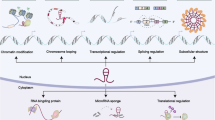
Abstract
With the development of sequencing technology, a large number of long non-coding RNAs (lncRNAs) have been identified in addition to coding genes. LncRNAs, originally considered as junk RNA, are dysregulated in various types of cancer. Although protein-coding signaling pathways underlie various biological activities, and abnormal signal transduction is a key trigger and indicator for tumorigenesis and cancer progression, lncRNAs are sparking keen interest due to their versatile roles in fine-tuning signaling pathways. We are just beginning to scratch the surface of lncRNAs. Therefore, despite the fact that lncRNAs drive malignant phenotypes from multiple perspectives, in this review, we focus on important signaling pathways modulated by lncRNAs in cancer to demonstrate an up-to-date understanding of this emerging field.
Similar content being viewed by others
Facts
-
Long noncoding RNAs are >200 nt in length and have almost no protein-encoding function, but they can exert regulatory effects on transcriptional, post-transcriptional, and post-translational modifications of coding genes.
-
Signaling pathways are a series of enzymatic reactions in response to intracellular and extracellular stimuli and underlie almost every disease, including cancer.
-
LncRNAs participate in tumorigenesis and cancer progression by fine-tuning various cancer signaling pathways.
Open questions
-
Whether disparate signaling pathways regulated by an individual lncRNA converge on similar cancer hallmarks?
-
What are the mechanisms through which the crosstalk among diverse signaling pathways is established?
-
How can we target lncRNAs to fine-tune cancer signaling pathways?
Introduction
Recently, many noncoding RNAs have been discovered using high-throughput sequencing technology. Although they do not encode proteins, they have powerful functions, participating in transcriptional, post-transcriptional, and post-translational regulation of coding genes. Long noncoding RNAs (lncRNAs) are >200 nt in length and have almost no protein-encoding function [1]. LncRNAs are further divided into different subclasses, including intergenic transcripts, enhancer RNAs, and sense or antisense transcripts, according to their locations or relationship with protein-coding protein [2]. LncRNAs perform various functions at transcriptional and translational levels, acting in cis or trans [3]. They play important roles in many biological activities, such as genomic imprinting and X-chromosome inactivation, and participate in cellular behaviors, such as proliferation, differentiation, and survival [4, 5]. They are also implicated in cancer hallmarks and are involved in tumorigenesis and cancer progression [6, 7].
Protein is the executor of biological function. Protein-coding signaling pathways underlie almost every biological process, especially carcinogenesis and progression. Signaling pathways are a series of enzymatic reactions in response to intracellular and extracellular stimuli. They participate in the transmission of cellular signals and are important for biological reactions and gene expression. Signaling pathways are extremely complex, and many signaling molecules are involved in cascade reactions, which are often regulated, directly or indirectly, by various types of molecules [8]. Emerging evidence shows that abnormal signal transduction is a key trigger and indicator for tumorigenesis and cancer progression and that lncRNAs, as versatile regulators, can shape the malignant phenotypes by activating or suppressing different signaling pathways [9, 10]. In this review, we selectively discuss those signaling pathways that are fine-tuned by lncRNAs and well studied mechanistically in tumors, such as Wnt/β-catenin, Hippo, Notch, and nuclear factor (NF)-κB pathway. While we did our best to cover this topic, some additional publications may also help get the full picture [10,11,12].
The Wnt/β-catenin signaling pathway
Wnt/β-catenin signaling pathway is the canonical pathway of the Wnt signaling cascade. Its dysregulation is implicated in tumorigenesis, including proliferation, invasion, metastasis, and apoptosis, and has been the focus of cancer research [13,14,15,16,17]. The Wnt/β-catenin signaling pathway starts from Wnt ligands binding to a receptor complex, comprising Frizzled (Fz), and its co-receptor, low-density lipoprotein receptor-related protein 5/6. When a ligand attaches, the receptor is activated and the protein disheveled homolog (DVL) is phosphorylated, resulting in deactivation of the destruction complex, which comprises AXIN, adenomatous polyposis coli, glycogen synthase kinase 3β (GSK3β), and casein kinase 1α (CK1α). Non-phosphorylated β-catenin accumulates in the cytoplasm and migrates to the nucleus, where it interacts with T cell-specific factor (TCF)/lymphoid enhancer-binding factor transcription factors and triggers the expression of downstream genes, such as those encoding c-Myc, cyclin D1, and CDKN1A [16, 18, 19]. Under the inactivation of Wnt signaling, GSK3β and CK1α within the degradation complex phosphorylate β-catenin, which then undergoes ubiquitin-mediated proteolysis [20, 21].
LncRNAs play a tumor-suppressive or tumor-promoting role via interacting with key factors in the Wnt pathway. For instance, in lung adenocarcinoma, lncRNA LINC00673 enhances the interaction between DDX3 and CK1ε to phosphorylate DVL, thereby activating the Wnt /β-catenin signaling pathway and promoting cancer progression [22]. LncRNAs can inhibit β-catenin phosphorylation by interacting with key enzymes in the destruction complex, thus activating the Wnt/β-catenin signaling pathway. The binding of LncRNAs CYTOR and SLCO4A1-AS1 to β-catenin impedes CK1- and GSKβ-mediated phosphorylation of β-catenin, respectively, leading to β-catenin accumulation and translocation to the nucleus. Activation of the Wnt/β-catenin signaling pathway facilitates colorectal cancer growth and metastasis [23, 24]. As the central protein of the Wnt/β-catenin signaling pathway, the function of β-catenin is closely related to its stability and interacting proteins. Linc01354 contributes to the stability of β-catenin mRNA by interacting with hnRNP-D and the activated Wnt/β-catenin signaling pathway aggravates colorectal cancer [25]. Linc00210 interacts with CTNNBIP1 and blocks its inhibitory role in Wnt/β-catenin activation, playing a tumor-promoting role in liver tumorigenesis [26]. LncRNAs can also inhibit Wnt/β-catenin signaling. Linc01197 impedes the β-catenin–TCF4 interaction by occupying the β-catenin-binding site, thus restraining Wnt/β-catenin signaling and inhibiting pancreatic adenocarcinoma progression [27]. Besides, lncRNAs affect downstream gene expression by regulating transcription factors in Wnt/β-catenin signaling. LincTCF7 regulates TCF7 expression by recruiting the SWI/SNF complex to its promoter, causing activation of Wnt signaling, and thus promoting liver cancer stem cell self-renewal [28].
Collectively, lncRNAs can regulate multiple molecules of the Wnt/β-catenin signaling pathway, ultimately facilitating malignant phenotypes (Fig. 1).
“Wnt ON”: When Wnt ligands bind to a receptor complex, including Frizzled (Fz), and its co-receptor, LRP5/6, the receptor is activated and DVL phosphorylation occurs. This leads to the deactivation of the destruction complex, which is composed of AXIN, APC, GSK3b, and CK1a. Subsequently, stabilized β-catenin migrates to the nucleus to interact with TCF/LEF and trigger the downstream gene expression. “Wnt OFF”: In the absence of Wnt ligand, GSK3β and CK1α phosphorylate β-catenin, resulting in ubiquitin-mediated proteolysis of β-catenin.
The Hippo signaling pathway
Hippo signaling is an evolutionarily highly conserved pathway that regulates multiple biological processes including cell proliferation and differentiation. Disruption of the Hippo pathway may trigger tumorigenesis. However, mutations in Hippo pathway genes are not common. Molecular events that interfere with the Hippo pathway might be an important mechanism in cancer initiation [29]. The Hippo pathway is mainly regulated by different cell- and tissue-level characteristics, such as cell-to-cell adhesion, basement membrane adhesion, planar cell polarity, and mechanical forces perceived by the actin cytoskeleton [30]. After receiving the upstream signals, the core kinase combination, including LATS1, LATS2, MST1, and MST2, and the SAV1 and MOB1 adaptor proteins, are activated. They cooperate to phosphorylate transcriptional coactivators yes associated protein (YAP) and TAZ [30] and promote cytoplasmic retention of YAP and TAZ by creating a binding site on 14-3-3 proteins [31]. Additionally, re-phosphorylation of YAP/TAZ results in proteasomal degradation [32]. When dephosphorylation of YAP and TAZ occurs, they translocate to the nucleus and promote tissue growth by interacting with transcription factors, such as TEADs and SMADs [33].
As mentioned above, molecular events might play an important role in regulating the Hippo pathway. Many lncRNAs are involved in regulating this pathway. LncRNAs can interact with the core kinase, thus affecting the downstream cascade. LncRNA MIR100HG binds to the histone methylation regulator EZH2, acting as a scaffold that epigenetically silences LATS1/2. MIR100HG plays an oncogenic role in osteosarcoma through inactivating the Hippo pathway [34]. LncRNA MAYA participates in the ROR1/HER3-LLGL2-MAYA-NSUN6 signaling axis, mediating the NSUN6-dependent methylation of MST1, which abolishes kinase activity and activates the HIPPO pathway. This might provide a new therapeutic direction for bone metastasis [35]. LncRNAs can also target the important downstream executor, YAP, in the Hippo pathway. LncRNA RP11‑323N12.5 is upregulated in gastric cancer and exerts its tumor-promoting and immunosuppression role by activating the Hippo pathway. RP11‑323N12.5 promotes the transcription of YAP1 by binding to c-MYC on the YAP1 promoter. In turn, YAP1/TAZ/TEADs can also regulate RP11-323N12.5 transcription by forming a positive feedback loop [36]. Moreover, LncRNAs can affect the localization of YAP. LncRNA UCA1, which is overexpressed in pancreatic cancer, can inhibit the phosphorylation of YAP by forming shielding composites with MOB1, LATS1, and YAP. Consequently, YAP accumulates in the nucleus and activates the downstream genes of the Hippo signaling pathway [37].
Thus, lncRNAs modulate the Hippo signaling pathway in diverse manners, eventually leading to the activation or suppression of downstream cancer-related genes (Fig. 2).
After receiving the upstream signals, the core kinase combination, including LATS1, LATS2, MST1, MST2, SAV1, and MOB1, are activated and cooperate to phosphorylate YAP and TAZ. Phosphorylated YAP/TAZ underdo retention by 14-3-3 proteins or proteasomal degradation. When dephosphorylation of YAP and TAZ occurs, they translocate to the nucleus and interact with TEADs and SMADs to promote downstream gene transcription.
The Notch signaling pathway
Notch signaling pathway is involved in various biological processes, including cell differentiation, proliferation, and apoptosis. Notch signaling also plays pleiotropic roles in carcinogenesis and tumor progression. Four receptors (Notch 1, 2, 3, and 4) and five ligands (DLL 1, 3, and 4; JAG 1 and 2) have been identified in Notch signaling [38]. The receptor–ligand interactions trigger sequential Notch receptor cleavage by the ADAM family of metalloproteases and γ-secretase, leading to the release of the Notch intracellular domain (NICD). NICD translocates to the nucleus and binds to the CBF-1/Su(H)/LAG1 transcription factor, displacing co-repressors, and recruiting co-activators such as Mastermind-like (MAML), thus inducing Notch target gene transcription [39]. Depending on the strength and dynamics of juxtacrine signaling, Notch pathway activates different programs that determine cell fate [40]. Notch signaling is involved in the regulation of tumor-related genes, such as those encoding MYC, cyclin-D1, or p21 [41].
LINC01152 is overexpressed in glioblastoma multiforme, functioning as an oncogene. It activates the Notch signaling pathway by upregulating the expression of transcriptional co-activator 2 (MAML2). LINC01152 serves as a competing endogenous RNA via sponging microRNA miR-466. Moreover, LINC01152 can prevent MAML2 from shearing via recruiting SRSF1, thus maintaining the level of MAML2. Besides, MAML2 could, in turn, promote the expression of LINC01152 via regulating the Notch pathway [42]. LncRNA CEBPA-AS1 acts as a tumor suppressor in osteosarcoma by inhibiting Notch signaling pathway activity. LncRNA CEBPA-AS1 binds to miR-10b-5p and regulates the activity of NCOR2, which is a nuclear receptor co-repressor. Taken together, CEBPA-AS1 overexpression inhibits osteosarcoma progression by suppressing the Notch signaling pathway via upregulating NCOR2 expression [43].
Although the Notch signaling pathway has a relatively simple framework, it plays an important role in cell proliferation, differentiation, and apoptosis (Fig. 3). As mentioned above, lncRNAs are involved in various steps of this pathway. However, further effort is required to determine the details.
After Jagged-1 ligand binding to the Notch receptor, the Notch receptor is cleaved by ADAM family of metalloproteases and γ-secretase, leading to the release of NICD. NICD translocates to the nucleus and binds to CBF-1/Su(H)/LAG1 (CSL) transcription factor, displacing co-repressors (Co-Rs), and recruiting co-activators (Co-As), to induce target gene transcription.
The NF-κB signaling pathway
Since the NF-κB subunits were recognized >20 years ago as being homologous to the viral oncogene v-Rel, the role of these proteins in tumorigenesis has been accepted. However, early research on NF-κB mainly focused on the regulation of the immune response. With the deepening of the research on NF-κB, the role of the inflammatory response in tumor development gradually became prominent [44, 45]. NF-κB activation enhances the hallmarks of cancer (such as proliferation, invasion, metastasis, angiogenesis, and prevention of apoptosis), thus supporting tumorigenesis [46].
In mammals, NF-κB complexes comprise five proteins, including RelA (p65), RelB, c-Rel, p50 (p105 precursor), and p52 (p100 precursor), which all share an N-terminal Rel homology domain that is required for dimerization, DNA binding, and inhibitory IκB protein interaction [47]. There are two distinct modes: canonical (activated by tumor necrosis factor (TNF), interleukin-1 (IL-1), and Toll-like receptor ligands) and non-canonical (activated by TNF superfamily receptors, B cell activating factor receptor, and receptor-activated NF-κB ligand) [48]. In the canonical pathway, generally, NF-κB is inhibited in an inactive state in the cytoplasm by a family of inhibitor of NF-κB (IκB) proteins, which consists of IκBα, IκBβ, and IκBε [49]. Following exposure to an inducing stimulus, IκB is phosphorylated by activated IKK2, part of the IκB kinase (IKK) complex, which consists of two catalytic subunits, IKKα (IKK1) and IKKβ (IKK2), and the non-catalytic regulatory protein NEMO or IKKγ. The phosphorylated IκB then undergoes ubiquitination and proteasome-mediated degradation, together with NF-κB nuclear localization [50]. In the non-canonical pathway, IKKα is phosphorylated by NF-κB-inducing kinase following receptor ligation. Activated IKKα induces p100 phosphorylation, which undergoes partial proteolysis to p52 [51]. Various NF-κB dimers released by either the canonical or non-canonical pathways translocate to the nucleus and interact with transcription factors and cofactors (such as TBP, TFIIB, p300, and CBP) to trigger target gene transcription [52].
As an important signaling pathway, NF-κB signaling is modulated by multiple regulatory mechanisms. Increasing evidence shows that lncRNAs are involved in the regulation of the NF-κB signaling pathway. LncRNAs can directly interact with functional signaling proteins involved in NF-κB signaling, serving as modulators. LncRNA NKILA inhibits IKK-induced IκB phosphorylation by masking the phosphorylation motifs of IκB and impeding NF-κB activation. NKILA prevents breast cancer metastasis by inhibiting over-activation of the NF-κB pathway [53]. In gastric cancer, lncRNA KRT19P3 could suppress cancer cell proliferation and metastasis by regulating the expression of COPS7A. COPS7A can promote deubiquitinylation of IκBα, which inactivates the NF-κB signaling pathway [54]. In addition, lncRNA SChLAP1 interacts with HNRNPL, forming a complex, which stabilizes ACTN4 by preventing its proteasomal degradation. The stabilization of ACTN4 increases the p65 subunit nuclear localization and activates NF-κB signaling. Therefore, SChLAP1 acts as a growth-promoting lncRNA in glioblastoma [55].
LncRNAs are emerging as important modulators in the management of NF-κB signaling, gradually unraveling the mystery of cancer development and progression (Fig. 4).
NF-κB consists of five subunits, RelA (p65), RelB, c-Rel, p50 (p105 precursor), and p52 (p100 precursor). In the canonical pathway: followed by exposure to an inducing stimulus, IκB is phosphorylated by activated IKKβ. The phosphorylated IκB then undergoes ubiquitination and proteasome-mediated degradation along with NF-κB nuclear localization. In the non-canonical pathway: IκB kinase is phosphorylated by NF-κB-inducing kinase following receptor ligation. Activated IKKα induces p100 phosphorylation, which undergoes partial proteolysis to p52 and translocates to the nucleus.
The Hedgehog (HH) signaling pathway
The HH signaling pathway is essential for many fundamental processes, including embryogenesis, stem cell maintenance, and tissue regeneration [56]. Some studies indicated that HH signaling plays a broader role in carcinogenesis, such as in breast cancer, gastric cancer, and prostate cancer [56,57,58,59]. Here we present a brief overview of the HH signaling pathway. In mammals, there are three HH ligands (SHH, DHH, and IHH), one 12-pass transmembrane receptor PTCH1, one G protein-coupled receptor (GPCR)-like seven-pass transmembrane protein, SMO, and three transcription factors (GLI1, GLI2, and GLI3) in the HH signaling cascade. All components of the HH signal pathway were discovered in the primary cilia [60]. In the absence of HH ligands, PTCH1 suppresses the accumulation of SMO, thus repressing its activity [61]. Transcription factors GLI2 and GLI3 are trapped in the cytoplasm by SUFU and are consequently phosphorylated by protein kinase A, CK1, and GSK3β. Phosphorylated GLI2 and GLI3 are degraded by the proteasome into repressor forms (GLI2R and GLI3R) [62]. GLIR binds to the promoters of HH signaling target genes and silences their transcription [63]. Once HH binds to PTCH1, SMO inhibition is relieved. Activated SMO relieves the suppression of GLI2 and GLI3 by SUFU. The GLIs evade phosphorylation and the activated forms (GLI2A and GLI3A) translocate to the nucleus to activate HH target gene expression [64, 65].
LncRNAs, as small-molecule regulators, may be involved in modulating the HH signaling pathway at different levels and might be related to tumorigenesis and development. LncRNA HOTAIR expression is dysregulated in various cancers and affects cancer development through multiple mechanisms [66]. In renal cell carcinoma, highly expressed HOTAIR interacts with the androgen receptor (AR) and cooperatively promotes GLI2 transcription by binding to its promoter. Increased GLI2 activates the HH signaling pathway and promotes the expression of its downstream genes, which are associated with tumor angiogenesis and cancer stemness [67]. LncHDAC2 is highly expressed in liver cancer and drives cancer stem cell self-renewal and tumor propagation. Mechanistically, lncHDAC2 inhibits PTCH1 expression via recruiting the NuRD complex to its promoter, resulting in activation of HH signaling [68] (Fig. 5). Further study of the regulation of HH signaling pathway by lncRNAs in cancer will clarify the regulatory network between them.
“Hedgehog OFF”: In the absence of Hedgehog ligands, PTCH1 suppresses the accumulation of SMO, thus repressing its activity. GLI2 and GLI3 are trapped in the cytoplasm by SUFU and consequently phosphorylated by PKA, CK1, and GSK3β. Phosphorylated GLI2 and GLI3 are degraded by the proteasome into repressor forms (GLI2R and GLI3R), which inhibit downstream gene expression. “Hedgehog ON”: Once Hedgehog binds to PTCH1, SMO inhibition is relieved. Activated SMO relieves the suppression of GLI2 and GLI3 by SUFU. The GLIs evade phosphorylation and the activated forms (GLI2A and GLI3A) translocate to the nucleus to activate Hedgehog target gene expression.
The transforming growth factor (TGF)-β signaling pathway
The TGF-β signaling pathway consists of TGF-β ligands, receptors, and downstream signaling molecules. The pathway participates in organismic homeostasis, controlling cell proliferation, motility, and differentiation [69, 70]. In normal tissues, the TGF-β pathway maintains homeostasis, and once tumorigenesis has progressed, TGF-β is involved in promoting tumor malignancy [71,72,73]. The cascade initiates from the binding of TGF-β ligands to TGF-β type II receptors (TGFBR2), subsequently recruiting and phosphorylating TGF-β type I (TGFBR1). In the canonical pathway, phosphorylated TGFBR1, in turn, phosphorylates the signaling proteins SMAD2 and SMAD3. Following phosphorylation, they form a complex with SMAD4. Subsequently, the complex translocates to the nucleus and regulates downstream gene expression, together with cofactors and transcription factors [74]. Meanwhile, another class of SMAD proteins functions as inhibitors of the TGF-β signaling pathway, called I-SMADs, comprising SMAD6 and SMAD7. They attach directly to TGFBR1, thereby interrupting subsequent phosphorylation of SMADs or hindering the formation of the SMAD2,3/SMAD-4 complex [75]. In addition to SMAD proteins, TGF-β receptors also induce other signal transducers, including mitogen-activated protein kinase, phosphoinositide-3 kinase (PI3K)/AKT, Janus-activated kinase (JAK)/signal transducer and activator of transcription factor (STAT), and c-Jun N-terminal kinase/p38 in non-canonical pathway [76, 77].
Many studies have confirmed that lncRNAs are involved in the regulation of the TGF-β signaling pathway, thus affecting tumorigenesis and cancer progression. Lnc-TSI exerts a tumor-suppressor role in clear cell renal cell carcinoma by inhibiting metastasis. Lnc-TSI impedes the phosphorylation of SMAD3 by blocking the interaction between SMAD3 and TGFBR1 at the MH2 domain. Consequently, the TGF-β/SMADs signaling pathway is inhibited, which suppresses epithelial–mesenchymal transition [78]. LncRNAs can also regulate the TGF-β signaling pathway by modulating its co-factors. LncRNA TGFB2-AS1 assists repressive histone H3K27me3 modifications at the promoters of TGF-β-target genes by binding to the EED adaptor of the Polycomb repressor complex 2 (PRC2) via the 3’ terminal region of TGFB2-AS1. Thus, TGFB2-AS1 exerts an inhibitory role on TGF-β signaling downstream genes [79]. Additionally, lncRNA MEG3 regulates the activity of TGF-β genes by forming an RNA–DNA triplex structure through GA-rich binding sites. The RNA–DNA triplex might be a general characteristic whereby lncRNAs recognize their target genes [80] (Fig. 6).
TGF-β ligands bind to TGF-β type II receptors (TGFBR2), subsequently recruiting and phosphorylating TGF-β type I (TGFBR1). The phosphorylated TGFBR1 phosphorylates the signaling proteins SMAD2 and SMAD3. Following phosphorylation, they form a complex with SMAD4. The complex translocates to the nucleus and regulates downstream gene expression. Inhibitors, termed I-SMADs, attach directly to TGFBR1 and intercept the phosphorylation of SMADs or hinder the formation of the SMAD2, 3/SMAD-4 complex.
Although a large number of lncRNAs have been reported to be related to TGF-β signaling pathway, the exact mechanisms of their mutual interactions are mostly unknown. Further research into the lncRNA–TGF-β signaling network will expand our understanding of cancer pathogenesis and provide novel insight into therapeutic strategies.
Other signaling pathways
In addition to the signaling pathways mentioned above, lncRNAs also play regulatory roles in other signaling pathways during tumorigenesis. One of these is the JAK-STAT signaling pathway, which is necessary for homeostasis and development, mainly mediating inflammation and immunity [81]. Disturbance of the JAK-STAT signaling pathway might be involved in the cancer hallmarks, such as evasion of immune surveillance and promoting or suppressing tumor progression. Activation of the pathway is initiated from the interaction of extracellular cytokines and other ligands with transmembrane receptors, and thus receptor-bound JAKs are activated. Subsequently, JAK mediates the phosphorylation STATs, which form homodimers or heterodimers and transfer into the nucleus to trigger downstream gene expression [82, 83]. LncRNA LINC00669 facilitates the malignancy of nasopharyngeal carcinoma by competitively binding to suppressor SOCS1 of the JAK/STAT signaling pathway. In this way, LINC00669 protects STAT1 from ubiquitination modification and stabilizes it. STAT1 translocates to the nucleus and promotes the transcription of target genes [84]. Meanwhile, tumor-suppressive lncRNAs, such as RP11-468E2.5, suppress the JAK/STAT signaling pathway by targeting STAT5 and STAT6, thereby inhibiting colorectal cancer cell proliferation and promoting apoptosis [85].
Another potent pathway is the PI3K/AKT signaling pathway, which consists of two main proteins, PI3K and AKT. Activated by growth factors through GPCRs or receptor tyrosine kinase receptors, PI3K activates AKT and downstream effectors through the lipid second messenger phosphatidylinsoitol-3,4,5-trisphosphate. The pathway is important for cell growth and differentiation. Disturbance of the PI3K/AKT pathway might be a critical event in the process of tumor development [86]. It has been demonstrated that the PI3K pathway is one of the most frequently activated pathways in human cancer [87]. LncRNAs serve as indispensable regulators in this signaling cascade. Overexpression of lncRNA DANCR correlates with poor prognosis and low survival rate in triple-negative breast cancer. Mechanically, DANCR interacts with RXRA and increases its phosphorylation via GSK3β, which alleviates the inhibition of PIK3CA transcription and subsequently leads to activation of the PI3K/AKT signaling pathway and tumorigenesis [88]. LncRNAs can indirectly regulate the PI3K signaling pathway by regulating growth factor ligand activity of the cascade. LncRNA PART1, which serves as a tumor suppressor in gastric cancer, epigenetically silences PDGFB (encoding platelet-derived growth factor B) via PLZF-mediated recruitment of EZH2, thus inhibiting PI3K/Akt signaling activation [89].
LncRNAs regulate signaling pathways of cancer
Deregulation of protein-coding signaling pathways is fundamental to tumorigenesis and cancer progression. However, lncRNAs, hidden in the signaling pathways, cannot be neglected for their fine-tuning functions. Notably, one signaling pathway can be fine-tuned by multiple lncRNAs. As mentioned previously, Linc01354, Linc00210, Linc01197, LincTCF7, and LINC00673 participate in activating or suppressing the Wnt/β-catenin signaling pathway. In another example, lncRNAs, as upstream mediators, have a dual function in the STAT3 signaling pathway in different cancers. NEAT1, SNHG3, and H19 induce STAT3 expression, while MEG3, PTCSC3, and NKILA inhibit STAT3 expression [90]. Not surprisingly, one lncRNA may also regulate several signaling pathways. For example, MALAT1, acting as an oncogene, can activate the Wnt/β-catenin signaling pathway through upregulation of β-catenin as well as downregulation of GSK3β [91]. Meanwhile, MALAT1 increases JAG1 (Jagged1) expression through inhibiting miR-124 and thus activates the Notch signaling pathway [92]. In addition to independently regulating different pathways, lncRNAs can connect different signaling pathways. In breast cancer, lncRNA BCAR4 connects the Hippo and HH signaling pathways. The downstream effector YAP of the Hippo pathway promotes BCAR4 expression, which activates HH signaling to facilitate the transcription of glycolysis activators HK2 and PFKFB3, and thus reprograms glucose metabolism [93]. Table 1 provides a brief overview of lncRNAs regulating the signaling pathways discussed above.
Conclusion
The regulation of signaling pathways by lncRNAs is complex and diverse. Studies have confirmed the importance of lncRNAs in the regulatory network, functioning to make the cascades operate more smoothly or with greater flexibility. LncRNAs may interact directly or indirectly with signaling molecules to regulate signaling pathways. Despite the support of advanced technologies, such as RNA precipitation, RNA immunoprecipitation, crosslinking immunoprecipitation, and RNA interference or CRISPR (clustered regularly interspaced short palindromic repeats) technologies, the complexity of signaling pathways makes research on the exact interaction mechanism between lncRNAs and signaling molecules a challenge. With the further development of experimental techniques, a deeper understanding of lncRNA-mediated signaling transduction will be gained.
Although numerous studies have shown that lncRNAs play significant roles in the regulation of signal networks and even in the development of tumors, the human body is a complex system, and no molecule can achieve the effect of affecting the whole body by one action; therefore, it would make sense to investigate intermolecular synergy. However, in the cascades of signaling pathways, a “butterfly effect” might also be a feature. In this complex network, many fields remain unexplored. Thus, further effort on the influence of lncRNAs on signaling pathways and even the whole biological process will shed light on the mystery of cancer. However, even our current limited knowledge has provided a novel and promising avenue for the fast-emerging field.
Data availability
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
References
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009;458:223–7.
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012;22:1775–89.
Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell 2018;172:393–407.
Mirhosseini SA, Sarfi M, Samavarchi Tehrani S, Mirazakhani M, Maniati M, Amani J. Modulation of cancer cell signaling by long noncoding RNAs. J Cell Biochem. 2019;120:12224–46.
Sun W, Shi Y, Wang Z, Zhang J, Cai H, Zhang J, et al. Interaction of long-chain non-coding RNAs and important signaling pathways on human cancers (Review). Int J Oncol. 2018;53:2343–55.
Wang J, Zhang X, Chen W, Hu X, Li J, Liu C. Regulatory roles of long noncoding RNAs implicated in cancer hallmarks. Int J Cancer. 2020;146:906–16.
Zhao S, Zhang X, Chen S, Zhang S. Natural antisense transcripts in the biological hallmarks of cancer: powerful regulators hidden in the dark. J Exp Clin Cancer Res. 2020;39:187.
Zhao H, Wu L, Yan G, Chen Y, Zhou M, Wu Y, et al. Inflammation and tumor progression: signaling pathways and targeted intervention. Signal Transduct Target Ther. 2021;6:263.
Liu SJ, Dang HX, Lim DA, Feng FY, Maher CA. Long noncoding RNAs in cancer metastasis. Nat Rev Cancer. 2021;21:446–60.
Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22:96–118.
Li J, Zhu Y, Wang H, Ji X. Targeting long noncoding RNA in glioma: a pathway perspective. Mol Ther Nucleic Acids. 2018;13:431–41.
Chew CL, Conos SA, Unal B, Tergaonkar V. Noncoding RNAs: master regulators of inflammatory signaling. Trends Mol Med. 2018;24:66–84.
Ge X, Wang X. Role of Wnt canonical pathway in hematological malignancies. J Hematol Oncol. 2010;3:33.
Wang W, Smits R, Hao H, He C. Wnt/beta-catenin signaling in liver cancers. Cancers. 2019;11:926.
Clara JA, Monge C, Yang Y, Takebe N. Targeting signalling pathways and the immune microenvironment of cancer stem cells - a clinical update. Nat Rev Clin Oncol. 2020;17:204–32.
Fuentes RG, Arai MA, Ishibashi M. Natural compounds with Wnt signal modulating activity. Nat Prod Rep. 2015;32:1622–8.
Zhang Y, Wang X. Targeting the Wnt/beta-catenin signaling pathway in cancer. J Hematol Oncol. 2020;13:165.
Koni M, Pinnaro V, Brizzi MF. The Wnt signalling pathway: a tailored target in cancer. Int J Mol Sci. 2020;21:7697.
Hu XY, Hou PF, Li TT, Quan HY, Li ML, Lin T, et al. The roles of Wnt/beta-catenin signaling pathway related lncRNAs in cancer. Int J Biol Sci. 2018;14:2003–11.
Wiese KE, Nusse R, van Amerongen R. Wnt signalling: conquering complexity. Development. 2018;145:dev165902.
Nusse R, Clevers H. Wnt/beta-catenin signaling, disease, and emerging therapeutic modalities. Cell 2017;169:985–99.
Guan H, Zhu T, Wu S, Liu S, Liu B, Wu J, et al. Long noncoding RNA LINC00673-v4 promotes aggressiveness of lung adenocarcinoma via activating WNT/beta-catenin signaling. Proc Natl Acad Sci USA. 2019;116:14019–28.
Yue B, Liu C, Sun H, Liu M, Song C, Cui R, et al. A positive feed-forward loop between lncRNA-CYTOR and Wnt/beta-catenin signaling promotes metastasis of colon cancer. Mol Ther. 2018;26:1287–98.
Yu J, Han Z, Sun Z, Wang Y, Zheng M, Song C. LncRNA SLCO4A1-AS1 facilitates growth and metastasis of colorectal cancer through beta-catenin-dependent Wnt pathway. J Exp Clin Cancer Res. 2018;37:222.
Li J, He M, Xu W, Huang S. LINC01354 interacting with hnRNP-D contributes to the proliferation and metastasis in colorectal cancer through activating Wnt/beta-catenin signaling pathway. J Exp Clin Cancer Res. 2019;38:161.
Fu X, Zhu X, Qin F, Zhang Y, Lin J, Ding Y, et al. Linc00210 drives Wnt/beta-catenin signaling activation and liver tumor progression through CTNNBIP1-dependent manner. Mol Cancer. 2018;17:73.
Ling J, Wang F, Liu C, Dong X, Xue Y, Jia X, et al. FOXO1-regulated lncRNA LINC01197 inhibits pancreatic adenocarcinoma cell proliferation by restraining Wnt/beta-catenin signaling. J Exp Clin Cancer Res. 2019;38:179.
Wang Y, He L, Du Y, Zhu P, Huang G, Luo J, et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell. 2015;16:413–25.
Harvey KF, Zhang X, Thomas DM. The Hippo pathway and human cancer. Nat Rev Cancer. 2013;13:246–57.
Kulkarni A, Chang MT, Vissers JHA, Dey A, Harvey KF. The Hippo pathway as a driver of select human cancers. Trends Cancer. 2020;6:781–96.
Han Y. Analysis of the role of the Hippo pathway in cancer. J Transl Med. 2019;17:116.
Calses PC, Crawford JJ, Lill JR, Dey A. Hippo pathway in cancer: aberrant regulation and therapeutic opportunities. Trends Cancer. 2019;5:297–307.
Zheng Y, Pan D. The Hippo signaling pathway in development and disease. Dev Cell. 2019;50:264–82.
Su X, Teng J, Jin G, Li J, Zhao Z, Cao X, et al. ELK1-induced upregulation of long non-coding RNA MIR100HG predicts poor prognosis and promotes the progression of osteosarcoma by epigenetically silencing LATS1 and LATS2. Biomed Pharmacother. 2019;109:788–97.
Li C, Wang S, Xing Z, Lin A, Liang K, Song J, et al. A ROR1-HER3-lncRNA signalling axis modulates the Hippo-YAP pathway to regulate bone metastasis. Nat Cell Biol. 2017;19:106–19.
Wang J, Huang F, Shi Y, Zhang Q, Xu S, Yao Y, et al. RP11-323N12.5 promotes the malignancy and immunosuppression of human gastric cancer by increasing YAP1 transcription. Gastric Cancer. 2021;24:85–102.
Zhang M, Zhao Y, Zhang Y, Wang D, Gu S, Feng W, et al. LncRNA UCA1 promotes migration and invasion in pancreatic cancer cells via the Hippo pathway. Biochim Biophys Acta Mol Basis Dis. 2018;1864:1770–82.
Kopan R, Ilagan MX. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell. 2009;137:216–33.
Takebe N, Nguyen D, Yang SX. Targeting notch signaling pathway in cancer: clinical development advances and challenges. Pharmacol Ther. 2014;141:140–9.
Meurette O, Mehlen P. Notch signaling in the tumor microenvironment. Cancer Cell. 2018;34:536–48.
Reicher A, Fosselteder J, Kwong LN, Pichler M. Crosstalk between the Notch signaling pathway and long non-coding RNAs. Cancer Lett. 2018;420:91–6.
Wu J, Wang N, Yang Y, Jiang G, Mu Q, Zhan H, et al. LINC01152 upregulates MAML2 expression to modulate the progression of glioblastoma multiforme via Notch signaling pathway. Cell Death Dis. 2021;12:115.
Xia P, Gu R, Zhang W, Sun YF. LncRNA CEBPA-AS1 overexpression inhibits proliferation and migration and stimulates apoptosis of OS cells via Notch signaling. Mol Ther Nucleic Acids. 2020;19:1470–81.
Ben-Neriah Y, Karin M. Inflammation meets cancer, with NF-kappaB as the matchmaker. Nat Immunol. 2011;12:715–23.
Grivennikov SI, Greten FR, Karin M. Immunity, inflammation, and cancer. Cell. 2010;140:883–99.
Naugler WE, Karin M. NF-kappaB and cancer-identifying targets and mechanisms. Curr Opin Genet Dev. 2008;18:19–26.
Tormanen CD. Comparison of the properties of purified mitochondrial and cytosolic rat kidney transamidinase. Int J Biochem. 1990;22:1243–50.
Patel M, Horgan PG, McMillan DC, Edwards J. NF-kappaB pathways in the development and progression of colorectal cancer. Transl Res. 2018;197:43–56.
Hayden MS, Ghosh S. Shared principles in NF-kappaB signaling. Cell. 2008;132:344–62.
Kanarek N, London N, Schueler-Furman O, Ben-Neriah Y. Ubiquitination and degradation of the inhibitors of NF-kappaB. Cold Spring Harb Perspect Biol. 2010;2:a000166.
Hoesel B, Schmid JA. The complexity of NF-kappaB signaling in inflammation and cancer. Mol Cancer. 2013;12:86.
Wan F, Lenardo MJ. The nuclear signaling of NF-kappaB: current knowledge, new insights, and future perspectives. Cell Res. 2010;20:24–33.
Liu B, Sun L, Liu Q, Gong C, Yao Y, Lv X, et al. A cytoplasmic NF-kappaB interacting long noncoding RNA blocks IkappaB phosphorylation and suppresses breast cancer metastasis. Cancer Cell. 2015;27:370–81.
Zheng J, Zhang H, Ma R, Liu H, Gao P. Long non-coding RNA KRT19P3 suppresses proliferation and metastasis through COPS7A-mediated NF-kappaB pathway in gastric cancer. Oncogene. 2019;38:7073–88.
Ji J, Xu R, Ding K, Bao G, Zhang X, Huang B, et al. Long noncoding RNA SChLAP1 forms a growth-promoting complex with HNRNPL in human glioblastoma through stabilization of ACTN4 and activation of NF-kappaB signaling. Clin Cancer Res. 2019;25:6868–81.
Katoh Y, Katoh M. Comparative genomics on Sonic hedgehog orthologs. Oncol Rep. 2005;14:1087–90.
Xu Y, Song S, Wang Z, Ajani JA. The role of hedgehog signaling in gastric cancer: molecular mechanisms, clinical potential, and perspective. Cell Commun Signal. 2019;17:157.
Wilkinson SE, Furic L, Buchanan G, Larsson O, Pedersen J, Frydenberg M, et al. Hedgehog signaling is active in human prostate cancer stroma and regulates proliferation and differentiation of adjacent epithelium. Prostate. 2013;73:1810–23.
Fattahi S, Pilehchian Langroudi M, Akhavan-Niaki H. Hedgehog signaling pathway: epigenetic regulation and role in disease and cancer development. J Cell Physiol. 2018;233:5726–35.
Michaud EJ, Yoder BK. The primary cilium in cell signaling and cancer. Cancer Res. 2006;66:6463–7.
Denef N, Neubuser D, Perez L, Cohen SM. Hedgehog induces opposite changes in turnover and subcellular localization of patched and smoothened. Cell. 2000;102:521–31.
Price MA, Kalderon D. Proteolysis of the Hedgehog signaling effector Cubitus interruptus requires phosphorylation by Glycogen Synthase Kinase 3 and Casein Kinase 1. Cell. 2002;108:823–35.
Goetz SC, Anderson KV. The primary cilium: a signalling centre during vertebrate development. Nat Rev Genet. 2010;11:331–44.
Wu F, Zhang Y, Sun B, McMahon AP, Wang Y. Hedgehog signaling: from basic biology to cancer therapy. Cell Chem Biol. 2017;24:252–80.
Skoda AM, Simovic D, Karin V, Kardum V, Vranic S, Serman L. The role of the Hedgehog signaling pathway in cancer: a comprehensive review. Bosn J Basic Med Sci. 2018;18:8–20.
Rajagopal T, Talluri S, Akshaya RL, Dunna NR. HOTAIR lncRNA: a novel oncogenic propellant in human cancer. Clin Chim Acta. 2020;503:1–18.
Bai JY, Jin B, Ma JB, Liu TJ, Yang C, Chong Y, et al. HOTAIR and androgen receptor synergistically increase GLI2 transcription to promote tumor angiogenesis and cancer stemness in renal cell carcinoma. Cancer Lett. 2021;498:70–9.
Wu J, Zhu P, Lu T, Du Y, Wang Y, He L, et al. The long non-coding RNA lncHDAC2 drives the self-renewal of liver cancer stem cells via activation of Hedgehog signaling. J Hepatol. 2019;70:918–29.
David CJ, Massague J. Contextual determinants of TGFbeta action in development, immunity and cancer. Nat Rev Mol Cell Biol. 2018;19:419–35.
Tzavlaki K, Moustakas A. TGF-beta signaling. Biomolecules. 2020;10:487.
Pardali K, Moustakas A. Actions of TGF-beta as tumor suppressor and pro-metastatic factor in human cancer. Biochim Biophys Acta. 2007;1775:21–62.
Batlle E, Massague J. Transforming growth factor-beta signaling in immunity and cancer. Immunity. 2019;50:924–40.
Hao Y, Baker D, Ten Dijke P. TGF-beta-mediated epithelial-mesenchymal transition and cancer metastasis. Int J Mol Sci. 2019;20:2767.
Yu Y, Feng XH. TGF-beta signaling in cell fate control and cancer. Curr Opin Cell Biol. 2019;61:56–63.
Zhang Y, Derynck R. Regulation of Smad signalling by protein associations and signalling crosstalk. Trends Cell Biol. 1999;9:274–9.
Derynck R, Zhang YE. Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature. 2003;425:577–84.
Zhang YE. Non-Smad signaling pathways of the TGF-beta family. Cold Spring Harb Perspect Biol. 2017;9:a022129.
Wang P, Chen W, Ma T, Lin Z, Liu C, Liu Y, et al. lncRNA lnc-TSI inhibits metastasis of clear cell renal cell carcinoma by suppressing TGF-beta-induced epithelial-mesenchymal transition. Mol Ther Nucleic Acids. 2020;22:1–16.
Papoutsoglou P, Tsubakihara Y, Caja L, Moren A, Pallis P, Ameur A, et al. The TGFB2-AS1 lncRNA regulates TGF-beta signaling by modulating corepressor activity. Cell Rep. 2019;28:3182.e11–98.e11.
Mondal T, Subhash S, Vaid R, Enroth S, Uday S, Reinius B, et al. MEG3 long noncoding RNA regulates the TGF-beta pathway genes through formation of RNA-DNA triplex structures. Nat Commun. 2015;6:7743.
Villarino AV, Kanno Y, O’Shea JJ. Mechanisms and consequences of Jak-STAT signaling in the immune system. Nat Immunol. 2017;18:374–84.
Owen KL, Brockwell NK, Parker BS. JAK-STAT signaling: a double-edged sword of immune regulation and cancer progression. Cancers. 2019;11:2002.
O’Shea JJ, Schwartz DM, Villarino AV, Gadina M, McInnes IB, Laurence A. The JAK-STAT pathway: impact on human disease and therapeutic intervention. Annu Rev Med. 2015;66:311–28.
Qing X, Tan GL, Liu HW, Li W, Ai JG, Xiong SS, et al. LINC00669 insulates the JAK/STAT suppressor SOCS1 to promote nasopharyngeal cancer cell proliferation and invasion. J Exp Clin Cancer Res. 2020;39:166.
Jiang L, Zhao XH, Mao YL, Wang JF, Zheng HJ, You QS. Long non-coding RNA RP11-468E2.5 curtails colorectal cancer cell proliferation and stimulates apoptosis via the JAK/STAT signaling pathway by targeting STAT5 and STAT6. J Exp Clin Cancer Res. 2019;38:465.
Noorolyai S, Shajari N, Baghbani E, Sadreddini S, Baradaran B. The relation between PI3K/AKT signalling pathway and cancer. Gene. 2019;698:120–8.
Martini M, De Santis MC, Braccini L, Gulluni F, Hirsch E. PI3K/AKT signaling pathway and cancer: an updated review. Ann Med. 2014;46:372–83.
Tang J, Zhong G, Zhang H, Yu B, Wei F, Luo L, et al. LncRNA DANCR upregulates PI3K/AKT signaling through activating serine phosphorylation of RXRA. Cell Death Dis. 2018;9:1167.
Han H, Wang S, Meng J, Lyu G, Ding G, Hu Y, et al. Long noncoding RNA PART1 restrains aggressive gastric cancer through the epigenetic silencing of PDGFB via the PLZF-mediated recruitment of EZH2. Oncogene. 2020;39:6513–28.
Ashrafizadeh M, Gholami MH, Mirzaei S, Zabolian A, Haddadi A, Farahani MV, et al. Dual relationship between long non-coding RNAs and STAT3 signaling in different cancers: new insight to proliferation and metastasis. Life Sci. 2021;270:119006.
Guo C, Wang X, Chen LP, Li M, Li M, Hu YH, et al. Long non-coding RNA MALAT1 regulates ovarian cancer cell proliferation, migration and apoptosis through Wnt/beta-catenin signaling pathway. Eur Rev Med Pharmacol Sci. 2018;22:3703–12.
Zhang TH, Liang LZ, Liu XL, Wu JN, Su K, Chen JY, et al. Long non-coding RNA MALAT1 interacts with miR-124 and modulates tongue cancer growth by targeting JAG1. Oncol Rep. 2017;37:2087–94.
Zheng X, Han H, Liu GP, Ma YX, Pan RL, Sang LJ, et al. LncRNA wires up Hippo and Hedgehog signaling to reprogramme glucose metabolism. EMBO J. 2017;36:3325–35.
Lin Z, Sun L, Xie S, Zhang S, Fan S, Li Q, et al. Chemotherapy-induced long non-coding RNA 1 promotes metastasis and chemo-resistance of TSCC via the Wnt/beta-catenin signaling pathway. Mol Ther. 2018;26:1494–508.
Tang T, Guo C, Xia T, Zhang R, Zen K, Pan Y, et al. LncCCAT1 promotes breast cancer stem cell function through activating WNT/beta-catenin signaling. Theranostics. 2019;9:7384–402.
Ma Y, Yang Y, Wang F, Moyer MP, Wei Q, Zhang P, et al. Long non-coding RNA CCAL regulates colorectal cancer progression by activating Wnt/beta-catenin signalling pathway via suppression of activator protein 2alpha. Gut. 2016;65:1494–504.
Luo Y, Chen JJ, Lv Q, Qin J, Huang YZ, Yu MH, et al. Long non-coding RNA NEAT1 promotes colorectal cancer progression by competitively binding miR-34a with SIRT1 and enhancing the Wnt/beta-catenin signaling pathway. Cancer Lett. 2019;440-441:11–22.
Zhang M, Weng W, Zhang Q, Wu Y, Ni S, Tan C, et al. The lncRNA NEAT1 activates Wnt/beta-catenin signaling and promotes colorectal cancer progression via interacting with DDX5. J Hematol Oncol. 2018;11:113.
Fan X, Song J, Zhao Z, Chen M, Tu J, Lu C, et al. Piplartine suppresses proliferation and invasion of hepatocellular carcinoma by LINC01391-modulated Wnt/beta-catenin pathway inactivation through ICAT. Cancer Lett. 2019;460:119–27.
Liang WC, Ren JL, Wong CW, Chan SO, Waye MM, Fu WM, et al. LncRNA-NEF antagonized epithelial to mesenchymal transition and cancer metastasis via cis-regulating FOXA2 and inactivating Wnt/beta-catenin signaling. Oncogene. 2018;37:1445–56.
Wang G, Zhang ZJ, Jian WG, Liu PH, Xue W, Wang TD, et al. Novel long noncoding RNA OTUD6B-AS1 indicates poor prognosis and inhibits clear cell renal cell carcinoma proliferation via the Wnt/beta-catenin signaling pathway. Mol Cancer. 2019;18:15.
Zhang Y, Fang Z, Guo X, Dong H, Zhou K, Huang Z, et al. lncRNA B4GALT1-AS1 promotes colon cancer cell stemness and migration by recruiting YAP to the nucleus and enhancing YAP transcriptional activity. J Cell Physiol. 2019;234:18524–34.
Hao Q, Zong X, Sun Q, Lin YC, Song YJ, Hashemikhabir S, et al. The S-phase-induced lncRNA SUNO1 promotes cell proliferation by controlling YAP1/Hippo signaling pathway. Elife. 2020;9:e55102.
Wu DM, Wang S, Wen X, Han XR, Wang YJ, Shen M, et al. LncRNA SNHG15 acts as a ceRNA to regulate YAP1-Hippo signaling pathway by sponging miR-200a-3p in papillary thyroid carcinoma. Cell Death Dis. 2018;9:947.
Lv B, Zhang L, Miao R, Xiang X, Dong S, Lin T, et al. Comprehensive analysis and experimental verification of LINC01314 as a tumor suppressor in hepatoblastoma. Biomed Pharmacother. 2018;98:783–92.
Wang Y, Wu P, Lin R, Rong L, Xue Y, Fang Y. LncRNA NALT interaction with NOTCH1 promoted cell proliferation in pediatric T cell acute lymphoblastic leukemia. Sci Rep. 2015;5:13749.
Zhou S, Yu L, Xiong M, Dai G. LncRNA SNHG12 promotes tumorigenesis and metastasis in osteosarcoma by upregulating Notch2 by sponging miR-195-5p. Biochem Biophys Res Commun. 2018;495:1822–32.
Yang X, Duan B, Zhou X. Long non-coding RNA FOXD2-AS1 functions as a tumor promoter in colorectal cancer by regulating EMT and Notch signaling pathway. Eur Rev Med Pharmacol Sci. 2017;21:3586–91.
Guo Q, Qian Z, Yan D, Li L, Huang L. LncRNA-MEG3 inhibits cell proliferation of endometrial carcinoma by repressing Notch signaling. Biomed Pharmacother. 2016;82:589–94.
Ren X, Chen C, Luo Y, Liu M, Li Y, Zheng S, et al. lncRNA-PLACT1 sustains activation of NF-kappaB pathway through a positive feedback loop with IkappaBalpha/E2F1 axis in pancreatic cancer. Mol Cancer. 2020;19:35.
Liao Z, Zhao J, Yang Y. Downregulation of lncRNA H19 inhibits the migration and invasion of melanoma cells by inactivating the NFkappaB and PI3K/Akt signaling pathways. Mol Med Rep. 2018;17:7313–8.
Lu Z, Li Y, Wang J, Che Y, Sun S, Huang J, et al. Long non-coding RNA NKILA inhibits migration and invasion of non-small cell lung cancer via NF-kappaB/Snail pathway. J Exp Clin Cancer Res. 2017;36:54.
Wang H, Liang L, Dong Q, Huan L, He J, Li B, et al. Long noncoding RNA miR503HG, a prognostic indicator, inhibits tumor metastasis by regulating the HNRNPA2B1/NF-kappaB pathway in hepatocellular carcinoma. Theranostics. 2018;8:2814–29.
Zhou M, Hou Y, Yang G, Zhang H, Tu G, Du YE, et al. LncRNA-Hh strengthen cancer stem cells generation in twist-positive breast cancer via activation of Hedgehog signaling pathway. Stem Cells. 2016;34:55–66.
Yuan Z, Xiu C, Liu D, Zhou G, Yang H, Pei R, et al. Long noncoding RNA LINC-PINT regulates laryngeal carcinoma cell stemness and chemoresistance through miR-425-5p/PTCH1/SHH axis. J Cell Physiol. 2019;234:23111–22.
Guo L, Zhou Y, Chen Y, Sun H, Wang Y, Qu Y. LncRNA ASAP1-IT1 positively modulates the development of cholangiocarcinoma via hedgehog signaling pathway. Biomed Pharmacother. 2018;103:167–73.
Li L, Ma TT, Ma YH, Jiang YF. LncRNA HCG18 contributes to nasopharyngeal carcinoma development by modulating miR-140/CCND1 and Hedgehog signaling pathway. Eur Rev Med Pharmacol Sci. 2019;23:10387–99.
Qiu S, Chen G, Peng J, Liu J, Chen J, Wang J, et al. LncRNA EGOT decreases breast cancer cell viability and migration via inactivation of the Hedgehog pathway. FEBS Open Bio. 2020;10:817–26.
Sakai S, Ohhata T, Kitagawa K, Uchida C, Aoshima T, Niida H, et al. Long noncoding RNA ELIT-1 acts as a Smad3 cofactor to facilitate TGFbeta/Smad signaling and promote epithelial-mesenchymal transition. Cancer Res. 2019;79:2821–38.
Wang X, Lai Q, He J, Li Q, Ding J, Lan Z, et al. LncRNA SNHG6 promotes proliferation, invasion and migration in colorectal cancer cells by activating TGF-beta/Smad signaling pathway via targeting UPF1 and inducing EMT via regulation of ZEB1. Int J Med Sci. 2019;16:51–9.
Luo K, Geng J, Zhang Q, Xu Y, Zhou X, Huang Z, et al. LncRNA CASC9 interacts with CPSF3 to regulate TGF-beta signaling in colorectal cancer. J Exp Clin Cancer Res. 2019;38:249.
Zhao B, Lu YL, Yang Y, Hu LB, Bai Y, Li RQ, et al. Overexpression of lncRNA ANRIL promoted the proliferation and migration of prostate cancer cells via regulating let-7a/TGF-beta1/Smad signaling pathway. Cancer Biomark. 2018;21:613–20.
Wu N, Jiang M, Liu H, Chu Y, Wang D, Cao J, et al. LINC00941 promotes CRC metastasis through preventing SMAD4 protein degradation and activating the TGF-beta/SMAD2/3 signaling pathway. Cell Death Differ. 2021;28:219–32.
Zhang Y, Zhu M, Sun Y, Li W, Wang Y, Yu W. Upregulation of lncRNA CASC2 suppresses cell proliferation and metastasis of breast cancer via inactivation of the TGF-beta signaling pathway. Oncol Res. 2019;27:379–87.
Zhang B, Yu L, Han N, Hu Z, Wang S, Ding L, et al. LINC01116 targets miR-520a-3p and affects IL6R to promote the proliferation and migration of osteosarcoma cells through the Jak-stat signaling pathway. Biomed Pharmacother. 2018;107:270–82.
Yan D, Jin F, Lin Y. lncRNA HAND2-AS1 inhibits liver cancer cell proliferation and migration by upregulating SOCS5 to inactivate the JAK-STAT pathway. Cancer Biother Radiopharm. 2020;35:143–52.
Wu Q, Ma J, Wei J, Meng W, Wang Y, Shi M. FOXD1-AS1 regulates FOXD1 translation and promotes gastric cancer progression and chemoresistance by activating the PI3K/AKT/mTOR pathway. Mol Oncol. 2021;15:299–316.
Huang JL, Cao SW, Ou QS, Yang B, Zheng SH, Tang J, et al. The long non-coding RNA PTTG3P promotes cell growth and metastasis via up-regulating PTTG1 and activating PI3K/AKT signaling in hepatocellular carcinoma. Mol Cancer. 2018;17:93.
Li T, Xiao Y, Huang T. HIF1alphainduced upregulation of lncRNA UCA1 promotes cell growth in osteosarcoma by inactivating the PTEN/AKT signaling pathway. Oncol Rep. 2018;39:1072–80.
Hu J, Shan Y, Ma J, Pan Y, Zhou H, Jiang L, et al. LncRNA ST3Gal6-AS1/ST3Gal6 axis mediates colorectal cancer progression by regulating alpha-2,3 sialylation via PI3K/Akt signaling. Int J Cancer. 2019;145:450–60.
Funding
This work was financially supported by the China Postdoctoral Science Foundation (No. 2020M670095ZX) and the Natural Science Foundation of Liaoning Province, China (No. 2020-BS-108).
Author information
Authors and Affiliations
Contributions
S. Zhao and S. Zhang performed substantial contributions to conception and design and drafted the article. X.Z. and S.C. designed the figure and revised the article. All authors have read and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, S., Zhang, X., Chen, S. et al. Long noncoding RNAs: fine-tuners hidden in the cancer signaling network. Cell Death Discov. 7, 283 (2021). https://doi.org/10.1038/s41420-021-00678-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-021-00678-8
This article is cited by
-
Biological roles of SLC16A1-AS1 lncRNA and its clinical impacts in tumors
Cancer Cell International (2024)
-
Dyads of GGC and GCC form hotspot colonies that coincide with the evolution of human and other great apes
BMC Genomic Data (2024)
-
Long non-coding RNA-derived peptides are immunogenic and drive a potent anti-tumour response
Nature Communications (2023)
-
Characterization of lncRNAs involved in drought response in Betula platyphylla
Trees (2023)