Abstract
The dopamine D2 receptor (D2R) is involved in food reward and compulsive food intake. The present study developed a virtual screening (VS) method to identify food components, which may modulate D2R signalling. In contrast to their common applications in drug discovery, VS methods are rarely applied for the discovery of bioactive food compounds. Here, databases were created that exclusively contain substances occurring in food and natural sources (about 13,000 different compounds in total) as the basis for combined pharmacophore searching, hit-list clustering and molecular docking into D2R homology models. From 17 compounds finally tested in radioligand assays to determine their binding affinities, seven were classified as hits (hit rate = 41%). Functional properties of the five most active compounds were further examined in β-arrestin recruitment and cAMP inhibition experiments. D2R-promoted G-protein activation was observed for hordenine, a constituent of barley and beer, with approximately identical ligand efficacy as dopamine (76%) and a Ki value of 13 μM. Moreover, hordenine antagonised D2-mediated β-arrestin recruitment indicating functional selectivity. Application of our databases provides new perspectives for the discovery of bioactive food constituents using VS methods. Based on its presence in beer, we suggest that hordenine significantly contributes to mood-elevating effects of beer.
Similar content being viewed by others
Introduction
Homeostatic food intake is regulated mainly by the hypothalamus and caudal brainstem by integration of various peripheral and central signals resulting in an ingestive behaviour which counterbalances the energy expenditure1. However, it is also well established that certain food stimuli induce hedonic food intake in the state of satiety, leading to an overconsumption of calories and, thus, eventually to obesity. Dopaminergic pathways are heavily involved in hedonic food intake by mediating reward, motivation and reinforcement2. Among the five dopamine receptor subtypes, in particular the dopamine D2 receptor (D2R) seems to be involved in food reward and compulsive food intake2,3. However, the molecular determinants of palatable food inducing non-homeostatic food intake are still not fully understood. Mixtures of carbohydrates and fat most efficiently induce hyperphagia in rats4,5,6, with a carbohydrate/fat ratio of about 45%: 35% having the most pronounced effect7. Although being very effective in inducing food intake, fat/carbohydrate mixtures have a lower impact on brain reward areas compared to palatable food items with the same fat/carbohydrate composition7,8. Thus, bioactive food components with the potential to modulate dopaminergic pathways may be able to change the rewarding properties of food. To date, only little is known about dopaminergic food components. Therefore, the aim of the present study was the identification of novel food-derived D2R ligands.
For the discovery of novel bioactive food components, food extracts or food compounds are either selected hypothesis-driven or even arbitrary and then tested by bioassays addressing the desired physiological effect. Subsequently, active food extracts are subjected to activity-guided fractionation to identify the bioactive components9. This approach, however, is rather time-consuming, has a low hit rate and tends to overlook promising novel food bioactives. Furthermore, only a limited number of food components are commercially available for testing or easily accessible. Therefore, in silico screening methods would be useful for prioritising the food compounds to be submitted to biological assays.
Virtual screening (VS) refers to the use of computational methods for the knowledge-based identification of compounds that exhibit a desired biological activity10. In drug-discovery research, this technique has been developed as a response to stagnating high-throughput screening (HTS) hit rates in combination with rising costs for HTS assays. Nowadays, VS methods are a common complementary technique to HTS to analyse large compound databases in order to prioritise a set of molecules for further experimental testing11,12,13. However, no VS applications in food science have been reported in the literature, possibly because of the lack of specific food-compound databases, since pharmaceutical companies focus on drug-like compounds and natural products. In the present study, we developed a VS protocol consisting of pharmacophore screening, hit-list clustering and molecular docking to search for possible D2R-ligands in a newly assembled database containing 13,000 components from foods and some other natural sources. The most promising compounds were evaluated in biological assays. This approach allowed for the first time the unbiased identification of novel food bioactives by virtual screening of a novel food-compound database.
Results and Discussion
Molecular properties of FCDB and PhyDB compared to other VS databases
Our first aim was the generation of an in silico 3D food-compound database. Besides this database (FCDB; 12,579 compounds), which we constructed by selecting natural food constituents from the Dictionary of Food Compounds14, we also generated a small natural products database (PhyDB; 987 compounds) based on the catalogue from the vendor PhytoLab Vestenbergsgreuth, Germany (available at http://www.phytolab.com/de/phytolab.html). The databases are part of the Supplementary information.
For the characterisation of our newly generated databases, we calculated molecular property distributions for FCDB and PhyDB and for samples containing 10,000 randomly selected compounds from the UNPD database15, the ZINC biogenic compounds subset (ZBC) and the ZINC all purchasable subset (ZAP)16. Thus, we could compare our databases with established freely available VS databases that contain natural products (UNPD database and ZBC) and drug-like compounds (ZAP). Detailed data are available in the Supplementary information, Fig. S1.
The calculations revealed that the FCDB and PhyDB databases are substantially different to a typical drug-like library (ZAP). While the compounds in drug-like libraries tend to comply with Lipinski’s rule-of-five17 resulting in Gaussian-like molecular property distributions with distinct maxima, the molecules in FCDB, PhyDB, and UNPD exhibit very broad molecular-property distributions. Hence, the compounds in these databases tend to have higher structural diversity, which is typical for natural compound libraries18. Despite their distinct similarity, differences between FCDB/PhyDB and UNPD still exist, especially in terms of molecular weight. The natural product library ZBC turned out rather to possess drug-like than natural-product-like properties, probably because the authors of ZINC (http://zinc.docking.org/subsets/zbc) took their information about the natural character of a compound from vendor catalogues. The vendors often also include synthetic derivatives of natural products in their catalogues to make them more attractive for medicinal chemists, as also observed by Manallack et al.19. In summary, it turned out that our generated screening databases represent a new type of screening library, which is much more similar to databases containing natural products than drug-like compounds.
Virtual screening process
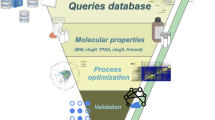
The next goal was the search for nutritive or natural D2R-ligands in the generated VS databases. D2R is a target of great interest for the pharmaceutical industry, for which agonists have been developed as drugs for treating Parkinson’s disease and antagonists for treating schizophrenia20,21. Although no X-ray structure of D2R has been published yet, a wide variety of ligands has been synthesised that facilitates the effective use of ligand-based VS methods such as pharmacophore searching11. The GPCR Ligand Library22 provides a comprehensive collection of known D2R-ligands compiled from the GLIDA database23. However, since the majority of D2R-ligands are synthetic in origin, the ligand collections contain primarily drug-like compounds, in contrast to those contained in FCDB and PhyDB. Therefore, the selected VS methods needed to be capable of finding VS hits in FCDB and PhyDB that are structurally different to the input ligands. Therefore, we did not only apply VS methods that rely on molecular similarity but rather used pharmacophore searching in combination with hit-list clustering and molecular docking. The overall VS workflow is summarised in Fig. 1.
The numbers on the left and on the right side represent the number of compounds that passed through the respective VS step depicted in the centre. Due to 230 compounds that are present in both FCDB and PhyDB, the absolute number of different compounds is smaller than the sum of the compounds in the two databases.
Pharmacophore modelling
By selecting relatively large and structurally diverse training sets for both D2R-agonists and -antagonists, we ensured that the resulting pharmacophore models only retain the most critical features for binding. The training sets are depicted in the Supplementary information, Figs S2 and S3. Training sets containing only large and structurally similar compounds would probably have resulted in very specific pharmacophore models with a large number of features, which would not be able to find any hits in FCDB and PhyDB. Additionally, we did not consider excluded volumes in the pharmacophore models to avoid obtaining small hit lists.
It is a prerequisite for an alignment of the ligands to be a good approximation to their binding mode that the template compound for the alignment provides an orientation close to its bioactive conformer. In addition to molecular docking and subsequent molecular-dynamics refinement of the ligand conformers as techniques for approaching the bioactive conformation, we selected some ligands, namely # 21 and # 27 from the agonist training set, Fig. S2, and # 32 from the antagonist training set, Fig. S3. We used the conformers that are stored in the GPCR Ligand Library directly, because the conformational space in low-energy regions is sparsely occupied for structurally restrained ligands. The bioactive conformer has been shown to be often located near the global minimum24 and a low-energy conformer for a restrained ligand can thus be a good approximation. The training set ligand conformers that possess the maximum similarity compared to the template were then determined by ParaAlign (see Methods and Supplementary information). For each ligand, this conformer was extracted and used as input for the HipHop25 pharmacophore model generation algorithm. Since we generated up to 25 pharmacophore models per run, the variation of the conformer types and the templates that were used for ParaAlign resulted in a large number of models, which had to be validated subsequently.
The validation process involved large test databases compiled from the GPCR Ligand Library and GPCR Decoy Database22 in order to determine the overall best pharmacophore models. We found two very similar pharmacophore models with almost identical excellent VS performance in model validation for the D2R-agonists. Both models contained hydrogen-bond donor, aromatic ring and positive ionisable features. We therefore decided to use only the model that gave a slightly better receiver operating characteristic (ROC) curve (Fig. 2b).
The overall best D2R-agonist pharmacophore and its inter-feature distances (a), depicted together with its ROC curve that results from VS of the test database (b). The features are hydrogen bond donor (purple, HBD), aromatic ring (orange, R) and positive ionisable (red, P). The dashed red line in the ROC plot indicates a random selection.
A comparison between this D2R-agonist pharmacophore model (Fig. 2a) and models reported in the literature revealed a high degree of similarity. Both Malo et al.26 and Chidester et al.27 reported D2R-agonist pharmacophore models that also contain hydrogen-bond donor, aromatic ring and positive ionisable features with similar relative orientations. The only difference is an additional hydrophobic feature close to the positive ionisable feature present in their models. However, since not all our training set ligands contain such a feature, we conclude that the hydrophobic feature is able to increase binding affinity but that it is not critical for binding.
We also obtained two pharmacophore models for the D2R-antagonists that exhibited the best VS performance in the validation step (Fig. 3c). In contrast to the agonists, these two models did not show a high degree of similarity concerning shape and features (Fig. 3a,b). While the first D2R-antagonist model contained a hydrogen-bond acceptor, an aromatic ring and a positive ionisable feature, the other model possessed a hydrophobic feature together with the aromatic ring and positive ionisable feature. The relative orientation of the features was also different. However, this observation is not surprising, since our D2R-antagonist training set covered ligands with considerably different shapes (e.g. ligands # 35 and 39, Fig. S3). Such diverse ligands probably adopt different binding modes and, hence, it is not surprising that we obtained two diverse pharmacophore models.
The overall best D2R-antagonist pharmacophore models and their inter-feature distances (a and b) depicted together with the corresponding ROC curves that result from VS of the test database (c). The features are hydrogen bond acceptor (green, HBA), aromatic ring (orange, R), positive ionisable (red, P) and hydrophobic (blue, H). The dashed red line in the ROC plot indicates a random selection. The blue curve corresponds to model a) and the green curve to model b).
The models further resemble a ligand classification for pharmacophore model generation reported by Klabunde and Evers28. They divided their D2R-antagonist training set into two different classes and built different pharmacophore models from the two subsets. Our first D2R-antagonist pharmacophore model (Fig. 3a) resembles their class I model, whereas our second model (Fig. 3b) shows some similarity to their class II model. As our validation database contains a substantial number of actives from both ligand classes, similar VS performance of the pharmacophore models was not surprising. The validation performance is not as good as for the D2R-agonist models, probably because, on the one hand, either model is recognising class I-like actives or class II-like actives better and, on the other hand, hydrophobic and hydrogen-bond acceptor features are not as selective as a hydrogen-bond donor feature. For this study, we decided to use both models for database searching in FCDB and PhyDB and, subsequently, to combine the two hit lists to avoid missing any type of antagonist.
The 3D database-searching step using the selected D2R-agonist pharmacophore model selected 1,441 compounds from FCDB and PhyDB. In contrast, the two selected D2R-antagonist pharmacophore models gave a combined hit list with 1,866 compounds. Note that we treated tautomeric compounds and different ionisation states as different database entries in this step, which led to a lower number of chemically different hit compounds.
Hit-list clustering and compound prioritisation
The next goal of our study was to select structurally diverse compounds for experimental testing. Therefore, we performed a clustering analysis of the hit lists in Discovery Studio 3.1 (Accelrys Inc., San Diego, CA) to divide them into groups that contain structurally different classes (Fig. 1). Moreover, the possible corresponding tautomers and ionisation states for a given hit compound could be pooled as they were assigned to the same compound cluster. After variation of the number of generated clusters, we found that values of 80 clusters for the D2R-agonist hits and 100 clusters for the D2R-antagonist hits represent good values for characterising the hit-list compounds.
Selecting compounds from different clusters ensured that we chose structurally diverse molecules for the following molecular-docking step. The choice was further based on the fit value to the pharmacophore models and on visual inspection. For both D2R-agonist and -antagonist screening, we derived a cut-off limit for the fit value from the pharmacophore validation step, in which databases consisting of known actives and putative inactives were mapped to the models. From the actives mapped to the D2R-agonist model, all but one gave a fit value higher than 2.2. Hence, we used this value as a cut-off limit in compound prioritisation. By contrast, most of the actives mapped to the D2R-antagonist models showed fit values of 1.9 and 2.3 for model A and model B, respectively. We retained pharmacophore searching hits with larger fit values than these cut-off limits for at least one of the models.
Hit-list clustering and compound prioritisation reduced the number of VS hits to 125 and 69 different compounds for the D2R-agonist and -antagonist screening, respectively. Thus, we selected less compounds from the D2R-antagonist screening for the molecular docking step, although the pharmacophore hit list and the number of clusters was larger than for the agonists. However, many compounds in the antagonist hit-list did not pass the fit-value cut-off. Additionally, visual inspection of the hit-list molecules revealed that large and bulky compounds tended to have very high fit values. These molecules were rejected because they are unlikely to fit into the binding pocket.
Molecular docking and final compound selection
The combination of pharmacophore searching with molecular docking enabled an initial check of our database molecules for the required molecular features and their relative orientations. Then we examined more thoroughly, whether the candidates could fit well into the binding pocket and establish important protein-ligand interactions. A D2R inactive-state homology model used successfully in a previous docking study29 served as docking target. The compounds that passed the clustering and prioritisation step were docked together with their corresponding tautomers and ionisation states using AutoDockVina30 with high exhaustiveness and maximum conformer values for detailed examination. For each compound, we carefully inspected both the value of the scoring function and the resulting receptor-ligand complexes. An interaction between a positively charged moiety and the carboxylate group of Asp1143.32 was mandatory for a compound to be regarded as hit in the docking step.
Eventually, we obtained 53 hit compounds and searched for corresponding possible activity data concerning dopamine receptors in the literature. We excluded hits with known bioactivity at D2R and checked the commercial availability of the remaining compounds. Finally, 17 compounds of different origin (Fig. 4a,b, Table 1) were purchased and prepared for experimental testing.
Structures of the experimentally tested D2R-agonist (a) and -antagonist (b) VS hits. The 17 compounds can be classified into natural food constituents (delphinidin, hordenine, kukoamine A, pyrraline, salsolinol, roquefortine C), further compounds from natural sources (fumigaclavine A, leonurine, muscimol, ajmalicine, dihydroberberine, emetine), and food contaminants (clenbuterol, fenpropimorph, halofuginone, robenidine, sarafloxacin).
We also examined, whether a 2D molecular similarity search would have been sufficient to extract these molecules from the VS databases. Therefore, we calculated Extended Connectivity Fingerprint 4 (ECFP4)-based Tanimoto coefficients (Tc) for the nine D2R-agonist screening hits and the eight D2R-antagonist screening hits compared to the respective training set ligands. For the D2R-agonist hits, the Tc values varied from 0.094 (pyrraline) to 0.324 (salsolinol), while we obtained Tc values between 0.110 (fenpropimorph) and 0.345 (ajmalicine) for the D2R-antagonist hits. When comparing all our screening database compounds to the training set ligands, the top 100 hits using the D2R-agonist training set as a reference gave Tc values of at least 0.234, while the Tc values of the top 100 hits using the D2R-antagonist training set as a reference were at least 0.243. Thus, when applying the 2D molecular similarity search, only three of our selected compounds, ajmalicine (Tc = 0.345), salsolinol (Tc = 0.324) and fumigaclavine A (Tc = 0.288) would have been present among the top-ranked compounds. For details, please refer to the Supplementary information, Table S1.
Experimental evaluation
To confirm the VS predictions, we conducted radioligand binding assays to estimate the binding affinity of our hit compounds at D2R. As we performed VS to identify both nutritive D2R-agonists and -antagonists, we were interested in the ability of the VS hits to activate the receptor or to antagonise the effects of dopamine. Hence, functional assays were carried out to evaluate both G protein- and β-arrestin-mediated signalling pathways.
Ligand binding experiments
The affinity profiles of our VS hits towards the human dopamine D2L receptor isoform were compared with those of the reference compound dopamine by radioligand binding experiments (Table 1).
The binding data revealed Ki values ≤ 50 μM for five of nine tested D2R-agonist VS hits, representing a hit rate of 56%. Two out of eight tested D2R-antagonist VS hits gave Ki values ≤ 50 μM corresponding to a 25% hit rate. All observed binding affinities were in the micromolar range, which is typical for active compounds found by VS31. Fumigaclavine A, an alkaloid with an ergoline-like structure produced by different Aspergillus species32, showed the highest binding affinity with a Ki value that was only seven times higher than for the endogenous ligand dopamine. Furthermore, the coccidiostat and possible food contaminant robenidine33, as well as salsolinol and hordenine bind in the low micromolar range. Salsolinol, for which controversial activity profiles for D2 receptors have been reported34,35,36, is a constituent of cocoa and chocolate37, whereas hordenine is present in barley38 and beer39. The anthocyanidin delphinidin, which has a considerably different molecular scaffold to the remaining VS hits, gave a Ki value of 26 μM. Most active compounds contain an aliphatic amine as a positively charged moiety at physiological pH to enable a salt bridge to Asp1143.32. Only robenidine, which contains a guanidinium group, and delphinidin, which features cationic oxygen, are different.
Functional experiments
Finally, we examined the ability of our VS workflow not only to identify compounds that bind to D2R, but also to predict the compound efficacy. Hence, we performed functional experiments for the five compounds that showed the best binding affinities, namely fumigaclavine A, robenidine, salsolinol, hordenine, and delphinidin. As it has been shown that ligands can induce G protein-mediated signalling and β-arrestin recruitment independently40,41, the individual pathways were investigated separately (Table 2).
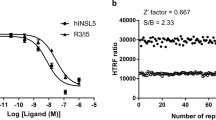
Among the five investigated VS hits, three compounds showed significant intrinsic activity in the cAMP accumulation assay. The EC50 values of fumigaclavine A and hordenine are both in the extended range of the Ki values determined in the radioligand binding experiments. Conversely, probably because of a very flat dose-response curve, the EC50 value of salsolinol is considerably lower than its Ki value. For hordenine, we could observe almost full activation (Emax = 76%) compared to the reference compound quinpirole and to dopamine (Fig. 5a). This is particularly interesting, because hordenine did not show significant β-arrestin recruitment (Fig. 5b). In fact, titration curves using a constant concentration of 300 nM of the D2R-agonist quinpirole with hordenine confirm its antagonistic activity (Fig. 5c). Fumigaclavine A and salsolinol also did not show β-arrestin recruitment, but the maximum response of these compounds in the cAMP accumulation assay was well below 50%, categorising them as weak partial agonists.
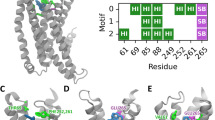
Although the G protein-biased D2R-agonist hordenine shares structural similarities to the balanced agonist dopamine, receptor–ligand interactions obtained after docking and energy minimisation in presence of a D2R homology model appear to be different. Hordenine lacks a meta-hydroxyl group compared to dopamine, rendering it unable to form hydrogen bonds to both residues Ser1935.42 and Ser1975.46 as dopamine does42 (Fig. 6). The receptor–ligand complex that results from docking and energy minimisation cannot explain whether double methylation of the nitrogen atom enables hordenine to form additional hydrophobic interactions compared to dopamine.
We used an active-state homology model of D2R and performed MD simulations with the endogenous ligand dopamine42. Dopamine was removed from the model, hordenine was docked into the binding pocket and the resulting receptor-ligand complex was subjected to energy minimisation in a water box. Whereas dopamine is able to form two hydrogen bonds with both Ser1935.42 and Ser1975.46 in the D2UpR model42, our VS hit hordenine forms only a single hydrogen bond to Ser1975.46 due to the lack of a second hydroxyl group.
Taken together, we could observe partial or almost full agonist activity in the cAMP inhibition assays for three of four tested D2R-agonist VS hits. For the D2R-antagonist VS hit robenidine, we could not observe any activation in either assay. These results confirm the pharmacophore models and VS workflow.
Thus, the VS workflow proved to be able to identify hordenine as almost full D2R-agonist in Gi/o activation and simultaneous antagonism in β-arrestin recruitment, indicating that this substance acts as a G protein-biased agonist at D2R. Additionally, two VS hits with partial agonism in Gi/o activation, fumigaclavine A and salsolinol, were identified. The D2R-antagonist robenidine did not show any activation in either assay. The VS method based on the newly generated VS food database can now be applied to other targets for unbiased identification of novel bioactive compounds in food.
Hordenine is a natural constituent of barley and, in particular, beer, a food that is often linked to alcohol abuse. In the present study, hordenine was able to bind to D2R with a Ki value in the low micromolar range. The activation profile of hordenine showed significant bias for G protein-promoted activation over β-arrestin recruitment compared to functionally balanced D2R-agonists like the endogenous ligand dopamine. Given that β-arrestin recruitment leads to desensitisation and receptor internalisation, hordenine-promoted D2R activation may be stronger and more sustainable than activation with the balanced neurotransmitter dopamine. Further studies are necessary to clarify the possible influence of hordenine on the dopaminergic system and food reward.
Methods
Database generation
From the 40,000 entries (natural food constituents, food contaminants, food additives, nutraceuticals) in the Dictionary of Food Compounds14 only those molecules were selected for FCDB that were reported in the literature to occur in food. We excluded compounds that were heavier than 750 Da or possessed more than two sugar moieties, because they are unlikely to be resorbed in the gastrointestinal tract. The Supplementary information includes further details on the database generation procedure.
Molecular property calculations
For comparison of FCDB and PhyDB with other established database types, we selected the UNPD database15, the ZINC biogenic compounds (ZBC), and the ZINC all purchasable (ZAP) subset from the ZINC database16. From these three databases, we extracted a sample of 10,000 random compounds each. By contrast, FCDB and PhyDB were used in their entirety for the calculation of molecular properties. The molecular properties (molecular weight, AlogP, numbers of rotatable bonds, hydrogen bond donors and acceptors) were calculated in Discovery Studio 3.1.
Pharmacophore modelling
For pharmacophore model generation, we selected structurally diverse training sets containing 11 D2R-agonists and 12 D2R-antagonists collected from the GPCR Ligand Library22. To obtain low-energy conformers for the training set ligands, we generated conformational sets with up to 255 conformers using the Catalyst BEST conformer generation algorithm (once including and once without a conformer minimisation step using the CHARMm force field implemented in Discovery Studio 3.1.) The generated ligand conformers were aligned to a template compound, which is assumedly present in its bioactive conformation.
To obtain template conformers for the rigid-body alignment algorithm ParaAlign (detailed description of ParaAlign is given in the Supplementary information), we submitted each training set ligand to a molecular docking step into an in-house D2R homology model using AutoDock Vina30 followed by molecular dynamics refinement of the ligand conformers in the receptor-ligand complex using AMBER10 programme package (University of California). The refined docking conformation of each training set ligand was subsequently used as template for ParaAlign. Because our training sets also contained structurally restrained ligands, we additionally used the ligand conformers for these compounds that are stored in the GPCR Ligand Library directly as templates for ParaAlign. Two agonists (# 21 and 27, Fig. S2) and one antagonist (# 32, Fig. S3) underwent this procedure. Using the ParaAlign algorithm, we could then determine and extract the respective training set conformer for each alignment that possessed the highest similarity to the template conformer. Subsequently, pharmacophore models were generated for those training set conformers using the Common Feature Pharmacophore (HipHop) algorithm in Discovery Studio 3.1. The applied parameters are given in the Supplementary information.
In a validation step consisting of a database search with test databases that contain a large number (136 agonists, 493 antagonists, and 39 decoys for each) of chemically diverse ligands and suitable corresponding decoys compiled from the GPCR Ligand Library and GPCR Decoy Database, we evaluated all generated pharmacophore models. The overall best D2R-agonist and -antagonist pharmacophore models were extracted for the pharmacophore screening in FCDB and PhyDB using the best flexible 3D database search in Discovery Studio 3.1.
Hit-list preparation and molecular docking
The hit lists resulting from the pharmacophore searches were merged and duplicate structures were removed. In the following, we applied the ligand-clustering tool in Discovery Studio 3.1 using MDL public key fingerprints as clustering properties to divide the hit lists into groups of structurally similar molecules. Thus, a total number of 80 clusters for the D2R-agonist and 100 clusters for the D2R-antagonist screening hits were generated.
To obtain structurally diverse screening hits for the subsequent molecular docking step, we picked out single cluster representatives based on the pharmacophore-model fit values and on visual inspection. For chiral compounds, we checked if the configuration generated by CORINA corresponded to the natural stereoisomer. If this was not the case, we produced the natural stereoisomer in ChemDraw 12.0, converted it to a SMILES string and to a 3D conformer by CORINA and submitted it to the ligand-preparation steps previously applied to all database compounds. Ligand conformers were generated and mapped to the pharmacophore models to check, if a match was still present.
Subsequently, the selected pharmacophore hits were docked into an in-house D2R-homology model previously used in molecular docking studies29 using AutoDock Vina30. We applied a search space of 26 × 24 × 24 Å and an exhaustiveness value of 32 and obtained up to 20 conformations of each ligand. For the final selection of screening hits to be tested in vitro, the resulting protein-ligand complexes were subjected to an accurate visual inspection. We retained only poses, where the most important receptor-ligand interactions such as the conserved salt bridge between Asp1143.32 and a positively charged moiety were present. After checking the final hit lists for commercial availability, 17 compounds were purchased. The VS hit hordenine was further docked into a D2R-homology model used in molecular-dynamics studies with the endogenous ligand dopamine42. The docking procedure is described in detail in the Supporting information.
VS hit compounds
Ajmalicine, delphinidine chloride, dihydroberberine, emetine dihydrochloride, hordenine, leonurine, and muscimol were purchased from PhytoLab GmbH (Vestenbergsgreuth, Germany). Clenbuterol hydrochloride, fenpropimorph, halofuginone hydrobromide, robenidine hydrochloride, and sarafloxacin hydrochloride hydrate were obtained from Sigma Aldrich (Taufkirchen, Germany). Fumigaclavine A was purchased from AdipoGen AG (Liestal, Switzerland), and kukoamine A from AChemTek, Inc. (Worcester, MA, USA). Pyrraline was provided by PolyPeptide Group (Strasbourg, France). Roquefortine C was obtained from Cfm Oskar Tropitzsch GmbH (Marktredwitz, Germany) and salsolinol hydrochloride from ABCAM biochemical (Cambridge, U.K.).
Radioligand binding assays
Receptor binding studies were carried out as described previously43,44. In brief, preparations of membranes from CHO cells that stably express the human D2L receptor were used together with [3H]spiperone (specific activity 73 Ci/mmol, Biotrend, Cologne, Germany) at a final concentration of 0.15 nM. The assays were carried out at a protein concentration of 6 μg/assay tube, showing KD values of 0.052–0.10 nM and corresponding Bmax values of 950–1500 fmol/mg.
cAMP Inhibition assay
HEK293T cells were transiently co-transfected with pcDNA3L-His-CAMYEL45 (purchased from ATCC, Manassas, VA via LGC Standards, Wesel, Germany) and D2s receptor, respectively, and the assay was performed according to literature46. Twenty-four hours after transfection cells were split into white half-area 96-well plates at 2 × 105 cells/well and grown overnight in a phenol-red free medium supplemented with serum (Invitrogen, Darmstadt, Germany). On the day of measurement, the medium was removed and replaced by phosphate buffered saline (PBS). The cells were serum-starved for 1 h before treatment. The assay was started by adding 10 μL of coelenterazine-h (Promega, Mannheim, Germany) to each well to yield a final concentration of 5 μM. After 5 min of incubation, test compounds were added in PBS containing forskolin at a final concentration of 10 μM. After an incubation time of 15 min, BRET measurement was performed with a CLARIOstar plate reader (BMG LabTech, Ortenberg, Germany). Emission signals from Renilla luciferase and YFP were measured simultaneously using a BRET1 filter set (475–30 nm/535–30 nm). BRET ratios (emission at 535 ± 30 nm/emission at 475 ± 30 nm) were calculated and dose-response curves were fitted by nonlinear regression analysis using the algorithm of PRISM 6.0. Curves were normalised to basal BRET ratio obtained from dPBS (0%) and the maximum effect of the reference ligand quinpirole (100%).
β-Arrestin recruitment assay
The measurement of β-arrestin-2 recruitment stimulated by D2R activation was performed using the PathHunter® assay purchased from DiscoveRx (Birmingham, U.K.) according to the manufacturer’s protocol as described previously47. To determine the antagonist properties, different concentrations of the test compounds (10−10–10−3 M) were pre-incubated for 30 min before adding 300 nM (final concentration) quinpirole to start the standard incubation period.
Data analysis
The resulting competition curves of the receptor binding experiments were analysed by nonlinear regression using the algorithms in PRISM 6.0 (GraphPad Software, San Diego, CA). The data were initially fit using a sigmoid model to provide an IC50 value, representing the concentration corresponding to 50% of maximal inhibition. IC50 values were transformed to Ki values according to the equation of Cheng and Prusoff48.
Data from cAMP measurements were analysed by normalising the BRET ratios with 0% for the unstimulated receptor and 100% for the full effect of the reference ligand quinpirole. Dose-response curves were calculated by nonlinear regression using the algorithms of PRISM 6.0.
The amount of recruitment of β-arrestin was derived from the agonist-induced increase of chemiluminescence, which was expressed in counts per second (cps). Dose-response curves were normalised to basal cps stimulated by buffer (=0%) and the effect of the maximum effect of the reference compound quinpirole (=100%).
From each curve, a set of mean values was derived and pooled to result in an average curve showing the EC50 and Emax value, respectively.
Additional Information
How to cite this article: Sommer, T. et al. Identification of the Beer Component Hordenine as Food-Derived Dopamine D2 Receptor Agonist by Virtual Screening a 3D Compound Database. Sci. Rep. 7, 44201; doi: 10.1038/srep44201 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Lenard, N. R. & Berthoud, H. R. Central and peripheral regulation of food intake and physical activity: Pathways and genes. Obesity 16, 11–22 (2008).
Wang, G. J., Volkow, N. D., Thanos, P. K. & Fowler, J. S. Imaging of brain dopamine pathways implications for understanding obesity. J. Addict. Med. 3, 8–18 (2009).
Baik, J. H. Dopamine signaling in food addiction: role of dopamine D2 receptors. Bmb Rep. 46, 519–526 (2013).
Lucas, F. & Sclafani, A. Hyperphagia in rats produced by a mixture of fat and sugar. Physiol. Behav. 47, 51–55 (1990).
Ramirez, I. & Friedman, M. I. Dietary hyperphagia in rats: Role of fat, carbohydrate, and energy content. Physiol. Behav. 47, 1157–1163 (1990).
Hoch, T., Pischetsrieder, M. & Hess, A. Snack food intake in ad libitum fed rats is triggered by the combination of fat and carbohydrates. Front. Psychol. 5, 250 (2014).
Hoch, T., Kreitz, S., Gaffling, S., Pischetsrieder, M. & Hess, A. Fat/carbohydrate ratio but not energy density determines snack food intake and activates brain reward areas. Sci. Rep. 5, 10041 (2015).
Hoch, T., Kreitz, S., Gaffling, S., Pischetsrieder, M. & Hess, A. Manganese-enhanced magnetic resonance imaging for mapping of whole brain activity patterns associated with the intake of snack food in ad libitum fed rats. Plos One 8, e55354 (2013).
Schwager, J., Mohajeri, M. H., Fowler, A. & Weber, P. Challenges in discovering bioactives for the food industry. Curr. Opin. Biotech. 19, 66–72 (2008).
Koeppen, H., Kriegl, J., Lessel, U., Tautermann, C. S. & Wellenzohn, B. Ligand-based virtual screening In Virtual Screening: Principles, Challenges, and Practical Guidelines Methods and Principles in Medicinal Chemistry (ed C. Sotriffer ) 61–85 (Wiley-VCH, 2011).
Walters, W. P., Stahl, M. T. & Murcko, M. A. Virtual screening - an overview. Drug Discov. Today 3, 160–178 (1998).
Cummings, M. D. et al. Preparing and filtering compound databases for virtual and experimental screening In Virtual Screening: Principles, Challenges, and Practical Guidelines Methods and Principles in Medicinal Chemistry (ed C. Sotriffer ) 35–59 (Wiley-VCH, 2011).
Shoichet, B. K. Virtual screening of chemical libraries. Nature 432, 862–865 (2004).
Yannai, S. Dictionary of Food Compounds with CD-ROM. 2nd edn, (CRC Press, 2012).
Gu, J. Y. et al. Use of natural products as chemical library for drug discovery and network pharmacology. Plos One 8, e62839 (2013).
Irwin, J. J., Sterling, T., Mysinger, M. M., Bolstad, E. S. & Coleman, R. G. ZINC: A free tool to discover chemistry for biology. J. Chem. Inf. Model. 52, 1757–1768 (2012).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliver. Rev. 23, 3–25 (1997).
Rollinger, J., Stuppner, H. & Langer, T. Virtual screening for the discovery of bioactive natural products In Natural Compounds as Drugs Volume I Vol. 65 Progress in Drug Research (eds Frank Petersen & René Amstutz ) 211–249 (Birkhäuser, 2008).
Manallack, D. T. et al. The acid/base profile of the human metabolome and natural products. Mol. Inform. 32, 505–515 (2013).
Hisahara, S. & Shimohama, S. Dopamine receptors and Parkinson’s disease. Int. J. Med. Chem. 403039 (2011).
Seeman, P. & Vantol, H. H. M. Dopamine-receptor pharmacology. Trends Pharmacol. Sci. 15, 264–270 (1994).
Gatica, E. A. & Cavasotto, C. N. Ligand and decoy sets for docking to G protein-coupled receptors. J. Chem. Inf. Model. 52, 1–6 (2012).
Okuno, Y. et al. GLIDA: GPCR--ligand database for chemical genomics drug discovery–database and tools update. Nucleic Acids Res. 36, D907–912 (2008).
Kirchmair, J., Laggner, C., Wolber, G. & Langer, T. Comparative analysis of protein-bound ligand conformations with respect to catalyst’s conformational space subsampling algorithms. J. Chem. Inf. Model. 45, 422–430 (2005).
Barnum, D., Greene, J., Smellie, A. & Sprague, P. Identification of common functional configurations among molecules. J. Chem. Inf. Comp. Sci. 36, 563–571 (1996).
Malo, M., Brive, L., Luthman, K. & Svensson, P. Selective pharmacophore models of dopamine D(1) and D(2) full agonists based on extended pharmacophore features. ChemMedChem 5, 232–246 (2010).
Chidester, C. G., Lin, C. H., Lahti, R. A., Haadsma-Svensson, S. R. & Smith, M. W. Comparison of 5-HT1A and dopamine D2 pharmacophores. X-ray structures and affinities of conformationally constrained ligands. J. Med. Chem. 36, 1301–1315 (1993).
Klabunde, T. & Evers, A. GPCR antitarget modeling: pharmacophore models for biogenic amine binding GPCRs to avoid GPCR-mediated side effects. Chembiochem 6, 876–889 (2005).
Hiller, C. et al. Functionally selective dopamine D-2/D-3 receptor agonists comprising an enyne moiety. J. Med. Chem. 56, 5130–5141 (2013).
Trott, O. & Olson, A. J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Ripphausen, P., Nisius, B., Peltason, L. & Bajorath, J. Quo vadis, virtual screening? A comprehensive survey of prospective applications. J. Med. Chem. 53, 8461–8467 (2010).
Spilsbury, J. F. & Wilkinson, S. The isolation of festuclavine and two new clavine alkaloids from Aspergillus fumigatus Fres. J. Chem. Soc. 2085–2091 (1961).
Zulalian, J., Champagne, D. A., Wayne, R. S. & Blinn, R. C. Absorption, excretion, and metabolism of 1,3-bis(p-chlorobenzylideneamino) guanidine hydrochloride (Robenz robenidine hydrochloride) in the chicken. J. Agric. Food Chem. 23, 724–730 (1975).
Britton, D. R., Rivier, C., Shier, T., Bloom, F. & Vale, W. In vivo and in vitro effects of tetrahydroisoquinolines and other alkaloids on rat pituitary function. Biochem. Pharmacol. 31, 1205–1211 (1982).
Antkiewicz-Michaluk, L., Michaluk, J., Romanska, I., Papla, I. & Vetulani, J. Antidopaminergic effects of 1,2,3,4-tetrahydroisoquinoline and salsolinol. J. Neural Transm. 107, 1009–1019 (2000).
Melzig, M. F., Putscher, I., Henklein, P. & Haber, H. In vitro pharmacological activity of the tetrahydroisoquinoline salsolinol present in products from Theobroma cacao L. like cocoa and chocolate. J. Ethnopharmacol. 73, 153–159 (2000).
Riggin, R. M. & Kissinger, P. T. Identification of salsolinol as a phenolic component in powdered cocoa and cocoa-based products. J. Agric. Food Chem. 24, 900 (1976).
Leger, E. Hordenine: A new alkaloid extracted from the germ of barley. J. Pharm. Chim. 23, 177–181 (1906).
Brauers, G., Steiner, I. & Daldrup, T. Quantification of the biogenic phenethylamine alkaloid hordenine by LC-MS/MS in beer. Toxichem Krimtech 80, 323–326 (2013).
Violin, J. D. & Lefkowitz, R. J. β-Arrestin-biased ligands at seven-transmembrane receptors. Trends Pharmacol. Sci. 28, 416–422 (2007).
Shonberg, J. et al. Biased agonism at G protein-coupled receptors: The promise and the challenges—a medicinal chemistry perspective. Med. Res. Rev. 34, 1286–1330 (2014).
Kling, R. C., Lanig, H., Clark, T. & Gmeiner, P. Active-state models of ternary GPCR complexes: Determinants of selective receptor-G-protein coupling. Plos One 8, e67244 (2013).
Tschammer, N. et al. Highly potent 5-aminotetrahydropyrazolopyridines: Enantioselective dopamine D3 receptor binding, functional selectivity, and analysis of receptor−ligand interactions. J. Med. Chem. 54, 2477–2491 (2011).
Hübner, H., Haubmann, C., Utz, W. & Gmeiner, P. Conjugated enynes as nonaromatic catechol bioisosteres: Synthesis, binding experiments, and computational studies of novel dopamine receptor agonists recognizing preferentially the D3 subtype. J. Med. Chem. 43, 756–762 (2000).
Jiang, L. I. et al. Use of a cAMP BRET sensor to characterize a novel regulation of cAMP by the sphingosine 1-phosphate/G13 pathway. J. Biol. Chem. 282, 10576–10584 (2007).
Espinoza, S. et al. Functional interaction between trace amine-associated receptor 1 and dopamine D2 receptor. Mol. Pharmacol. 80, 416–425 (2011).
Weichert, D. et al. Molecular determinants of biased agonism at the dopamine D2 receptor. J. Med. Chem. 58, 2703–2717 (2015).
Cheng, Y.-C. & Prusoff, W. H. Relationship between the inhibition constant (KI) and the concentration of inhibitor which causes 50 per cent inhibition (I50) of an enzymatic reaction. Biochem. Pharmacol. 22, 3099–3108 (1973).
Prather, I. D., Brown, D. E., North, P. & Wilson, J. R. Clenbuterol - a substitute for anabolic-steroids. Med. Sci. Sport. Exer. 27, 1118–1121 (1995).
Afaq, F. et al. Delphinidin, an anthocyanidin in pigmented fruits and vegetables, protects human HaCaT keratinocytes and mouse skin against UVB-mediated oxidative stress and apoptosis. J. Invest. Dermatol. 127, 222–232 (2006).
Parr, A. J., Mellon, F. A., Colquhoun, I. J. & Davies, H. V. Dihydrocaffeoyl polyamines (kukoamine and allies) in potato (Solanum tuberosum) tubers detected during metabolite profiling. J. Agric. Food Chem. 53, 5461–5466 (2005).
Yeung, H. W., Kong, Y. C., Lay, W. P. & Cheng, K. F. The structure and biological effect of leonurine. A uterotonic principle from the Chinese drug, I-mu Ts’ao. Planta Med. 31, 51–56 (1977).
Müller, G. F. R. & Eugster, C. H. Muscimol, ein pharmakodynamisch wirksamer Stoff aus Amanita muscaria. Helv. Chim. Acta 48, 910–926 (1965).
Belitz, H.-D., Grosch, W. & Schieberle, P. Lehrbuch der Lebensmittelchemie. 290–292 (Springer, 2007).
Siddiqui, S. & Siddiqui, R. H. Chemical examination of the roots of Rauwolfia serpentina, Benth. J. Indian Chem. Soc. 8, 667–680 (1931).
Chatterjee, R. & Maiti, P. C. Plant alkaloids. VII. Lambertine and berlambine. J. Indian Chem. Soc. 32, 609–610 (1955).
Lachman, M. F., Romeo, R. & Mccomb, R. B. Emetine identified in urine by HPLC, with fluorescence and ultraviolet-diode array detection, in a patient with cardiomyopathy. Clin. Chem. 35, 499–502 (1989).
Himmele, W. & Pommer, E.-H. 3-Phenylpropylamines, a new class of systemic fungicides. Angew. Chem., Int. Ed. 19, 184–189 (1980).
Beier, R. C. et al. Detection of halofuginone residues in chicken liver tissue by HPLC and a monoclonal-based immunoassay. J. Agric. Food Chem. 46, 1049–1054 (1998).
Ohmomo, S., Utagawa, T. & Abe, M. Identification of roquefortine C produced by Penicillium roqueforti . Agric. Biol. Chem. 41, 2097–2098 (1977).
Chu, D. T. W. et al. Synthesis and structure-activity relationships of novel arylfluoroquinolone antibacterial agents. J. Med. Chem. 28, 1558–1564 (1985).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: T.S., H.H., A.E.K., M.P., T.C. Performed the experiments: T.S., H.H., A.E.K. Analysed the data: T.S., H.H. Interpreted the data: T.S., P.G., M.P., T.C. Contributed reagents/materials/analysis tools: A.E.K., P.G., T.C. Wrote the paper: T.S., H.H., A.E.K., P.G., M.P., T.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sommer, T., Hübner, H., El Kerdawy, A. et al. Identification of the Beer Component Hordenine as Food-Derived Dopamine D2 Receptor Agonist by Virtual Screening a 3D Compound Database. Sci Rep 7, 44201 (2017). https://doi.org/10.1038/srep44201
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep44201
This article is cited by
-
Rational design of agonists for bitter taste receptor TAS2R14: from modeling to bench and back
Cellular and Molecular Life Sciences (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.