Abstract
Natural killer (NK) cells play a central role during innate immune responses by eliminating pathogen-infected or tumorigenic cells. In the microenvironment, NK cells encounter not only target cells but also other cell types including non-target bystander cells. The impact of bystander cells on NK killing efficiency is, however, still elusive. In this study we show that the presence of bystander cells, such as P815, monocytes or HUVEC, enhances NK killing efficiency. With bystander cells present, the velocity and persistence of NK cells were increased, whereas the degranulation of lytic granules remained unchanged. Bystander cell-derived H2O2 was found to mediate the acceleration of NK cell migration. Using mathematical diffusion models, we confirm that local acceleration of NK cells in the vicinity of bystander cells reduces their search time to locate target cells. In addition, we found that integrin β chains (β1, β2 and β7) on NK cells are required for bystander-enhanced NK migration persistence. In conclusion, we show that acceleration of NK cell migration in the vicinity of H2O2-producing bystander cells reduces target cell search time and enhances NK killing efficiency.
Similar content being viewed by others
Introduction
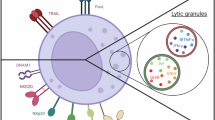
Natural killer (NK) cells play a key role in eliminating virus-infected or tumorigenic cells without prior exposure to antigen for their activation1,2. The interaction between MHC class I molecules on target cells and NK inhibitory receptors plays a major role in regulating NK cell activation. Down-regulated expression of MHC class I molecules on pathogenic cells, following infection by certain virus or neoplastic transformations, renders those cells susceptible to NK cell attack3,4,5. Upon recognition, NK cells form a tight junction with a target cell, which is called immunological synapse (IS)6. Lytic granules (LG) containing perforin and granzymes are then deployed, which constitutes the major mechanism to induce target death7. Upon IS formation, LG are accumulated and released exclusively at the IS to avoid damage of surrounding non-target bystander cells6.
NK cells constantly patrol peripheral organs as essential effectors of immune surveillance8. NK cells can be rapidly recruited to inflammatory sites9 and infiltrate into tumors10. Gradients of chemokines are favorable as directional cues to guide immune cells11 towards or away from anatomically stable structures such as lymphatic vessels12 or bone marrow13. NK cell trafficking and recruitment are mainly regulated by G-protein coupled chemotactic receptors8,14. Extracellular messengers, such as reactive oxygen species (ROS), could also play a role to guide NK cells to their destination. Previous studies have shown that hydrogen peroxide (H2O2), a relatively stable form of ROS, can recruit leukocytes to wounded sites15 or oncogene-transformed cells16.
In a pathological scenario, not all cells in a given NK-patrolling area are necessarily target cells. For example, NK cells encounter stromal cells17, infiltrated immune cells18 as well as malignant cells with expression of MHC class I molecules. These bystander cells pose a challenge to NK cells to efficiently identify their targets in a complex microenvironment. Whether and how the presence of bystander cells can affect the efficiency for NK cells to find and kill their targets has not yet been investigated.
In this study we show that the presence of non-target bystander cells unexpectedly enhanced the killing efficiency as well as NK cell migration. The presence of bystander cells accelerates NK cell migration via H2O2. We establish three mathematical diffusion models and confirmed that local acceleration of NK cells in the presence of bystander cells can decrease search time, and thus increase killing efficiency. We also show that the surface molecule β-integrin on NK cells is involved in mediating bystander-enhanced NK persistence. Together, our findings unravel a novel regulation mechanism between the microenvironment and NK cells.
Results
Presence of bystander cells increases killing efficiency and enhances NK cell migration
We first hypothesized that in the presence of bystander cells, NK cells would require more time to identify their pathologic target cells, due to the need for NK cells to examine each cell they encounter. This in turn should result in an overall reduced killing efficiency. To test this, we used a real-time killing assay, where the cells of interest, normally target cells, were fluorescently labeled with calcein. When target cells are killed by primary NK cells, calcein is released into the supernatant, resulting in a reduction in fluorescence intensity19. We first used P815 cells as bystander cells. Unexpectedly, the presence of P815 cells increased rather than decreased the efficiency of target cell lysis by NK cells (Fig. 1a,b). We further confirmed that P815 cells did not trigger NK killing, with (Supplementary Fig. 1a, P815 as bystanders) or without the presence of target cells (Supplementary Fig. 1a, P815).
(a,b) Target lysis in the presence of bystander cells is analyzed by the real-time killing assay. K562 cells were used as targets for primary human NK cells with an effector to target (E:T) ratio of 5:1. Target cells were plated at 25 × 103 cells/well. P815 cells were used as bystanders at a density of 25 × 103, 50 × 103 or 75 × 103 cells/well, respectively. One representative donor is shown in (a). Quantification from three donors is shown in (b). (c) Bystander primary monocytes enhance NK killing efficiency. Autologous monocytes (auto mono) and NK cells were isolated from the same donors. Non-autologous monocytes (non-auto mono) were from the other donor. K562 and NK cells were plated as described in a. Monocytes were plated at a density of 75 × 103 cells/well. (d,e) The presence of monocytes does not enhance LG release upon target recognition. Primary monocytes and NK cells are from the same donors. The density of monocytes, NK and K562 cells were the same as described in c. Samples were incubated at 37 °C with 5% CO2 for 4 hours before flow cytometry analysis. Quantification from four donors is shown in (e). One representative donor is shown in (d). (f,g) Cell-sized beads do not enhance NK cell killing. K562 cells were used as targets (25 × 103 cells/well) for primary NK cells with an E:T ratio of 5:1. Bystander P815 cells or beads were plated at a density of 75 × 103 cells/well. One representative donor out of four is shown in (f). Quantification from four donors is shown in (g).
To test whether the bystander-enhanced NK killing is restricted to P815 cells or indeed a more general mechanism, we used primary human peripheral monocytes from the same donor (autologous) or from a different donor (non-autologous) as bystanders. As shown in Fig. 1c, both autologous and non-autologous monocytes increased the killing efficiency of primary NK cells. Furthermore, we tested human umbilical vein endothelial cells (HUVEC) as bystander cells. Results from three donors show that the presence of HUVEC increased NK killing efficiency substantially in two donors (Donor 1 and 3) and marginally in one donor (Donor 2) (Supplementary Fig. 2). These findings suggest that bystander-enhancement of NK cell killing efficiency is a general mechanism.
To pinpoint the mechanisms whereby bystander cells enhance NK cell killing, we first examined whether bystander cells increase lytic granule (LG) release of NK cells. LG express the lysosome-associated membrane protein 1 (LAMP1), also known as CD107a, on their membranes. Following target cell recognition by NK cells, lytic granules are released at the immunological synapse resulting in the integration of CD107a into the plasma membrane of NK cells. CD107a is widely used as a functional marker to identify LG release in NK cells20. We found that the surface staining of CD107a on NK cells triggered by target cells was unchanged in the absence or presence of monocytes (Fig. 1d,e) or P815 (Supplementary Fig. 1b,c), indicating that bystanders do not change LG release.
We next investigated whether space occupancy played a role in bystander-enhanced NK killing. For this purpose, we used cell-sized polystyrene beads to replace live bystander cells. The results from our real-time killing assay shows that unlike bystander cells, the polystyrene beads do not enhance the killing efficiency of NK cells (Fig. 1f,g).
In sum, we conclude that the bystander effect on NK cell-mediated cytotoxicity is not a result of increased LG release from NK cells or the mere space occupancy by bystander cells.
Next we examined migration properties of NK cells, which are relevant for efficient NK cell search time to localize target cells and thus influence the killing efficiency. Target cells (K562) and bystander cells (P815) were distinguished by calcein-green and calcein-red, respectively (Fig. 2a, inset). To avoid any possible interference of NK cell functions by a fluorescent dye, we tracked NK cells using bright field with no cell staining. Within the same time frame, NK cells in the presence of bystanders migrated further away from their starting point (Fig. 2a). Our analyses show that the presence of bystander cells significantly enhanced NK cell velocity (Fig. 2b) as well as NK persistence (Fig. 2c), defined as the displacement between the start and end point divided by total length of the trajectory. Quantification of the step size of NK cells revealed higher values in the presence of bystander cells (Fig. 2d). We also found that NK cell do not avoid bystander cells, in fact they appear to approach bystanders on their path to finding target cells (Fig. 2e and Supplementary Movie 1). In contrast, cell-sized polystyrene beads did not enhance NK migration velocity (Fig. 2f). These findings suggest that the increase in target eradication by NK cells in the presence of bystander cells is likely induced by accelerated and more persistent NK cell migration.
(a–e) Primary human NK cell migration on coverslips was visualized with the cell observer at 37 °C. Target K562 cells and bystander P815 cells were labeled with calcein-AM (green) and calcein-red-AM (red), respectively. Trajectories of NK cells were tracked manually and 20 randomly chosen trajectories are shown in (a). Velocity (b), persistence (c) and step sizes of killer cell migration (d) were analyzed. Results are from three donors. One exemplary migrating NK cell is marked by the yellow circle in e. Scale bars are 20 μm. (f) Cell-sized beads do not change migration velocity of NK cells. Migration of primary human NK cells on coverslips was visualized with the cell observer at 37 °C. Target K562 cells were loaded with calcein-AM. Beads or calcein-red-AM loaded P815 cells were used as bystanders. NK cells were plated with K562 cells only (Ctrl), K562 and P815 cells (P815) or K562 and beads (Beads). Results are from three independent experiments. (g,h) NK cells are attracted to bystander cells. The probability that NK cells bypassed or were attracted to nearby (closer to 10 μm) bystander cells was analyzed as shown in (g). To avoid bias, only NK cells that encountered at least 8 bystander cells were analyzed. Quantification from six NK cells is shown in (h). (i,j) NK cells are accelerated in the vicinity of bystanders. As depicted in (i), NK cells attracted to bystanders (P815 or beads) were analyzed for the duration of approaching (0–1), touching (1–2), leaving (2–3) and full trajectory from approaching to leaving (0–3). Quantification from three experiments is shown in (j). (k,l) Lightsheet microscopy was used to analyze NK cell migration in 3D at 37 °C. Migration velocity is accelerated by bystander cells. For imaging, primary NK cells, target K562 cells and monocytes were loaded with CellTracker™ Deep Red, Calcein-AM and CellMask™ Orange, respectively. Matrigel was polymerized at 37 °C with 5% CO2 for 30 minutes before imaging. NK cells were automatically tracked and analyzed by Imaris. Results are from two donors.
We further quantified how often NK cells were attracted to bystander cells using the analysis shown in Fig. 2g. When an NK cell approaches a bystander cell (at 10 μm distance), this NK cell can either make contact with this bystander (defined as ‘attracted’) or pass by without contact (defined as ‘passing by’). We found that in most cases (more than 80%) NK cells were attracted to bystander cells (Fig. 2h).
To investigate whether bystander cells influence NK cell migration locally, we analyzed the velocity of NK cells in the vicinity of bystander cells. As shown in Fig. 2i, the trajectory of an NK cell near a bystander was separated into three phases: approaching (0–1), touching (1–2) and leaving (2–3). Using this classification we compared P815 cells and cell-sized polystyrene beads as bystanders. NK cells did not distinguish between P815 and the beads regarding the contact phase, indicating that NK cells need comparable times to examine different objects. On the other hand, approaching and leaving times were clearly accelerated in the case of P815 bystanders compared to the beads (Fig. 2j). Adding up the times needed for approaching, touching and leaving, NK cells altogether need less time to examine P815 bystanders compared to beads (Fig. 2j, All). This is consistent with an increased migration velocity in the vicinity of P815 bystanders. We conclude that bystander cells attract NK cells and locally accelerate NK cell migration.
In most cases, in vivo NK cells migrate in a three dimensional (3D) environment. We thus examined by lightsheet microscopy whether NK cell migration in 3D matrices could be affected by the presence of bystander cells. Primary human NK cells and monocytes were isolated from the PBMCs of the same healthy donors. We observed that the velocity of NK cells was enhanced by the presence of bystander monocytes, whereas the persistence of NK cells was not altered (Fig. 2k,l). In conclusion, NK cells are also accelerated by bystander cells in a 3D environment. Different from the 2D scenario, however, in 3D collagen matrices the persistence of NK cells was not affected by the bystander cells.
The increased motility of NK cells near bystander cells decreases search time for target cells
Considering the results described so far, we hypothesized that the local acceleration of NK cells close to bystanders would decrease search time to locate target cells and thereby enhance NK killing efficiency. We tested this hypothesis using a mathematical model for the random search process performed by the NK cells in a two-dimensional environment. The model consists of: 1) Nk disk-like particles performing random Brownian motion with diffusion constant D in a plane of size L × L (representing the NK cells searching for target cells), 2) Nt disk-like target cells which are placed at random positions in the plane and do not move (target cells also barely move in our in vitro experiments), and 3) the number of remaining target cells as a function of time, Nt(t). Whenever a killer disk touches a target disk, the target disk is removed and Nt(t) decreases by one. Moreover, the model contains No randomly placed immobile disk-like obstacles of radius robs representing inactive NK cells. Their effect is only to exclude areas where the killers can go. Finally, the model contains Nb randomly placed immobile disk-like particles with radius R representing bystanders. There are two effects of bystander cells on the motion of the NK cells in the model: 1) they exclude the area occupied by the bystander itself, and 2) they change the diffusion constant for the killers from D to Dacc > D in a circular region of radius Δ > R around the center of the bystander cell. The increased diffusion constant Dacc mimics the experimentally observed acceleration of the killer motion close to the bystander cells.
With the help of computer simulations (see Materials and Methods) we calculated for various parameter sets (Nk, Nt(t = 0), No, Nb, robs, Dacc, Δ): 1) the average number of surviving targets at time t, <Nt(t) >, and 2) the average half-time t1/2, the average time at which half the targets were found and removed. The average half time (t1/2) is a measure for the killing efficiency of the NK cells in this model framework with the given parameters. For the model calculations, it is convenient to use dimensionless units; we used the diameter of the NK cells as the unit of length, i.e. set R = 0.5, and fixed the time unit by setting the diffusion constant of the NK cells to D = 1. Moreover, we fixed the system size to L = 50 and chose the particle numbers such that particle densities in the model match the cell densities in the experiments.
First we analyzed a scenario that would arise if bystanders were obstacles with no accelerating effect on NK cells (formally: Nb = 0). The corresponding model is shown in Fig. 3a. Note that the center of the randomly moving NK cells (black) cannot approach the obstacles (grey) closer than the concentric circle surrounding them. Figure 3b shows the fraction of killed targets up to time t, 1-<Nt(t) >/Nt(t = 0), as a function of time for different obstacle numbers No of radius robs = 0.5 and using a fixed number of killer and target cells Nk = Nt(t = 0) = 20. The model suggests that the higher the number of obstacles the slower the killing rate. This observation is precisely quantified by the average half-time t1/2 as a function of the number of obstacles No for different obstacle radii robs, once for 20 killers (left) and once for only one killer (right) (Fig. 3c): t1/2 increases monotonically with No, and the larger the obstacles the faster NK cells kill. Consequently, bystanders in the experiments cannot simply be considered passive obstacles.
(a) Sketch of the mathematical model of a Brownian (diffusive) disk-like particle performing a random search for disk-like targets among disk-like obstacles: the black disk represents one randomly moving NK cell, the black wiggly lines its diffusive motion with diffusion constant D, grey disks the immobile obstacles of radius robs, and green disks immobile targets of radius R. Targets that are hit, here at time tkill, are removed upon contact with the NK cells. (b) The ratio of killed targets is depicted as a function of time for Nk = 20 killers, Nt(0) = 20 targets, different numbers of obstacles No, and robs = 0.5. (c) The average half time t1/2 is given as a function of the number of obstacles No for different values of robs for the same number of killers and targets (left) and for the case of only one killer (right). (d) Same as (a) but now with bystanders (blue). Diffusion of the NK cells is accelerated by an increased diffusion constant, Dacc > D, within the circles with radius Δ around bystanders, as indicated by red portions of the wiggly lines representing the random killer motion. Otherwise the symbols are similar to (a). (e) The ratio of killed targets for the model shown in (d) as a function of time for Nk = 20 killers, Nt(0) = 20 targets, No = 200 obstacles, Dacc = 4 and Δ = 3 and different numbers of obstacles. (f) Average half time t1/2 as a function of Nb for Nk = 20 killers, Nt(0) = 20 targets, No = 200 obstacles and Dacc = 4 and different values of Δ. (g) Average half time t1/2 as a function of Dacc for Nk = 20 killers, Nt(0) = 20 targets, No = 200 obstacles and Δ = 3 and different numbers of obstacles.
Next, we studied the effect of bystanders accelerating the NK cells in their proximity, represented in the model by Nb > 0 as depicted in Fig. 3d. Note that the red parts of the killer trajectory lie within a circle of radius Δ > R around a bystander, where the random motion of the killer is governed by a larger diffusion constant Dacc > D. To limit the parameter space, we fixed robs = R here. Figure 3e shows our results for the fraction of killed targets up to time t, 1- < Nt(t) >/Nt(t = 0), as a function of time for different bystander numbers Nb and a fixed number of killer and target cells Nk = Nt(t = 0) = 20 and obstacles No = 200. The increased diffusion constant is Dacc = 4 and the acceleration radius Δ = 3. The data demonstrate that the higher the number of bystanders, the faster the killing proceeds. Furthermore, the half-time t1/2 is depicted as a function of the number of obstacles Nb for different acceleration radii Δ and fixed Nk, Nt, No, and Dacc (Fig. 3f). Now t1/2 decreases monotonically with Nb, and the larger the acceleration radius the faster NK cells kill. Figure 3g shows that t1/2 also decreases monotonically with the diffusion constant Dacc.
Finally, we examined the robustness of our results using two different discrete lattice-based “random walk” models. In the discrete Model 1, the bystanders are placed in this lattice and all cells could move (Supplementary Fig. 3a). In the discrete Model 2, the bystanders are always immobile and placed between the lattices (Supplementary Fig. 3b). In both discrete models the hopping rates of NK cells are dependent on the neighboring cells. Different combinations for the motility of the cell types in Model 1 (Supplementary Fig. 3c) and various hopping possibilities of NK cells in Model 2 (Supplementary Fig. 3d) were investigated. From both Models, the half time t1/2 decreases with an increase of bystander cell numbers (Supplementary Fig. 3c and d). The discrete Model 1 also predicts that the more bystander cells, the faster target cells are found by NK cells (Supplementary Fig. 3e) and the half time t1/2 as a function of q (hopping rate to an allowed direction, details see Supplementary Method) decreases with an increase of bystander cell numbers (Supplementary Fig. 3f). Thus, both the continuous model and the discrete models predict that accelerating NK cells by bystanders enhances killing efficiency.
Bystander cell-generated H2O2 plays a key role in the acceleration of NK cell migration and enhanced killing efficiency
We next searched for molecular mechanisms responsible for accelerating NK cells in the vicinity of bystander cells. We tested H2O2, which has been shown to influence immune cell migration21,22,23. Analyzing H2O2 production by different bystander cell types, we found that P815 bystander cells (Fig. 4a) and primary monocytes (Supplementary Fig. 4) but not K562 target cells or NK cells themselves produced H2O2 (Fig. 4a). To test if environmental H2O2 is relevant for the bystander effect seen for NK cell migration, we applied the naturally occurring H2O2 scavenger catalase, which depletes H2O2 (as confirmed in Supplementary Fig. 5). We then analyzed NK cell migration in the presence of P815 bystander cells with or without catalase. The bystander cell-induced increase in NK cell migration velocity was inhibited by catalase (Fig. 4b,c), whereas LG release of NK cells was not affected (Fig. 4d,e), suggesting that H2O2 acts specifically on migration but not LG release. We conclude that bystander-produced H2O2 accelerates NK cell migration and thereby enhances NK killing efficiency.
(a) H2O2 production by NK, K562 and P815 cells. As for real-time killing assay, target K562, bystander P815 cells and primary NK cells were plated at a density of 25 × 103, 75 × 103 and 25 × 104 cells/well, respectively. Production of H2O2 was determined by the Amplex UltraRed assay at 37 °C every 4 minutes for 4 hours. (b,c) Depletion of H2O2 by catalase attenuates bystander cell-enhanced migration velocity of NK cells. NK cell migration on coverslips was visualized at 37 °C using a 10× objective. Primary NK cells were incubated with target K562 cells and bystander P815 cells with or without catalase (10 units/ml). Randomly chosen NK trajectories are shown in (b). NK velocity is shown in (c). (d,e) Depletion of H2O2 by catalase does not affect degranulation. Primary NK, target K562 and bystander P815 cells were plated at a density of 12.5 × 104, 25 × 103 and 75 × 103 cells/well, respectively. Samples were incubated at 37 °C with 5% CO2 for 4 hours before flow cytometry analysis. One representative donor out of five is shown in (d). Quantification of all five donors is shown in (e). (f–h) Addition of H2O2 enhances velocity but not persistence of NK cell migration. NK cell migration in the presence of vehicle or H2O2 (2 μM) was visualized at 37 °C with the cell observer. Trajectories, velocity and persistence of NK cell migration are shown in (f,g and h), respectively. (i) NK population-depleted PBMCs (LO) produces minimal amount of H2O2. Bystander cells (P815 or LO) and target K562 cells were plated at 75 × 103 and 25 × 103 cells/well, respectively. Production of ROS was measured by the Amplex UltraRed assay. Arbitrary fluorescence units (AFU) were assigned following the removal of fluorescence background from the medium. Average AFU of triplicates is shown. (j,k) NK population-depleted PBMCs (LO) do not increase NK killing efficiency. The real-time killing assay was used to determine target lysis. Bystander cells (P815 or LO) and target K562 cells were plated at 75 × 103 and 25 × 103 cells/well, respectively. One representative experiment is shown in (j). Quantification from four experiments is shown in (k).
To further verify if the observed bystander-effects in target killing are due to increased H2O2, we added 2 μM of external H2O2 into the growth medium, which is in the range of H2O2 produced by P815 bystander cells in vitro (Supplementary Fig. 6). We found that 2 μM H2O2 specifically increased the velocity of NK cells (Fig. 4f,g), whereas migration persistence was not affected (Fig. 4h). LG release was also not changed by the addition of H2O2 (Supplementary Fig. 7), confirming the specific action of H2O2 on NK cell migration. To further test the importance of H2O2 for the bystander-enhanced NK killing, we applied NK population-depleted PBMC (LO) as bystander cells, which produce very low levels of H2O2 (Fig. 4i). P815 cells were used as positive controls. Real-time killing assay analysis shows that NK population-depleted PBMC could not enhance NK cell killing efficiency (Fig. 4j,k). Taken together, these data suggest that the alteration of H2O2 in the microenvironment generated by bystander cells is responsible for accelerating NK cell migration and the enhanced killing efficiency.
Beta-integrins on NK cells are responsible for the bystander-enhanced NK cell persistence
As NK cells clearly make contact with bystanders (shown in Fig. 2e), we examined whether cell adhesion molecules on NK cells are involved in the decreased search time. Integrins are essential for NK cell migration24,25,26. From the expression profile of integrins in NK cells determined by microarrays (Supplementary Table 1), we found that integrin β1, β2 and β7 are the three dominantly expressed integrin β-chains in NK cells. We thus applied a cocktail of antibodies targeting β1, β2 and β7 integrin to functionally block the interaction of integrins with other effector molecules27,28,29. The migration persistence was impaired by the elimination of β-integrin function as shown in the trajectories and the analysis (Fig. 5a,b), whereas the accelerated NK cell migration by bystander cells was not affected (Fig. 5c).
Primary NK cell migration was visualized at 37 °C by the cell observer with a 20× objective every 20 seconds for 30 min. K562 and P815 cells were used as target and bystander cells, respectively. Antibody cocktail targeting integrin β1, β2 and β7 (anti-ITGB) were applied in (a). Twenty randomly chosen NK trajectories are shown in (a). Analyses of persistence and velocity from three donors are shown in (b and c). (d–f) An antibody against LFA-1 α chain (anti-ITGAL) was applied. Twenty randomly chosen NK trajectories are shown in (d). Analyses of persistence and velocity from three donors are shown in (e and f).
The other predominantly expressed integrin identified by microarray analysis is integrin α-L chain (ITGAL, Supplementary Table 1), the distinct α chain for lymphocyte function-associated antigen 1 (LFA-1) involved in asymmetrical NK cell spreading and migration30. A neutralizing antibody targeting LFA-1 α chain (ITGAL) changed neither the velocity nor the persistence of NK cells in the presence of bystander cells (Fig. 5d–f). Taken together, bystander-enhanced NK velocity is regulated by H2O2; whereas bystander-enhanced NK persistence is regulated by β-integrins (β1, β2 and β7) not by the LFA-1 α chain on NK cells.
Discussion
NK cell migration velocity is enhanced in the vicinity of non-target bystander cells, which increases NK cell dependent target cell killing. This effect is mediated by H2O2 production of bystander cells. In addition bystander cells enhance NK cell migration persistence in an integrin dependent manner in a 2D but not 3D environment.
An obvious important question is whether bystander cells can promote killing efficiency in vivo. Unfortunately, this cannot easily be resolved; however, our results indicate that potentially, all H2O2-producing non-target cells could act as bystanders. Leukocytes produce large amounts of ROS to fight pathogens at inflammation sites31, making them good candidates as bystander cells. We indeed found that H2O2-producing monocytes can enhance killer cell velocity. In addition, HUVEC, a human endothelial cell line, is able to act as bystander cells (Supplementary Fig. 2) and that P815 cells, a mastocytoma cell line, enhance both NK migration and killing efficiency. Together, these data suggest that the bystander effect is a general phenomenon. Considering that H2O2 enhances leukocyte recruitment to wounded sites in vivo in zebrafish15 and that bystanders increase migration velocity in human NK cells in a 3D matrigel environment, it appears likely that the bystander effect operates in vivo in humans.
Our hypothesis is that a local increase in killer cell motility, when in contact with or close to bystander cells, is sufficient to decrease search time. This is experimentally very difficult to verify. Since theoretical models have been proven to be a powerful tool to untangle complex cell behavior from experimental data32, we analyzed three mathematical models for random search in a two-dimensional confined area. Although upon first examination it appears counterintuitive that without providing a directional cue towards target cells, local acceleration of NK cells in the vicinity of bystander cells can result in a more efficient search process, the models predict that search time decreases systematically with increasing density of bystander cells and with increasing local acceleration. The explanation for this is that in a random search the ensemble of possible search trajectories is not changed by local accelerations and thus the net search time only depends on the velocity along those trajectories. However, bystander cells themselves also represent disk-like obstacles of radius R and we have demonstrated that diffusion in an archipelago of obstacles is slowed down, depending on the density of obstacles, see also33. Consequently, there is a competition between diffusion acceleration in the vicinity of bystander cells and the diffusion slowing down due to the bystanders themselves being obstacles. As we have shown the first effect wins when the acceleration region around bystanders and the increased diffusion constant is sufficiently large in these regions and a local motility enhancement through the bystanders is indeed sufficient to increase killing efficiency.
H2O2 can be generated as a by-product of cellular metabolism34,35,36 or more specifically by specialized enzymes like NADPH oxidases37,38. Increased ROS production has been observed in various tissues, for instance lung tissues39, hepatocytes40 and kidney41. Studies also indicate that in the kidney NADPH oxidases are expressed in a regional and cell-specific manner41. Thus it is expected that the bystander effect depends on the microenvironment generated by the interaction of tumor or inflammation with a specific organ or tissue. This in turn suggests that in different tissues or microenvironments of cancer or inflammation we could have different cell types as bystanders. On the other hand, too much H2O2 in the microenvironment, i.e. higher than 10 μM, induces the apoptosis of NK cells, especially in cytotoxic CD56dim NK cells42,43. Therefore appropriate H2O2 levels may be of great importance to optimize the effect of immune surveillance.
Our results show that in 2D bystander cells enhance both velocity and persistence of NK cells (Fig. 2b,c). However, in a 3D scenario, only the velocity of NK is increased by the presence of bystander cells (Fig. 2i). It is reported that in vivo, the motility of murine leukocytes is largely independent of integrin44,45. Together, this suggests that in 3D, primary human NK cells could also employ more integrin-independent amoeboid movement.
We have identified that β1, β2 and β7 integrins on NK cells are responsible for regulating the bystander-enhanced NK persistence. Since we showed that H2O2 is involved in the bystander-enhanced velocity of NK migration but not in the persistence change, it is unlikely that integrins on NK cells are the effector molecules of environmental H2O2. There is evidence that in CD4+ T cells ROS can regulate several molecules in T cell receptor signaling pathways including protein tyrosine phosphatase, Src-family kinases23 and Orai1 channels46. These proteins may be promising candidates to mediate the H2O2-dependent increase in NK cell migration velocity. Regardless of the molecular mechanism of H2O2-mediated bystander-enhanced migration and killing efficiency of NK cells, H2O2 is an important factor how the microenvironment influences tumor progression and elimination by killer cells.
Materials and Methods
Reagents and antibodies
All chemicals not specifically mentioned were from Sigma (highest grade). Calcein-AM, CellMask™ Orange, CellTracker™ Deep Red, CellTraceTM and calcein red-orange AM were purchased from ThermoFisher Scientific. The antibodies used in our experiments include: antibody against Integrin-α-L (ITGAL, LFA-1) (Antibodies-online GmbH), human Integrin β1 (R&D Systems) and human Integrin β2 (R&D Systems) polyclonal antibodies and antibody against human/mouse Integrin β7 (Biolegend). Polybead® Polystyrene Microspheres (15 μm, 18328–5) were purchased from Polysciences. Cultrex® Reduced Growth Factor Basement Membrane Extract and PathClear® was purchase from Trevigen.
Cell culture
Raji, K562 and P815 cells were cultured in RPMI-1640 medium (ThermoFisher Scientific) supplemented with 10% FCS at 37 °C with 5% CO2. Peripheral blood mononuclear cells (PBMC) were obtained from healthy donors as previously described19. Primary human NK cells and monocytes were negatively isolated from PBMC from the same donors using Dynabeads® Untouched™ Human NK Cells Kits and Dynabeads® Untouched™ Human Monocytes Kits, respectively (ThermoFisher Scientific). Before use, the isolated primary NK cells and monocytes were cultured overnight at 37 °C with 5% CO2 in AIMV medium with 10% FCS at a density of 2 × 106 and 1.5 × 106 cells/ml, respectively.
Real time killing assay
The real-time killing assay was carried out as described before19. Briefly, target cells were loaded with calcein-AM and plated on a Falcon® 96-well black with clear flat bottom plates (Corning) at a density of 2.5 × 104 cells/well. NK cells were subsequently added and the fluorescence was measured by a GENios Pro micro-plate reader (TECAN) every 10 minutes for 4 hours at 37 °C. Bystander cells were plated together with target cells.
Time-lapse imaging and migration analysis
Target cells were loaded with calcein-AM and bystander cells were loaded with CellTrace calcein red-orange-AM. Afterwards 75 × 103 bystander cells and 25 × 103 target cells were plated for each well in 96-well plates. 1.25 × 105 NK cells were then added to each well. NK cell migration was visualized at 37 °C using the cell observer microscope (Zeiss) equipped with a 10× objective (Fluar, NA 0.50). The green channel (Ex 470 nm/Em 525 nm) and the red channel (Ex 550 nm/Em 600 nm) as well as wide field images were acquired at an interval of 16.8–38.2 seconds according to the experimental conditions. Cells were tracked manually with ImageJ 1.45 s plug-in Speckle TrackerJ. Immobile NK cells were discarded from statistical analysis. For testing the function of adhesion molecules, NK cells were pre-incubated with the neutralizing antibody or vehicle at 37 °C for 30 minutes before being added to target cells. The cocktail of antibodies against integrins contains 10 μg/ml anti-integrin β1, 20 μg/ml anti-integrin β2 and 10 μg/ml anti-integrin β7. The antibody against LFA1 was used at a concentration of 10 μg/ml.
3D live cell imaging with lightsheet microscopy
Target K562 cells were loaded with calcein-AM, bystander cells (monocytes) were loaded with CellMask™ Orange, and NK cells were loaded with CellTracker™ Deep Red. Target cells and NK cells were mixed in AIMV with 10% FCS with or without bystander cells. Afterwards, cell suspension was mixed with growth factor reduced Matrigel to an end concentration of 8.1 mg/ml. This cell/matrigel mix was transferred into a capillary (BRAND) and kept at 37 °C with 5% CO2 for 30 minutes for polymerization. Final density of NK cells, bystander monocytes and target K562 cells were 1 × 107, 1 × 107 and 5 × 106 cells/ml, respectively. Afterwards, the migration of cells was visualized with the lightsheet micoscopy (5× objective) at 37 °C with a z-step size of 1 μm and time interval of around 30 seconds. NK cells were tracked and analyzed using Imaris 8.1.2 (containing Imaris, ImarisTrack, ImarisMeasurementPro, ImarisVantage; Bitplane AG, software available at http://bitplane.com).
Amplex UltraRed Assay
To determine the concentration of H2O2 generated during the killing process, we carried out a direct peroxidase assay using Amplex UltraRed (ThermoFisher Scientific). Target cells (2.5 × 104) alone or together with bystander cells (7.5 × 104) were seeded in 96-well plates in 100 μl phenol red free DMEM-F12 supplemented with 10 mM HEPES. Twenty minutes after the cells were seeded, NK cells in 10 μl DMEM-F12 medium were added carefully on top of the pre-seeded cells. DMEM-F12 alone was used as the background control. Freshly mixed 100 μM Amplex UltraRed regent containing 10 units/ml SOD and 0.1 units/ml HRP ± 10 units/ml catalase in the phenol red free DMEM were added into wells immediately after the NK cell addition. Plates were immediately measured with a Genios Pro micro-plate reader (TECAN) at an excitation/emission of 535 nm/590 nm at 37 °C every 4 or 6 minutes for 4 hours.
Degranulation assay
NK cells were settled in 96-well plates with target cells and/or bystander cells as indicated in the figures. Cells were kept in AIMV medium containing Alexa488 conjugated anti-human CD107a antibody (Biolegend) and GolgiStopTM (BD Biosciences). Cells were incubated at 37 °C with 5% CO2 for 4 hours. Afterwards cells were washed twice and stained with APC/Cy7 conjugated anti-human CD16 Antibody (Biolegend) to identify NK cells. Data were acquired using a FACSVerse™ flow cytometer (BD Biosciences) and were analyzed with FlowJo v10 (FLOWJO, LLC). Only CD16+ cells in lymphocytes population were gated for further analysis.
Microarray measurement and data analysis
Preparation of the templates and microarray analysis were performed as described before47. Using the Agilent Feature Extraction Software the raw values from the scanned image files were extracted. In the next step, the R software was used to normalize the data (quantile normalization) and to perform a log2 transformation. The normalized data were used to compare the expression differences between the different samples.
Computational analysis of the mathematical model
A kinetic Monte Carlo scheme was implemented to generate the Brownian motion of disk-like particles in a square geometry; disks represent targets, obstacles, and bystanders. A very efficient First Passage Kinetic Monte Carlo algorithm (FPKMC) was used as documented in the literature48,49,50, also denoted as GFRD MC (Greens function reaction diffusion Monte Carlo). For each parameter set 40.000 samples of the random trajectories and the random times were generated when targets were hit and removed. Movies 2 and 3 in the Supplementary Material show example trajectories for the generated search processes without and with bystanders, respectively. A subsequent average over these 40.000 samples produced the results for the average number of remaining targets at time t, <Nt(t) >, for each parameter set, which is presented and discussed in the main text.
Statistical analysis
Data are presented as mean ± S.E.M unless otherwise indicated. The differences between two groups were tested using the student’s t-test. For multiple comparisons between groups, the significance was first examined using the One way ANOVA test, followed by Kruskal-Wallis test. *P < 0.05; **P < 0.01; ***P < 0.001; n.s., not significant.
Ethical considerations
Research carried out for this study with human material (leukocyte reduction system chambers from human blood donors) is authorized by the local ethic committee (declaration from 16.4.2015 (84/15; Prof. Dr. Rettig-Stürmer)).
Additional Information
How to cite this article: Zhou, X. et al. Bystander cells enhance NK cytotoxic efficiency by reducing search time. Sci. Rep. 7, 44357; doi: 10.1038/srep44357 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Long, E. O., Kim, H. S., Liu, D., Peterson, M. E. & Rajagopalan, S. Controlling natural killer cell responses: integration of signals for activation and inhibition. Annu Rev Immunol 31, 227–258 (2013).
Vivier, E., Tomasello, E., Baratin, M., Walzer, T. & Ugolini, S. Functions of natural killer cells. Nat Immunol 9, 503–510 (2008).
Karre, K., Ljunggren, H. G., Piontek, G. & Kiessling, R. Selective rejection of H-2-deficient lymphoma variants suggests alternative immune defence strategy. Nature 319, 675–678 (1986).
Ljunggren, H. G. et al. Empty MHC class I molecules come out in the cold. Nature 346, 476–480 (1990).
Hoglund, P. & Brodin, P. Current perspectives of natural killer cell education by MHC class I molecules. Nat Rev Immunol 10, 724–734 (2010).
Orange, J. S. Formation and function of the lytic NK-cell immunological synapse. Nat Rev Immunol 8, 713–725 (2008).
Krzewski, K. & Coligan, J. E. Human NK cell lytic granules and regulation of their exocytosis. Front Immunol 3, 335, 10.3389/fimmu.2012.00335 (2012).
Walzer, T. & Vivier, E. G-protein-coupled receptors in control of natural killer cell migration. Trends Immunol 32, 486–492 (2011).
Luster, A. D., Alon, R. & von Andrian, U. H. Immune cell migration in inflammation: present and future therapeutic targets. Nat Immunol 6, 1182–1190 (2005).
Waldhauer, I. & Steinle, A. NK cells and cancer immunosurveillance. Oncogene 27, 5932–5943 (2008).
Schumann, K. et al. Immobilized chemokine fields and soluble chemokine gradients cooperatively shape migration patterns of dendritic cells. Immunity 32, 703–713 (2010).
Weber, M. et al. Interstitial dendritic cell guidance by haptotactic chemokine gradients. Science 339, 328–332 (2013).
Poznansky, M. C. et al. Active movement of T cells away from a chemokine. Nat Med 6, 543–548 (2000).
Bernardini, G., Sciume, G. & Santoni, A. Differential chemotactic receptor requirements for NK cell subset trafficking into bone marrow. Front Immunol 4, 12, 10.3389/fimmu.2013.00012 (2013).
Niethammer, P., Grabher, C., Look, A. T. & Mitchison, T. J. A tissue-scale gradient of hydrogen peroxide mediates rapid wound detection in zebrafish. Nature 459, 996–999 (2009).
Feng, Y., Santoriello, C., Mione, M., Hurlstone, A. & Martin, P. Live imaging of innate immune cell sensing of transformed cells in zebrafish larvae: parallels between tumor initiation and wound inflammation. PLoS Biol 8, e1000562, 10.1371/journal.pbio.1000562 (2010).
Turley, S. J., Cremasco, V. & Astarita, J. L. Immunological hallmarks of stromal cells in the tumour microenvironment. Nat Rev Immunol 15, 669–682 (2015).
Gajewski, T. F., Schreiber, H. & Fu, Y. X. Innate and adaptive immune cells in the tumor microenvironment. Nat Immunol 14, 1014–1022 (2013).
Kummerow, C. et al. A simple, economic, time-resolved killing assay. Eur J Immunol 44, 1870–1872 (2014).
Alter, G., Malenfant, J. M. & Altfeld, M. CD107a as a functional marker for the identification of natural killer cell activity. Journal of immunological methods 294, 15–22 (2004).
Matheny, H. E., Deem, T. L. & Cook-Mills, J. M. Lymphocyte migration through monolayers of endothelial cell lines involves VCAM-1 signaling via endothelial cell NADPH oxidase. J Immunol 164, 6550–6559 (2000).
Wittmann, C. et al. Hydrogen peroxide in inflammation: messenger, guide, and assassin. Adv Hematol 2012, 541471, 10.1155/2012/541471 (2012).
Simeoni, L. & Bogeski, I. Redox regulation of T-cell receptor signaling. Biol Chem 396, 555–568 (2015).
Somersalo, K. et al. Activation of natural killer cell migration by leukocyte integrin-binding peptide from intracellular adhesion molecule-2 (ICAM-2). J Biol Chem 270, 8629–8636 (1995).
Somersalo, K., Tarkkanen, J., Patarroyo, M. & Saksela, E. Involvement of beta 2-integrins in the migration of human natural killer cells. J Immunol 149, 590–598 (1992).
Stabile, H. et al. Impaired NK-cell migration in WAS/XLT patients: role of Cdc42/WASp pathway in the control of chemokine-induced beta2 integrin high-affinity state. Blood 115, 2818–2826 (2010).
Naci, D. & Aoudjit, F. Alpha2beta1 integrin promotes T cell survival and migration through the concomitant activation of ERK/Mcl-1 and p38 MAPK pathways. Cell Signal 26, 2008–2015 (2014).
Morrison, V. L. et al. The beta2 integrin-kindlin-3 interaction is essential for T-cell homing but dispensable for T-cell activation in vivo . Blood 122, 1428–1436 (2013).
Kelly, K. A. et al. The combination of the gastrointestinal integrin (alpha4beta7) and selectin ligand enhances T-Cell migration to the reproductive tract during infection with Chlamydia trachomatis. Am J Reprod Immunol 61, 446–452 (2009).
Culley, F. J. et al. Natural killer cell signal integration balances synapse symmetry and migration. PLoS Biol 7, e1000159, 10.1371/journal.pbio.1000159 (2009).
Mittal, M., Siddiqui, M. R., Tran, K., Reddy, S. P. & Malik, A. B. Reactive oxygen species in inflammation and tissue injury. Antioxid Redox Signal 20, 1126–1167 (2014).
Maiuri, P. et al. Actin flows mediate a universal coupling between cell speed and cell persistence. Cell 161, 374–386 (2015).
Saxton, M. J. Lateral diffusion in an archipelago. Distance dependence of the diffusion coefficient. Biophys J 56, 615–622 (1989).
Murphy, M. P. How mitochondria produce reactive oxygen species. Biochem J 417, 1–13 (2009).
Ott, M., Gogvadze, V., Orrenius, S. & Zhivotovsky, B. Mitochondria, oxidative stress and cell death. Apoptosis 12, 913–922 (2007).
Turrens, J. F. Mitochondrial formation of reactive oxygen species. J Physiol 552, 335–344 (2003).
Bedard, K. & Krause, K. H. The NOX family of ROS-generating NADPH oxidases: physiology and pathophysiology. Physiol Rev 87, 245–313 (2007).
Bogeski, I. et al. Redox regulation of calcium ion channels: chemical and physiological aspects. Cell Calcium 50, 407–423 (2011).
Al-Mehdi, A. B., Shuman, H. & Fisher, A. B. Intracellular generation of reactive oxygen species during nonhypoxic lung ischemia. Am J Physiol 272, L294–300 (1997).
Young, T. A., Cunningham, C. C. & Bailey, S. M. Reactive oxygen species production by the mitochondrial respiratory chain in isolated rat hepatocytes and liver mitochondria: studies using myxothiazol. Arch Biochem Biophys 405, 65–72 (2002).
Sedeek, M., Nasrallah, R., Touyz, R. M. & Hebert, R. L. NADPH oxidases, reactive oxygen species, and the kidney: friend and foe. J Am Soc Nephrol 24, 1512–1518 (2013).
Harlin, H. et al. The CD16- CD56(bright) NK cell subset is resistant to reactive oxygen species produced by activated granulocytes and has higher antioxidative capacity than the CD16+ CD56(dim) subset. J Immunol 179, 4513–4519 (2007).
Thoren, F. B., Romero, A. I., Hermodsson, S. & Hellstrand, K. The CD16-/CD56bright subset of NK cells is resistant to oxidant-induced cell death. J Immunol 179, 781–785 (2007).
Woolf, E. et al. Lymph node chemokines promote sustained T lymphocyte motility without triggering stable integrin adhesiveness in the absence of shear forces. Nat Immunol 8, 1076–1085 (2007).
Lammermann, T. et al. Rapid leukocyte migration by integrin-independent flowing and squeezing. Nature 453, 51–55 (2008).
Bogeski, I. et al. Differential redox regulation of ORAI ion channels: a mechanism to tune cellular calcium signaling. Sci Signal 3, ra24, 10.1126/scisignal.2000672 (2010).
Bhat, S. S. et al. Syntaxin 8 is required for efficient lytic granule trafficking in cytotoxic T lymphocytes. Biochim Biophys Acta 1863, 1653–1664 (2016).
Oppelstrup, T. et al. First-passage kinetic Monte Carlo method. Phys Rev E Stat Nonlin Soft Matter Phys 80, 066701, 10.1103/PhysRevE.80.066701 (2009).
van Zon, J. S. & ten Wolde, P. R. Green’s-function reaction dynamics: a particle-based approach for simulating biochemical networks in time and space. J Chem Phys 123, 234910; 10.1063/1.2137716 (2005).
Schwarz, K. & Rieger, H. Efficient kinetic Monte Carlo method for reaction-diffusion problems with spatially varying annihilation rates. J Comput Phys 237, 396–410 (2013).
Acknowledgements
We thank the Institute for Clinical Hemostaseology and Transfusion Medicine for providing donor blood; Carmen Hässig, Cora Hoxha and Gertrud Schwär for excellent technical help, Eckart Meese and Petra Leidinger for microarray measurements, Zhuo Xu for helping the analysis of NK migration, and Adina Vultur for her careful and critical reading. We are grateful to Markus Hoth for constant support, advice regarding writing of the manuscript and for interpretion of the mathematical models. This project was funded by Sonderforschungsbereich 1027 project A2 (to B.Q.), A3 (to H.R.), C3 (to V.H.) and C4 (I.B.). The lightsheet microscope was funded by DFG (GZ: INST 256/4 19-1 FUGG).
Author information
Authors and Affiliations
Contributions
X.Z. performed all real-time killing assays, 2D migration assays, Amplex assays and the corresponding analyses; R.Z. generated all the NK degranulation data and analyzed NK migration in 3D; E.C.S., M.M. and V.H. produced microarray data and the corresponding analysis; I.B. supervised ROS detection experiments; K.S., M.M. and H.R. formulated the mathematical model and performed computer simulations; B.Q. generated concepts, designed experiments, and wrote the manuscript. All authors contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhou, X., Zhao, R., Schwarz, K. et al. Bystander cells enhance NK cytotoxic efficiency by reducing search time. Sci Rep 7, 44357 (2017). https://doi.org/10.1038/srep44357
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep44357
This article is cited by
-
Detecting long-range interactions between migrating cells
Scientific Reports (2021)
-
On the efficiency of chemotactic pursuit - Comparing blind search with temporal and spatial gradient sensing
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.