Abstract
Long non-coding RNAs (lncRNAs) have been demonstrated to play essential roles in diverse cellular processes and biological functions. Exploring the functions associated with lncRNAs may help provide insight into their underlying biological mechanisms. The current methods primarily focus on investigating the functions of individual lncRNAs; however, essential biological functions may be affected by the combinatorial effects of multiple lncRNAs. Here, we have developed a novel computational method, LncRNAs2Pathways, to identify the functional pathways influenced by the combinatorial effects of a set of lncRNAs of interest based on a global network propagation algorithm. A new Kolmogorov–Smirnov-like statistical measure weighted by the network propagation score, which considers the expression correlation among lncRNAs and coding genes, was used to evaluate the biological pathways influenced by the lncRNAs of interest. We have described the LncRNAs2Pathways methodology and illustrated its effectiveness by analyzing three lncRNA sets associated with glioma, prostate and pancreatic cancers. We further analyzed the reproducibility and robustness and compared our results with those of two other methods. Based on these analyses, we showed that LncRNAs2Pathways can effectively identify the functional pathways associated with lncRNA sets. Finally, we implemented this method as a freely available R-based tool.
Similar content being viewed by others
Introduction
Long non-coding RNAs (lncRNAs) are non-protein-coding transcripts >200 nucleotides that have been reported to play essential roles in diverse cellular processes and biological functions, such as transcriptional regulation, chromatin modification, cell differentiation, epigenetic regulation and immune responses1,2,3,4,5. More importantly, emerging evidence suggests that the dysregulation of lncRNAs is associated with the development and progression of a variety of human diseases, including cancer and other immune and neurological disorders6,7,8. Therefore, exploring the biological functions influenced by lncRNAs may help provide insight into the underlying mechanisms of lncRNAs in human diseases. With the development of next-generation sequencing technologies, tens of thousands of lncRNAs have been identified. As lncRNAs are generally weakly conserved in their primary sequences and interaction data between lncRNAs and other molecules are lacking9, the functions of most lncRNAs in complex diseases have not been widely studied.
Several recent methods of lncRNA function identification have been proposed. These methods can be classified into experimental and computational methods9. Although experimental methods can give more reliable lncRNA functions than computational methods, they are expensive and time consuming. Computational analysis to identify the probable functions of lncRNAs is promising as a way to guide further studies on lncRNAs. Guttman et al.10 assigned functions to ~1600 mouse long-intervening non-coding RNAs (lincRNAs) identified by chromatin-state maps based on coding–non-coding gene co-expression relationships extracted from custom-designed tiling array data. Liao et al.11 constructed a coding–non-coding gene co-expression network from re-annotated mouse microarray data and predicted the functions of 340 mouse lncRNAs based on topological or other network characteristics. Guo et al.12 developed a lncRNA global function predictor by integrating coding–non-coding co-expression data and protein interaction data. In this method, an information flow algorithm was applied, and 1625 mouse lncRNAs were functionally characterized. The LncRNAdb database included comprehensive functions for 280 eukaryotic cell lncRNAs by collecting curated literature evidence13. Linc2Go was developed as a functional annotation resource for lincRNA14. It integrated microRNA-mRNA and microRNA-lincRNA interaction data to generate comprehensive functional annotations for human long intergenic non-coding RNA based on the competing endogenous RNA hypothesis. The LncRNAtor platform collected 208 RNA-Seq datasets and identified co-expressed protein-coding genes of lncRNAs, which were then subjected to functional enrichment analysis15. Despite accumulating insights into lncRNA functions, the above studies primarily focused on investigating individual lncRNA functions and did not consider the combinatorial effects of multiple lncRNAs. However, recent evidence has demonstrated that essential cellular processes and biological functions may be affected by a set of lncRNAs, such as differentially expressed lncRNAs between cancer and normal samples16,17.
To identify the important functional terms affected by a set of lncRNAs, Jiang et al.18 developed a web interface named LncRNA2Function. LncRNA2Function first investigates the expression correlation between lncRNAs and protein-coding genes across the RNA-Seq data of 19 human normal tissues and then performs the hypergeometric test to functionally annotate a set of lncRNAs with significantly enriched functional terms among the protein-coding genes co-expressed with the lncRNAs. Zhao et al.19 introduced Co-LncRNA, a web-based computational tool that provides enrichment analyses of lncRNAs for Gene Ontology (GO) annotations and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways. Co-LncRNA collects 241 publicly available human RNA-Seq datasets and identifies the co-expressed protein-coding genes associated with multiple lncRNAs. The combinatorial effects of lncRNAs in the modulation of a given functional term are investigated by the simultaneous analysis of multiple lncRNAs. Although both LncRNA2Function and Co-LncRNA helped us to explore the combinatorial effects of a set of lncRNAs, only directly co-expressed protein-coding genes of lncRNAs were exploited, and their downstream genes were neglected. Typically, protein-coding and non-coding genes co-operate as a biological system. The network-based strategy has been successfully used for protein function annotation20 and tumor biomarker identification21, and thus, this strategy may be a promising way to address the combinatorial effects of a set of lncRNAs.
Here, we developed a novel computational method, LncRNAs2Pathways, to identify the functional pathways influenced by the combinatorial effects of a set of interesting lncRNAs based on a global network propagation algorithm. In this study, we first used 28 independent RNA-Seq datasets under different experimental conditions to extract gene co-expressed relationships of coding–coding, coding–non-coding and non-coding–non-coding. The co-expressed relationships were integrated with protein–protein interaction data to construct a coding–non-coding gene correlation (CNC) network, in which the nodes represent protein-coding and non-coding genes, and the edges are co-expression and protein–protein interactions. We then obtained a set of interesting lncRNAs, such as differentially expressed lncRNAs between disease and normal samples, and mapped them to the CNC network as source nodes. When a protein-coding gene is located closer to the lncRNAs, the gene may be more likely to be regulated. We used a global network propagation algorithm, random walk with restart (RWR), to calculate the propagation scores of protein-coding genes, which reflect the extent of genes influenced by the lncRNAs. A list of protein-coding genes was formed by ranking the protein-coding genes according to their propagation scores. Finally, we used a new Kolmogorov–Smirnov-like statistical measure weighted by the propagation scores to evaluate each pathway by mapping the genes in the pathway to the ranked gene list. We applied LncRNAs2Pathways to three sets of lncRNAs associated with prostate cancer, glioma, and pancreatic cancer. We then analyzed the reproducibility and robustness and compared our method with two other function analysis methods for lncRNAs. Our results indicate that LncRNAs2Pathways can produce biologically meaningful outcomes.
Materials and Methods
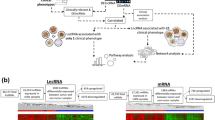
LncRNAs2Pathways was developed to identify the biological pathways influenced by the combinatorial effects of a set of lncRNAs of interest. Figure 1 depicts the flow diagram of the LncRNAs2Pathways. The main steps consist of (1) constructing the CNC network by integrating RNA-Seq data and protein–protein interaction data, (2) estimating the extent of protein-coding genes influenced by the set of lncRNAs of interest based on a global network propagation algorithm, and (3) calculating pathway enrichment scores (ESs) to evaluate the biological pathways. LncRNAs2Pathways has been implemented as a freely available R-based tool (https://cran.r-project.org/web/packages/LncPath/). The user inputs a set of lncRNAs of interest, and the biological pathways influenced by the lncRNAs are then returned.
Step 1. RNA-Seq data and protein–protein interaction data are integrated to construct a CNC network. Step 2. A set of lncRNAs of interest are mapped to the gene correlation network, and the global network propagation algorithm is used to calculate the propagation scores of protein-coding genes, which reflect the extent of the genes influenced by the lncRNAs. A ranked protein-coding gene list is constructed according to the propagation scores. Step 3. Protein-coding genes in a given pathway are mapped to the ranked protein-coding gene list, and the ES(P) is calculated by walking down the list. The permutation test is performed to identify the statistically significant pathways.
Constructing the CNC network
We collected 28 human RNA-Seq datasets covering a wide range of experimental and physiological conditions from the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) database22 (see Supplementary Table S1 for the list of datasets). Each dataset was required to contain at least six samples. For each dataset, RNA-Seq reads were mapped to the human genome by TopHat v2.0.1323, and the expression values of human lncRNAs and protein-coding genes were quantified by Cufflinks v2.2.124. Both TopHat and Cufflinks were performed with the default parameters. The gene expression values were measured as fragments per kilobase of exon per million fragments mapped (FPKM). For this study, annotations of all human lncRNAs and protein-coding genes were downloaded from GENCODE v2225.
We then extracted coding–coding, coding–non-coding and non-coding–non-coding gene co-expression relationships based on the 28 RNA-Seq datasets. For each dataset, the data were used as follows:
-
1
Genes (lncRNAs and protein-coding genes) with an average FPKM >115 and the variance of FPKM ranked in the top 75% of all genes in the dataset were retained11.
-
2
The Pearson correlation coefficient (Pcc) between the FPKM values of each gene pair was calculated. We applied Fisher’s z transform method26 to convert each correlation coefficient r to the z-statistic by z = 0.5[ln(1 + r)−ln(1−r)]. The result was approximately normal with a standard error of
 , where N is the sample size of the dataset. For each gene, we standardized the z-scores to enforce zero mean and unit variance, and then, a set of Pcc p-values was calculated for each gene pair. Bonferroni multiple test correction was implemented to adjust the p-values.
, where N is the sample size of the dataset. For each gene, we standardized the z-scores to enforce zero mean and unit variance, and then, a set of Pcc p-values was calculated for each gene pair. Bonferroni multiple test correction was implemented to adjust the p-values. -
3
Gene pairs in the given dataset were considered co-expressed if they had a Pcc value ranked in the top or bottom 0.1% for each gene and an adjusted p-value < 0.01.
According to the sign of the Pcc, gene co-expression relationships can be classified as positive correlation or negative correlation (Pcc > 0 or Pcc < 0). To derive reliable co-expressed relationships, we only retained co-expressed gene pairs with the same correlation direction (i.e., positive or negative) in three or more datasets. Thus, 114,006 co-expression relationships of coding–coding, coding–lncRNA and lncRNA–lncRNA genes were obtained (Supplementary Table S2).
To model real and comprehensive biological processes, we further integrated gene co-expression relationships with protein–protein interactions. Protein interaction data for humans were downloaded from four public available databases (Human Protein Reference Database [HPRD]27, the Database of Interacting Proteins [DIP]28, the Molecular INTeraction database [MINT]29 and Reactome30). We merged gene co-expression relationships with protein–protein interactions to construct a CNC network. The CNC network consists of 28,613 nodes (11,391 lncRNAs and 17,222 protein-coding genes) and 295,698 edges (104,391 gene co-expressions, 181,692 protein–protein interactions and 9,615 both gene co-expression and protein–protein interaction) (see Supplementary Table S2 for details about the CNC network). By detecting the degree distribution of the CNC network (Supplementary Figure S1), we found that it follows a power law distribution (P(k)~k−γ, γ = 1.71 and fitted line R-squared = 0.9185), which means that it is a scale-free network.
Estimating the extent of protein-coding genes influenced by lncRNAs of interest
We mapped a set of lncRNAs of interest, such as differentially expressed lncRNAs between disease and normal samples, to the CNC network as source nodes. The CNC network was constructed based on gene co-expression and protein–protein interaction, and the closer a protein-coding gene is located to the source nodes, the more likely this gene may be to be regulated. We used a global network propagation algorithm, RWR, to estimate the extent of protein-coding genes influenced by the source nodes. The RWR algorithm31 simulates an iterative walker that transitions from its current source node (or a set of source nodes simultaneously) to a randomly selected neighbor or returns to the source node(s) with a given probability. It can be used to compute the network-proximity of a node to the source node(s). This algorithm has been used to prioritize candidate disease genes32,33. Here, we applied the RWR algorithm to prioritize the protein-coding genes influenced by lncRNAs of interest in the CNC network. Formally, the RWR is defined as follows:

where M is the column-normalized adjacency matrix of the CNC network graph; pt is the vector of nodes at time step t, and its i-th element holds the probability of being at node i at time step t; and p0 is the initial probability vector of nodes, which was constructed by assigning to source nodes with 1 and other nodes with 0 and then normalized to a unit vector. This is equivalent to letting the random walker begin from each source node with equal probability. The parameter r is the restart probability of the walk in every time step at the source nodes. Kohler et al.32 proposed that r has only a slight effect on the results of the RWR algorithm when r varies from 0.1 to 0.9. In this study, the parameter r was set at 0.7.
The probability vector pt will converge to a unique steady state p∞after certain steps. This was achieved at query time by iterating until the L1-norm between pt and pt+1 fell below 10−10. p∞provides a measure of the extent of the genes in the CNC network influenced by the source nodes. At the steady state p∞, the protein-coding genes were assigned with the probabilities at their corresponding nodes in the CNC network graph. A protein-coding gene with a larger probability indicates that the gene locates closer to the source nodes and, thus, may be influenced by the source nodes to a greater extent. We then normalized the probabilities of protein-coding genes to their square roots and defined them as propagation scores. Finally, a protein-coding gene list L = < g1, g2,… gn> was generated by ranking the protein-coding genes in the CNC network according to their propagation scores.
Calculating pathway ESs to evaluate the biological pathways
We downloaded pathways from the KEGG databases34,35. Pathways with fewer than 15 protein-coding genes or more than 500 protein-coding genes in the CNC network were filtered out to avoid overly narrow or broad functional categories.
For each pathway, we mapped the protein-coding genes to the ranked gene list L. If the protein-coding genes in the pathway occur toward the top of the list L, the pathway will tend to be influenced by the combinatorial effects of the inputted lncRNAs. Inspired by Gene Set Enrichment Analysis (GSEA)36, we used a Kolmogorov–Smirnov-like statistic weighted by the propagation score to calculate an ES, which reflects the degree to which the pathway is overrepresented at the top of the ranked gene list L. The ES was calculated by walking down the list L. In detail, at a given position i in the ranked gene list L = < g1, g2,… gn>, we evaluated the fraction of genes in the pathway (FInP) weighted by their propagation scores and the fraction of genes not in the pathway (FNotP) as follows:


where tj is the propagation score of gene j, NNotP is the number of genes in list L not in the pathway, and p is used to weight the propagation scores of the genes in the pathway and was set to p = 1 in this study. With position i walking down the list L, a running-sum statistic FInP−FNotP was calculated by increasing it when we encounter a gene in the pathway and decreasing it when we encounter genes not in the pathway. The ES of the pathway (ES(P)) was defined as follows:

The ES(P) will be relatively high if the genes in the pathway are concentrated at the top of list L, but if the genes are randomly distributed in L, then the ES(P) will be correspondingly small.
We performed a permutation test to estimate the statistical significance (empirical p-value) of the ES(P). Specifically, we randomly selected a set of lncRNAs including the same number of lncRNAs with source nodes and mapped them to the CNC network. The ES(P) was then recomputed. A null distribution for the ES (designated as ESNULL) was generated after performing N permutations. The empirical p-value of the observed ES was calculated by comparing it with the set of scores in ESNULL, that is, p-value = M/N, where M is the number of ESNULL values greater than the observed ES(P). In this study, for the examples, the number of permutations N was set at 1000. To correct for multiple comparisons, we adjusted the empirical p-values using the false discovery rate (FDR)37.
In fact, not all of the members of a pathway will be influenced by the combinatorial effects of the inputted lncRNAs. Therefore, it is meaningful to extract the core members of high-scoring pathways that contribute to the ES(P). Here, we defined the core genes of the pathway to be the genes in the pathway that appear in the ranked gene list L at or before the point where the ES(P) is obtained. The core genes of a pathway may account for the influence from the inputted lncRNAs.
Results
Identification of pathways influenced by dysregulated lncRNAs in prostate cancer
Our first case was a set of differentially expressed lncRNAs in prostate cancer. We downloaded RNA-Seq data on prostate cancer retrieved from the NCBI SRA database (SRA ID: SRP006731).These data sequenced the transcriptome of seven LNCaP prostate cancer cell lines and four normal prostate epithelial cell lines38. Raw RNA-Seq data were used to produce transcriptome assemblies by applying the “TopHat v2.0.1323+ Cufflinks v2.2.124” pipeline with the default parameters. The expression values of lncRNAs and protein-coding genes were quantified using FPKM.
We applied Cuffdiff239 to identify differentially expressed lncRNAs between the tumors and normal samples. An lncRNA was considered to be differentially expressed when the adjusted p-value of the lncRNA in Cuffdiff2 was less than 0.01. A total of 60 differentially expressed lncRNAs were identified in the CNC network (see Supplementary Table S3 for details). We subsequently inputted these lncRNAs into LncRNAs2Pathways to identify KEGG pathways. With an FDR < 0.01 pathway significance threshold, LncRNAs2Pathways yielded 20 statistically significant pathways (Table 1). Most of these pathways are readily interpreted in terms of the current knowledge of prostate cancer. For instance, the inhibition of oxidative phosphorylation may trigger the reactive oxygen species-mediated death of human prostate cancer cells40; chemokine receptors are associated with enhanced adhesive and invasive activities41; cell signaling and regulators of the cell cycle are proposed to be molecular targets for prostate cancer prevention42; the suppression of focal adhesion kinase activity is shown to precede the induction of apoptosis of prostate cancer cells43; gap junctional intercellular communication is decreased in prostate cancer44; and regulation of the actin cytoskeleton is proposed to be associated with cancer cell migration and invasion45.
We take the chemokine signaling pathway as an example to explain the rationale underlying LncRNAs2Pathways. Specifically, we mapped the differentially expressed lncRNAs to the CNC network and calculated the propagation scores of protein-coding genes using the RWR algorithm. A protein-coding gene list L was formed by ranking the genes according to their propagation scores, which reflect the degree of influence of each of the differentially expressed lncRNAs. The genes in the chemokine signaling pathway were mapped to the ranked list L, and 186 genes were obtained (Fig. 2A). The goal of LncRNAs2Pathways is to determine whether the genes in the pathway tend to occur toward the top of the ranked gene list L, in which case the pathway may be influenced by the combinatorial effects of differentially expressed lncRNAs. We calculated an ES that reflects the degree to which the pathway genes cluster toward the top of the ranked list L. A running-sum statistic was calculated by walking down the ranked list L, increasing it when we encounter a gene in the pathway and decreasing it when we encounter genes not in the pathway (Fig. 2A). The ES is the maximum value of the statistic. The protein-coding genes that contributed to the ES were defined as core genes of the pathway, and 131 core genes were obtained (Supplementary Table S4).
(A) A running-sum statistic is calculated by walking down the protein-coding gene list, and the statistic’s maximum deviation from zero is used as the ES(P). (B) Chemokine signaling pathway in the KEGG database34,35. The gene products that correspond to the core protein-coding genes are annotated in red. (C) Heatmap of the expression levels of protein-coding genes in the pathway. Differentially expressed core protein-coding genes are marked in red.
We then mapped the core genes to the pathway graph in KEGG database, and a number of gene products were annotated (Fig. 2B). Most of these genes have been reported to be associated with the progression of prostate cancer. Activated cell division cycle 42 (CDC42)-associated kinase Ack1 promotes prostate cancer progression46. Ras homolog family member A (RHOA) regulates clinically relevant androgen action in prostate cancer cells47. Protein kinase C, alpha (PRKCA) mediates epidermal growth factor receptor transactivation in human prostate cancer cells48. Interestingly, these protein-coding genes are located close to the differentially expressed lncRNAs in the CNC network. Specifically, the shortest distances from CDC42 and RHOA to LncRNA metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) are all two, and the shortest distances from PRKCA to lncRNAs nuclear paraspeckle assembly transcript 1 (NEAT1), LINC00963 and MALAT1 are all three. More importantly, these lncRNAs are reported to be associated with the development of prostate cancer. MALAT1 down-regulation by siRNA inhibits prostate cancer cell growth, invasion and migration49; NEAT1 drives prostate cancer growth by altering the epigenetic landscape of target gene promoters to favor transcription50; and LINC00963 is involved in the transition of prostate cancer from androgen dependent to androgen independent51. These results indicate that the core genes in this pathway may be influenced by the dysregulated lncRNAs in the development of prostate cancer.
To further validate our results, we tested the expression changes of core genes in the significant pathways between prostate tumors and normal samples. We applied the same RNA-Seq data of prostate cancer and quantified the protein-coding gene expression levels with FPKM. The fold-change (FC) method was used to evaluate the changes in gene expression levels between tumors and normal samples. A coding gene was considered to be differentially expressed when the |log2 (FC)| value of the gene exceeded 1 (i.e., FC > 2 or FC < 0.5). For the chemokine signaling pathway, more than 40% (54/131) of its core genes are differentially expressed (Fig. 2C). This significant pathway was identified by applying the LncRNAs2Pathways method to the set of differentially expressed lncRNAs and observing that the expressions of a number of core genes in the pathways changed accordingly. Thus, the coding genes in the pathway may be influenced by the differentially expressed lncRNAs in the progression of prostate cancer. We also tested other significant pathways, such as cell cycle and gap junction pathways, and found that 43.6% (41/94) and 46.2% (30/65) of their core genes are differentially expressed in these pathways, respectively (Supplementary Figures S2 and S3).
Identification of pathways influenced by the lncRNAs in glioma
Our second case was a set of lncRNAs associated with glioma, which was downloaded from Lnc2Cancer, a manually curated database of experimentally supported cancer-associated lncRNAs52. We obtained 11 glioma-associated lncRNAs in the CNC network (Supplementary Table S5). We inputted these lncRNAs to LncRNAs2Pathways, and 24 significant pathways (FDR < 0.01) were identified (Table 2). Most of these pathways are clearly related to glioma. For instance, the activation of the mitogen-activated protein kinase (MAPK) signaling pathway is associated with the chemosensitivity of glioma cells53; the neuregulin-1/ERBB receptor signaling cascade contributes to enhancing the survival of human astrocytic glioma cells54; impairment of the cell cycle is associated with the growth inhibition of human glioma cells55; blocking the focal adhesion pathway has the potential to be an efficacious treatment for human gliomas56; and neurotrophin signaling could be a target for the combinatorial treatment of malignant glioma57. The glioma pathway was also found to be significant. We took this pathway as an example to illustrate how the significant pathways were identified by LncRNAs2Pathways.
The 11 glioma-associated lncRNAs were mapped to the CNC network. The propagation scores of the protein-coding genes, which reflect the degree of coding genes influenced by the inputted lncRNAs, were calculated by the RWR algorithm. A coding gene list was constructed by ranking the coding genes according to their propagation scores. The genes in the glioma pathway were mapped to the ranked coding gene list. If the coding genes in the pathway cluster at the top of the ranked list, the pathway will be regulated by the combinatorial effects of the lncRNA set. We calculated a running-sum statistic by walking down the list (Fig. 3A), and the maximum value of the statistic was used as the ES(P), which reflects the extent to which the pathway is overrepresented at the top of the ranked gene list. The core genes of the pathway that contribute to the ES were extracted and annotated on the original pathway graph (Fig. 3B). In the pathway, almost all the annotated genes were reported to be associated with the initiation and progression of glioma. Epidermal growth factor receptor (EGFR) promotes the malignant potential of glioma cells by interacting with the functional subunit of the cysteine/glutamate transporter xc-system (xCT) at the cell surface58; the expression of insulin-like growth factor 1 receptor (IGF1R) was found to be associated with the proliferation, migration, invasion, and tumorigenesis of glioma cells59. The phosphoinositol phosphatase activity of phosphatase and tensin homolog (PTEN) mediates serum-sensitive G1 growth arrest in glioma cells60. Neuroblastoma RAS viral oncogene homolog (NRAS) was identified as promoting oncogenesis in glioma stem cells61, and the upregulation of growth factor receptor bound protein 2 (GRB2)-associated binder 2 was demonstrated to be correlated with glioma62. Moreover, we extracted the connected sub-network among core protein-coding genes and inputted lncRNAs in the CNC network, which revealed that none of the core genes in the pathway were direct neighbors of the inputted lncRNAs. However, they were located close to the inputted lncRNAs (Fig. 3C). For instance, the average shortest distance from EGFR, IGF1R, PTEN, NRAS or GRB2 to the inputted lncRNAs did not exceed three. This result suggests that the core genes may be regulated by the lncRNAs. If we focus only on the co-expressed protein-coding genes of the lncRNAs, this pathway would be neglected. Our LncRNAs2Pathways method, which applies the global network propagation algorithm, successfully identified this pathway.
(A) A running-sum statistic is calculated by walking down the protein-coding gene list, and the statistic’s maximum deviation from zero is used as the ES(P). (B) Glioma pathway in the KEGG database34,35. The gene products that correspond to the core protein-coding genes are annotated in red. (C) Connected sub-network of core protein-coding genes (marked with blue) and inputted lncRNAs (marked with red) in the CNC network.
Identification of pathways influenced by lncRNAs in pancreatic cancer
The third case we used was a set of lncRNAs associated with pancreatic cancer. We downloaded this set from the Lnc2Cancer database, and a total of nine pancreatic-cancer associated lncRNAs were identified in the CNC network (Supplementary Table S6). With FDR < 0.01, LncRNAs2Pathways obtained seven statistically significant pathways (Table 3), examples of which include the following: arachidonic acid metabolism, in which lipoxygenases, which are the key constituents, play a critical role in pancreatic cancer cell proliferation63; oxidative phosphorylation, whose KCa3.1 channel was identified as a regulator in a subset of pancreatic carcinoma cell lines64; the reduction of retinoids and their receptors, which is associated with pancreatic cancer patient survival65; and the overexpression of ribosomal proteins, which can promote tumorigenesis by interacting with the p53 tumor suppressor66. For the significant pathways, such as arachidonic acid metabolism, 14 core genes were identified (Supplementary Figure S4A). Because the core genes contributed to the ES(P), they might tend to be regulated by the pancreatic-cancer associated lncRNAs in the development of cancer. By mapping the core genes to the pathway graph (Supplementary Figure S4B), we found that 10 enzymes corresponding to the catalytic reaction of arachidonate were annotated. Interestingly, arachidonate has been reported to play a key role in carcinogenesis67. These results indicate that dysfunction of the arachidonic acid metabolism pathway may be regulated by the combinatorial effects of lncRNAs associated with pancreatic cancer.
Comparing two studies of prostate cancer
We applied LncRNAs2Pathways to two independent lncRNA sets associated with prostate cancer to test whether reproducible results could be obtained. The two lncRNA sets are as follows: (i) 60 differentially expressed lncRNAs (calculated by Cuffdiff239, adjusted p-value < 0.01) derived from RNA-Seq data proposed by Kim et al. (SRA ID: SRP006731)38, which was used in our first case; (ii) 8 differentially expressed lncRNAs derived from RNA-Seq data proposed by Kannan et al. (SRA ID: SRP002628)68. Only one lncRNA was shared between the two lncRNA sets. Our goal was to examine whether LncRNAs2Pathways could identify consistent significant pathways. With FDR < 0.01, 20 and 19 pathways were identified by LncRNAs2Pathways in the two sets, respectively. We found that over 50% of the statistically significant pathways (12 pathways) were shared between the two studies (Fig. 4A). The shared pathways, such as the oxidative phosphorylation, chemokine signaling, focal adhesion, and gap junction pathways, have been proposed to be clearly related to the progression of prostate cancer40,41,43,44. To explain why the same significant pathways were identified using different lncRNA sets, we further compared the core genes in the pathways. For example, in the chemokine signaling pathway, 131 and 137 core genes were found in the two studies, and 121 were shared between them. The large overlap between the core genes in the two studies indicates that although the lncRNAs in the two sets are different, they may regulate consistent functional pathways associated with prostate cancer.
(A) Venn diagram of the overlapping significant pathways identified by two prostate cancer-associated lncRNA sets derived from the SRP006731 and SRP002628 datasets. (B–D) The mean recalled ratio of the significant pathways calculated by randomly deleting 5%, 10%, 15%, 20%, 25% and 30% of edges in the CNC network for the three lncRNA sets associated with prostate cancer, glioma and pancreatic cancer, respectively.
We then introduced a third prostate cancer-associated lncRNA set downloaded from the Lnc2Cancer database, which includes 28 lncRNAs. To provide a more general comparison, we compared significant pathways other than those that satisfied FDR < 0.01. Specifically, the top scoring 30 pathways determined by each of the three lncRNA sets were considered. Interestingly, half of these pathways (15/30) were shared across the three studies. The above analysis shows that LncRNAs2Pathways could obtain reproducible results across the three lncRNA sets associated with prostate cancer.
Robustness analysis for the CNC network
The robustness analysis was performed by deleting a portion of the edges in the CNC network. Specifically, we randomly deleted 5%, 10%, 15%, 20%, 25% and 30% of the edges in the CNC network and repeated the LncRNAs2Pathways method 20 times for each deletion. We then calculated the mean recalled ratio of the original significant pathways for each deletion separately. We performed robustness analysis on the three aforementioned lncRNA sets of interest, namely, the differentially expressed lncRNA set in prostate cancer, glioma-associated lncRNA set and pancreatic cancer-associated lncRNA set. For the lncRNA set in prostate cancer, 20 statistically significant pathways were identified (FDR < 0.01). With 5%, 10%, 15%, and 20% of edges deleted, the mean recalled ratio of the significant pathways fell slowly, and the mean recalled ratio of the significant pathways exceeded 70%. With 25% and 30% edges deleted, the results were slightly inferior, but the mean recalled ratio of the significant pathways remained above 55% (Fig. 4B). We repeated this operation on the glioma-associated lncRNA set and pancreatic cancer-associated lncRNA set, obtaining similar results (Fig. 4C and D). The results of these experiments show that LncRNAs2Pathways is robust against edge deletion in the CNC network. This finding could be explained by the fact that the global network propagation algorithm used by our method can propagate the effects of lncRNAs effectively, even if the CNC network is incomplete.
Comparison of LncRNAs2Pathways with other methods
To determine whether LncRNAs2Pathways can provide new insight into the identification of the pathways regulated by a given lncRNA set, we compared the results of LncRNAs2Pathways with those of two other methods: LncRNA2Function18 and Co-LncRNA19. We applied all three methods to identify significant pathways based on the differentially expressed lncRNAs in prostate cancer and the lncRNAs associated with glioma. For the differentially expressed lncRNAs in prostate cancer, a total of 38 statistically significant pathways were identified by the three methods using the default threshold for each method (Supplementary Table S7). Specifically, LncRNA2Function found 18 statistically significant pathways (Benjamini-Hochberg (BH)‘s FDR < 0.01), Co-LncRNA found 3 statistically significant pathways (Bonferroni adjusted p-value < 0.01), and LncRNAs2Pathways identified 20 significant pathways (BH’s FDR < 0.01). By comparing these results, we found that 17 pathways identified by LncRNAs2Pathways were missed by both other methods (Supplementary Table S7). Some of the missed pathways, such as the oxidative phosphorylation, cell cycle and focal adhesion pathways, have been well documented as being related to prostate cancer40,42,43. The pathways identified by LncRNAs2Pathways but missed by the other two methods may be attributed to the fact that LncRNA2Function and Co-LncRNA mainly use the hypergeometric test to identify significant pathways by annotating the protein-coding genes that are significantly co-expressed with the lncRNAs in the pathways. In contrast, LncRNAs2Pathways identifies significant pathways using a global network propagation algorithm. In LncRNAs2Pathways, the RWR algorithm was applied to calculate the propagation score for each protein-coding gene, which reflects the average network-proximity of the coding gene to the inputted lncRNAs. A coding gene located close to the lncRNAs will be assigned a larger propagation score. The Kolmogorov–Smirnov-like statistic weighted by the propagation score was used to evaluate the combinatorial effect of the lncRNAs in the biological pathways. Thus, LncRNAs2Pathways could identify new pathways whose genes are not co-expressed with the inputted lncRNAs but are close to the lncRNAs. We also applied the three methods to the set of lncRNAs associated with glioma, and a total of 37 statistically significant pathways were identified (Supplementary Table S8). Interestingly, LncRNAs2Pathways exclusively identified 20 significant pathways. The above results indicate that LncRNAs2Pathways may uncover some new biological pathways associated with a given lncRNA set and may therefore complement currently used methods.
Discussion
LncRNAs have been reported to be involved in a wide range of biological processes1,2,3,4,5 and complex human diseases6,7,8. From a systems biology perspective, the LncRNAs generally perform their biological functions jointly rather than individually16,17. Thus, the identification of biological functions influenced by the combinatorial effects of a set of lncRNAs of interest is indispensable. Our study is the first to predict probable functions regulated by the combinatorial effects of a set of lncRNAs of interest based on a global network propagation strategy. In this study, we first constructed a CNC network by integrating RNA-Seq datasets and protein–protein interactions. Then, a set of lncRNAs was mapped to the network as source nodes, and the RWR algorithm was applied to evaluate the extent of protein-coding genes influenced by the combinatorial effects of the source nodes. Finally, we used a Kolmogorov–Smirnov-like statistic weighted by the influenced extent to prioritize the functional pathways. The results show that LncRNAs2Pathways can effectively identify the functional pathways associated with the lncRNA sets. To make the method more broadly applicable, we have implemented LncRNAs2Pathways as an R-based tool, which is freely available on the Comprehensive R Archive Network (CRAN; https://cran.r-project.org/web/packages/LncPath/). The users input an interesting lncRNA set, and the significant pathways are then returned. It is expected that LncRNAs2Pathways could facilitate the study of lncRNAs and further guide experimental design for biologists.
We collected 28 human RNA-Seq datasets under different biological conditions. For each dataset, we extracted the co-expression relationships among genes (both lncRNAs and protein-coding genes) through a strict computational pipeline (see the Methods section). The co-expressed gene pairs with the same correlation direction (i.e., positive or negative) in more than a certain number of datasets were retained for further analysis. To determine the number of datasets, we analyzed several network parameters with different dataset number cutoffs (from two to nine) (Supplementary Table S9). The size of the network naturally decreased as the cutoff value increased. The co-expression networks were confirmed with cutoffs of three or more datasets whose degree distributions approximately obey a power law distribution (Supplementary Figure S5), as observed for many biological networks and other types of networks69,70. Other studies of co-expression among genes also consider two genes to be co-expressed based on at least three datasets11,71. In this study, based on the size and quality of the networks, we only retained co-expressed gene pairs with a cutoff of three datasets for further analysis. Alternatively, the co-expression relationships among genes can also be extracted by combining p-values or effect sizes of Pcc across multiple datasets72. Because our method is flexible, researchers could also apply the algorithm to their network of interest using our software package.
In our study, to model real biological processes as accurately as possible, we further integrated gene co-expression relationships with protein–protein interactions to construct the CNC network. The CNC network consists of 28,613 nodes (11,391 lncRNAs and 17,222 protein-coding genes) and 295,698 edges (see Supplementary Table S2 for details about the CNC network). Previously, the integration of gene co-expression relationships with protein–protein interactions was performed by Guo et al.12 to predict probable functions for lncRNAs individually; however, we used this strategy to identify the function pathways influenced by the combinatorial effects of a set of lncRNAs. Moreover, the method used by Guo et al. used re-annotated microarray data corresponding to only 1,713 lncRNAs to extract gene co-expression relationships; in contrast, the CNC network covers 11,391 lncRNAs.
To investigate the influence of the CNC network, we performed data removal tests by randomly deleting 5%, 10%, 15%, 20%, 25% and 30% of edges in the CNC network. For each deletion, the LncRNAs2Pathways method was repeated 20 times. For the lncRNA set that was differentially expressed in prostate cancer, the mean recalled ratio of the original significant pathways remained above 55%, even after the deletion of up to 30% of edges in the CNC network (Fig. 4B). For the glioma-associated lncRNA set and pancreatic cancer-associated lncRNA set, we obtained similar results (Fig. 4C and D). These findings are attributable to the global network propagation algorithm, RWR31, used in our method. As long as the network is connected, this algorithm can propagate the functional effects of source nodes (i.e., the lncRNA set of interest) on the CNC network iteratively, even when the network is not comprehensive. Kohler et al. used this algorithm to prioritize candidate disease genes by mapping known disease genes to the protein interaction network32. We applied this algorithm to evaluate the extent of protein-coding genes influenced by the given set of lncRNAs in the CNC network.
Our method was designed to identify the pathways influenced by the combinatorial effects of a set of lncRNAs of interest, such as differentially expressed lncRNAs in a disease, and thus, the pathways identified by our method may be specifically regulated by the dysregulated lncRNAs in the disease state. To explain the effectiveness of LncRNAs2Pathways, we applied the method to three separate sets of lncRNAs. For the lncRNAs associated with prostate cancer, LncRNAs2Pathways identified 20 statistically significant pathways (FDR < 0.01, Table 1). Most of these pathways have been reported to be associated with the progression of prostate cancer. Because the pathways were obtained by mapping the differentially expressed lncRNAs in prostate cancer to the CNC network, they may be regulated by the combinatorial effects of the set of lncRNAs. We took the chemokine signaling pathway as an example to validate our results by testing the expression changes of core genes in significant pathways between tumors and normal samples. Interestingly, we found that more than 40% (54/131) of the core genes are differentially expressed (Fig. 2C). The pathway was identified by the differentially expressed lncRNAs, and the expression levels of the core genes in the pathways changed accordingly. Therefore, the pathway may indeed be regulated by the differentially expressed lncRNAs. For the lncRNAs associated with glioma, 24 significant pathways were identified (FDR < 0.01, Table 2), and the glioma pathway was identified as significant. By detecting the distances between the core genes of the pathway and inputted lncRNAs in the CNC network, we found that the core genes were not direct neighbors of the inputted lncRNAs but were located near the inputted lncRNAs in the CNC network (Fig. 3C). These results suggested that if we focused only on the protein-coding genes that are significantly co-expressed with the lncRNAs, this pathway might be neglected. The pathway obtained by LncRNAs2Pathways may be ascribed to the Kolmogorov–Smirnov-like statistic weighted by the propagation score used in our method.
Moreover, LncRNAs2Pathways can also prioritize the pathways influenced by a single lncRNA. For example, maternally expressed 3 (MEG3) is a maternally expressed, imprinted lncRNA gene that acts as a growth suppressor in tumor cells73. We inputted this lncRNA into LncRNAs2Pathways and obtained five statistically significant pathways with FDR < 0.01: (i) ribosome, (ii) focal adhesion, (iii) extracellular matrix (ECM)-receptor interaction, (iv) regulation of actin cytoskeleton, and (v) pathways in cancer. Interestingly, the changes in the ribosome biogenesis pathway reflect an increased incidence of tumor onset74; focal-adhesion kinase mediates cell proliferation, cell survival and cell migration75; the ECM-receptor interaction contributes to the venous metastases of hepatocellular carcinoma76; and regulation of the actin cytoskeleton is associated with cancer cell migration and invasion45. These results indicate that the significant pathways may be regulated by MEG3 in the beginning and during the progression of cancer.
Although a global network propagation strategy was successfully exploited in LncRNAs2Pathways for the functional annotation of a set of lncRNAs, our method can be improved in the following two ways. First, the cancer-associated lncRNA sets used in the study may be incomplete. In this work, we collected the lncRNAs associated with glioma and pancreatic cancers from the Lnc2Cancer database, and obtained 11 and 9 lncRNAs, respectively (Supplementary Tables S5 and S6). Although this database collected all experimentally supported cancer-associated lncRNAs from recent studies, a number of lncRNAs associated with the cancers remain to be validated. For prostate cancer, with an adjusted p-value < 0.01 in Cuffdiff239, 60 differentially expressed lncRNAs were identified, but several moderately cancer-associated lncRNAs may have been missed. If the data on cancer-associated lncRNAs are improved, the specificity of LncRNAs2Pathways will be enhanced. Second, our method regarded the lncRNAs in the set of interest as equally important and did not consider the extent of the association between lncRNAs and cancer. Other network-based prediction algorithms should be incorporated to achieve better performance. Taken together, LncRNAs2Pathways not only provides a function predictor for lncRNA sets but also an open computational framework for the study of the combinatorial effects of lncRNAs.
Additional Information
How to cite this article: Han, J. et al. LncRNAs2Pathways: Identifying the pathways influenced by a set of lncRNAs of interest based on a global network propagation method. Sci. Rep. 7, 46566; doi: 10.1038/srep46566 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Yang, J. H., Li, J. H., Jiang, S., Zhou, H. & Qu, L. H. ChIPBase: a database for decoding the transcriptional regulation of long non-coding RNA and microRNA genes from ChIP-Seq data. Nucleic Acids Res 41, D177–187 (2013).
Bernard, D. et al. A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. Embo J 29, 3082–3093 (2010).
Lee, J. T. Epigenetic regulation by long noncoding RNAs. Science 338, 1435–1439 (2012).
Fatica, A. & Bozzoni, I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet 15, 7–21 (2013).
Batista, P. J. & Chang, H. Y. Long noncoding RNAs: cellular address codes in development and disease. Cell 152, 1298–1307 (2013).
Mitra, S. A., Mitra, A. P. & Triche, T. J. A central role for long non-coding RNA in cancer. Front Genet 3, 17 (2012).
Bhan, A. & Mandal, S. S. Long noncoding RNAs: emerging stars in gene regulation, epigenetics and human disease. ChemMedChem 9, 1932–1956 (2014).
Wapinski, O. & Chang, H. Y. Long noncoding RNAs and human disease. Trends Cell Biol 21, 354–361 (2011).
Guo, X. et al. Advances in long noncoding RNAs: identification, structure prediction and function annotation. Brief Funct Genomics 15, 38–46 (2016).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Liao, Q. et al. Large-scale prediction of long non-coding RNA functions in a coding-non-coding gene co-expression network. Nucleic Acids Res 39, 3864–3878 (2011).
Guo, X. et al. Long non-coding RNAs function annotation: a global prediction method based on bi-colored networks. Nucleic Acids Res 41, e35 (2013).
Quek, X. C. et al. lncRNAdb v2.0: expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res 43, D168–173 (2015).
Liu, K., Yan, Z., Li, Y. & Sun, Z. Linc2GO: a human LincRNA function annotation resource based on ceRNA hypothesis. Bioinformatics 29, 2221–2222 (2013).
Park, C., Yu, N., Choi, I., Kim, W. & Lee, S. lncRNAtor: a comprehensive resource for functional investigation of long non-coding RNAs. Bioinformatics 30, 2480–2485 (2014).
Hu, Y. et al. A long non-coding RNA signature to improve prognosis prediction of colorectal cancer. Oncotarget 5, 2230–2242 (2014).
Li, J. et al. LncRNA profile study reveals a three-lncRNA signature associated with the survival of patients with oesophageal squamous cell carcinoma. Gut 63, 1700–1710 (2014).
Jiang, Q. et al. LncRNA2Function: a comprehensive resource for functional investigation of human lncRNAs based on RNA-seq data. BMC Genomics 16 Suppl 3, S2 (2015).
Zhao, Z. et al. Co-LncRNA: investigating the lncRNA combinatorial effects in GO annotations and KEGG pathways based on human RNA-Seq data. Database (Oxford) 2015 (2015).
Sharan, R., Ulitsky, I. & Shamir, R. Network-based prediction of protein function. Mol Syst Biol 3, 88 (2007).
Zhang, F. et al. A network medicine approach to build a comprehensive atlas for the prognosis of human cancer. Brief Bioinform (2016).
Leinonen, R., Sugawara, H. & Shumway, M. The sequence read archive. Nucleic Acids Res 39, D19–21 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14, R36 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28, 511–515 (2010).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 22, 1775–1789 (2012).
Li, W. et al. High-resolution functional annotation of human transcriptome: predicting isoform functions by a novel multiple instance-based label propagation method. Nucleic Acids Res 42, e39 (2014).
Goel, R., Harsha, H. C., Pandey, A. & Prasad, T. S. Human Protein Reference Database and Human Proteinpedia as resources for phosphoproteome analysis. Mol Biosyst 8, 453–463 (2012).
Salwinski, L. et al. The Database of Interacting Proteins: 2004 update. Nucleic Acids Res 32, D449–451 (2004).
Licata, L. et al. MINT, the molecular interaction database: 2012 update. Nucleic Acids Res 40, D857–861 (2012).
Croft, D. et al. Reactome: a database of reactions, pathways and biological processes. Nucleic Acids Res 39, D691–697 (2011).
Can, T., Camoglu, O. & Singh, A. K. Analysis of ProteinProtein Interaction Networks Using Random Walks. Proceedings of the 5th international workshop on Bioinformatics page 61–68 (2005).
Kohler, S., Bauer, S., Horn, D. & Robinson, P. N. Walking the interactome for prioritization of candidate disease genes. Am J Hum Genet 82, 949–958 (2008).
Han, J. et al. A novel dysregulated pathway-identification analysis based on global influence of within-pathway effects and crosstalk between pathways. J R Soc Interface 12, 20140937 (2015).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40, D109–114 (2012).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45, D353–D361 (2017).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550 (2005).
Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society, Series B (Methodological) 57 (1), 289–300 (1995).
Kim, J. H. et al. Deep sequencing reveals distinct patterns of DNA methylation in prostate cancer. Genome Res 21, 1028–1041 (2011).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat Biotechnol 31, 46–53 (2013).
Xiao, D. et al. Phenethyl isothiocyanate inhibits oxidative phosphorylation to trigger reactive oxygen species-mediated death of human prostate cancer cells. J Biol Chem 285, 26558–26569 (2010).
Wang, J. et al. The role of CXCR7/RDC1 as a chemokine receptor for CXCL12/SDF-1 in prostate cancer. J Biol Chem 283, 4283–4294 (2008).
Agarwal, R. Cell signaling and regulators of cell cycle as molecular targets for prostate cancer prevention by dietary agents. Biochem Pharmacol 60, 1051–1059 (2000).
Kyle, E., Neckers, L., Takimoto, C., Curt, G. & Bergan, R. Genistein-induced apoptosis of prostate cancer cells is preceded by a specific decrease in focal adhesion kinase activity. Mol Pharmacol 51, 193–200 (1997).
Habermann, H., Ray, V., Habermann, W. & Prins, G. S. Alterations in gap junction protein expression in human benign prostatic hyperplasia and prostate cancer. J Urol 167, 655–660 (2002).
Yamaguchi, H. & Condeelis, J. Regulation of the actin cytoskeleton in cancer cell migration and invasion. Biochim Biophys Acta 1773, 642–652 (2007).
Mahajan, N. P. et al. Activated Cdc42-associated kinase Ack1 promotes prostate cancer progression via androgen receptor tyrosine phosphorylation. Proc Natl Acad Sci USA 104, 8438–8443 (2007).
Schmidt, L. J. et al. RhoA as a mediator of clinically relevant androgen action in prostate cancer cells. Mol Endocrinol 26, 716–735 (2012).
Stewart, J. R. & O’Brian, C. A. Protein kinase C-{alpha} mediates epidermal growth factor receptor transactivation in human prostate cancer cells. Mol Cancer Ther 4, 726–732 (2005).
Ren, S. et al. Long noncoding RNA MALAT-1 is a new potential therapeutic target for castration resistant prostate cancer. J Urol 190, 2278–2287 (2013).
Chakravarty, D. et al. The oestrogen receptor alpha-regulated lncRNA NEAT1 is a critical modulator of prostate cancer. Nat Commun 5, 5383 (2014).
Wang, L. et al. Linc00963: a novel, long non-coding RNA involved in the transition of prostate cancer from androgen-dependence to androgen-independence. Int J Oncol 44, 2041–2049 (2014).
Ning, S. et al. Lnc2Cancer: a manually curated database of experimentally supported lncRNAs associated with various human cancers. Nucleic Acids Res 44, D980–985 (2016).
Wang, Q. et al. Silence of bFGF enhances chemosensitivity of glioma cells to temozolomide through the MAPK signal pathway. Acta Biochim Biophys Sin (Shanghai) 48, 501–508 (2016).
Ritch, P. S., Carroll, S. L. & Sontheimer, H. Neuregulin-1 enhances survival of human astrocytic glioma cells. Glia 51, 217–228 (2005).
Otsuka, G. et al. Inhibition of nuclear factor-kappaB activation confers sensitivity to tumor necrosis factor-alpha by impairment of cell cycle progression in human glioma cells. Cancer Res 59, 4446–4452 (1999).
Liu, T. J. et al. Inhibition of both focal adhesion kinase and insulin-like growth factor-I receptor kinase suppresses glioma proliferation in vitro and in vivo . Mol Cancer Ther 6, 1357–1367 (2007).
Lawn, S. et al. Neurotrophin signaling via TrkB and TrkC receptors promotes the growth of brain tumor-initiating cells. J Biol Chem 290, 3814–3824 (2015).
Tsuchihashi, K. et al. The EGF Receptor Promotes the Malignant Potential of Glioma by Regulating Amino Acid Transport System xc(-). Cancer Res 76, 2954–2963 (2016).
Shi, Z. M. et al. MiRNA-181b suppresses IGF-1R and functions as a tumor suppressor gene in gliomas. Rna 19, 552–560 (2013).
Furnari, F. B., Huang, H. J. & Cavenee, W. K. The phosphoinositol phosphatase activity of PTEN mediates a serum-sensitive G1 growth arrest in glioma cells. Cancer Res 58, 5002–5008 (1998).
Gong, W. et al. Knockdown of NEAT1 restrained the malignant progression of glioma stem cells by activating microRNA let-7e. Oncotarget (2016).
Liu, H. et al. Combined detection of Gab1 and Gab2 expression predicts clinical outcome of patients with glioma. Med Oncol 31, 77 (2014).
Ding, X. Z., Iversen, P., Cluck, M. W., Knezetic, J. A. & Adrian, T. E. Lipoxygenase inhibitors abolish proliferation of human pancreatic cancer cells. Biochem Biophys Res Commun 261, 218–223 (1999).
Kovalenko, I. et al. Identification of KCa3.1 Channel as a Novel Regulator of Oxidative Phosphorylation in a Subset of Pancreatic Carcinoma Cell Lines. PLoS One 11, e0160658 (2016).
Bleul, T. et al. Reduced retinoids and retinoid receptors’ expression in pancreatic cancer: A link to patient survival. Mol Carcinog 54, 870–879 (2015).
Shenoy, N. et al. Alterations in the ribosomal machinery in cancer and hematologic disorders. J Hematol Oncol 5, 32 (2012).
Yarla, N. S. et al. Targeting arachidonic acid pathway by natural products for cancer prevention and therapy. Semin Cancer Biol (2016).
Kannan, K. et al. Recurrent chimeric RNAs enriched in human prostate cancer identified by deep sequencing. Proc Natl Acad Sci USA 108, 9172–9177 (2011).
Barabasi, A. L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Bhan, A., Galas, D. J. & Dewey, T. G. A duplication growth model of gene expression networks. Bioinformatics 18, 1486–1493 (2002).
Lee, H. K., Hsu, A. K., Sajdak, J., Qin, J. & Pavlidis, P. Coexpression analysis of human genes across many microarray data sets. Genome Res 14, 1085–1094 (2004).
Nguyen, T., Diaz, D., Tagett, R. & Draghici, S. Overcoming the matched-sample bottleneck: an orthogonal approach to integrate omic data. Sci Rep 6, 29251 (2016).
Zhang, X. et al. A pituitary-derived MEG3 isoform functions as a growth suppressor in tumor cells. J Clin Endocrinol Metab 88, 5119–5126 (2003).
Montanaro, L., Trere, D. & Derenzini, M. Changes in ribosome biogenesis may induce cancer by down-regulating the cell tumor suppressor potential. Biochim Biophys Acta 1825, 101–110 (2012).
McLean, G. W. et al. The role of focal-adhesion kinase in cancer - a new therapeutic opportunity. Nat Rev Cancer 5, 505–515 (2005).
Zhang, H. et al. Comparative transcriptome analysis reveals that the extracellular matrix receptor interaction contributes to the venous metastases of hepatocellular carcinoma. Cancer Genet 208, 482–491 (2015).
Acknowledgements
We thank all the research staff for their contributions to this project. We would like to thank the GSEA team for their research. We thank the KEGG Support Team for providing the copyright permission of KEGG imagery. We also thank the editor and anonymous reviewers sincerely for their time and helpful comments. This work was supported in part by the National Natural Science Foundation of China (grant nos 31401127, 91439117, 81472367, 61603116, 31501074), the Heilongjiang Province Education Department of Scientific Research Projects (grant no. 12541474), the China Postdoctoral Science Foundation (2016M601443), the Postdoctoral Foundation of Heilongjiang Province (LBH-Z16130), the National Program on Key Basic Research Project (973 program, grant no. 2014CB910504).
Author information
Authors and Affiliations
Contributions
J.H., C.L. and X.L. conceived and designed the study. S.L., Y.Z. and Y.X. developed software. Z.S., F.Z. and C.Z. analyzed the data and implemented the methodology. H.R. and H.Y. revised the manuscript. D.S. and F.S. provided constructive discussions. J.H. and C.L. drafted the manuscript. All the authors read and agreed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Han, J., Liu, S., Sun, Z. et al. LncRNAs2Pathways: Identifying the pathways influenced by a set of lncRNAs of interest based on a global network propagation method. Sci Rep 7, 46566 (2017). https://doi.org/10.1038/srep46566
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep46566
This article is cited by
-
Immune-related 3-lncRNA signature with prognostic connotation in a multi-cancer setting
Journal of Translational Medicine (2022)
-
Comprehensive analysis of LncRNAs expression profiles in an in vitro model of steatosis treated with Exendin-4
Journal of Translational Medicine (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.




 , where N is the sample size of the dataset. For each gene, we standardized the z-scores to enforce zero mean and unit variance, and then, a set of Pcc p-values was calculated for each gene pair. Bonferroni multiple test correction was implemented to adjust the p-values.
, where N is the sample size of the dataset. For each gene, we standardized the z-scores to enforce zero mean and unit variance, and then, a set of Pcc p-values was calculated for each gene pair. Bonferroni multiple test correction was implemented to adjust the p-values.

