Abstract
The Lonely Guy (LOG) protein has been identified as a crucial enzyme involved in the production of cytokinins, which are important phytohormones, in plants and plant-interacting organisms. However, C. glutamicum has an isoform (Cg1261) of LOG that contains an extended N-terminal region compared to those of known LOGs, and this type of isoforms are also found in a variety of organisms. Nevertheless, these proteins are considered as lysine decarboxylases, without their functional characterization. To investigate the function of Cg1261, we determined its crystal structure at a resolution of 1.95 Å. Unlike known dimeric LOGs, Cg1261 was found to form a hexamer. The overall shape of the hexamer resembles a trillium flower, in which a twisted dimer constitutes each petal. The dimeric petal is well superposed with known LOG dimers, and its active site conformation is similar to those of LOG dimers, suggesting that the hexameric LOG-like protein also acts as a LOG. Biochemical and in vivo cytokinin production studies on Cg1261 confirms that Cg1261 functions as a cytokinin-activating protein. Phylogenetic tree analysis using 123 LOG-like proteins suggest that the LOG-like proteins can be categorized to the dimeric type-I LOG and the hexameric type-II LOG.
Similar content being viewed by others
Introduction
Cytokinins are plant hormones crucial for promoting cell division and differentiation in plants1. These molecules are chemically known as purine bases, such as isopentenyladenine (iP), trans-zeatin, and 6-benzylaminopurine, in which the N6 atom is modified with isoprenoids or aromatic rings1. Cytokinins often appear as conjugated forms with sugar moieties such as nucleotides, nucleosides, and glucosides, which are biologically less active or inactive for plant receptors2. In cytokinin biosynthesis, the Lonely Guy (LOG) protein has been recently identified as a phosphoribohydrolase that finally releases cytokinins3. LOG proteins produce active cytokinins via dephosphoribosylation, directly hydrolyzing the bond between N6-substituted bases and ribose 5′-monophosphates in conjugated forms. Importantly, the LOG domain is conserved in a wide range of organisms3,4,5, and the majority of LOG proteins are from prokaryotic organisms. However, bacterial LOGs have been especially poorly understood because a LOG protein was originally characterized as a phytohormone-activating plant enzyme, while LOG-like proteins have been considered possible lysine decarboxylases (LDCs) so far.
Corynebacterium glutamicum is widely known for its advantage in production of amino acids, nucleotides, and vitamins6. Although the relationships of C. glutamicum with plant species have not yet been reported, we have previously shown a phosphoribohydrolase activity of misannotated Cg2612 and proposed it as CgLOG based on the structural and biochemical characteristics, which may indicate as yet undiscovered interactions of this microorganism with plants7. Interestingly, we have found that C. glutamicum has an isoform (Cg1261) of CgLOG. Cg1261 contains an extended N-terminal region compared to those of known LOGs, including CgLOG, and this type of isoforms have also been found in a variety of organisms. However, a protein from Thermus thermophilus HB8 (Tt1465), which is homologous to Cg1261, was once assigned as a possible LDC before the discovery of its LOG identity8, and these proteins have also been considered as LDCs, without their functional characterization9,10,11. Thus, the function of this new LOG-like protein from C. glutamicum remains totally unknown. Therefore, despite the high similarity of Cg1261 to other known LOGs, it is not clear whether Cg1261 is actually a LOG protein.
To investigate the function of Cg1261, we determined its crystal structure at a resolution of 1.95 Å and revealed its hexameric oligomerization involving the extended N-terminal region. Based on the biochemical and in vivo cytokinin production experiments, we propose that Cg1261 is a novel type of LOG and belongs to type II LOGs (CgLOGII). Comparative analysis of 123 LOG-like proteins also suggested that LOG proteins could be divided into two different types, dimeric type I LOGs and hexameric type II LOGs.
Results
Monomeric and dimeric structure of Cg1261
To elucidate the function and molecular mechanism of Cg1261, we determined its crystal structure at a resolution of 1.95 Å (Table 1). The asymmetric unit contained three molecules (Molecules I, II, and III), and the crystal volume per unit of protein mass was approximately 1.91 Å3·Da−1, which corresponds to a solvent content of approximately 35.76%. Molecules I, II, and III were modeled as visible residues 42–135 and 138–248, 41–133 and 137–249, and 40–135 and 138–249, respectively. Interestingly, the electron density maps of the N-terminal region (Met1–His39) were invisible in all three molecules, indicating that this region is highly disordered (hereinafter referred to as disordered N-terminal region, DNR). The monomeric structure of Cg1261 showed an overall fold similar to that of CgLOG, and these two proteins have an amino acid identity of 26% (Fig. 1a)7. The Cg1261 monomer forms a Rossmann α/β structure and is composed of eight α-helices and seven parallel β-strands (Fig. 1b). The root-mean-square deviation (RMSD) values among these three monomeric structures were under 0.4, indicating that the three monomers have quite similar structures. Molecules I and II form a dimeric structure, which is similar to the dimeric structure of CgLOG (Fig. 1c). The dimerization is mainly mediated by interactions of hydrophobic patches located at the α1, α5, and α6 helices, and the polar interactions Lys156–Glu181 and Asp46–Lys161 additionally aid the dimerization. Based on the PISA software12 computation, the buried interface area was 2,288 Å per monomer, and the percentage of participating residues was 28.5%.
(a) Amino acid sequence alignment of LOGs. The figure of aligned sequences was generated by ESPript 3.0 software. The secondary structure elements are drawn based on the structure of CgLOGII. Residues involved in enzyme catalysis, AMP binding, and prenyl-group binding are indicated by red, blue, and orange-colored triangles, respectively. The PGGxGTxxE motif and the additional α-helix from type-II LOGs are indicated with green- and orange-colored dotted rectangle, respectively. The disordered N-terminal region (DNR) is labeled. CgLOGII, TtLOGII and AtLOGII are abbreviations of the type-II LOGs from C. glutamicum and T. thermophiles and A. thaliana, respectively. MmLOG, CgLOG and AtLOG3 are abbreviations of the type-I LOGs from M. marinum, C. glutamicum and A. thaliana, respectively. (b) Monomeric structure of CgLOGII. The monomeric structure of CgLOGII is presented as a cartoon diagram. Secondary structure elements are labeled. The additional α-helix located at the N-terminal region is distinguished with an orange color and labeled. (c) Dimeric structure of CgLOGII. The dimeric structure of CgLOGII is presented as a cartoon diagram. Two additional α-helices from two monomers are indicated with an orange-colored circle and labeled. The right figure is the left figure rotated horizontally by 90°.
Hexameric structure of Cg1261
Although the dimeric structure of Cg1261 resembles that of the CgLOG dimer, Cg1261 forms a hexameric structure. The C2221 crystallographic symmetry generated a hexameric structure (Fig. 2a), which was consistent with the size-exclusion chromatography results (Fig. 2b). The hexameric structure was formed by oligomerization of three dimeric structures, and the overall shape of the hexamer looked like a trillium flower, in which each petal was made up of a twisted dimer. The α1 helices of six monomers mainly contribute to hexamerization of Cg1261. The helices form an antiparallel helix bundle through a mixture of hydrogen bonds, salt bridges, and hydrophobic interactions (Fig. 2c). A detailed explanation of the α1 helix and the structural comparison with CgLOG will be provided later. Using the PISA software12, the buried surface of the six helices was computed to be 2,409 Å. In order to confirm the oligomeric status of Cg1261 in solution and to investigate the relative position of DNR in the Cg1261 hexamer, we performed small-angle X-ray scattering (SAXS) analysis in solution using full-length (Cg1261F) and DNR-truncated (Cg1261ΔDNR) Cg1261 proteins (Supplementary Fig. S1). Consistent with the crystallographic and size-exclusion chromatography results, the SAXS results suggested that Cg1261 existed as a hexamer in solution. Because both Cg1261F and Cg1261ΔDNR proteins formed a hexamer in solution, we suspected that DNR is not a main contributor in hexamerization (Fig. 2b and Supplementary Fig. S1). We then performed three-dimensional (3D) reconstructed structure modeling using the SAXS data. Our Cg1261 crystal structure fit well to a 3D reconstructed model of Cg1261ΔDNR. Compared with the SAXS model of Cg1261ΔDNR, that of Cg1261F showed a bulged structure at both sides of the Cg1261ΔDNR model (Fig. 2d,e). These results suggest that DNRs from three monomers are located on the top of the hexameric disc and those from the other three monomers are on the bottom of the disc (Fig. 2d,e).
(a) Hexameric structure of CgLOGII. The hexameric structure of CgLOGII is presented as a cartoon diagram. The additional α-helices from six different monomers are distinguished with different colors. The right figure is the left figure rotated horizontally by 90°. (b) Size-exclusion chromatography of CgLOGII. The full-length (CgLOGIIF) and the DNR-truncated form (CgLOGIIΔDNR) of CgLOGII are eluted as a hexameric form, and CgLOG is as a dimeric form. 1, 2, and 3 indicate standard samples of Ferritin (440 kDa), Aldolase (158 kDa), and Ovalbumin (44 kDa), respectively. (c) Hexamerization mode of CgLOGII. Three monomers are presented as an electrostatic potential surface model and the other three monomers are as a cartoon diagram. The right figure is the left figure rotated horizontally by 90°. (d) (e) SAXS 3-D reconstructed models of CgLOGIIF (d) and CgLOGIIΔDNR (e), respectively. The right figures are the left figure rotated vertically by 90°. The DNR region in CgLOGIIF is indicated with a red-colored dotted rectangle.
Structural comparison of Cg1261 with LOG homologs
Because Cg1261 is an uncharacterized LOG isoform, comparisons between Cg1261 and other LOGs may elucidate the functional and structural implications of Cg1261. First, we compared the structure of Cg1261 with that of CgLOG. When we superposed these two structures, the monomeric structures of these two proteins showed a similar overall fold between each other (Fig. 3a, Supplementary Fig. S2). The most conspicuous distinction between these proteins was observed at their N-terminal regions (Figs 1a and 3a). Compared with CgLOG, Cg1261 contained an extended N-terminal region, which formed an α1 helix (Asp46–Leu64) (Fig. 3a). Since, as described above, the α1 helix is the main contributor to the hexameric architecture of Cg1261 (Fig. 3b), the additional helix is a determinant of the oligomeric state of Cg1261. The α1 helix is located distally to the active site (Fig. 3a), and we speculated that the addition of the α1 helix and hexamerization of Cg1261 would not influence the enzymatic activity of the protein. When we compared the dimeric structures of Cg1261 and CgLOG, noticeable differences were also observed (Fig. 3a, Supplementary Fig. S2). First, the β3–β4 loop (Ile131–Leu145) was highly disordered in Cg1261, whereas the corresponding region (Thr74–Glu89) of CgLOG formed a one-turn helix (Fig. 3c). Because this region is quite diverse in various LOGs and is located in the vicinity of the ribose ring of the substrate7, we suspect that the stabilization mode of the ribose ring may be somewhat different between these two proteins. Second, the β1–α2 loop (Arg78–His83) of Cg1261 was flipped 180° compared with the corresponding region (Ala20–Ser25) of CgLOG (Fig. 3d). The region is located distally to the active site, and the difference does not seem to influence the enzyme activity.
(a) Structural comparison of the LOG monomers. Four LOGs such as CgLOGII, TtLOGII, CgLOG, and MmLOG are superposed (PDB code 5QW3, 1WEK, 5ITS, and 3SBX, respectively). The additional α-helices from CgLOGII and TtLOGII are indicated with a red-colored circle and distinguished with colors of cyan and orange, respectively. Two structurally variable regions, connecting loops of β1-α2 and β3-β4, are also distinguished with different colors. (b) The hexameric forms of CgLOGII and TtLOGII are superposed with the dimeric forms of CgLOG and MmLOG. The additional α-helices from CgLOGII and TtLOGII are indicated with a red-colored circle and distinguished with colors of cyan and orange, respectively. (c) Connecting loops of β3-β4 in LOGs. Four LOGs such as CgLOGII, TtLOGII, CgLOG, and MmLOG are superposed, and the connecting loops of these LOGs are distinguished with colors of cyan, orange, magenta, and green, respectively. The bound AMP in MmLOG is presented as a stick model with a yellow color. The secondary structure elements of β3 and β4 are labeled. (d) Connecting loop of β1-α2 in LOGs. Four LOGs such as CgLOGII, TtLOGII, CgLOG, and MmLOG are superposed, and the connecting loops of these LOGs are distinguished with the same color scheme as in (C). The secondary structure elements of β1 and α2 are labeled.
The results of Dali analysis13 showed that the most structurally similar protein to Cg1261 was Tt1465 (Table 2). Although the function of Tt1465 has not been experimentally defined yet, the protein with a hexameric structure was reported as a potential LDC enzyme8. To compare the structure of Cg1261 with that of Tt1465, we superposed the structures of these two proteins (Supplementary Fig. S2). Interestingly, the hexamerization modes of these two proteins were almost identical to each other. Similar to Cg1261, the additional α-helices at the N-terminal regions of six monomers formed a six-helical bundle at the core of the Tt1465 hexamer (Fig. 3b). Moreover, the dimeric structure and active site formation of these proteins were also almost identical to each other, indicating that Cg1261 and Tt1465 might have a similar function. One noticeable difference between these two proteins, observed at the N-terminal region, was that unlike Cg1261, which has a DNR with 40 amino acids at the N-terminal region, Tt1465 lacked the DNR region (Fig. 1a). Because the DNR region is located distal from the active site, we suspect that the difference is irrelevant to the function of the proteins. The Dali analysis results also showed that the other proteins of high structural similarity to Cg1261 were cytokinin-related LOGs from organisms such as Arabidopsis thaliana, Claviceps purpurea, Mycobacterium marinum, and C. glutamicum (Table 2, Supplementary Fig. S2). Except the existence of the additional α1 helix at the N-terminal region and the hexameric oligomerization by the helix, Cg1261 formed a dimeric component with a mode similar to that in known dimer-forming LOGs (Fig. 3b). Moreover, the active site formation of Cg1261 was also quite similar to that of LOGs, which will be described later. These structural analysis data imply that the Cg1261 and Tt1465 hexamers have a function similar to that of known LOG dimers.
Active site of Cg1261
Superposition of Cg1261 with a LOG protein from M. marinum (MmLOG) in complex with AMP (Protein Data Bank code 3SBX) revealed the active site conformation of Cg1261. As observed in other LOGs7, the active site of Cg1261 is located near the PGGxGTxxE motif. The motif has been known to serve as a monophosphate nucleotide-binding structure, and the amino acid residues 170PGGFGTLDE178 constitute the motif (Fig. 1a). The phosphate moiety was found to be hydrogen-bonded with the main-chain Gly171, Phe173, and Gly174 and the side-chain Thr175 (Fig. 4a). The ribose moiety was mainly stabilized by hydrogen bond interactions between Arg155 and two hydroxyl groups of the ribose moiety (Fig. 4a). To stabilize the adenine moiety, a mixture of hydrophobic and hydrophilic residues, such as Phe152, Lys156, and Asp177, formed the adenine-binding site (Fig. 4a). Two catalytic residues, Arg155 and Glu178, are located near the covalent bond between adenine-N9 and ribose-C1, which is hydrolyzed by the enzyme. In addition, based on a previous prediction7, the Phe152, Phe153, Lys156, Glu181, and Met185 residues seemed to form a prenyl group-binding site (Fig. 4b). Detailed structural comparison of the active site of Cg1261 with those of other LOGs and Tt1465 revealed that the residues constituting the modified adenine-binding site were somewhat variable, whereas those involved in the stabilization of the phosphoribose moiety were mostly conserved (Fig. 4a,b). The Cg1261 and Tt1465 hexamers utilize Phe152 and Asp177 to stabilize the adenine moiety, whereas known LOG dimers contain methionine and glutamate residues at the corresponding positions (Fig. 4b). The residues constituting the prenyl group-binding site are more variable in different proteins. Instead of Phe152 and Phe153 present in Cg1261, the AtLOG3 and CgLOG dimers contain methionine and histidine, respectively, at the corresponding positions, whereas Tt1465 also has phenylalanine residues at the same positions (Fig. 4b). At the position of Met185 in Cg1261, a leucine residue is located in Tt1465 and a tryptophan residue is in the AtLOG3 and CgLOG dimers. However, most importantly, two catalytic residues, Arg155 and Glu178, are located at the same positions in all four proteins, including Cg1261. Taken together, these structural observations led us to propose that the hexameric form of LOG-like proteins such as Cg1261 and Tt1465 might have a function similar to that of the dimeric form of LOGs such as AtLOG3 and CgLOG. The structural difference at the prenyl group-binding site also suggests that, compared to the dimeric LOGs, the hexameric form of LOG-like proteins may accommodate a modified AMP, with a slightly different prenyl group, as a substrate.
CgLOGII is shown as a cartoon diagram in cyan color scheme. The CgLOG structure is superposed with LOGs such as TtLOGII, CgLOG, and AtLOG3, and those LOGs are presented as cartoon diagrams in orange, magenta, and green, respectively. The AMP molecule was prepared as in Fig. 3 and shown as a stick model in yellow. Secondary structure elements are labeled. (a) AMP binding site. Residues involved in the constitution of the AMP binding site are shown as line models. (b) Prenyl-group binding site. Residues involved in the constitution of the AMP binding site are shown as line models.
Cg1261 has a LOG function
To confirm that the Cg1261 and Tt1465 hexamers function as LOG proteins, rather than LDCs as suggested by structural observations, we performed LDC and phosphoribohydrolase activity assays using these proteins and CgLOG. As expected, Cg1261 and Tt1465 exhibited no LDC activity (Fig. 5a), indicating that, contrary to the previous annotation, neither protein is an LDC. However, the phosphoribohydrolase activity assays showed that both Cg2612 and Tt1465 hydrolyzed AMP into an adenine base and ribose 5-phosphate, and the hydrolase activities tended to increase with the reaction time (Fig. 5b). Although the conversion of AMP to adenine and ribose 5-phosphate requires a long incubation time, the levels of the hydrolyzing activities of these proteins were quite similar to that of CgLOG7. In addition, involvement of the suggested residues in enzyme catalysis and substrate binding was also confirmed by site-directed mutagenesis experiments (Fig. 5c). As expected, substitutions of the Arg155, Lys156, Thr175, and Glu178 residues with alanine resulted in a complete loss of phosphoribohydrolase activity, and a Asp177Ala substitution resulted in an almost complete loss of the activity. However, the proteins with their prenyl group-binding residues, such as Phe152, Phe153, and Met185, substituted with alanine still showed the activity because we performed the assay using AMP as a substrate. These results indicate that the hexameric form of LOG-like proteins such as Cg2612 and Tt1465 has a LOG function with a similar substrate-binding mode. It is worth noting that Cg1261ΔDNR showed the same level of phosphoribohydrolase activity as Cg1261 (Fig. 5b), indicating that the DNR region is not involved in the enzyme reaction, as we previously suggested based on the structural observations.
(a) Lysine decarboxylase activity assay of EcCadA, CgLOGII, and CgLOG. The lysine decarboxylase activity of EcCadA, CgLOGII, and CgLOG are measured with or without PLP. All experiments are performed in triplicates. (b) Phosphoribohydrolase activity of CgLOGIIF, CgLOGIIΔDNR and TtLOGII. The activities of CgLOGII and TtLOGII were measured at 30 and 65 °C, respectively. The AMP and adenine standards are indicated at the right side of the figure. (c) Site-directed mutagenesis experiments of CgLOGII. The residues involved in the enzyme catalysis, prenyl-group binding site and the AMP binding were replaced by alanine. The reaction mixture containing each mutant was incubated for 6 hours.
In cytokinin biosynthesis, the most important enzyme is isopentenyltransferase (IPT), which transfers the prenyl group of dimethylallyl pyrophosphate to adenylate or transfer RNA (tRNA). We have previously expected that C. glutamicum may produce cytokinins through a tRNA-mediated pathway and that the IPT (CgIPT) encoded by the Cg2130 gene is involved in the pathway7. To further confirm the LOG function of Cg1261, we performed cytokinin (iP) production experiment in Escherichia coli and monitored the iP production by a liquid chromatography–tandem mass spectrometry method (Fig. 6a–f). No noticeable iP production was detected in cultures of an E. coli strain without a heterologous gene and a strain expressing only the CgIPT-coding gene (EcCgIPT) (Fig. 6b,c,f). However, a significant amount of iP was detected in a culture of an E. coli strain expressing both the CgIPT- and Cg1261-coding genes (EcCgIPT/Cg1261) (Fig. 6d,f). The results confirm that Cg1261 has a cytokinin-activating function. To compare iP production levels between Cg1261 and CgLOG, we also monitored the production of iP using an E. coli strain expressing the CgLOG-coding gene (EcCgIPT/CgLOG) instead of the Cg1261-coding gene. The EcCgIPT/CgLOG strain produced 2~3 times more iP than the EcCgIPT/Cg1261 strain (Fig. 6e,f). The difference in the iP production between Cg1261 and CgLOG could be caused by differences in the protein expression levels in E. coli and/or by those in amino acid residues at the active site of these two enzymes. Taken together, based on the structural and biochemical observations on Cg1261, we propose that hexameric LOG-like proteins such as Cg1261 and Tt1465 have the same cytokinin-activating function as known dimeric LOG proteins.
(a) Linear calibration curve of iP standard. Trend line equation, R2, and the standard curve range were indicated. (b–e) The production of cytokinin in E. coli strains expressing no extra gene (b), CgIPT (c), CgIPT/CgLOGII (d) and CgIPT/CgLOG (e) were selectively monitored using high pressure liquid chromatography-tandem mass spectrometry.<N/A is not available because it deviated from the credible standard curve range. (f) Histograms of detected iP concentration from (b) to (e) were presented with standard deviations.
Classification of LOG proteins
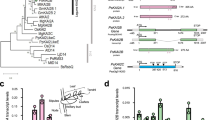
Our study revealed that the proteins such as Cg1261 and Tt1465 are not LDCs but rather a novel type of cytokinin-activating proteins. Because, compared with known LOGs, this type of LOG family proteins shows differences in the oligomeric state and in the residues at the prenyl group-binding site, classification of LOG proteins is required. Thus, we analyzed 123 LOG-like proteins from various phylogenetically diverse organisms found in multifarious habitats and constructed a maximum-likelihood phylogenetic tree (Supplementary Fig. S3). The majority of the sequences were well matched with the LOG motif based on our multiple alignment data. As expected, the LOG-like proteins were categorized into two clusters with high bootstrap values (Fig. 7a). One cluster contained dimeric LOGs, including CgLOG and AtLOG3, and the other contained hexameric LOGs, including Cg1261 and Tt1465. Therefore, we classified the cluster containing the dimeric LOGs as type I LOGs and that containing the hexameric LOGs as type II LOGs. Based on this classification, we refer to the Cg1261 and Tt1465 proteins as CgLOGII and TtLOGII, respectively. Interestingly, the phylogenetic analysis suggested that the type I LOGs could be divided into two subgroups, type Ia and type Ib (Fig. 7a). Type Ia included dimeric LOGs from most organisms such as A. thaliana, C. purpurea, and C. glutamicum. Type Ib included dimeric LOGs from the Actinomycetales, including mammalian pathogens such as Mycobacterium tuberculosis, Nocardia asteroides, and Rhodococcus equi. The phylogenetic analysis also suggested that the type II LOGs could be divided into two subgroups, type IIa and type IIb (Fig. 7a). Type IIa included hexameric LOGs from most organisms, except higher plants, which could be categorized as type IIb.
(a) Unrooted Maximum Likelihood tree of LOG proteins. The phylogenetic tree was drawn as a circle model. Bootstrap values are shown at each node as percentage of 100 replicates. Four subgroups are labeled as different color schemes. Typical LOGs referred in the manuscript are indicated by stars. (b) Amino acid sequence alignment of key residues in LOGs. The key residues involved in the PGGxGTxxE motif, enzyme catalysis, and substrate binding in four subgroups of LOGs are aligned.
These four subgroups showed their own synapomorphies in the residues at the active site (Fig. 7b). The residues involved in the enzyme catalysis in CgLOGII, Arg155 and Glu178, were found to be completely conserved throughout all LOG subgroups (Fig. 7b), indicating that the function of all LOG family enzymes involves the same catalytic mechanism. The PGGxGTxxE motif is also highly conserved in all subgroups (Fig. 7b). One noticeable difference was found in position 177, which is occupied in CgLOGII by aspartate, a residue involved in the binding of the adenine ring moiety. The type II LOGs contained aspartate residues at the corresponding position, while the majority of type I LOGs contained glutamate residues (Fig. 7b). However, the residues involved in substrate binding were somewhat variable among the different types and subgroups. In particular, different residues were found in the positions occupied by Phe152, Phe153, and Met186 in CgLOGII (Fig. 7b). These residues are all involved in the constitution of the prenyl group-binding site, and we suspect that LOGs from the different types and subgroups may produce cytokinin compounds with somewhat different modifications at the position of the prenyl group.
Discussion
None of the type II LOG proteins has been functionally characterized so far, and our structural and biochemical studies of CgLOGII revealed that this new type of LOG proteins exists in a variety of organisms, in addition to known type I LOGs. LOGs have been known to function in plants and plant-interacting organisms as phytohormone producers. Because C. glutamicum is a soil bacterium, it can be speculated that the microorganism also utilizes cytokinins produced by its LOGs to interact with plants. However, the fact that a variety of microorganisms, including mammalian pathogens, have LOG-coding genes raises two hypotheses about the function of LOGs. First, LOGs in some organisms may be involved in the production of different forms of cytokinins, not phytohormone cytokinins. Second, cytokinins may have cellular functions other than those of phytohormones in a variety of microorganisms. Thus, investigations on cellular functions of cytokinins and their analogs in bacterial cells are required.
When we analyzed the operons containing type II LOG-coding genes and their neighboring genes, genes encoding for succinyl-diaminopimelate desuccinylase (DapE), dihydropteroate synthase, glucosyl-3-phosphoglycerate synthase, and a methyltransferase were found to be located close to the type II LOG-coding genes. It is interesting that DapE, an enzyme in the lysine biosynthetic pathway, is located close to the type II LOG-coding genes, which seems to be one of the reasons for misannotation of the LOG-like protein as LDC. More interestingly, the phytopathogen Rhodococcus fascians has genes encoding both types of LOGs, which are located in tandem, indicating that both types of LOGs have similar cellular functions. Most importantly, A. thaliana contains the coding gene (At2g50575) for a type IIb LOG, as well as those for known dimeric forms of LOGs. Further investigations are crucial to reveal whether the type IIb LOG protein is another enzyme for the production of phytohormones.
Molybdenum cofactor carrier proteins (MCPs), known to transfer a molybdenum cofactor to molybdenum enzymes14,15,16, are considered LOG-like proteins because of their structural similarity to LOGs. However, the active site conformation of MCPs is completely different from that of LOG proteins, and the residues involved in the enzyme catalysis and constitution of the PGGxGTxxE motif are not conserved in MCPs. Especially, the arginine residue involved in the LOG enzyme catalysis is replaced by alanine in MCPs. Based on these differences between MCPs and LOG proteins, we suggest that MCPs are excluded from the LOG family of proteins.
Materials and Methods
Cloning, expression, and purification
The genes corresponding for Cg1261 (CgLOGII) from Corynebacterium glutamicum ATCC 13032 was amplified from genomic DNA of C. glutamicum by polymerase chain reaction (PCR) with primers: forward, 5-GCGC CATATG GCTCCTAAACAAACTCCCAGC-3, and reverse, 5-GCGC CTCGAG ATTGTGGCGACGCGCTACGTCC-3. The PCR product was then subcloned using restriction endonucleases NdeI and XhoI into pET30a vector (Merck Millipore) with 6xHis tag at the C-terminus. The resulting expression vectors pET30a: CgLOGII was transformed into E. coli BL21 (DE3) strain and the cell were grown on LB medium containing 100 mgl−1 kanamycin at 37 °C to OD600 of 0.6. The cell was induced by adding 1.0 mM Isopropyl 1-thio-β-D-galactopyranoside (IPTG) for 20 h at 18 °C and harvested by centrifugation at 4000 rpm for 20 minute. Harvested cells was resuspended in ice-cold lysis buffer (40 mM Tris-HCl, pH 8.0) and disrupted by ultrasonication. The cell debris was removed by centrifugation at 11,000 × g for 1 h, and the supernatant was loaded on to Ni-NTA agarose column (QIAGEN). After washing with lysis buffer containing 18 mM imidazole, the bound proteins were eluted with 300 mM imidazole in lysis buffer. Further purification was carried out by applying the HiTrap Q ion exchange chromatography and size exclusion chromatography using Sephacryl-300 (320 ml, GE Healthcare). The purified proteins were concentrated to 32 mg ml−1 in 40 mM Tris–HCl, pH 8.0, and stored at −80 °C for crystallization trials. Site-directed mutagenesis experiments were performed using the QuikChange site-directed mutagenesis kit (Stratagene). The production and purification of the CgLOGII mutants were carried out by the same procedures as described for the wild-type protein. Tt1465 from Thermus thermophilus (TtLOGII) was prepared by the procedure similar to CgLOGII. CgLOG and CadA from E. coli (EcCadA) were prepared as described in ref. 7.
Crystallization, Data collection, and Structure determination
Crystallization of the purified proteins were initially performed by the hanging-drop vapor-diffusion method at 20 °C using commercially available sparse-matrix screens from Hampton Research and Emerald BioSystems. Each experiment consisted of mixing 1.0 μl protein solution with 1.0 μl reservoir solution and then equilibrating it against 0.5 ml of the reservoir solution The CgLOGII crystals were observed from several crystallization screening conditions. After several optimization steps using the hanging-drop vapor-diffusion method, the best-quality crystals appeared in 11 day using a reservoir solution consisting of 0.2 M Lithium chloride and 26% PEG 3350 and reached maximal dimensions of approximately 0.6 × 0.5 × 0.1 mm. For the cryo-protection of the crystals, glycerol of 30% glycerol in reservoir solution was used. Data were collected at 100 K at 7 A beamline of the Pohang Accelerator Laboratory (Pohang, Korea) using a Quantum 270 CCD detector (San Diego, CA, USA). The data were then indexed, integrated, and scaled using the HKL2000 program17. Crystals of CgLOGII belonged to the C-centered orthorhombic space group C2221, with unit cell constants of a = 98.7 Å, b = 173.6, Å c = 79.9 Å. Assuming three molecules of CgLOGII per asymmetric unit, the crystal volume per unit of protein mass was approximately 1.99 Å3·Da−1, which corresponds to a solvent content of approximately 38.38%18. To solve the structure of CgLOGII, phasing was carried out by molecular replacement method. The molecular replacement was performed by MOLREP19 using the structure of possible lysine decarboxylase Tt1465 from T. thermophilus HB8 (PDB code 1WEK) approaching 49% amino acid identity as a search model. The model building was performed using the program WinCoot20 and the refinement was performed with REFMAC521. The data statistics are summarized in Table 1. The refined model of CgLOGII was deposited in the Protein Data Bank (PDB code 5WQ3).
Size-exclusion chromatographic analysis
To investigate the oligomerization of CgLOGIIF and CgLOGIIΔDNR, analytical size-exclusion chromatography was performed using a Superdex 200 10/300 column (GE Healthcare) at NaCl concentrations of 150 mM. Protein samples of 1 mL with concentration of 3 mg/ml were analyzed. The molecular weights of the eluted samples were calculated based on the calibration curve of standard samples.
Solution SAXS measurements
Small-angle X-ray scattering (SAXS) measurements were carried out using the 4 C SAXS II beamline of the Pohang Accelerator Laboratory (Pohang, Korea) with 3 GeV power. A light source from an In-vacuum Undulator 20 (IVU20: 1.4 m length, 20 mm period) of the Pohang Light Source II storage ring was focused with a vertical focusing toroidal mirror coated with rhodium and monochromatized with a Si (111) double crystal monochromator (DCM), yielding an X-ray beam wavelength of 0.734 Å. The X-ray beam size at the sample stage was 0.1 (V) × 0.3 (H) mm2. A two-dimensional (2D) charge-coupled detector (Mar USA, Inc.) was employed. A sample-to-detector distance (SDD) of 4.00 m and 1.00 m for SAXS were used. The magnitude of scattering vector, q = (4π/λ) sinθ, was 0.1 nm−1 < q < 6.50 nm−1, where 2θ is the scattering angle and λ is the wavelength of the X-ray beam source. The scattering angle was calibrated with polystyrene-b-polyethylene-b-polybutadiene-b-polystyrene (SEBS) block copolymer standard. We used quartz capillary with an outside diameter of 1.5 mm and wall thickness of 0.01 mm, as solution sample cells. All scattering measurements were carried out at 4 °C by using a FP50-HL refrigerated circulator (JULABO, Germany). The SAXS data were collected in six successive frames of 0.1 min each to monitor radiation damage. Measurements of LOG protein solutions were carried out over a small concentration range 0.5 ~4.5 mg/mL. Each 2D SAXS pattern was radial averaged from the beam center and normalized to the transmitted X-ray beam intensity, which was monitored with a scintillation counter placed behind the sample. The scattering of specific buffer solutions were used as the experimental background. The Rg,G (radius of gyration) values were estimated from the scattering data using Guinier analysis22. The molecular mass (MM) was calculated from the scattering curve based on the QR method23. The pair distance distribution p(r) function was obtained through the indirect Fourier transform method using the program GNOM24.
Construction of 3D structural models
To reconstruct the molecular shapes, the ab initio shape determination program DAMMIF25 was used. For each model reconstruction, five independent models were selected, and the averaged aligned using the program DAMAVER26. The SAXS curves were calculated from the atomic models using the program CRYSOL27. For comparison of the overall shapes and dimensions, the ribbon diagrams of the atomic crystal models were superimposed onto the reconstructed dummy atom models using the program SUPCOMB28.
Lysine decarboxylase activity assay
Lysine decarboxylase activity assay was performed as described in ref. 7. The activity of LDC was determined by measuring residual concentration of L-lysine using lysine oxidase and peroxidase. After LDC reaction, lysine oxidase converts remaining lysine into 6-amino-2-oxohexanoate, NH3, and H2O2 and then the hydrogen peroxide is reduced by peroxidase with 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS). The oxidized ABTS is detected by spectrophotometric method in absorbance at 412 nm. The assay was performed at 30 °C in a total volume 200 μl, containing 100 mM potassium phosphate, pH 6.0, 0.1 M L-lysine, 0.2 mM pyridoxal-5-phosphate, and 25 μg of purified enzymes. The reaction was stopped by heating the reaction mixture at 100 °C for 5 min. After centrifugation at 13,500 × g for 1 min, 2X reaction solution that contains 0.1 unit ml−1 lysine oxidase and 1 unit ml−1 peroxidase in potassium phosphate buffer is added to the reaction mixture.
Phosphoribohydrolase activity assay
Phosphoribohydrolase activity assay was performed as described in7. The activity was determined by detecting adenine ring compounds separated by thin layer chromatography (TLC) method. Enzyme reactions were carried out in the mixture of 20 mM AMP, 36 mM Tris-HCl, pH 8.0, and 23 μM purified enzymes at 30/65 °C and then the reactions were stopped by heating the mixture at 95 °C for 1.5 min. The reaction mixtures were then dotted on PEI-cellulose-F plastic TLC sheet (Merck Millipore). The mobile phase was 1 M sodium chloride. After development in the TLC chamber, the sheet was dried completely. Adenine ring-including compounds were detected by UV lamp (290 nm).
HPLC-MS/MS analysis
E. coli BL21 (DE3) strains containing cytokinin synthesis genes were grown on LB medium at 37 °C to OD600 of 0.6. After induction with IPTG, the cells were grown for 12 h at 18 °C and harvested by centrifugation at 4000 rpm for 20 minute. 2 g of wet cells were resuspended in ice-cold extracted in Bieleski buffer (60% methanol, 25% CHCl3, 10% HCOOH and 5% H2O)29 of 15 ml and disrupted by ultrasonication. The cell debris was removed by centrifugation at 11,000 × g for 1 h, and 2 μL aliquot of the supernatant was directly injected into the chromatographic system. Prepared samples were analyzed with a reversed-phase Kinetex XB-C18 column (2.1 × 100 mm, 2.6 μm particle size; Phenomenex, USA) in the Nexera XR system (Shimadzu, Japen). The mobile phase for the HPLC system was solvent A and B; (A) 0.1% formic acid in water and (B) 0.1% formic acid in acetonitrile. The HPLC system was interfaced to a TSQ vantage triple quadrupole mass spectrometer equipped with Xcalibur version 1.1.1 (Thermo, Waltham, USA) operating selected reaction monitoring (SRM) Turbo Ion spray mode in the positive ion as the following m/z transitions: 204 → 136 for iP; 277 → 175 for internal standard (ISTD), a chlorpropamide. To avoid contamination by particles, the mobile phase was filtered via a 0.45 μm filter device (PEEK, Germany) before use. Quantifications of iP were carried out by the calibration curve of standard iP (Sigma-aldrich) ranging 1 to 20000 nM. After comparison between specific amount and calculated amount of iP standards from the curve, quantities in calibration range under 20 nM that showed over 10% of the standard differences were discarded. All experiments are performed in triplicates.
Phylogenetic tree analysis
Iterative searching for LOG-like proteins was performed by Basic Local Alignment Search Tool (BLAST) in National Center for Biotechnology Information (NCBI) server using position-specific iterated BLAST (PSI-BLAST) method30, and also selected using protein sequence and name in BlastKOALA (KEGG Orthology And Links Annotation, http://www.kegg.jp/blastkoala), and the Uniprot protein database (http://www.uniprot.org/). Multiple alignment was performed by Clustal omega31. Evolutionary analyses were conducted in MEGA732. The evolutionary history was inferred by using the Maximum Likelihood method based on the Le_Gascuel_2008 model33. The tree with the highest log likelihood (−25391.3439) is shown. The percentage of trees in which the associated taxa clustered together is shown next to the branches. Initial tree(s) for the heuristic search were obtained automatically by applying Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using a JTT model, and then selecting the topology with superior log likelihood value. A discrete Gamma distribution was used to model evolutionary rate differences among sites (5 categories (+G, parameter = 1.8372)). The rate variation model allowed for some sites to be evolutionarily invariable ([+I], 2.6471% sites). The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. The analysis involved 123 amino acid sequences. All positions with less than 95% site coverage were eliminated. That is, fewer than 5% alignment gaps, missing data, and ambiguous bases were allowed at any position. There were a total of 170 positions in the final dataset.
Additional Information
How to cite this article: Seo, H. and Kim, K.-J. Structural basis for a novel type of cytokinin-activating protein. Sci. Rep. 7, 45985; doi: 10.1038/srep45985 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Mok, D. W. & Mok, M. C. Cytokinin metabolism and action. Annu Rev Plant Physiol Plant Mol Biol 52, 89–118 (2001).
Sakakibara, H. Cytokinins: activity, biosynthesis, and translocation. Annu Rev Plant Biol 57, 431–49 (2006).
Kurakawa, T. et al. Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature 445, 652–5 (2007).
Dzurova, L. et al. The three-dimensional structure of “Lonely Guy” from Claviceps purpurea provides insights into the phosphoribohydrolase function of Rossmann fold-containing lysine decarboxylase-like proteins. Proteins 83, 1539–46 (2015).
Samanovic, M. I. et al. Proteasomal control of cytokinin synthesis protects Mycobacterium tuberculosis against nitric oxide. Mol Cell 57, 984–94 (2015).
Vertes, A. A., Inui, M. & Yukawa, H. Manipulating corynebacteria, from individual genes to chromosomes. Appl Environ Microbiol 71, 7633–42 (2005).
Seo, H. et al. Structural basis for cytokinin production by LOG from Corynebacterium glutamicum. Sci Rep 6, 31390 (2016).
Kukimoto-Niino, M. et al. Crystal structures of possible lysine decarboxylases from Thermus thermophilus HB8. Protein Sci 13, 3038–42 (2004).
Michel, A., Koch-Koerfges, A., Krumbach, K., Brocker, M. & Bott, M. Anaerobic Growth of Corynebacterium glutamicum via Mixed-Acid Fermentation. Applied and environmental microbiology 81, 7496–7508 (2015).
Jander, G. & Joshi, V. Aspartate-derived amino acid biosynthesis in Arabidopsis thaliana. The Arabidopsis Book, e0121 (2009).
Bermudez, L. et al. A candidate gene survey of quantitative trait loci affecting chemical composition in tomato fruit. Journal of Experimental Botany 59, 2875–2890 (2008).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. Journal of molecular biology 372, 774–797 (2007).
Holm, L. & Sander, C. Touring protein fold space with Dali/FSSP. Nucleic Acids Res 26, 316–9 (1998).
Aguilar, M., Kalakoutskii, K., Cárdenas, J. & Fernández, E. Direct transfer of molybdopterin cofactor to aponitrate reductase from a carrier protein in Chlamydomonas reinhardtii. FEBS letters 307, 162–163 (1992).
Witte, C.-P., Igeño, M. I., Mendel, R., Schwarz, G. & Fernández, E. The Chlamydomonas reinhardtii MoCo carrier protein is multimeric and stabilizes molybdopterin cofactor in a molybdate charged form. FEBS letters 431, 205–209 (1998).
Fischer, K. et al. Function and structure of the molybdenum cofactor carrier protein from Chlamydomonas reinhardtii. Journal of Biological Chemistry 281, 30186–30194 (2006).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods in enzymology 276, 307–326 (1997).
Matthews, B. W. Solvent content of protein crystals. Journal of molecular biology 33, 491–497 (1968).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallographica Section D: Biological Crystallography 66, 22–25 (2009).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallographica Section D: Biological Crystallography 60, 2126–2132 (2004).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallographica Section D: Biological Crystallography 53, 240–255 (1997).
Glatter, O. & Kratky, O. Small angle X-ray scattering, (Academic press, 1982).
Rambo, R. P. & Tainer, J. A. Accurate assessment of mass, models and resolution by small-angle scattering. Nature 496, 477–481 (2013).
Semenyuk, A. & Svergun, D. GNOM-a program package for small-angle scattering data processing. Journal of Applied Crystallography 24, 537–540 (1991).
Franke, D. & Svergun, D. I. DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. Journal of applied crystallography 42, 342–346 (2009).
Volkov, V. V. & Svergun, D. I. Uniqueness of ab initio shape determination in small-angle scattering. Journal of applied crystallography 36, 860–864 (2003).
Svergun, D., Barberato, C. & Koch, M. CRYSOL-a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. Journal of Applied Crystallography 28, 768–773 (1995).
Kozin, M. B. & Svergun, D. I. Automated matching of high-and low-resolution structural models. Journal of applied crystallography 34, 33–41 (2001).
Bieleski, R. L. The problem of halting enzyme action when extracting plant tissues. Analytical biochemistry 9, 431–442 (1964).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic acids research 25, 3389–3402 (1997).
Sievers, F. et al. Fast, scalable generation of high‐quality protein multiple sequence alignments using Clustal Omega. Molecular systems biology 7, (2011).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Molecular biology and evolution, msw054 (2016).
Le, S. Q. & Gascuel, O. An improved general amino acid replacement matrix. Molecular biology and evolution 25, 1307–1320 (2008).
Acknowledgements
This work was supported by a National Research Foundation of Korea (NRF) grant funded by the Korean Government (MSIP) (NRF-2014M1A2A2033626) and by the Advanced Biomass R&D Center (ABC) of Global Frontier Project (NRF-2011-0031361).
Author information
Authors and Affiliations
Contributions
The authors have made the following declarations about their contributions: Conceived and designed the experiments: K.J.K. Performed the experiments: H.S. Analyzed the data: H.S., K.J.K. Wrote the paper: H.S., K.J.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Seo, H., Kim, KJ. Structural basis for a novel type of cytokinin-activating protein. Sci Rep 7, 45985 (2017). https://doi.org/10.1038/srep45985
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep45985
This article is cited by
-
Seed yield as a function of cytokinin-regulated gene expression in wild Kentucky bluegrass (Poa pratensis)
BMC Plant Biology (2024)
-
Suppressing a phosphohydrolase of cytokinin nucleotide enhances grain yield in rice
Nature Genetics (2023)
-
A cytokinin-activation enzyme-like gene improves grain yield under various field conditions in rice
Plant Molecular Biology (2020)
-
A LONELY GUY protein of Bordetella pertussis with unique features is related to oxidative stress
Scientific Reports (2019)
-
Cytokinins, the Cinderella of plant growth regulators
Phytochemistry Reviews (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.