Abstract
The aim of this study was to investigate the effects of three Lactobacillus plantarum strains on in-vivo small intestinal barrier function and gut mucosal gene transcription in human subjects. The strains were selected for their differential effects on TLR signalling and tight junction protein rearrangement, which may lead to beneficial effects in a stressed human gut mucosa. Ten healthy volunteers participated in four different intervention periods: 7-day oral intake of either L. plantarum WCFS1, CIP104448, TIFN101 or placebo, proceeded by a 4 weeks wash-out period. Lactulose-rhamnose ratio (an indicator of small intestinal permeability) increased after intake of indomethacin, which was given as an artificial stressor of the gut mucosal barrier (mean ratio 0.06 ± 0.04 to 0.10 ± 0.06, p = 0.001), but was not significantly affected by the bacterial interventions. However, analysis in small intestinal biopsies, obtained by gastroduodenoscopy, demonstrated that particularly L. plantarum TIFN101 modulated gene transcription pathways related to cell-cell adhesion with high turnover of genes involved in tight- and adhesion junction protein synthesis and degradation (e.g. actinin alpha-4, metalloproteinase-2). These effects were less pronounced for L. plantarum WCFS1 and CIP104448. In conclusion, L. plantarum TIFN101 induced the most pronounced probiotic properties with specific gene transcriptional effects on repair processes in the compromised intestine of healthy subjects.
Similar content being viewed by others

Introduction
Impairment of intestinal barrier function has been implicated as an early event in the pathogenesis of various intestinal and systemic disorders. It may lead to increased permeation of substances present in the gut lumen, such as bacteria and their products (e.g. lipopolysaccharides), into the mucosal layer and may result in local and systemic inflammatory responses1,2,3. This process is assumed to play a role in the pathophysiology of organic gastro-intestinal (GI) disorders such as inflammatory bowel disease4,5,6, and celiac disease7, but also of functional GI disorders, such as irritable bowel syndrome8,9. Furthermore, it has been associated with systemic disorders, such as diabetes mellitus10, atopic eczema11, liver disease12, as well as the use of medication, such as non-steroidal anti-inflammatory drugs (NSAIDs)13,14. NSAIDs are frequently consumed worldwide but have also been used in research as a model to stress the intestinal mucosal layer and barrier function in healthy subjects15. Up to now, no therapeutic agents have been developed that are able to successfully restore the intestinal mucosal barrier and thereby influence disease outcome16. Positive effects on intestinal barrier function have been attributed to probiotic bacteria17,18,19,20,21,22,23. However, further research is needed to prove these beneficial effects in humans and to gain further insight into the mechanisms through which live bacterial organisms improve the human gut barrier function24,25.
In a healthy intestinal homeostasis commensal non-pathogenic bacteria interact with host mucosa and thereby presumably co-regulate mucosal barrier function. Lactobacillus plantarum, a commensal bacterium present in humans, has been reported to reinforce the intestinal barrier and to reduce intestinal permeability in animal studies18,19,20. In a previous study by our consortium, the administration of L. plantarum WCFS1 directly into the duodenum of healthy human subjects beneficially contributed to the organization of the epithelial tight junctions17. Next to L. plantarum WCFS1, we have selected two more L. plantarum strains, i.e. CIP104448 (CIP48), and TIFN101 (in previous studies referred to as CIP104450), to be tested in the current study. They were selected based on research wherein these bacterial strains were shown to have differential effects in toll-like receptor (TLR) signalling in vitro and to affect both the innate and specific immune system both in vitro and in vivo26,27,28. Although it has recently been demonstrated in an in vitro model that TLR signalling is involved in regulation of the intestinal barrier function29, it is unknown whether these three L. plantarum strains have positive effects on intestinal mucosal barrier in human subjects after ingestion, or through which mechanisms these live bacterial organisms may improve the human gut barrier function.
We hypothesized that daily oral administration of L. plantarum WCFS1, CIP48, and TIFN101 over a 7-day period (i) will have positive effects on compromised (due to intake of NSAIDs) intestinal barrier function in healthy human subjects, with regard to intestinal permeability, tight junction protein- and gene expression, and furthermore (ii) will induce alterations in the transcription of pathways involved in mucosal structure and cell function, as measured in duodenal biopsy material.
Aim of the present randomized, double-blind, placebo-controlled trial was to assess the effects of oral intake of the three L. plantarum strains over a 7-day period on small intestinal permeability, duodenal epithelial tight junction protein expression and mucosal gene transcription, in healthy human subjects.
Results
Ten healthy volunteers, 7 female and 3 male, with the age between 19 and 48 years (mean 26.3 ± 10.1) and a BMI between 16 and 24 kg/m2 (mean 21.8 ± 2.40) were included in the trial and underwent four intervention periods in a double blind randomised cross over design (Fig. 1). None of the participants reported discomfort or possible side effects during the test and follow-up period, nor have we found any significant differences in the outcomes of the self-report questionnaires, i.e. GSRS and daily symptom diary, before and after supplementation or between the placebo and the three L. plantarum test periods (data not shown). One subject did not undergo two of the four gastroduodenoscopies (during the L. plantarum TIFN101 test period) due to anticipation of discomfort. These data were considered as missing during analyses.
Three and one-day prior to start of supplementation period, small intestinal permeability (L/R ratio) was assessed; at day-3 without, and at day 0 with indomethacin induced mucosal stress. Subsequently, a 7-day supplementation period followed, with oral intake of placebo or one of the three L. plantarum strains twice a day. At day 7 the permeability test with intake of indomethacin was repeated and biopsies were obtained using flexible gastroduodenoscopy.
Intestinal permeability
L/R ratio of the baseline test with indomethacin stressor (mean 0.098 ± 0.056) was significantly higher compared to baseline L/R ratio without stressor (0.058 ± 0.037, p < 0.001) (Fig. 2), particularly due to a strong increase in urinary lactulose excretion. Comparing L/R ratio between these two time points per intervention period, the L/R ratio increased during three of the four periods (Table 1 and Suppl. Figure I). For the L. plantarum WCFS1 test period this difference was not statistically significant. Furthermore, there was no statistically significant difference between L/R ratios at baseline after the indomethacin induced stress compared to this ratio assessed after 7-day supplementation with either placebo or one of the three L. plantarum strains, measured after the indomethacin induced stress. This was also true for delta L/R ratio differences between test periods (Fig. 3).
Study participants were subjected to a sugar-based permeability test. Ratio of lactulose and rhamnose (L/R ratio) was determined in 5 hr urine samples of subjects without indomethacin stressor (day-3) or with stressor (day 0) before intake of the permeability test. Mean ± SD of L/R ratio without stressor = 0.058 ± 0.037 and with stressor = 0.098 ± 0.056. Differences tested with Linear Mixed Models, adjusting for repeated measurements for each subject.
Immunofluorescence labelling: ZO-1 and occludin expression
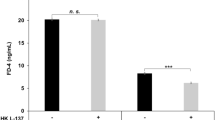
Immunofluorescent staining of the duodenal mucosa tissue samples showed expression of ZO-1 and occludin proteins at the apical part of epithelial cells, colocalized along the villous epithelium. Although the expression of both proteins seemed to increase after the interventions when compared to placebo (Fig. 4), the differences did not reach statistically significance.
Transcriptional responses of tight junction associated genes
Microarray analysis of single gene transcriptome with focus on the tight junctions and supporting proteins (Suppl. Table I) showed a significant down-regulation of the claudin 5 gene by L. plantarum WCFS1 (mean fold change (FC): −1.168, p = 0.019) and the claudin 19 gene by CIP48 (−1.146, p = 0.036), and a significant up-regulation of the actinin alpha 4 gene by TIFN101 (1.105, p = 0.032). RT-qPCR (Suppl. Figure II) was conducted for five genes, which in line microarray data, showed no statistically significant differences for claudin 3, claudin 4, MLCK, ZO-1 and occludin gene transcription between bacterial strains and placebo.
Transcriptional responses: Ingenuity Pathway Analysis
To further explore possible effects of the L. plantarum strains on the mucosal barrier, we focused on canonical pathways related to mucosal structure, enterocyte energy supply, DNA-repair, stress response and cell activity. All three interventions induced changes in transcriptional responses in the indomethacin stressed duodenal mucosa when compared to placebo.
TIFN101 demonstrated regulation of pathways involved in remodelling of the mucosal structure. Epithelial-mesenchymal transition pathway (Fig. 5), involved in disruption and remodelling of desmosomal adhesion, and pathways that might be involved in restructuring of cell-cell adhesions (i.e. macropinocytosis signalling pathway, Fig. 6) and cytoskeleton rearrangement (i.e. integrin signalling pathway, Fig. 7) were upregulated. Moreover, we observed a combined up-regulation of matrix metalloproteinase 2 (MMP2) gene (1.297, p = 0.006), involved in degradation of extracellular structures, and its inhibitors, so-called tissue inhibitors of metalloproteinase (TIMP) 1 (1.362, p = 0.006), and TIMP3 (1.243, p = 0.028). TIFN101 also upregulated the muc2 gene (1.286, p = 0.031), encoding for the mucin 2 protein. Furthermore, the glutamate-ammonia ligase (GLUL) gene (1.155, p = 0.022), involved in glutamine biosynthesis pathways, was upregulated, which may enhance energy release based on glutamine biosynthesis of mucosal cells. On the other hand this strain downregulated expression of genes involved in the tricarboxylic acid (TCA) cycle II pathway (Suppl. Figure IIIa), which may lead to reduced adenosine triphosphate (ATP) production.
WCFS1 and CIP48 downregulated nitric oxide synthase 3 gene (NOS3) (−1.113, p = 0.039 and −1.162, p = 0.018, respectively), involved in cell-cell adhesion, and MMP9 (−1.178, p = 0.045 and −1.218, p = 0.025, respectively), involved in breakdown of the extracellular matrix. CIP48 upregulated genes related to inositol phosphates biosynthesis, components of the cytosolic side of the cell membrane, such as myotubularin related protein 9 (MTMR9) (1.132, p = 0.014), MTMR8 (1.182, p = 0.043), MTMR2 (1.131, p = 0.006), phospholipase-C β4 (PLCB4) (1.131, p = 0.014), and protein tyrosine phosphatase, non-receptor type 2 (PTPN2) (1.187, p = 0.018).
WCFS1 demonstrated regulation of genes and pathways related to cell division processes and DNA repair (Suppl. Figure IIIb and c), with involvement of BRCA1 (1.225, p = 0.022) and BRCA2 genes (1.199, p = 0.018), and upregulation of protection of telomere1 (POT1) gene (1.161, p = 0.004). CIP48 showed similar effects, with upregulation of BRCA1 (1.182, p = 0.043) and 2 (1.199, p = 0.018) genes and a pathway involved in cell division and DNA-repair (Suppl. Figure IIId). Finally, CIP48 also inhibited expression of the GADD45B gene (−1.223, p = 0.034) in the glutathione-mediated detoxification pathway, involved in cell stress response, and the neuropeptide Y (NPY) gene (−1.159, p = 0.022), a protein that influences the fat metabolism.
Discussion
In the current study we investigated the effects of oral intake of three L. plantarum strains over a 7-day period on small intestinal barrier function under a stressed condition, induced by intake of NSAIDs, in healthy volunteers. Small intestinal permeability was increased after intake of indomethacin, but was not significantly affected by any of the treatments. However, the different interventions did modulate gut mucosal gene transcription related to mucosal structure, enterocyte energy supply, DNA-repair, stress response and cell activity pathways, when compared to placebo, revealing new insights into the mechanisms by which live potentially probiotic organisms may modulate human small intestinal mucosa.
Intake of indomethacin at baseline and after the intervention period resulted in a significant increase in small intestinal permeability, as reflected by the increased urinary excretion of lactulose versus rhamnose. This confirmed that the NSAID is a potent stressor of small intestinal barrier function, and that this model is well suited to investigate potential beneficial effects of bioactive substances in healthy individuals. The 7-day supplementation period of the different bacterial strains did not significantly affect small intestinal permeability. Previously, Karczewski et al. demonstrated that a continuous intraduodenal administration of L. plantarum WCFS1 over a 6-hr period in healthy subjects (without intake of NSAIDs) increased tight junction-linked localisation of occludin and ZO-1 proteins17. Our results do not confirm these findings which should be explained by pertinent differences in design such as application of indomethacin as stressor and differences in administration of bacteria. The influence of indomethacin on epithelial barrier may have been too pronounced to detect beneficial functional effects induced by the bacterial interventions. However, also differences in route of administration (i.e. oral versus intraduodenal) and exposure time (i.e. 7 days versus 6 hours) may account for the observed difference between these studies.
Although the interventions did not affect indomethacin induced changes in small intestinal permeability, the three tested L. plantarum strains did influence intrinsic repair processes of the small intestinal mucosa on gene transcription level. Differential effects on transcription of genes and pathways involved in mucosal structure maintenance have been observed. The strongest modulatory effects in this respect were demonstrated by L. plantarum TIFN101 when compared to placebo. Intake of this bacterial strain influenced the transcriptional response related to the turnover of adhesion molecules and homeostasis of the mucus layer. This is exemplified by alterations of pathways involved in cytoskeleton rearrangement, and upregulation of genes such as actinin alpha 4, several MMP genes but also their inhibitors (TIMP genes). Moreover, the up-regulation of the macropinocytosis signalling pathway, which is one of the mechanisms involved in disruption of intercellular contacts of adherens and tight junctions30,31, may lead to increased recycling of adhesive molecules. L. plantarum TIFN101 also modulated genes that may be involved in inducing a shift of energy supply of the mucosal cells from oxidative phosphorylation (by TCA II pathway downregulation) towards glutamine production (by increased transcription of glutamine biosynthesis pathways). Glutamine is an important energy source for intestinal epithelial cells and it promotes cell differentiation and proliferation32. Moreover, glutamine deprivation in intestinal epithelial cells was associated with a loss of tight junction proteins which may lead to impaired paracellular permeability33. On the other hand, glutamine supplementation has been reported to reinforce the intestinal epithelial mucosal barrier34,35. Therefore, the effect of L. plantarum TIFN101 on the glutamine biosynthesis pathways may be considered beneficial for the intestinal barrier function. Finally, the mucin 2 gene is upregulated by TIFN101. Mucin 2 is produced by the goblet cells and together with related mucin proteins is polymerized into insoluble mucus, which is an important part of the intestinal barrier36,37. As the number of goblet cells, and thereby mucin production, is increasing along the proximal to distal axis of the intestine, the effect of TIFN101 on intestinal barrier might also be stronger when it reaches more distal parts of the intestine.
L. plantarum WCFS1 showed some negative effects on the transcription of genes and pathways involved in mucosal barrier homeostasis, by downregulation of genes such as claudin 538 and NOS3. The latter gene is linked to maintenance of cytoskeletal proteins, e.g. α-tubulin, actinin and vimentin, and tight junction integral membrane proteins, e.g. occludin39. Contradictory, L. plantarum WCFS1 downregulated MMP9 gene, which leads to inhibitory effects on breakdown of extracellular matrix40, suggesting beneficial effects of L. plantarum WCFS1 supplementation on mucosal integrity. L. plantarum CIP48 demonstrated similar modest effects on gene transcription. NOS3, MMP9, and claudin 19 genes were downregulated after the intervention. Furthermore, genes involved in inositol phosphates biosynthesis, important for cell membrane structure and membrane potential41, was upregulated by L. plantarum CIP48. Thereby, both strains, L. plantarum WCFS1 and CIP48, seem to be involved in maintaining a balance between enhancement and disruption of cell-cell adhesion and mucosal structure, with no direct functional consequences for intestinal permeability and tight junction protein expression.
We identified more mechanisms by which the three tested L. plantarum strains affect the human host on transcriptome level. L. plantarum WCFS1 increased the transcription of genes such as BRCA1 and 2 and POT1, which are involved in cell division processes, DNA repair and telomere length regulation, suggesting that this bacterial strain may be involved in the enhancement of epithelial healing after indomethacin induced mucosal stress. The transcription of the NPY gene, which encodes for a neuro-hormone in the leptin signalling pathway was inhibited by L. plantarum CIP48. Antagonising NPY-ergic activity has been reported to reduce food intake42. However, no other effects on satiety related mechanisms were observed in the transcriptome analysis, and therefore the effects on satiety are probably minimal.
In summary, L. plantarum TIFN101 demonstrated regulation of genes and pathways involved in cell-cell adhesion and turnover of adhesion molecules. Furthermore, this strain seems to be able to induce gene transcriptions involved in energy supply of the mucosal cells by affecting both oxidative phosphorylation- and glutamine production pathways. L. plantarum WCFS1 modulated the transcription of genes related to DNA repair, possibly based on mucosal tissue healing upon NSAID induced stress, and showed minor negative effects on gene transcription related to mucosal structure. L. plantarum CIP48 demonstrated similar effects with respect to DNA repair mechanisms and minor positive effects on gene transcription and pathways related to mucosal structure. Overall our study demonstrates that the effect of bacterial supplementation on gene transcription is highly strain dependent and differs considerably within the same species. In the current study the transcriptome analysis was performed on duodenal biopsies. The effects might even be more pronounced or at least different in more distal parts of the intestine, as the mucosal function along the longitudinal axis of the intestine does vary. Further research is needed to investigate this hypothesis.
The cross-over design of the present study was preferred over a parallel arms design, to eliminate potential influences of inter-subject differences. An implicit potential disadvantage of a cross-over study is that previous interventions can influence effects of the next tests. To minimize such an influence a washout period of four weeks was chosen, and furthermore the order of the interventions was randomly assigned to each study participant.
In the present study the significantly regulated genes in the microarray data have not been remeasured by RT-qPCR, which may be a limitation with regard to the interpretation of the transcriptome data. In addition, future research should include in-vitro experiments to knock-down regulated genes and pathways in the present study to test the intrinsic processes described above. The difference in dose between the three L. plantarum strains used may be a potential limitation of the study. The number of colony forming unites (cfu) is determined by bacterial survival during freeze-drying of the supplements, which differed per bacterial strain. Although a dose-response is possible, it is unlikely that differences observed between interventions are affected by the difference in doses used in the current study. Furthermore, the dose consumed by study subjects per shot was for all three bacterial strains in the range of 1010, which are dosages recommended to reach clinical usefulness43.
In previous studies L. plantarum WCFS1 was found to modulate multiple pathways in duodenal mucosa, related to lipid biosynthesis, cellular proliferation, cell-death and immune responses28,44. In the present study, we were able to confirm that L. plantarum WCFS1 does affect some similar pathways related to cell function; however, most transcriptional responses were dissimilar. The main difference between the current and previous research was the intake of indomethacin in the present study. While in previous research the effects of L. plantarum WCFS1 on healthy intestinal function were investigated, we assessed effect of the three bacterial strains on small intestinal mucosa under a stressed condition. Furthermore, Troost et al. administrated a load of L. plantarum WCFS1 using oro- and nasogastric catheters during 1 and 6 hours44, and Van Baarlen et al. administered supplements of the same bacterium by oral intake, and assessed effects after 6 hours28. Therefore, besides the difference in administration route44, we measured long term effects (i.e. after a 7-day administration period), compared to the short term effects (i.e. after 1 and 6 hours) assessed in the previous studies. Troost et al. demonstrated that different supplementation periods lead to different transcriptional responses even between one and six hour administration of the bacteria44.
Comparison with other studies involving transcriptomics in the duodenum reveals that different bacterial species (i.e. L. acidophilus, L. casei, L. rhamnosus) induce very distinct gene-regulatory networks and pathways45, which were among others related to cell proliferation, hormonal regulation of blood pressure and ion homeostasis. These were not observed in the presented study, in which we used different L. plantarum strains. The described previous findings28,44,45 showed that the selection of the bacterial species as well as well as route and time period of administration of viable microbes may induce different effects on human health. In addition, taking into account these findings, our data indicate that also the selected bacterial strains and the presence of intestinal mucosal distress of the consumer may add to differences in effects of live bacteria on the intestinal mucosa. This pinpoints the need for further research in this area, but also underlines the care in selecting bacteria when testing potential management options of gastrointestinal disorders. The current study contributes to further understanding of complex host-microbe interactions and human intestinal homeostasis, providing leads for future research in this field.
Conclusion
Application of the NSAID indomethacin induced a profound increase in small intestinal permeability, which was not significantly affected by the intake of the three L. plantarum strains. However, the tested bacterial strains did modulate intrinsic repair processes on mucosal gene transcription level. L. plantarum TIFN101 demonstrated the most pronounced beneficial effects, by among others modulation of gene-transcriptions that are related to mucosal structure. L. plantarum WCFS1 and CIP48 induced more moderate transcriptional effects with respect to mucosal integrity. This illustrates the strain dependency with respect to inducing specific effects on gene transcription in human intestinal mucosa.
Materials and Methods
The study protocol has been approved by the Maastricht University Medical Center+ Committee of Ethics, was in compliance with the Declaration of Helsinki (64th WMA General Assembly, Fortaleza, Brazil, 2013), and has been registered in the US National Library of Medicine (http://www.clinicaltrials.gov, NCT01456767).
Study participants
Healthy subjects were enrolled via public advertising, and were screened, involving a standardized general physical examination. All study participants gave written informed consent prior to inclusion. Exclusion criteria were current GI symptoms, a history of severe disease or major abdominal surgery, use of any medication within 14 days or investigational drugs within 180 days prior to inclusion, blood donation within 3 months before the study period, pregnancy, lactation, obesity, smoking, use of drugs or excessive alcohol consumption, and known allergy/hypersensitivity towards intake of NSAIDs, sweeteners or pre- or probiotic supplements of any kind.
Sample size was determined using OpenEpi Sample Size Calculation for Cross-Sectional, Cohort, and Clinical Trials according to Fleiss46. Considering the primary outcome of the present cross-over study, which is lactulose and rhamnose recovery (the ratio is an indicator of intestinal permeability), based on previous studies22, taking into account an estimated effect size of 20% alteration in intestinal permeability between the interventions (three L. plantarum strains and placebo), a power of 80%, and a significance level of 5%, the study required eight completers. We included ten subjects in this study, anticipating drop-outs due to complexity and test/time burden for subjects.
Bacterial growth conditions and definition of placebo
The bacterial strains L. plantarum WCFS147,48, CIP48, and TIFN10126,27 were cultured overnight at 37 °C in Man, Rogosa and Sharpe (MRS) medium (Merck, New Jersey, USA), to obtain stationary-phase cultures. Maltodextrin and glucose were added to a final concentration of 20% and 2% (wt/vol), respectively, to obtain bacterial preparations. Detailed protocols for culturing, harvesting, freeze-drying, storing, and viable count determining of Lactobacillus species has been published previously28. Placebo controls only contained maltodextrin and glucose at similar final concentrations as bacterial supplements, ensuring a similar appearance and taste. Encoded and non-transparent vials containing the supplements were provided by NIZO Food Research (Ede, NL) to ensure the double-blind design.
Study design
This human study was designed and conducted as a randomized double-blind placebo-controlled cross-over trial. The effects of three potentially beneficial bacterial strains were investigated in each subject during four randomized test periods (Suppl. Table II). During each test period (Fig. 1), one of the following supplements has been orally ingested twice a day, during breakfast and dinner, for 7 consecutive days: L. plantarum WCFS1 (2.6 × 1010 colony forming units (cfu) per time point), L. plantarum CIP48 (2.4 × 1010 cfu/time point), L. plantarum TIFN101 (5.9 × 1010 cfu/time point) or placebo. Each test period was followed by a washout period of four weeks. There was no standardised diet for the study participants; however, subjects were instructed to consume on the last day of each test period the same diet as they did on the last day of the first period.
Baseline permeability measurements were performed three days and one day prior to the intake of the supplements, in the absence and presence of the intestinal mucosal barrier stressor indomethacin, respectively. Subsequently, subjects consumed one of the three L. plantarum strains or placebo for seven consecutive days. Thereafter, the permeability test with addition of indomethacin (but not the test without indomethacin) was repeated, whereupon the same day standard flexible gastroduodenoscopy was performed to obtain six tissue samples from the duodenum (D2 section), at approximately 15 cm distal to the pylorus. The mucosal biopsies were immediately flash frozen and used for RNA isolation followed by genome-wide microarray, and tight junction protein analysis. In line with previous research17,28,44 investigating effects L. plantarum strains, we focussed on the proximal small bowel, with minimal interference of other microbial interaction. This region of the small intestine is relatively accessible by means of endoscopy.
Possible side effects of the ingestion of the bacterial strains, adverse reactions to intake of indomethacin, and overall feelings of well-being were assessed using the validated gastrointestinal symptom rating scale (GRSR)49 pre and post test period and during the intervention by a daily symptom diary9.
In vivo small intestinal permeability test
After an overnight fast, participants emptied their urinary bladder and consumed a drink (150 ml of tap water) containing 1 g lactulose (Centrafarm, Etten-Leur, the Netherlands) and 0.5 g L-rhamnose (Danisco, Copenhagen, Denmark). Participants were requested to remain fasted, with the exception of water intake, and to collect urine for 5 hours. Lactulose (disaccharide) can permeate the intestinal epithelial layer through the paracellular route and rhamnose (mono-saccharide) through the transcellular route. The ratio in 0–5 h urine (L/R ratio) represents small intestinal permeability, and is used to minimize the effect of differences in gut motility and kidney function on the results50. During and two days prior to the test period, excessive physical exercise and consumption of alcohol were prohibited for the study participants. Three days after this baseline permeability test and after the 7-day intervention period (Fig. 1), the test was repeated with ingestion of indomethacin, a NSAID, administered in two different dosages; i.e. 75 mg exactly 9 hours prior and 50 mg 1 hour prior to the intake of the sugars14. Levels of the urinary excreted sugar probes were measured under blinded conditions by isocratic ion-exchange high-pressure liquid chromatography with mass spectrometry (HPLC-MS) as described previously50.
RNA isolation, microarray processing and pathway analysis
Total RNA was isolated form snap frozen (in liquid nitrogen) duodenal biopsies, using 1 ml Trizol reagent (Invitrogen, Breda, NL). Subsequently it was purified using Qiagen RNeasy Micro kit (Qiagen, Venlo, NL). NanoDrop ND-1000 spectrophotometer (Isogen Life Science, De Meer, NL) was used for RNA quantification, and quality was checked by the Agilent 2100 bioanalyzer (Agilent Technologies, Amsterdam, NL). Samples were obtained from ten subjects under four different conditions: L. plantarum WCFS, CIP48, TIFN101 and placebo. Off one subject two biopsies were missing yielding a total of 38 microarrays. Samples were used for further analyses in case of intact bands corresponding to 18S and 28S ribosomal subunits and in the absence of chromosomal peaks or RNA degradation products. Whole transcript cDNA synthesis of total RNA (100 ng) material was conducted using Ambion WT expression kit (Life Technologies, Bleiswijk, NL), and was labeled using the Affymetrix GeneChip WT Terminal Labelling Kit (Affymetrix, Santa Clara, CA). Next, samples were hybridized to human whole genome Affymetrix GeneChip Human Gene 1.1 ST arrays, washed, stained, and scanned on an Affymetrix GeneTitan instrument, according to the User Guide for Expression Array Plates (P/N 702933 Rev. 2).
Changes in gene expression were calculated as fold changes between placebo and the different L. plantarum strains by applying MADMAX51 for statistical comparison analyses for multiple ~omics experiments. Functional interpretation of gene signatures (pairwise IBMT p-value < 0.05) was performed using canonical pathway analysis of Ingenuity Pathway Analysis (IPA) 3.0 (Qiagen Ingenuity Systems, Redwood City, CA). Canonical pathways demonstrating significant differential regulation (threshold p < 0.05) were considered for analysis. The datasets generated from microarray profiling experiments have been deposited to the publicly accessible database repository Gene Expression Omnibus, and are available under accession number GSE74988.
Furthermore, gene transcription of tight junction proteins, i.e. claudin 3, claudin 4, myosin light chain kinase (MLCK), zonula occludens 1 (ZO-1) and occludin, in duodenal mucosal biopsies was also evaluated by RT-qPCR, as described previously52.
Expression of tight junction proteins in duodenal mucosa samples
Immunofluorescent staining for ZO-1 and occludin was performed on tissue-tek embedded frozen sections of the duodenal mucosa biopsy specimens, as described elsewhere53. Leica TCS SPE confocal laser-scanning microscope equipped with a 50-mW Argon laser and a 1-mW HeNe laser (Leica Microsystems, Wetzlar, Germany), was used to examine the slides, and ImageJ software to process and analyse the confocal images, as described previously54.
Data and statistical analysis
Statistical analysis was performed using IBM SPSS Statistics version 22.0 (IBM Statistics for Windows, Armonk, NY). To compare all baseline L/R ratio measurements without and with indomethacin stressor (assessing the effect of the stressor irrespective of the test period) Linear Mixed Models were used to take into account multiple measurements (i.e. four) per subject. To compare the effect of the 7-day supplementation of the three Lactobacilli versus placebo on urinary L/R ratio, a delta score was calculated, i.e. the L/R ratio after 7-day supplementation and with stress test minus L/R ratio of baseline with stress test. Related samples Wilcoxon Signed Rank test was used to assess differences between the interventions versus placebo and to test differences in L/R ratio between the time points per test period. A two-sided p-value of <0.05 was considered statistically significant.
Additional Information
How to cite this article: Mujagic, Z. et al. The effects of Lactobacillus plantarum on small intestinal barrier function and mucosal gene transcription; a randomized double-blind placebo controlled trial. Sci. Rep. 7, 40128; doi: 10.1038/srep40128 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Turner, J. R. Intestinal mucosal barrier function in health and disease. Nature reviews. Immunology 9, 799–809, doi: 10.1038/nri2653 (2009).
Ahrne, S. & Hagslatt, M. L. Effect of lactobacilli on paracellular permeability in the gut. Nutrients 3, 104–117, doi: 10.3390/nu3010104 (2011).
Camilleri, M., Madsen, K., Spiller, R., Greenwood-Van Meerveld, B. & Verne, G. N. Intestinal barrier function in health and gastrointestinal disease. Neurogastroenterology and motility: the official journal of the European Gastrointestinal Motility Society 24, 503–512, doi: 10.1111/j.1365-2982.2012.01921.x (2012).
Salim, S. Y. & Soderholm, J. D. Importance of disrupted intestinal barrier in inflammatory bowel diseases. Inflammatory bowel diseases 17, 362–381, doi: 10.1002/ibd.21403 (2011).
Vivinus-Nebot, M. et al. Functional bowel symptoms in quiescent inflammatory bowel diseases: role of epithelial barrier disruption and low-grade inflammation. Gut, doi: 10.1136/gutjnl-2012-304066 (2013).
Deuring, J. J., de Haar, C., Kuipers, E. J., Peppelenbosch, M. P. & van der Woude, C. J. The cell biology of the intestinal epithelium and its relation to inflammatory bowel disease. The international journal of biochemistry & cell biology 45, 798–806, doi: 10.1016/j.biocel.2012.12.020 (2013).
Heyman, M., Abed, J., Lebreton, C. & Cerf-Bensussan, N. Intestinal permeability in coeliac disease: insight into mechanisms and relevance to pathogenesis. Gut 61, 1355–1364, doi: 10.1136/gutjnl-2011-300327 (2012).
Martinez, C. et al. Diarrhoea-predominant irritable bowel syndrome: an organic disorder with structural abnormalities in the jejunal epithelial barrier. Gut 62, 1160–1168, doi: 10.1136/gutjnl-2012-302093 (2013).
Mujagic, Z. et al. Small intestinal permeability is increased in diarrhoea predominant IBS, while alterations in gastroduodenal permeability in all IBS subtypes are largely attributable to confounders. Aliment Pharmacol Ther 40, 288–297, doi: 10.1111/apt.12829 (2014).
Bosi, E. et al. Increased intestinal permeability precedes clinical onset of type 1 diabetes. Diabetologia 49, 2824–2827, doi: 10.1007/s00125-006-0465-3 (2006).
Barba, A. et al. Intestinal permeability in patients with atopic eczema. The British journal of dermatology 120, 71–75 (1989).
Cariello, R. et al. Intestinal permeability in patients with chronic liver diseases: Its relationship with the aetiology and the entity of liver damage. Digestive and liver disease: official journal of the Italian Society of Gastroenterology and the Italian Association for the Study of the Liver 42, 200–204, doi: 10.1016/j.dld.2009.05.001 (2010).
Kerckhoffs, A. P. et al. Intestinal permeability in irritable bowel syndrome patients: effects of NSAIDs. Digestive diseases and sciences 55, 716–723, doi: 10.1007/s10620-009-0765-9 (2010).
Troost, F. J., Saris, W. H. & Brummer, R. J. Recombinant human lactoferrin ingestion attenuates indomethacin-induced enteropathy in vivo in healthy volunteers. European journal of clinical nutrition 57, 1579–1585, doi: 10.1038/sj.ejcn.1601727 (2003).
Bjarnason, I., Williams, P., Smethurst, P., Peters, T. J. & Levi, A. J. Effect of non-steroidal anti-inflammatory drugs and prostaglandins on the permeability of the human small intestine. Gut 27, 1292–1297 (1986).
Odenwald, M. A. & Turner, J. R. Intestinal permeability defects: is it time to treat? Clinical gastroenterology and hepatology: the official clinical practice journal of the American Gastroenterological Association 11, 1075–1083, doi: 10.1016/j.cgh.2013.07.001 (2013).
Karczewski, J. et al. Regulation of human epithelial tight junction proteins by Lactobacillus plantarum in vivo and protective effects on the epithelial barrier. American journal of physiology. Gastrointestinal and liver physiology 298, G851–859, doi: 10.1152/ajpgi.00327.2009 (2010).
Mao, Y. et al. The effects of Lactobacillus strains and oat fiber on methotrexate-induced enterocolitis in rats. Gastroenterology 111, 334–344 (1996).
White, J. S. et al. The probiotic bacterium Lactobacillus plantarum species 299 reduces intestinal permeability in experimental biliary obstruction. Letters in applied microbiology 42, 19–23, doi: 10.1111/j.1472-765X.2005.01800.x (2006).
Mennigen, R. et al. Probiotic mixture VSL#3 protects the epithelial barrier by maintaining tight junction protein expression and preventing apoptosis in a murine model of colitis. American journal of physiology. Gastrointestinal and liver physiology 296, G1140–1149, doi: 10.1152/ajpgi.90534.2008 (2009).
Mangell, P. et al. Lactobacillus plantarum 299v inhibits Escherichia coli-induced intestinal permeability. Dig Dis Sci 47, 511–516 (2002).
Zeng, J. et al. Clinical trial: effect of active lactic acid bacteria on mucosal barrier function in patients with diarrhoea-predominant irritable bowel syndrome. Alimentary pharmacology & therapeutics 28, 994–1002, doi: 10.1111/j.1365-2036.2008.03818.x (2008).
Wang, J. H., Bose, S., Kim, H. G., Han, K. S. & Kim, H. Fermented Rhizoma Atractylodis Macrocephalae alleviates high fat diet-induced obesity in association with regulation of intestinal permeability and microbiota in rats. Scientific reports 5, 8391, doi: 10.1038/srep08391 (2015).
Shanahan, F. & Quigley, E. M. Manipulation of the microbiota for treatment of IBS and IBD-challenges and controversies. Gastroenterology 146, 1554–1563, doi: 10.1053/j.gastro.2014.01.050 (2014).
Sokol, H. Probiotics and antibiotics in IBD. Dig Dis 32 Suppl 1, 10–17, doi: 10.1159/000367820 (2014).
Meijerink, M. et al. Identification of genetic loci in Lactobacillus plantarum that modulate the immune response of dendritic cells using comparative genome hybridization. PloS one 5, e10632, doi: 10.1371/journal.pone.0010632 (2010).
van Hemert, S. et al. Identification of Lactobacillus plantarum genes modulating the cytokine response of human peripheral blood mononuclear cells. BMC microbiology 10, 293, doi: 10.1186/1471-2180-10-293 (2010).
van Baarlen, P. et al. Differential NF-kappaB pathways induction by Lactobacillus plantarum in the duodenum of healthy humans correlating with immune tolerance. Proceedings of the National Academy of Sciences of the United States of America 106, 2371–2376, doi: 10.1073/pnas.0809919106 (2009).
Vogt, L. M. et al. Toll-like receptor 2 activation by beta2–>1-fructans protects barrier function of T84 human intestinal epithelial cells in a chain length-dependent manner. The Journal of nutrition 144, 1002–1008, doi: 10.3945/jn.114.191643 (2014).
Ivanov, A. I., Nusrat, A. & Parkos, C. A. Endocytosis of epithelial apical junctional proteins by a clathrin-mediated pathway into a unique storage compartment. Molecular biology of the cell 15, 176–188, doi: 10.1091/mbc.E03-05-0319 (2004).
Feick, P., Haas, S. R., Singer, M. V. & Bocker, U. Low-dose exposure of intestinal epithelial cells to formaldehyde results in MAP kinase activation and molecular alteration of the focal adhesion protein paxillin. Toxicology 219, 60–72, doi: 10.1016/j.tox.2005.11.004 (2006).
Rhoads, M. Glutamine signaling in intestinal cells. JPEN. Journal of parenteral and enteral nutrition 23, S38–40 (1999).
Li, N., Lewis, P., Samuelson, D., Liboni, K. & Neu, J. Glutamine regulates Caco-2 cell tight junction proteins. American journal of physiology. Gastrointestinal and liver physiology 287, G726–733, doi: 10.1152/ajpgi.00012.2004 (2004).
Vermeulen, M. A., de Jong, J., Vaessen, M. J., van Leeuwen, P. A. & Houdijk, A. P. Glutamate reduces experimental intestinal hyperpermeability and facilitates glutamine support of gut integrity. World journal of gastroenterology: WJG 17, 1569–1573, doi: 10.3748/wjg.v17.i12.1569 (2011).
Wang, B. et al. Glutamine and intestinal barrier function. Amino acids, doi: 10.1007/s00726-014-1773-4 (2014).
Gouyer, V. et al. Delivery of a mucin domain enriched in cysteine residues strengthens the intestinal mucous barrier. Scientific reports 5, 9577, doi: 10.1038/srep09577 (2015).
Shan, Y. S. et al. Suppression of mucin 2 promotes interleukin-6 secretion and tumor growth in an orthotopic immune-competent colon cancer animal model. Oncology reports 32, 2335–2342, doi: 10.3892/or.2014.3544 (2014).
Kim, C. Y. & Kim, K. H. Curcumin prevents leptin-induced tight junction dysfunction in intestinal Caco-2 BBe cells. The Journal of nutritional biochemistry 25, 26–35, doi: 10.1016/j.jnutbio.2013.08.011 (2014).
Lee, N. P. & Cheng, C. Y. Regulation of Sertoli cell tight junction dynamics in the rat testis via the nitric oxide synthase/soluble guanylate cyclase/3′,5′-cyclic guanosine monophosphate/protein kinase G signaling pathway: an in vitro study. Endocrinology 144, 3114–3129, doi: 10.1210/en.2002-0167 (2003).
Bhattacharyya, S. & Tobacman, J. K. Arylsulfatase B regulates colonic epithelial cell migration by effects on MMP9 expression and RhoA activation. Clinical & experimental metastasis 26, 535–545, doi: 10.1007/s10585-009-9253-z (2009).
Dakin, K. & Li, W. H. Cell membrane permeable esters of D-myo-inositol 1,4,5-trisphosphate. Cell calcium 42, 291–301, doi: 10.1016/j.ceca.2006.12.003 (2007).
Hanson, E. S. & Dallman, M. F. Neuropeptide Y (NPY) may integrate responses of hypothalamic feeding systems and the hypothalamo-pituitary-adrenal axis. Journal of neuroendocrinology 7, 273–279 (1995).
Van Niel, C. W., Feudtner, C., Garrison, M. M. & Christakis, D. A. Lactobacillus therapy for acute infectious diarrhea in children: a meta-analysis. Pediatrics 109, 678–684 (2002).
Troost, F. J. et al. Identification of the transcriptional response of human intestinal mucosa to Lactobacillus plantarum WCFS1 in vivo . BMC Genomics 9, 374, doi: 10.1186/1471-2164-9-374 (2008).
van Baarlen, P. et al. Human mucosal in vivo transcriptome responses to three lactobacilli indicate how probiotics may modulate human cellular pathways. Proceedings of the National Academy of Sciences of the United States of America 108 Suppl 1, 4562–4569, doi: 10.1073/pnas.1000079107 (2011).
Fleiss, J. L., Tytun, A. & Ury, H. K. A Simple Approximation for Calculating Sample Sizes for Comparing Independent Proportions. Biometrics 36, 343–346 (1980).
Kleerebezem, M. et al. Complete genome sequence of Lactobacillus plantarum WCFS1. Proc Natl Acad Sci USA 100, 1990–1995, doi: 10.1073/pnas.0337704100 (2003).
Vesa, T., Pochart, P. & Marteau, P. Pharmacokinetics of Lactobacillus plantarum NCIMB 8826, Lactobacillus fermentum KLD, and Lactococcus lactis MG 1363 in the human gastrointestinal tract. Alimentary pharmacology & therapeutics 14, 823–828 (2000).
Svedlund, J., Sjodin, I. & Dotevall, G. GSRS–a clinical rating scale for gastrointestinal symptoms in patients with irritable bowel syndrome and peptic ulcer disease. Dig Dis Sci 33, 129–134 (1988).
van Wijck, K., van Eijk, H. M., Buurman, W. A., Dejong, C. H. & Lenaerts, K. Novel analytical approach to a multi-sugar whole gut permeability assay. Journal of chromatography. B, Analytical technologies in the biomedical and life sciences 879, 2794–2801, doi: 10.1016/j.jchromb.2011.08.002 (2011).
Lin, K. et al. MADMAX - Management and analysis database for multiple ~omics experiments. Journal of integrative bioinformatics 8, 160, doi: 10.2390/biecoll-jib-2011-160 (2011).
Keszthelyi, D. et al. Serotonergic reinforcement of intestinal barrier function is impaired in irritable bowel syndrome. Alimentary pharmacology & therapeutics 40, 392–402, doi: 10.1111/apt.12842 (2014).
Elamin, E., Masclee, A., Dekker, J. & Jonkers, D. Ethanol disrupts intestinal epithelial tight junction integrity through intracellular calcium-mediated Rho/ROCK activation. American journal of physiology. Gastrointestinal and liver physiology 306, G677–685, doi: 10.1152/ajpgi.00236.2013 (2014).
Abramoff, M. D. Image processing with ImageJ. Biophotonics International 11, 36–42 (2004).
Acknowledgements
This work was supported by a project from the Top Institute Food and Nutrition, Wageningen, The Netherlands. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Z.M. executed study, collected and analysed data and wrote manuscript, C.G. and N.W. were involved in transcriptome data interpretation, P.A.B. prepared and supplied the microbial strains, H.J.H.M.P. was involved in data collection, M.V.B. executed transcriptome analysis and was involved in data interpretation, P.d.V., A.A.M.M. and F.J.T. were involved in study design, data interpretation, and constructive review of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Mujagic, Z., de Vos, P., Boekschoten, M. et al. The effects of Lactobacillus plantarum on small intestinal barrier function and mucosal gene transcription; a randomized double-blind placebo controlled trial. Sci Rep 7, 40128 (2017). https://doi.org/10.1038/srep40128
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep40128
This article is cited by
-
The Kynurenine Pathway in Gut Permeability and Inflammation
Inflammation (2024)
-
The panda-derived Lactiplantibacillus plantarum BSG201683 improves LPS-induced intestinal inflammation and epithelial barrier disruption in vitro
BMC Microbiology (2023)
-
Protective effects of Bifidobacterium bifidum FL-228.1 on dextran sulfate sodium-induced intestinal damage in mice
European Journal of Nutrition (2023)
-
NSAID-enteropathy and intestinal microbes
Inflammopharmacology (2021)
-
Whole genome and acid stress comparative transcriptome analysis of Lactiplantibacillus plantarum ZDY2013
Archives of Microbiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.