and probands. We used functional ontology analysis for these PPI networks to search for convergent pathways that are enriched in the ASD-PPI network. Fine mapping maternally and paternally inherited causal variants and DNVs in the context of their molecular interactions provides a precise view of how biological pathways are likely to be functionally compromised (Figure 4).
Clustering analysis in ASD cases. PPI networks were constructed using putative causal variants and their first-degree protein interactors that result in three types of affected networks. Two possible schemes (a and b) show how maternally inherited variants (yellow), paternally inherited variants (green) and de novo variants (blue) converge and cluster in unique functional pathways (gray). Functional ontology of three proband cases: (c) 06.s1, neuronal migration; (d) 30.s1, long-term potentiation, (e) 38.s1, axon guidance and inherited variants that merge with de novo variants that converge in (f) 18.s1, ankyrin and adhesive related proteins involved the synapse development and neurodevelopment. Of the 48 ASD cases, there were four unique combinations (pedigrees) of variants that cluster in the L1CAM interaction pathway (REACTOME:22205; see Supplementary Table 5 for details of gene names). ASD, autism spectrum disorder; PPI, protein–protein interaction.
We found that there were a total of 116 GO, 33 KEGG and 94 Reactome terms identified in convergent variant pathway analysis of the 48 ASD cases (Supplementary Table 5). Of note, there was an increased frequency of affected pathways associated with synaptic development and neurodevelopment (Table 1). Clearly, incorrect synapse development and erosion of synaptic function due to genetic variants are widely considered to be key contributors to ASD.47,48 These pathways encompass long-term potentiation (KEGG:04720), trafficking of AMPA receptors (REACTOME:18307) and activation of NMDA receptors upon glutamate binding and postsynaptic events (REACTOME:20563). These data again highlight neurodevelopment as an important functional hub associated with ASD pathogenesis. Multiple cases were shown to have affected pathways related to neuron migration (GO:0001764), neurotrophin signaling (KEGG:04722) and axon guidance (REACTOME:18266). These analyses also identified affected pathways more directly linked to clinical phenotypes, such as social behavior (GO:0035176) and vocalization behavior (GO:0071625).
Notably we found that there were a number of putative causal DNA variations associated with the neurexin trans-synaptic complex (NTSC) that has previously been associated with autism.49,50 There were two putative causal DNA variations found in neurexins (NRXNs), CNTNAP 3 and CNTNAP 5. We also found a number of variations among interacting molecules of the NTSC specifically those that interact with neuroligins (NLGN), SH3 and multiple ankyrin repeat domains (SHANKs) and NRXN. These include SNVs that occur in DLG4 (postsynaptic density protein 95), DLGAP2 (disks large-associated protein 2), LPHN2 (latrophilin-2), SPTAN1 (spectrin, alpha, non-erythrocytic 1), CASK (calcium/calmodulin-dependent serine protein kinase), PDZD2 (PDZ domain containing 2), DDX24 (DEAD (Asp-Glu-Ala-Asp) box helicase 24), SIPA1L1 (signal-induced proliferation-associated 1 like 1) and NXPH3 (neurexophilin 3). We found SNVs also occur in other potential NRXN interacting molecules include LRRTM4 (leucine-rich repeat transmembrane neuronal 4), CNTN3 (contactin 3) and CNTN4 (contactin 4). Aside from synapse development, another striking association was 4/48 cases involved convergence of DNA variants in the L1CAM interaction pathway. The L1CAM interaction pathway consists of the L1 family of cell adhesion molecules, which has an important role in neuronal migration, axon guidance and synaptic formation.51,52 Genetic variants in L1CAM have been associated with a wide range of neurodevelopmental and mental health disorders.53,54
Our results also show pathways involved in cellular assembly, circadian rhythm, immune response, protein catabolism/modification and various signal transductions are also associated with ASD (Supplementary Table 5), and may be involved in developmental regression, locomotion impairment, metabolic abnormalities and sleep disturbances.55
Discussion
We have previously shown that the AXAS model provides a hypothetical framework to analyze and profile the molecular basis of neurodevelopmental disorders such as ASD.1 This a priori molecular network provides a ‘biological assumption’ for analysis of WES genetic screening data from Australian ASD families. Although heterogeneity of causal DNA variants is an inherent property of the human genome,56 as well as in ASD,3,14,57 we show that the AXAS model can successfully associate DNA variations with ASD from WES cohort data. This is because the AXAS network approach provides a means to recognize patterns of candidate genes that are putatively overrepresented in functional processes and pathways associated with ASD. Notably, we used this approach to search for neurofunctional pathways associated with ASD among individual cases and parents by investigating higher-order interaction of combinations of DNA variants (Figure 4). We confirm that putative causal variations often cluster in functional pathways and represent a molecular convergence of mostly heterozygous weak alleles that likely erode biological processes. Fine mapping of causal variants may therefore create plausible links between genotype and phenotype.18,58
We examined IVs comparing BAP and non-BAP parents to trace combinations of genetic determinants associated with ASD in the probands. Previous studies have found broader ASD-like behavioral phenotypes or clinical subtypes in parents, siblings or relatives of individuals with ASD.22,59, 60, 61 Our results suggest that there is a very strong genetic association of IVs from BAP parents with ASD (Figure 3), confirming a direct relationship between subclinical behavioral and genetic phenotypes in parents whose children present with ASD. Given that IVs are subject to evolutionary selection and putatively segregate in a distribution of effect including a contribution to weak BAP phenotypes in parents, de novo DNA variations are not subject to selection. We found that DNVs detected in the probands also had a strong statistical association with ASD and, to a lesser extent, with X-linked intellectual disorder and attention deficit and hyperactivity disorder compared with controls (non-BAP parents; Figure 3). This is probably due to the random occurrence of DNVs in neurological genes that are comorbid between mental health disorders.1 Therefore future genetic investigations need to estimate the statistical significance of IVs from BAP and non-BAP parents with DNVs, which can only be achieved through genetic screening of families.
The WES approach has limitations as it only detects individual genetic variation based on SNPs and small indels associated with exonic regions of the genome. WES does not detect large copy number variations or genetic variants of regulatory sequences located in intergenic regions. Copy number variations are important genetic contributors that account for ~10% of ASD, including associated syndromes.55,62 We did, however, formally examine DNA variations that occur in noncoding regions associated with WES data (Figure 1), including variations in noncoding RNAs such as miRNAs. Mutations were detected in miRNA transcripts and these may have implications for miRNA precursor stability and targeting on a broad scale.63 There were also many SNVs in miRNA target binding sites and these potentially have a more specific effect on miRNA targeting activity.63,64 Future screening studies could more comprehensively include using high density-array genotyping for detection of copy number variations and SNPs associated with noncoding information, whereas whole genome sequencing remains an option.
Although WES cannot provide a full-genome accounting of ASD, it does detect polygenic DNA variations associated with coding information that often cluster and disrupt neurodevelopmental pathways. A closer examination of putative causal DNA variations associated with the L1CAM interaction pathway involved in axon guidance (Figure 4) highlights how combinations of gene variations could disrupt biological function. There is an overlap of putative causative variants between case 06.s1 (Figure 4c) and 30.s1 (Figure 4d), including a variant of NFASC (neurofascin; chr1:204944474) that occurs in 14 families. Regarding the first-degree interactors of NFASC, we find that DCX (doublecortin) and NrCAM (neuronal cell adhesion molecule) have been previously associated with ASD.65,66 Case 06.s1 also contains a genetic variant of SPRK2 (serine/threonine protein kinase 2; chr7:104909268) that interacts with VAV2 (guanine nucleotide exchange factor VAV2) that is involved in the L1CAM pathway. In cases 6.s1 and 30.s1, there is a genetic variant of NPEPPS (puromycin-sensitive aminopeptidase; chr17:45669359) that is more commonly found in 33 families. NPEPPS has previously been shown to interact with RPS6KA3 (ribosomal protein S6 kinase, polypeptide 3), and variants of NPEPPS have been associated with ASD and with epileptic encephalopathy.67 The more frequent occurrence of these NFASC and NPEPPS variants associated with the L1CAM pathway may therefore represent important genetic elements that predispose to ASD.
In case 38.s1 (Figure 4e), there is a rare variant of DLG4 (disks large homolog 4; chr17:7097689) that also occurs in family 39. DLG4 is involved in the L1CAM pathway and has many first-degree interactors, including another rare variant of ITGA4 (integrin, alpha 4; chr2:182347303). Like a number of other L1 family genes, ITGA4 has also previously been associated with ASD.68, 69, 70 In case 18.s1 (Figure 4f), we found a DNV (chr2:166231244) of SCN2A (sodium channel, voltage-gated, type II, alpha subunit). SCN2A is a well-known candidate ASD gene6,12 which interacts with ANK3 (ankyrin 3). Ankyrins are important because they link voltage-gated sodium and potassium channels to spectrin, L1 and NrCAM. In addition, we find genetic variants of ERBB2IP (erbb2 interacting protein; chr5:65317206) and MACF1 (microtubule actin cross-linking factor 1; chr1: 39851427) that are also associated with the L1CAM pathway. Taken together, rare genetic variants and DNVs that occur in combination in probands, as detailed above, may therefore increase genetic liability associated with key processes involved in neuronal development and plasticity.
In summary, the AXAS model is a tool that can be used to abstract a molecular basis of ASD, helping to engage genetic screening data with hypothesis-driven paradigms based on biological evidence. Our findings provide a means to identify the functional consequence of combinations of causal DNA variations associated with ASD. This study raises the intriguing prospect of how we might examine the unique pathogenesis of individual families with ASD and the potential application of targeted therapies and personalized medicine.
References
Cristino AS, Williams SM, Hawi Z, An JY, Bellgrove MA, Schwartz CE et al. Neurodevelopmental and neuropsychiatric disorders represent an interconnected molecular system. Mol Psychiatry 2014; 19: 294–301.
Autism and Developmental Disabilities Monitoring Network Surveillance Year 2008 Principal Investigators, Centers for Disease Control and Prevention. Prevalence of autism spectrum disorders—Autism and Developmental Disabilities Monitoring Network, 14 sites, United States, 2008. MMWR Surveill Summ 2012; 61: 1–19.
Devlin B, Scherer SW . Genetic architecture in autism spectrum disorder. Curr Opin Genet Dev 2012; 22: 229–237.
O'Roak BJ, Vives L, Girirajan S, Karakoc E, Krumm N, Coe BP et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 2012; 485: 246–250.
Neale BM, Kou Y, Liu L, Ma'ayan A, Samocha KE, Sabo A et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 2012; 485: 242–245.
Sanders SJ, Murtha MT, Gupta AR, Murdoch JD, Raubeson MJ, Willsey AJ et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 2012; 485: 237–241.
Iossifov I, Ronemus M, Levy D, Wang Z, Hakker I, Rosenbaum J et al. De novo Gene disruptions in children on the autistic spectrum. Neuron 2012; 74: 285–299.
Lim ET, Raychaudhuri S, Sanders SJ, Stevens C, Sabo A, MacArthur DG et al. Rare complete knockouts in humans: population distribution and significant role in autism spectrum disorders. Neuron 2013; 77: 235–242.
Chahrour MH, Yu TW, Lim ET, Ataman B, Coulter ME, Hill RS et al. Whole-exome sequencing and homozygosity analysis implicate depolarization-regulated neuronal genes in autism. PLoS Genet 2012; 8: e1002635.
Yu TW, Chahrour MH, Coulter ME, Jiralerspong S, Okamura-Ikeda K, Ataman B et al. Using whole-exome sequencing to identify inherited causes of autism. Neuron 2013; 77: 259–273.
Toma C, Torrico B, Hervas A, Valdes-Mas R, Tristan-Noguero A, Padillo V et al. Exome sequencing in multiplex autism families suggests a major role for heterozygous truncating mutations. Mol Psychiatry advance online publication, 3 September 2013; doi:10.1038/mp.2013.106.
Jiang YH, Yuen RK, Jin X, Wang M, Chen N, Wu X et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am J Hum Genet 2013; 93: 249–263.
Shi L, Zhang X, Golhar R, Otieno FG, He M, Hou C et al. Whole-genome sequencing in an autism multiplex family. Mol Autism 2013; 4: 8.
Betancur C . Etiological heterogeneity in autism spectrum disorders: more than 100 genetic and genomic disorders and still counting. Brain Res 2011; 1380: 42–77.
Li Y, Vinckenbosch N, Tian G, Huerta-Sanchez E, Jiang T, Jiang H et al. Resequencing of 200 human exomes identifies an excess of low-frequency non-synonymous coding variants. Nat Genet 2010; 42: 969–972.
Kryukov GV, Pennacchio LA, Sunyaev SR . Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. Am J Hum Genet 2007; 80: 727–739.
Bamshad MJ, Ng SB, Bigham AW, Tabor HK, Emond MJ, Nickerson DA et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat Rev Genet 2011; 12: 745–755.
Berg JM, Geschwind DH . Autism genetics: searching for specificity and convergence. Genome Biol 2012; 13: 247.
Voineagu I, Wang X, Johnston P, Lowe JK, Tian Y, Horvath S et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 2011; 474: 380–384.
Parikshak NN, Luo R, Zhang A, Won H, Lowe JK, Chandran V et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 2013; 155: 1008–1021.
Willsey AJ, Sanders SJ, Li M, Dong S, Tebbenkamp AT, Muhle RA et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell 2013; 155: 997–1007.
Taylor LJ, Maybery MT, Wray J, Ravine D, Hunt A, Whitehouse AJ . Brief report: do the nature of communication impairments in autism spectrum disorders relate to the broader autism phenotype in parents? J Autism Dev Disord 2013; 43: 2984–2989.
American Psychiatric Association, DSM-IV. APATFo. Diagnostic and statistical manual of mental disorders: DSM-IV. American Psychiatric Association: Washington, DC, USA, 1994.
Lord C, Petkova E, Hus V, Gan W, Lu F, Martin DM et al. A multisite study of the clinical diagnosis of different autism spectrum disorders. Arch Gen Psychiatry 2012; 69: 306–313.
Lord C, Risi S, Lambrecht L, Cook EH Jr ., Leventhal BL, DiLavore PC et al. The autism diagnostic observation schedule-generic: a standard measure of social and communication deficits associated with the spectrum of autism. J Autism Dev Disord 2000; 30: 205–223.
Baron-Cohen S, Wheelwright S, Skinner R, Martin J, Clubley E . The autism-spectrum quotient (AQ): evidence from Asperger syndrome/high-functioning autism, males and females, scientists and mathematicians. J Autism Dev Disord 2001; 31: 5–17.
Wheelwright S, Auyeung B, Allison C, Baron-Cohen S . Defining the broader, medium and narrow autism phenotype among parents using the Autism Spectrum Quotient (AQ). Mol Autism 2010; 1: 10.
Li H, Durbin R . Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 2009; 25: 1754–1760.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010; 20: 1297–1303.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 2011; 43: 491–498.
Nielsen R, Paul JS, Albrechtsen A, Song YS . Genotype and SNP calling from next-generation sequencing data. Nat Rev Genet 2011; 12: 443–451.
Wang K, Li M, Hakonarson H . ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 2010; 38: e164.
Liu X, Jian X, Boerwinkle E . dbNSFP: a lightweight database of human nonsynonymous SNPs and their functional predictions. Hum Mutat 2011; 32: 894–899.
Hu J, Ng PC . Predicting the effects of frameshifting indels. Genome Biol 2012; 13: R9.
Fuentes Fajardo KV, Adams D, Program NCS, Mason CE, Sincan M, Tifft C et al. Detecting false-positive signals in exome sequencing. Hum Mutat 2012; 33: 609–613.
Kozomara A, Griffiths-Jones S . miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 2011; 39 (Database issue): D152–D157.
Enright AJ, John B, Gaul U, Tuschl T, Sander C, Marks DS . MicroRNA targets in Drosophila. Genome Biol 2003; 5: R1.
Rehmsmeier M, Steffen P, Hochsmann M, Giegerich R . Fast and effective prediction of microRNA/target duplexes. RNA 2004; 10: 1507–1517.
Quinlan AR, Hall IM . BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 2010; 26: 841–842.
Smoot ME, Ono K, Ruscheinski J, Wang PL, Ideker T . Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 2011; 27: 431–432.
Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009; 25: 1091–1093.
Bi C, Wu J, Jiang T, Liu Q, Cai W, Yu P et al. Mutations of ANK3 identified by exome sequencing are associated with autism susceptibility. Hum Mutat 2012; 33: 1635–1638.
Heinzen EL, Depondt C, Cavalleri GL, Ruzzo EK, Walley NM, Need AC et al. Exome sequencing followed by large-scale genotyping fails to identify single rare variants of large effect in idiopathic generalized epilepsy. Am J Hum Genet 2012; 91: 293–302.
Need AC, McEvoy JP, Gennarelli M, Heinzen EL, Ge D, Maia JM et al. Exome sequencing followed by large-scale genotyping suggests a limited role for moderately rare risk factors of strong effect in schizophrenia. Am J Hum Genet 2012; 91: 303–312.
O'Roak BJ, Deriziotis P, Lee C, Vives L, Schwartz JJ, Girirajan S et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nat Genet 2011; 43: 585–589.
Ku CS, Polychronakos C, Tan EK, Naidoo N, Pawitan Y, Roukos DH et al. A new paradigm emerges from the study of de novo mutations in the context of neurodevelopmental disease. Mol Psychiatry 2013; 18: 141–153.
Krumm N, O'Roak BJ, Shendure J, Eichler EE . A de novo convergence of autism genetics and molecular neuroscience. Trends Neurosci 2013; 37: 95–105.
Won H, Mah W, Kim E . Autism spectrum disorder causes, mechanisms, and treatments: focus on neuronal synapses. Front Mol Neurosci 2013; 6: 19.
Sudhof TC . Neuroligins and neurexins link synaptic function to cognitive disease. Nature 2008; 455: 903–911.
Wright GJ, Washbourne P . Neurexins, neuroligins and LRRTMs: synaptic adhesion getting fishy. J Neurochem 2011; 117: 765–778.
Maness PF, Schachner M . Neural recognition molecules of the immunoglobulin superfamily: signaling transducers of axon guidance and neuronal migration. Nat Neurosci 2007; 10: 19–26.
Enneking EM, Kudumala SR, Moreno E, Stephan R, Boerner J, Godenschwege TA et al. Transsynaptic coordination of synaptic growth, function, and stability by the L1-type CAM Neuroglian. PLoS Biol 2013; 11: e1001537.
Cheng L, Lemmon V . Pathological missense mutations of neural cell adhesion molecule L1 affect neurite outgrowth and branching on an L1 substrate. Mol Cell Neurosci 2004; 27: 522–530.
Kudumala S, Freund J, Hortsch M, Godenschwege TA . Differential effects of human L1CAM mutations on complementing guidance and synaptic defects in Drosophila melanogaster. PloS One 2013; 8: e76974.
Geschwind DH . Advances in autism. Annu Rev Med 2009; 60: 367–380.
MacArthur DG, Balasubramanian S, Frankish A, Huang N, Morris J, Walter K et al. A systematic survey of loss-of-function variants in human protein-coding genes. Science 2012; 335: 823–828.
Eapen V . Genetic basis of autism: is there a way forward? Curr Opin Psychiatry 2011; 24: 226–236.
Geschwind DH . Genetics of autism spectrum disorders. Trends Cogn Sci 2011; 15: 409–416.
Georgiades S, Szatmari P, Zwaigenbaum L, Bryson S, Brian J, Roberts W et al. A prospective study of autistic-like traits in unaffected siblings of probands with autism spectrum disorder. JAMA Psychiatry 2013; 70: 42–48.
Losh M, Childress D, Lam K, Piven J . Defining key features of the broad autism phenotype: a comparison across parents of multiple- and single-incidence autism families. Am J Med Genet B Neuropsychiatr Genet 2008; 147B: 424–433.
Schwichtenberg AJ, Young GS, Hutman T, Iosif AM, Sigman M, Rogers SJ et al. Behavior and sleep problems in children with a family history of autism. Autism Res 2013; 6: 169–176.
Carter MT, Scherer SW . Autism spectrum disorder in the genetics clinic: a review. Clin Genet 2013; 83: 399–407.
Gong J, Tong Y, Zhang HM, Wang K, Hu T, Shan G et al. Genome-wide identification of SNPs in microRNA genes and the SNP effects on microRNA target binding and biogenesis. Hum Mutat 2012; 33: 254–263.
Bruno AE, Li L, Kalabus JL, Pan Y, Yu A, Hu Z . miRdSNP: a database of disease-associated SNPs and microRNA target sites on 3'UTRs of human genes. BMC Genomics 2012; 13: 44.
Pilz DT, Matsumoto N, Minnerath S, Mills P, Gleeson JG, Allen KM et al. LIS1 and XLIS (DCX) mutations cause most classical lissencephaly, but different patterns of malformation. Hum Mol Genet 1998; 7: 2029–2037.
Marui T, Funatogawa I, Koishi S, Yamamoto K, Matsumoto H, Hashimoto O et al. Association of the neuronal cell adhesion molecule (NRCAM) gene variants with autism. Int J Neuropsychopharmacol 2009; 12: 1–10.
Pippucci T, Parmeggiani A, Palombo F, Maresca A, Angius A, Crisponi L et al. A novel null homozygous mutation confirms CACNA2D2 as a gene mutated in epileptic encephalopathy. PloS One 2013; 8: e82154.
Ramoz N, Cai G, Reichert JG, Silverman JM, Buxbaum JD . An analysis of candidate autism loci on chromosome 2q24-q33: evidence for association to the STK39 gene. Am J Med Genet B Neuropsychiatr Genet 2008; 147B: 1152–1158.
Correia C, Coutinho AM, Almeida J, Lontro R, Lobo C, Miguel TS et al. Association of the alpha4 integrin subunit gene (ITGA4) with autism. Am J Med Genet B Neuropsychiatr Genet 2009; 150B: 1147–1151.
Conroy J, Cochrane L, Anney RJ, Sutcliffe JS, Carthy P, Dunlop A et al. Fine mapping and association studies in a candidate region for autism on chromosome 2q31-q32. Am J Med Genet B Neuropsychiatr Genet 2009; 150B: 535–544.
Acknowledgements
We thank the families that participated in our study and R Tweedale, J Reinhard and A Larkin for critical reading of the manuscript. We acknowledge the financial support of the Commonwealth Department of Social Services (formerly the Department of Families, Housing, Community Services and Indigenous Affairs) and the Cooperative Research Centre for Living with Autism Spectrum Disorders the Autism CRC, established and supported under the Australian Government's Cooperative Research Centres program. CC was supported by funding from the Australian Research Council (FT110100292) and CC and AJOW by funding from the National Health and Medical Research Council (APP1008125; APP1004065). JYA was supported by a University of Queensland PhD scholarship and SMW by an Australian Postgraduate Award. We also thank John Beilby and PathWest for facilitating collection of samples and patient data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Translational Psychiatry website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
An, J., Cristino, A., Zhao, Q. et al. Towards a molecular characterization of autism spectrum disorders: an exome sequencing and systems approach. Transl Psychiatry 4, e394 (2014). https://doi.org/10.1038/tp.2014.38
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tp.2014.38
This article is cited by
-
Oligogenic inheritance in severe adult obesity
International Journal of Obesity (2024)
-
Cntnap2-dependent molecular networks in autism spectrum disorder revealed through an integrative multi-omics analysis
Molecular Psychiatry (2023)
-
Dysregulation of the Wnt/β-catenin signaling pathway via Rnf146 upregulation in a VPA-induced mouse model of autism spectrum disorder
Experimental & Molecular Medicine (2023)
-
Identification of primary copy number variations reveal enrichment of Calcium, and MAPK pathways sensitizing secondary sites for autism
Egyptian Journal of Medical Human Genetics (2020)
-
Autism-associated miR-873 regulates ARID1B, SHANK3 and NRXN2 involved in neurodevelopment
Translational Psychiatry (2020)