Abstract
Recent studies involving connectome analysis including graph theory have yielded potential biomarkers for mental disorders. In this study, we aimed to investigate the differences of resting-state network between patients with social anxiety disorder (SAD) and healthy controls (HCs), as well as to distinguish between individual subjects using topological properties. In total, 42 SAD patients and the same number of HCs underwent resting functional MRI, and the topological organization of the whole-brain functional network was calculated using graph theory. Compared with the controls, the patients showed a decrease in 49 positive connections. In the topological analysis, the patients showed an increase in the area under the curve (AUC) of the global shortest path length of the network (Lp) and a decrease in the AUC of the global clustering coefficient of the network (Cp). Furthermore, the AUCs of Lp and Cp were used to effectively discriminate the individual SAD patients from the HCs with high accuracy. This study revealed that the neural networks of the SAD patients showed changes in topological characteristics, and these changes were prominent not only in both groups but also at the individual level. This study provides a new perspective for the identification of patients with SAD.
Similar content being viewed by others
Introduction
Social anxiety disorder (SAD) is the most common anxiety disorder, and it is characterized by fear and avoidance of social situations associated with being observed or evaluated by others or a fear of embarrassing oneself1. Recent evidence has suggested that abnormal cerebral functioning is involved in the pathogenesis of SAD. Neuroimaging studies have found an increase in the regional activity of the limbic and paralimbic regions including the amygdala and anterior cingulate, as well as reduced activity in the striatal and parietal areas both in groups2 and at the individual level3. For example, using a linear support vector machine (SVM), we identified regional homogeneity (ReHo) in the default mode network, dorsal attention network, self-referential network, and sensory networks, thus distinguishing the patients with SAD from the healthy controls with a diagnostic accuracy of 76.25%3.
However, the findings from the regional analysis were inconsistent and distributed across the entire cerebral region of the patients with SAD, which may be due to the effects of medication or sample heterogeneity. These inconsistent findings can also be attributed to subtle and widespread white matter deficits across the brain, which are not perfectly integrated by traditional regional or voxel-based analyses. Therefore, connectomics has been used to evaluate the whole brain as an interconnected network4.
Graph theory provides powerful mathematical tools to study the behaviour of complex systems of interacting elements5. It has been widely used to characterize local and distributed interactions in the brain6, and altered topological characteristics in functional brain networks have been observed in psychiatric disorders such as schizophrenia7,8, major depressive disorder9,10, and obsessive-compulsive disorder11. In particular, we found significant correlations between functional connectivity and disease severity in specific regions of resting-state networks (RSNs), including the medial and lateral prefrontal cortex, as well as the parietal and occipital regions, in SAD patients using independent component analysis12.
Previous studies of SAD have indicated that the human connectome can yield potential biomarkers for SAD. However, few studies have explored the topology of the functional network in SAD. Last year, we used functional connectivity strength (FCS) to examine the cortical hubs in SAD using the graph theory method and found that the patients with SAD had disrupted cortical hubs during resting states, a discovery that provides novel insights into the pathophysiological mechanisms of SAD13. In addition to cortical hubs, “small-world” parameters and network efficiency, which can indicate the organization of the global network, are also important in graph theory analysis. However, no studies have explored both the global and regional topological properties of SAD. Although there are many topological indexes in the graph theory method, it is unclear whether the topological characteristics are altered and whether these indexes can be used to identify SAD patients individually.
The present study analysed the network topology of the brain in patients with SAD using intrinsic functional networks. The topological organization of the functional brain network based on resting-state functional connectivity and the linkage between the characteristics of the brain networks and the patients’ clinical conditions were assessed in SAD patients, as well as healthy controls (HCs). We hypothesized that the global and regional topological properties could be used to distinguish SAD patients from HCs by multivariate pattern analysis (MVPA).
Materials and Methods
Subjects
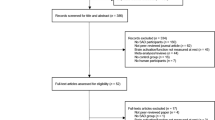
We initially recruited 47 SAD patients and 45 healthy controls, all Han Chinese, for this study. Data from five patients and three controls were discarded because of excessive head movement during the MRI scan. Therefore, 42 SAD patients (26 males and 16 females with a mean age of 27.33 ± 7.159 years) and 42 controls (26 males and 16 females with a mean age of 29.83 ± 8.748 years) were finally included in the statistical analysis. The psychiatric diagnostic classification of the participants was based on the Structured Clinical Interview for DSM-IV Axis I Disorders (SCID)14 conducted by two attending psychiatrists and a trained interviewer.
All of the subjects in the SAD group were in accordance with the DSM-IV criteria for SAD. Four of the patients had been taking a stable dosage of a selective serotonin reuptake inhibitor for at least 4 weeks (two patients were treated with paroxetine (20 mg/day), one patient was treated with paroxetine (10 mg/day), and one patient was treated with paroxetine (20 mg/day), as well as tandospirone (20 mg/day)). However, some of the patients had discontinued their use of psychotropic medications due to poor responses at least 2 weeks prior to the baseline MRI scan. We also recruited 45 controls who had never been diagnosed with SAD or any other psychiatric disorders.
The exclusion criteria included any history of neurological disease, bipolar disorder, alcohol and/or other substance abuse/dependence, major head injury involving loss of consciousness for more than 10 minutes, other DSM-IV axis I diagnoses, and mental retardation, as well as subjects with metal implants (e.g., surgical clips or pacemakers). According to the Structured Clinical Interview for DSM-IV14, the participants in the SAD group met the criteria for the following current comorbid diagnoses: major depression (N = 2) and generalized anxiety disorder (N = 1).
All of the participants were assessed using the Liebowitz Social Anxiety Scale Self-Report (LSAS), the Hamilton Rating Scale for Depression (HAMD-24), and the Hamilton Rating Scale for Anxiety (HAMA-14), and all of the patients provided informed consent. This study was approved by the Medical Ethics Committee of West China Hospital, Sichuan University, and all of the experiments were performed in accordance with the Declaration of Helsinki.
Image acquisition
We used a 3.0 T magnetic resonance scanner (Siemens 3.0 T Trio Tim, Germany) with a 12-channel phase array head coil. Each subject was positioned supine in the MRI scanner with foam padding to reduce head movements. The fMRI blood-oxygen-level-dependent (BOLD) images were acquired by a gradient-echo-planar imaging (EPI) sequence (TR/TE = 2,000/30 ms; flip angle = 90°). The slice thickness was 5 mm with a matrix size of 64 × 64, resulting in voxels of 3.75 × 3.75 × 5 mm3. The subjects were instructed to close their eyes, let their minds wander, and relax, but not to fall asleep during the scan, which lasted 6.8 min (205 volume), as described in our previous study15.
Data preprocessing
Functional image preprocessing and statistical analyses were conducted using the Data Processing Assistant for Resting-State fMRI (DPARSF_V4.0) in DPABI (http://rfmri.org/dpabi)16, which is based on Statistical Parametric Mapping (SPM8, http://www.fil.ion.ucl.ac.uk/spm) and the Matlab toolbox (http://www.brain-connectivity-toolbox.net). The DPARSF includes powerful and updated preprocessing steps and has been used in hundreds of research projects17. The first five time points were discarded for scanner calibration and acclimation of the subjects to the scanning conditions. Thus, 200 time points from the rest condition time series were used for network analysis. For each participant, the images were corrected for differences in the intravolume acquisition time between slices using sinc interpolation and then corrected for intervolume geometric displacement due to head movement using a six-parameter (rigid-body) spatial transformation10. Data from five patients and three healthy controls were discarded because their heads moved more than 3 mm of translation or 3 degrees of rotation in any direction. After these corrections, the realigned images were spatially normalized to the EPI template in SPM8, and each voxel was resampled to 3 × 3 × 3 mm3 without spatial filtering. For network metrics calculation, the images were smoothed using a 4-mm full-width half-maximum (FWHM) isotropic Gaussian filter18. For each run, the nuisance terms were regressed from the resting-state BOLD time series through multiple linear regressions. These nuisance regressors included the following: i) linear and quadratic trends, ii) a Friston 24-parameter autoregressive model that included current and past position parameters and adequately addressed head motion effects18, iii) white matter and CSF signals, which were masked based on SPM apriori, and iv) white matter and cerebrospinal fluid time series. The summary scalars of both the gross (maximum and root mean square) and micro (mean frame-wise displacement) head movements were matched between the two groups (all p > 0.15). Finally, the corrected BOLD time series were low-pass filtered using a cut-off frequency of 0.01–0.1 Hz to reduce the low frequency drift and high frequency noise.
Network construction
Node definition
A network is composed of nodes and edges between nodes. In the brain, the nodes represent the brain regions, and the edges represent the statistical relationships of BOLD signals across different regions. Since the atlas of Automated Anatomical Labeling19 (AAL) has been most commonly used in previous studies and is widely accepted in neuroimaging studies, we used the AAL, which includes 90 areas in the whole brain that represent 90 nodes of the brain network. The names and indexes of the 90 areas (45 for each hemisphere) are listed in Table S1 in the Supplementary Material.
Edge definition
To define the network edges, the interregional resting-state functional connectivity (RSFC) was calculated using Pearson correlations between the regional mean time series of all possible pairs of brain regions. The correlation coefficients were transformed to z-scores via Fisher’s transformation to improve normality20. We constructed a 90 × 90 correlation matrix for each subject.
Network analysis
Threshold Selection
Based on the 90 × 90 correlation matrix for each subject, we constructed a binary undirected graph, and a sparsity threshold was used to measure all of the correlation matrices. Due to the difficulty involved in selecting a single threshold, empirical studies9,21 were used with a wide range of 0.10 ≤ sparsity ≤ 0.34 (interval = 0.01), in which the “small-world” parameters could be properly estimated22 and the number of spurious edges was minimized23,24,25. According to the previous study, there were two criteria considered when the threshold was generated: 1) the average degree (see Supplementary Material for the definition) over all of the nodes of each thresholded network was larger than 2 × log(N), with N = 90 denoting the number of nodes; and 2) the small-worldness (see Supplementary Material for definition) of the thresholded networks was larger than 1.1 for all participants9. To make sure all/most of the nodes were connected for each subject after thresholding, we also used the largest component size (that is, the number of nodes in the largest connected component divided by the number of all of the available network nodes N26).
Due to the ambiguous biological explanation for negative correlations27,28, we restricted our analyses to positive correlations. We calculated both the global and node network metrics at each sparsity, and the area under the curve (AUC) for each network metric, which provides a summarized scalar for the topological characterization of brain networks (Fig. 1), was calculated.
Network Metrics
We analysed the global metrics in this study using the following parameters: (1) The “small-world” parameter clustering coefficients included the shortest path length, normalized characteristic path length (λ), normalized clustering coefficient (γ), and small-worldness (σ). The shortest path length is defined as the shortest mean distance from a particular vertex to all other vertices22,29. Thus, a smaller path length represents greater integration. The clustering coefficient is defined as the fraction of a vertex’s neighbours that are neighbours themselves29, while a larger clustering coefficient represents greater segregation30. The path length and clustering coefficient were normalized by the related mean metrics of the 100 random networks. These random networks had the same number of nodes, edges, and degree distributions as the real brain networks. (2) Network efficiency included the local efficiency of the whole network (Eloc), the global efficiency of the network (Eglob), the nodal global efficiency of the node (nodalEglob), and the nodal local efficiency of the node (nodalEloc). (3) Nodal centrality (the degree number of nodes (nodalDeg)) was the final parameter. These definitions and descriptions of the metrics are listed in Supplementary Material and are provided in a reference 29.
All of the network metrics were calculated using the GRaph thEoreTical Network Analysis (GRETNA) toolbox (https://www.nitrc.org/projects/gretna/)31, and this method of network construction and calculation has been used in previous studies of brain networks21,32,33. The brain networks were visualized using the BrainNet Viewer34.
Statistical analysis
Functional network connectivity analysis between the SAD patients and the HCs
We performed two-sample, two-tailed t-tests on all 7560 of the possible connections represented in the 90 × 90 correlation matrices between the patients and the controls35 using GRETNA, and the false discovery rate (FDR) procedure was applied to correct for multiple comparisons.
Group comparisons based on topological metrics
We used non-parametric permutation tests (10,000 permutations) adopting the Matlab language to test the intragroup differences in the brain network metrics, and gender and age were treated as the unconcerned covariates for comparisons. Furthermore, the problem of multiple comparisons was addressed by testing the graph-based metrics for survival using a Benjamini-Hochberg false discovery rate correction at the expected significance level of 0.0536. Spearman’s correlation coefficient was used to calculate the associations between the network metrics and the clinical variables in the SAD groups.
Multivariate pattern analysis
We used a multivariate pattern analysis (MVPA) to explore the role of these functional connectivity and network parameters in distinguishing the SAD patients from the healthy controls. To reduce the data dimensions and improve the performance of the classifier37,38, we used feature selection based on two-sample, two-tailed t-tests. Thus, the functional connectivity (FC) and network metrics with significant differences between groups were treated as discriminant features (p < 0.05, uncorrected). Maximum uncertainty linear discriminate analysis (MLDA) employing a maximum entropy covariance selection method instead of the within-class scatter matrix was used as the classifier39,40,41. The performance of the classifier was estimated using a leave-one-out cross-validation (LOOCV) approach, and these steps were supported by multi-modal imaging and multi-level characterization with multi-classifier (M3), which has been made publicly available at: http://www.nitrc.org/projects/pare/ 39. Finally, we used a permutation test to infer the significance of the classifier performance by random disruption of the label in all of the SAD patients and healthy controls, and the class validation procedure was repeated 1000 times. Thus, we obtained a distribution of the classifier performance with random labels, which was used to calculate the z-score and p value.
Results
Demographic data and clinical comparisons
There were no significant differences between the SAD group and the control group in terms of age, sex, and, education (p > 0.05; Table 1).
Disrupted Functional Network Connectivity in SAD
Compared with the HCs, 49 connections showed a significant decrease in positive connections in the SAD patients (p < 0.05, FDR-corrected). The most altered connections involved the frontal, occipital, parietal–(pre)motor, and temporal regions, and all of the altered connection pairs are listed in Fig. 2 and Supplement Table S2. Additionally, we used a stricter significance level of p < 0.01 (FDR-corrected) and found two decreased connections in the SAD patients, including the right superior frontal gyrus, medial–left posterior cingulate gyrus and the right superior frontal gyrus, medial–right precuneus. These results reveal that the abnormal correlations in the SAD patients are universal and more related to the default mode network.
There are 49 decreased connections which were significantly (FDR corrected p = 0.05) abnormal in the patients. All of 32 brain regions are marked by using different colored spheres (different colors represent distinct brain classification) and further mapped onto the cortical surfaces at the lateral, medial and top views, respectively, by using the BrainNet Viewer package (www.nitrc.org/projects/bnv). The size of the spheres represents the number of altered connections they involved. The color of the edges represents the t value of the comparisons, and the color bar is on the left side. ACG, anterior cingulate and paracingulate gyri; ANG, angular gyrus; CAL, calcarine fissure and surrounding cortex; CUN, cuneus; FFG, fusiform gyrus; IOG, inferior occipital gyrus; ITG, inferior temporal gyrus; MOG, middle occipital gyrus; MTG, middle temporal gyrus; ORBsuped, superior frontal gyrus, medial orbital; PCG, posterior cingulate gyrus; PCUN, precuneus; REC, gyrus rectus; SFGdor, superior frontal gyrus, dorsolateral; SFGmed, superior frontal gyrus, medial; TPOmid, temporal pole: middle temporal gyrus.
Global parameters of brain networks
To make sure that all/most of the nodes were connected after the strongest thresholding, we examined the largest component size at all thresholds (see Fig. 3a) and found that most of the nodes (0.944) were connected even at the strongest threshold (sparsity = 0.10).
Graphs show that in the defined threshold range, both the SAD and control groups exhibited (a) the number of nodes in the largest connected component divided by all the available nodes N, (b) the normalized clustering coefficient, (c) the normalized characteristic path length, and (d) small-world measure.
There were no significant differences between the SAD group and control group in local and global efficiencies, λ, γ, and σ (Fig. 4A). In both groups, the values of λ hovered at approximately 1, γ significantly exceeded 1, and σ exceeded 1, suggesting that both the patients with SAD and the controls showed “small-world” organization in resting states (Fig. 3b–d).
Part A: Topological properties of functional connectivity networks in SAD and controls are shown in different colors. The error bars represent standard deviation. ** Group difference is significant at the 0.01 level (FDR corrected) based on non-parametric permutation tests. Part B: Scatter plots in part B show significant correlation between the global topological characteristics and HAMD scores in the SAD group (p < 0.05). SAD, social anxiety disorder; HCs, health controls; AUC, area under the curve; Lp, shortest path length; Cp, clustering coefficient; γ, the normalized clustering coefficient; λ, the normalized characteristic path length; σ, small-world measure; Eglob, global efficiency; Eloc, local efficiency; aLp AUC of shortest path length; aCp, AUC of clustering coefficient; HAMD, Hamilton Rating Scale for Depression.
Compared with the HC group, an increased AUC of the shortest path length of the network (Lp) and a decreased clustering coefficient of the network (Cp) were observed in the patients with SAD (p < 0.01). A negative correlation was also found between the AUC of Cp (aCp) and the HAMD scores in the SAD group (r = 0.361, p = 0.026). (Fig. 4B). Nevertheless, the correlation was no longer significant after Bonferroni correction.
Regional parameters of brain networks
After FDR correction, no significant differences in the AUC of nodal degree (anodalDeg) and the AUC of nodal global efficiency (anodalEglob) were observed between the SAD and control groups.
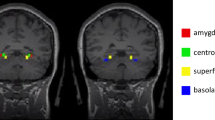
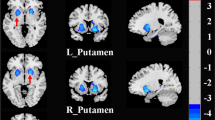
Regarding local efficiency, a decreased AUC of nodal local efficiency (anodalEloc) was observed in the left posterior cingulate gyrus (PCG) of the SAD patients compared with the controls, and the nodal local efficiency in the right putamen (PUT) was also increased (p < 0.05, FDR corrected) (see Fig. 5). The anodalEloc of the left PCG and the avoidance score on the LSAS showed a significant negative correlation in the SAD patients (r = −0.326, p = 0.035), but the correlation was no longer significant after Bonferroni correction.
The colored areas show the location of left posterior cingulate gyrus and the right putamen in the Automated Anatomical Labeling atlas. Scatter plots show significant correlation between the nodal local efficiency in left posterior cingulate gyrus and avoidance score of LSAS in the SAD group (p < 0.05). PUT.R, right putamen; PCG.L, left posterior cingulate gyrus; anodalEloc, area under the curve of nodal local efficiency; LSAS_avoid, avoidance score of Liebowitz Social Anxiety Scale.
Discriminant analysis
Only the altered FC and network metrics including the 49 decreased FCs, the AUCs of Lp and Cp, and the anodalEloc in the left PCG and right PUT were investigated to separately distinguish the SAD patients from the HCs. Because of the stricter inclusion criteria of the features before M3, all of the features we chose were retained after the looser feature selection of M3. These network metrics were abstracted individually and are shown in Fig. 6.
The network features which can discriminate SAD from healthy controls effectively were abstracted. (a) the AUC of Lp, (b) the AUC of Cp, (c) the anodalEloc of PCG.L and (d) the anodalEloc of PUT.R. SAD, social anxiety disorder; HCs, health controls; AUC, area under the curve; Lp, shortest path length; Cp, clustering coefficient; anodalEloc, area under the curve of nodal local efficiency; PCG.L, left posterior cingulate gyrus; PUT.R, right putamen.
Based on the 49 altered FC analysis findings, we found that the discrimination ability (accuracy = 0.667, sensitivity = 0.738, specificity = 0.595) was not significant after the nonparametric permutation test (z = 1.728). We also calculated the feature weights of all of the features, which represent the contribution to the classification39. The feature weights of all of the pairs in the M3 analysis are listed in Supplement Table S2, and the feature weights of the topological parameters are listed in the Table 2.
We found that the AUC of Lp and the AUC of Cp effectively discriminated between the two groups separately (aLp: accuracy = 0.988, sensitivity = 0.976, specificity = 1; aCp: accuracy = 0.964, sensitivity = 0.929, specificity = 1), which was significantly above the random level (aLp:z = 12.554; aCp: z = 12.783). Only a single patient was miscategorised as a control among the 84 subjects. Combining the AUC of Lp and the AUC of Cp yielded an accuracy rate that was similar to the AUC of Lp alone (accuracy = 0.988, sensitivity = 0.976, specificity = 1, z = 12.514). The classification performance and feature weights of the topological parameters are listed in the Table 2.
We used the anodalEloc in the left PCG and right PUT to distinguish the SAD patients from the HCs, and we also found that the local nodal efficiency of the left PCG and right PUT in the resting-state network significantly distinguished the two groups (accuracy = 0. 714, sensitivity = 0. 738, specificity = 0.691, z = 5.237).
Discussion
The present study used resting-state fMRI to explore changes in the brain topology of SAD patients. To our knowledge, this is the first study to investigate the “small-world” brain network in SAD patients. Compared with controls, the SAD patients had 49 decreased connections, which involved the frontal, occipital, parietal–(pre)motor, and temporal regions. In the global topological analysis, an increased AUC of Lp and a decreased AUC of Cpwere found in the patients with SAD. In the local regions, an increased anodalEloc of the right PUT was observed in the patients with SAD, while the anodalEloc of the left PCG was reduced. These findings reveal a shifting of the “small-world” properties in SAD patients during resting states and the critical role of the PUT and PCG in the pathogenesis of SAD. These altered topological metrics may be caused by a disruption of functional connectivity in SAD. Furthermore, these differences were used to effectively discriminate between the individual patients with SAD and the HCs with an accuracy of 98.8%.
In the functional network connectivity analysis, the major altered connections involved the frontal, occipital, parietal–(pre)motor, and temporal regions. Compared with the HCs, all of the changed connections were decreased, which may indicate that network efficiency is disrupted in patients with SAD. The attenuation of the FC between the frontal and occipital lobes was consistent with the results of our previous study of SAD, which had a small sample size42. The frontal, occipital, parietal–(pre)motor, and temporal regions are related to the functions of cognition, emotion, and memory, and these regions were examined in previous functional MRI studies of SAD43. Moreover, the areas in the default mode network (DMN)44,45,46 including the precuneus, posterior cingulate gyrus, angular gyrus, middle temporal gyrus, medial frontal gyrus, and superior frontal gyrus were obviously involved. The DMN is thought to be involved in episodic memory47, self-projection48, and social cognition49. Impairment of the DMN network in SAD might be relevant to the development of feelings of wariness concerning the judgement of others and may be related to the self-focused attention49. The cuneus and calcarine fissure in the occipital lobe are involved in converging facial expressions50,51 and contextual self-descriptions52. Therefore, the decreased correlations between the median prefrontal cortex and occipital lobe might indicate that SAD is related to abnormal processing of the nonverbal information relayed by human facial expressions42.
According to the currently fashionable idea, the brain network can be divided into the regular network, the random network, and the “small-world” network. The regular network is characterized by a higher clustering coefficient and a longer shortest path length. The random network has a lower clustering coefficient and a shorter shortest path length. The “small-world” network has a higher clustering coefficient similar to the regular network and a shortest path length that is similar to the random network22. In the current study, we found that both the patients with SAD and the controls had a λ ≈ 1, a γ ≫ 1, and a σ that was >1, indicating the presence of “small-world” brain networks in both groups. The functional connection was correlated with the structural connections in the brain, and “small-world” functional and structural networks occur in the human brain, as well as at the cellular level in other animals5. Numerous studies using electroencephalography (EEG)53, magnetoencephalography (MEG)54, and fMRI55 have reported “small-world” properties of whole-brain functional networks in humans. In the “small-world” brain network, functional integration and segregation are the two major organizational principles. An optimal brain requires a balance between global integration and local specialization of brain functional activity56 and achieves a maximal communication speed with minimal energy consumption57. Among these topological indices, Cp and Lp represent segregation and integration, respectively. Cp is equivalent to the fraction of nodal neighbours that are also neighbours to each other22, which reflects the local efficiency or fault tolerance of a network58. Thus, a decrease in Cp indicates local efficiency or fault tolerance, which decreases functional segregation when disrupted in the brains of patients with SAD. Lp is a global feature that indicates the information-carrying capacity of the brain. A lower Lp value ensures global information transmission capacity and provides a reliable base of functional integration in the brain59,60. The increase in the AUC of Lp in our study suggests a disruption of global information transmission in the brains of patients with SAD. Although the brain networks of the SAD patients and controls showed “small-world” characteristics, the increased AUC of Cp and decreased AUC of Lp represent disruption in the brain networks in terms of functional integration and separation in the SAD patients, which may reflect an imbalance of global integration and local specialization. This imbalance suggested a disruption in the energy cost of spontaneous brain activity and was associated with the severity of depression in SAD patients, which was compatible with major depressive disorder61.
Among these exploratory correlation analyses with clinical measurements, we found that the AUC of Cp was significantly associated with depression but not social (or general) anxiety. Since patients with depression also suffer from an imbalance of global integration and local specialization61, it is not clear if this abnormality is more a function of subclinical depressive symptoms rather than the chosen diagnosis. As the correlation with clinical measures involved exploratory analyses, which did not show significance after multiple comparison corrections, the reproducibility of the correlation between the AUC of Cp and HAMD needs to be verified in future studies. Furthermore, it is common for patients with SAD to suffer from more depressive symptoms. Thus, it was hard to disengage anxiety from depression in this study due to the small sample size. Even though the recruitment of SAD patients with few or no depressive symptoms may solve this problem, the application of the results would be restricted in the real world.
In terms of the regional parameters of the brain networks, the SAD group showed alterations of the anodalEloc in the left posterior cingulate gyrus (PCG) and right putamen (PUT) compared with the HCs, and the reduction in the anodalEloc of the left PCG was negatively associated with the avoidance score of the LSAS in the exploratory analysis. The nodalEloc is a summarized scalar reflecting the transmission efficiency of the network at the local level, and the posterior cingulate has been associated with self-referential functions62,63, self-focused attention (awareness of self-referent information)64, evaluation of self-emotional states65, and social behaviour66. Neuroimaging studies have suggested that the PCG is attenuated during task conditions49,64. Furthermore, functional imaging has revealed a reduction in rCBF in the PCG of SAD patients during resting states67, and a resting-state fMRI study that used functional connectivity suggested that attenuated functional connectivity in the amygdala and the PCG/precuneus is correlated with higher anxiety68. The altered nodal efficiency of the left PCG in our study demonstrated low efficiency of local information transfer and processing around the left PCG, which might be related to avoidance behaviour in SAD patients. The putamen is regarded as one of the sectors of the striatum that is the “emotion guarder”, and it is an important terminal for receiving sensory and emotional information from the prefrontal areas69. Striatal dysfunction in SAD patients has been observed in previous neuroimaging studies, and it may be linked to the information processing biases found in SAD patients, including negative interpretation of social events, detection of and obsession with negative responses from other people, and selective recall of negative aspects of social interactions70,71. The higher AUC of the local nodal efficiency of the PUT in the SAD patients reflects a reduction in regional integration of information in the right PUT, which may represent an information processing bias in SAD patients. Although the power of the nodal local efficiency in the left PCG and right PUT was lower than the AUC of Lp and Cp, it indicates an important role for these two brain areas in SAD, especially in avoidance symptoms.
In this study, we used discriminant analysis to distinguish SAD patients from controls, and we found that SAD patients were accurately identified using the AUC of Lp and the AUC of Cp. The remarkable accuracy of the discriminant analysis may be attributed to the huge disparity in the AUCs of Lp and Cp between the two groups and at the individual level. Thus, these attributes may facilitate diagnosis, suggesting the need for additional studies.
The study limitations were as follows. First, only the leave-one-out cross-validation (LOOCV) approach was supported in the M3 code. Even though this method has been widely used in research39,72,73,74, it is sub-optimal and has been shown to result in inaccurate estimates of the prediction error75. It is more appropriate to apply n-fold validation when running the primary analyses on a portion of the patients and healthy controls and then apply the primary results to classify the rest of the subjects. Because the sample size in this study was relatively small and because of the huge disparity in the AUCs of Lp and Cp between the two groups and at the individual level (see Fig. 6), we believe that these results should be independently verified in more samples. Second, compared with other studies, we only explored the network based on the AAL brain atlas, which was the most commonly used in previous studies. Although this is widely accepted in neuroimaging studies, new brain atlases have frequently been used in graph theory, for example, the Power 264-region atlas72,76, the Harvard-Oxford Structural atlas77, and the Dosenbach’s 160 functional atlas78,79. Future studies should verify our results using these atlases in the analyses of different brain networks. Third, several patients with comorbid conditions were included in the current study. Although SAD was their main diagnosis, comorbid conditions such as depression and generalized anxiety may have affected the results. Therefore, separate analyses of SAD patients with and without comorbid conditions are desirable.
Conclusion
In summary, using graph theory, the current study found that both the global and regional topological characteristics of the neural networks of SAD patients were less effective. This is the first study to investigate the “small-world” properties of SAD, and the changes were prominent at the group and individual levels, providing a new perspective for distinguishing patients from healthy individuals. However, future studies that focus on these topological attributes and brain areas in different samples are needed.
Additional Information
How to cite this article: Zhu, H. et al. Altered Topological Properties of Brain Networks in Social Anxiety Disorder: A Resting-state Functional MRI Study. Sci. Rep. 7, 43089; doi: 10.1038/srep43089 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Stein, M. B. & Stein, D. J. Social anxiety disorder. Lancet 371, 1115–1125 (2008).
Freitas-Ferrari, M. C. et al. Neuroimaging in social anxiety disorder: a systematic review of the literature. Prog. Neuropsychopharmacol. Biol. Psychiatry 34, 565–580 (2010).
Zhang, W. et al. Diagnostic Prediction for Social Anxiety Disorder via Multivariate Pattern Analysis of the Regional Homogeneity. Biomed Res Int 2015, 763965 (2015).
Pannekoek, J. N. et al. Resting-state functional connectivity abnormalities in limbic and salience networks in social anxiety disorder without comorbidity. Eur Neuropsychopharmacol 23, 186–195 (2013).
Bullmore, E. & Sporns, O. Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198 (2009).
Bassett, D. S. & Bullmore, E. T. Human brain networks in health and disease. Curr. Opin. Neurol. 22, 340–347 (2009).
Yu, Q., Sui, J., Rachakonda, S., He, H., Pearlson, G. & Calhoun, V. D. Altered small-world brain networks in temporal lobe in patients with schizophrenia performing an auditory oddball task. Front Syst Neurosci 5, 7 (2011).
Lynall, M. E. et al. Functional connectivity and brain networks in schizophrenia. J. Neurosci. 30, 9477–9487 (2010).
Zhang, J. et al. Disrupted brain connectivity networks in drug-naive, first-episode major depressive disorder. Biol. Psychiatry 70, 334–342 (2011).
Guo, H., Cao, X., Liu, Z., Li, H., Chen, J. & Zhang, K. Machine learning classifier using abnormal brain network topological metrics in major depressive disorder. Neuroreport 23, 1006–1011 (2012).
Gottlich, M., Kramer, U. M., Kordon, A., Hohagen, F. & Zurowski, B. Decreased limbic and increased fronto-parietal connectivity in unmedicated patients with obsessive-compulsive disorder. Hum Brain Mapp 35, 5617–5632 (2014).
Liao, W. et al. Selective aberrant functional connectivity of resting state networks in social anxiety disorder. Neuroimage 52, 1549–1558 (2010).
Liu, F. et al. Disrupted cortical hubs in functional brain networks in social anxiety disorder. Clin Neurophysiol 126, 1711–1716 (2015).
First, M. B., Spitzer, R. L. & Williams, J. B. Structured clinical interview for DSM-IV axis I disorders SCID-I: clinician version, administration booklet (American Psychiatric Pub, 1997).
Zhu, H. et al. Altered spontaneous neuronal activity in chronic posttraumatic stress disorder patients before and after a 12-week paroxetine treatment. J Affect Disord 174, 257–264 (2015).
Yan, C. G., Wang, X. D., Zuo, X. N. & Zang, Y. F. DPABI: Data Processing & Analysis for (Resting-State) Brain Imaging. Neuroinformatics (2016).
Chao-Gan, Y. & Yu-Feng, Z. DPARSF: A MATLAB Toolbox for “Pipeline” Data Analysis of Resting-State fMRI. Front Syst Neurosci 4, 13 (2010).
Zhu, H. et al. Altered spontaneous neuronal activity of visual cortex and medial anterior cingulate cortex in treatment-naive posttraumatic stress disorder. Compr Psychiatry 55, 1688–1695 (2014).
Tzourio-Mazoyer, N. et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 15, 273–289 (2002).
Lowe, M. J., Mock, B. J. & Sorenson, J. A. Functional connectivity in single and multislice echoplanar imaging using resting-state fluctuations. Neuroimage 7, 119–132 (1998).
Lei, D. et al. Disrupted Functional Brain Connectome in Patients with Posttraumatic Stress Disorder. Radiology 276, 818–827 (2015).
Watts, D. J. & Strogatz, S. H. Collective dynamics of ‘small-world’ networks. Nature 393, 440–442 (1998).
He, Y., Chen, Z. & Evans, A. Structural insights into aberrant topological patterns of large-scale cortical networks in Alzheimer’s disease. J. Neurosci. 28, 4756–4766 (2008).
Achard, S. & Bullmore, E. Efficiency and cost of economical brain functional networks. PLoS Comput. Biol. 3, e17 (2007).
Bassett, D. S., Bullmore, E., Verchinski, B. A., Mattay, V. S., Weinberger, D. R. & Meyer-Lindenberg, A. Hierarchical organization of human cortical networks in health and schizophrenia. J. Neurosci. 28, 9239–9248 (2008).
Hayasaka, S. & Laurienti, P. J. Comparison of characteristics between region-and voxel-based network analyses in resting-state fMRI data. Neuroimage 50, 499–508 (2010).
Fox, M. D., Zhang, D., Snyder, A. Z. & Raichle, M. E. The global signal and observed anticorrelated resting state brain networks. J. Neurophysiol. 101, 3270–3283 (2009).
Murphy, K., Birn, R. M., Handwerker, D. A., Jones, T. B. & Bandettini, P. A. The impact of global signal regression on resting state correlations: are anti-correlated networks introduced. Neuroimage 44, 893–905 (2009).
Rubinov, M. & Sporns, O. Complex network measures of brain connectivity: uses and interpretations. Neuroimage 52, 1059–1069 (2010).
Newman, M. E. J. The structure and function of complex networks. 45, 167–256 (2003).
Wang, J., Wang, X., Xia, M., Liao, X., Evans, A. & He, Y. GRETNA: a graph theoretical network analysis toolbox for imaging connectomics. Front Hum Neurosci 9, 386 (2015).
Jung, W. H., Kang, D. H., Kim, E., Shin, K. S., Jang, J. H. & Kwon, J. S. Abnormal corticostriatal-limbic functional connectivity in obsessive-compulsive disorder during reward processing and resting-state. Neuroimage Clin 3, 27–38 (2013).
Wang, Y. et al. Disrupted Resting-State Functional Connectivity in Nonmedicated Bipolar Disorder. Radiology 280, 529–536 (2016).
Xia, M., Wang, J. & He, Y. BrainNet Viewer: a network visualization tool for human brain connectomics. PLoS ONE 8, e68910 (2013).
Liu, Y. et al. Disrupted small-world networks in schizophrenia. Brain 131, 945–961 (2008).
Y. Benjamini & Y. H. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society 57, (1995).
De Martino F., Valente, G., Staeren, N., Ashburner, J., Goebel, R. & Formisano, E. Combining multivariate voxel selection and support vector machines for mapping and classification of fMRI spatial patterns. Neuroimage 43, 44–58 (2008).
Pereira, F., Mitchell, T. & Botvinick, M. Machine learning classifiers and fMRI: a tutorial overview. Neuroimage 45, S199–209 (2009).
Dai, Z. et al. Discriminative analysis of early Alzheimer’s disease using multi-modal imaging and multi-level characterization with multi-classifier (M3). Neuroimage 59, 2187–2195 (2012).
Thomaz, C. E. et al. A multivariate statistical analysis of the developing human brain in preterm infants. Image Vis Comput 25, 981–994 (2007).
Thomaz, C. E. et al. Whole brain voxel-based analysis using registration and multivariate statistics.
Ding, J. et al. Disrupted functional connectivity in social anxiety disorder: a resting-state fMRI study. Magn Reson Imaging 29, 701–711 (2011).
Mathew, S. J., Coplan, J. D. & Gorman, J. M. Neurobiological mechanisms of social anxiety disorder. Am J Psychiatry 158, 1558–1567 (2001).
Raichle, M. E., MacLeod, A. M., Snyder, A. Z., Powers, W. J., Gusnard, D. A. & Shulman, G. L. A default mode of brain function. Proc. Natl. Acad. Sci. USA 98, 676–682 (2001).
Fox, M. D., Snyder, A. Z., Vincent, J. L., Corbetta, M., Van Essen D. C. & Raichle, M. E. The human brain is intrinsically organized into dynamic, anticorrelated functional networks. Proc. Natl. Acad. Sci. USA 102, 9673–9678 (2005).
Greicius, M. D., Krasnow, B., Reiss, A. L. & Menon, V. Functional connectivity in the resting brain: a network analysis of the defaultmode hypothesis. Proc. Natl. Acad. Sci. USA 100, 253–258 (2003).
Vincent, J. L. et al. Coherent spontaneous activity identifies a hippocampal-parietal memory network. J. Neurophysiol. 96, 3517–3531 (2006).
Buckner, R. L. & Carroll, D. C. Self-projection and the brain. Trends Cogn. Sci. (Regul. Ed.) 11, 49–57 (2007).
Gentili, C. et al. Beyond amygdala: Default Mode Network activity differs between patients with social phobia and healthy controls. Brain Res. Bull. 79, 409–413 (2009).
Rämä, P., Martinkauppi, S., Linnankoski, I., Koivisto, J., Aronen, H. J. & Carlson, S. Working memory of identification of emotional vocal expressions: an fMRI study. Neuroimage 13, 1090–1101 (2001).
Streit, M. et al. Disturbed facial affect recognition in patients with schizophrenia associated with hypoactivity in distributed brain regions: a magnetoencephalographic study. Am J Psychiatry 158, 1429–1436 (2001).
Chiao, J. Y. et al. Neural basis of individualistic and collectivistic views of self. Hum Brain Mapp 30, 2813–2820 (2009).
Smit, D. J., Stam, C. J., Posthuma, D., Boomsma, D. I. & de Geus E. J. Heritability of “small-world” networks in the brain: a graph theoretical analysis of resting-state EEG functional connectivity. Hum Brain Mapp 29, 1368–1378 (2008).
Stam, C. J. Functional connectivity patterns of human magnetoencephalographic recordings: a ‘small-world’ network. Neurosci. Lett. 355, 25–28 (2004).
Salvador, R., Suckling, J., Coleman, M. R., Pickard, J. D., Menon, D. & Bullmore, E. Neurophysiological architecture of functional magnetic resonance images of human brain. Cereb. Cortex 15, 1332–1342 (2005).
Tononi, G., Edelman, G. M. & Sporns, O. Complexity and coherency: integrating information in the brain. Trends Cogn. Sci. (Regul. Ed.) 2, 474–484 (1998).
Tomasi, D., Wang, G. J. & Volkow, N. D. Energetic cost of brain functional connectivity. Proc. Natl. Acad. Sci. USA 110, 13642–13647 (2013).
Brust, M. R. Is the clustering coefficient a measure for fault tolerance in wireless sensor networks, 2012.
Tononi, G., Sporns, O. & Edelman, G. M. A measure for brain complexity: relating functional segregation and integration in the nervous system. Proc. Natl. Acad. Sci. USA 91, 5033–5037 (1994).
Sporns, O. & Zwi, J. D. The small world of the cerebral cortex. Neuroinformatics 2, 145–162 (2004).
Wang, Z., Yuan, Y., Bai, F., You, J. & Zhang, Z. Altered topological patterns of brain networks in remitted late-onset depression: a resting-state fMRI study. J Clin Psychiatry 77, 123–130 (2016).
Pessoa, L., Padmala, S. & Morland, T. Fate of unattended fearful faces in the amygdala is determined by both attentional resources and cognitive modulation. Neuroimage 28, 249–255 (2005).
Saleem, K. S., Kondo, H. & Price, J. L. Complementary circuits connecting the orbital and medial prefrontal networks with the temporal, insular, and opercular cortex in the macaque monkey. J. Comp. Neurol. 506, 659–693 (2008).
Nakao, T. et al. fMRI of patients with social anxiety disorder during a social situation task. Neurosci. Res. 69, 67–72 (2011).
Wright, P., Albarracin, D., Brown, R. D., Li, H., He, G. & Liu, Y. Dissociated responses in the amygdala and orbitofrontal cortex to bottom-up and top-down components of emotional evaluation. Neuroimage 39, 894–902 (2008).
Adolphs, R. Cognitive neuroscience of human social behaviour. Nat. Rev. Neurosci. 4, 165–178 (2003).
Warwick, J. M., Carey, P., Jordaan, G. P., Dupont, P. & Stein, D. J. Resting brain perfusion in social anxiety disorder: a voxel-wise whole brain comparison with healthy control subjects. Prog. Neuropsychopharmacol. Biol. Psychiatry 32, 1251–1256 (2008).
Hahn, A. et al. Reduced resting-state functional connectivity between amygdala and orbitofrontal cortex in social anxiety disorder. Neuroimage 56, 881–889 (2011).
Friedman, A. et al. A Corticostriatal Path Targeting Striosomes Controls Decision-Making under Conflict. Cell 161, 1320–1333 (2015).
Clark, D. M. & McManus, F. Information processing in social phobia. Biol. Psychiatry 51, 92–100 (2002).
Sareen, J. et al. Striatal function in generalized social phobia: a functional magnetic resonance imaging study. Biol. Psychiatry 61, 396–404 (2007).
Zhang, D., Wang, J., Liu, X., Chen, J. & Liu, B. Aberrant Brain Network Efficiency in Parkinson’s Disease Patients with Tremor: A Multi-Modality Study. Front Aging Neurosci 7, 169 (2015).
Fan, Y., Resnick, S. M., Wu, X. & Davatzikos, C. Structural and functional biomarkers of prodromal Alzheimer’s disease: a high-dimensional pattern classification study. Neuroimage 41, 277–285 (2008).
Sun, D. et al. Elucidating a magnetic resonance imaging-based neuroanatomic biomarker for psychosis: classification analysis using probabilistic brain atlas and machine learning algorithms. Biol. Psychiatry 66, 1055–1060 (2009).
Kohavi, R. A study of cross-validation and bootstrap for accuracy estimation and model selection. IJCAI'95 Proceedings of the 14th international joint conference on Artificial intelligence 2, 1137–1143 (1995).
Power, J. D. et al. Functional network organization of the human brain. Neuron 72, 665–678 (2011).
Lord, L. D. et al. Characterization of the anterior cingulate’s role in the at-risk mental state using graph theory. Neuroimage 56, 1531–1539 (2011).
Dosenbach, N. U. et al. Prediction of individual brain maturity using fMRI. Science 329, 1358–1361 (2010).
Brier, M. R. et al. Functional connectivity and graph theory in preclinical Alzheimer’s disease. Neurobiol. Aging 35, 757–768 (2014).
Acknowledgements
This study was supported by National Natural Science Foundation (Grant Nos. 81371484, 81671664 and 81371527), National Key Technologies R&D Program (Program No: 2012BAI01B03) of China and National Youth Top-notch Talent Support Program of China to S.L. These funding sources had no further role in the study design, collection, analysis, interpretation of the data, writing of the report, or the decision to submit the paper for publication. The authors gratefully acknowledge the contributions of Zhengjia Dai, Meng Gao, and Cui Yuan.
Author information
Authors and Affiliations
Contributions
Hongru Zhu, Changjian Qiu, Mengya Jing, Su Lui, Qiyong Gong and Wei Zhang designed the study and wrote the protocol. Hongru Zhu, Mengya Jing, Minlan Yuan, Yan Zhang, Zhengjia Ren, and Yuchen Li recruited participants and treated patients. Xiaoqi Huang and Su Lui conducted the scanning sessions and helped with data collection. Hongru Zhu, Xiaoqi Huang and Su Lui analyzed the fMRI data and wrote the manuscript. Su Lui, Qiyong Gong and Wei Zhang provided critical review and revisions of the manuscript, and obtained research funding. All authors contributed to and have approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhu, H., Qiu, C., Meng, Y. et al. Altered Topological Properties of Brain Networks in Social Anxiety Disorder: A Resting-state Functional MRI Study. Sci Rep 7, 43089 (2017). https://doi.org/10.1038/srep43089
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep43089
This article is cited by
-
Age-dependent alterations in the coordinated development of subcortical regions in adolescents with social anxiety disorder
European Child & Adolescent Psychiatry (2024)
-
Alterations in the topological organization of the default-mode network in Tourette syndrome
BMC Neurology (2023)
-
Resting-state neuroimaging in social anxiety disorder: a systematic review
Molecular Psychiatry (2022)
-
Disrupted fronto-temporal function in panic disorder: a resting-state connectome study
Brain Imaging and Behavior (2022)
-
Altered structural balance of resting-state networks in autism
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.