Abstract
Aβ amyloid fibrils, which are related to Alzheimer’s disease, have a cross-β structure consisting of two β-sheets: β1 and β2. The Aβ peptides are thought to be serially arranged in the same molecular conformation along the fibril axis. However, to understand the amyloid extension mechanism, we must understand the amyloid fibril structure and fluctuation at the fibril end, which has not been revealed to date. Here, we reveal these features by all-atom molecular dynamics (MD) simulations of Aβ42 and Aβ40 fibrils in explicit water. The structure and fluctuation were observed to differ between the two ends. At the even end, the Aβ peptide always took a closed form wherein β1 and β2 were closely spaced. The Aβ peptide fluctuated more at the odd end and took an open form wherein the two β-sheets were well separated. The differences are attributed to the stronger β-sheet formation by the β1 exposed at the even end than the β2 exposed at the odd end. Along with the small fluctuations at the even end, these results explain why the fibril extends from one end only, as observed in experiments. Our MD results agree well with recent observations by high-speed atomic force microscopy.
Similar content being viewed by others
Introduction
Amyloid fibrils, insoluble fibrous aggregates of misfolded proteins or peptides, are associated with approximately 40 human neurodegenerative diseases1,2,3,4. For example, Alzheimer’s disease is related to amyloid-β (Aβ) peptides, Huntington’s disease is caused by polyglutamine tracts, and dialysis-related amyloidosis is caused by β2-microglobulin. The Aβ peptide has 40–43 amino acid residues, and assembles into amyloid fibrils with a cross-β structure comprising two β-sheets, β1 and β2, as shown in Fig. 1(a) 5,6,7. The Aβ peptides arrange in an orderly array with the same confirmation along the amyloid fibril axis. Because the H and O atoms of the odd-numbered (even-numbered) residues in β1 are exposed at the right (left) side of this arrangement, this end is called the odd (even) end8. Although each end is known to expose a different side of the Aβ peptide to the solvent6, the different molecular structures and kinetics between the odd and even ends have not yet been reported.
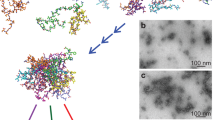
(a) Experimental conformation of the Aβ42 fibril (PDB: 2BEG). (b) Final conformation of one MD simulation. Side views of the Aβ42 monomer at (c) the even end and (d) the odd end. Residues D1 to K16 are not shown in the side views. The figures were created using PyMOL60. See Movie 1.
An amyloid fibril extends by adding another peptide to its end9,10. To understand the molecular mechanism underlying amyloid fibril elongation, we must reveal the atomic structure of the fibril ends. This knowledge is particularly important for drug design as blocking the fibril ends would prevent fibril elongation. The fibril end forms the interface between the amyloid fibril and solution and may generally adopt a different molecular structure and character from the peptides in the bulk region. The scenario is similar to the surface reconstruction of crystals such as silicon or gold in solution11 and polarization on water surface12. In an amyloid fibril, the fibril ends interface with the solution, while the bulk comprises the center region of the fibril far from the ends. The atomic conformation in the bulk region has been experimentally elucidated by solid-state NMR spectroscopy. In contrast, the conformation at the fibril end has not been experimentally revealed because amyloid fibrils are essentially one-dimensional with zero-dimensional (vanishingly small) ends. Therefore, the fibril-end structure must be clarified by alternative techniques such as molecular dynamics (MD) simulation.
Several MD simulation studies have revealed the aggregation13,14,15,16,17,18,19,20,21,22,23,24 and disaggregation25,26,27 of Aβ amyloid fibrils, and a few MD studies have reported easy deformation of the Aβ molecules at the two fibril ends under high temperature (398 K) conditions28,29. However, the structural and fluctuational differences between the odd and even ends have not been investigated. To reveal these differences, the present study conducts MD simulations on Aβ amyloid fibrils composed of 20 Aβ42 peptides and 20 Aβ40 peptides in explicit water. We report, for the first time, the structural and fluctuational differences between the two ends of the Aβ amyloid fibril.
Surface science has become well established in solid-state physics and solution chemistry, but not in amyloid fibril research. In this study, we predict the Aβ molecular structures at the fibril ends, which have not been experimentally determined, from a surface-science perspective.
Results and Discussion
Structure and fluctuation of Aβ amyloid fibril
Movie 1 of Supplementary Information (SI) shows a typical MD simulation of the Aβ42 amyloid fibril. In the initial conformation, the two β-sheets were closely spaced. Some time later, the N- and C-termini opened at the odd end but remained closed at the even end (panels (b,c and d) of Fig. 1). In all simulations, the odd end often opened, whereas the even end never opened.
Figure 2(b) shows time series of the root mean square deviations (RMSDs) of the Aβ42 peptides at both ends and in the center region. A typical snapshot is shown in Fig. 2(a). Fluctuations were largest at the odd end and slight at the center region and the even end. Figure 2(c and d) plot the average RMSDs over the nine initial conditions for Aβ42 and Aβ40, respectively, at times ranging from 100 to 200 ns. The fluctuations were clearly larger at the odd end than at the even end and were smallest in the center region. These differences were statistically significant and appeared in both Aβ42 and Aβ40 fibrils.
(a) Snapshot of a Aβ42 amyloid fibril. (b) Time series of RMSD of an Aβ42 peptide from the NMR conformation at both ends and in the center region, obtained from one MD trajectory. Average RMSD of (c) Aβ42 and (d) Aβ40 amyloid fibrils. (a) was created using PyMOL60.
Figure 3(b) shows the time series of the Cα-Cα distance between A21 and V36 at both ends and in the center region, calculated during the MD trajectory of Fig. 2(b). The pair of Cα atoms of A21 and V36 is illustrated in Fig. 3(a). The Cα-Cα distance notably increased with time at the odd end, fluctuated with no distinct temporal trend at the even end, and was essentially constant in the center region. Panels (c) and (d) of Fig. 3 plot the averages of three Cα-Cα distances along the peptide lengths. At the even end, these distances were indistinguishable from those in the center region, indicating that the two β-sheets were closely spaced. At the odd end, these distances were noticeably greater, indicating that the β-sheets were well separated. Consistent with this finding, the N- and C-termini at the odd end were far apart, as shown in Fig. 1(d).
(a) Side view of the experimental conformation of the Aβ42 amyloid fibril (chain C of model 1 of PDB: 2BEG). (b) Time series of Cα-Cα distance between A21 and V36 at both ends and in the center region. Average Cα-Cα distances of the (c) Aβ42 and (d) Aβ40 amyloid fibrils between F19 and G38 (orange), A21 and V36 (purple), and D23 and L34 (green). (a) was created using PyMOL60.
Free-energy landscape
The two-dimensional free-energy landscape  was calculated as a function of the Cα-Cα distance
was calculated as a function of the Cα-Cα distance  between A21 and V36 and the angle
between A21 and V36 and the angle  formed by three Cα atoms of K28, A30, and I32, as shown in Fig. 4. The angle
formed by three Cα atoms of K28, A30, and I32, as shown in Fig. 4. The angle  , which indicates the swelling of the loop between the two β-sheets, was 68° in the NMR conformation (Fig. 3(a)). In the center region, the highly probable conformation was distributed around the experimental conformation of
, which indicates the swelling of the loop between the two β-sheets, was 68° in the NMR conformation (Fig. 3(a)). In the center region, the highly probable conformation was distributed around the experimental conformation of  Å and
Å and  , as shown in Fig. 4(b and b’). Although some probability distribution appears in the wide-angle area of
, as shown in Fig. 4(b and b’). Although some probability distribution appears in the wide-angle area of  , the probability is higher at both ends. The even end shows a similar vertically elongated landscape as the center region with high probability distributed throughout the wide-angle area of
, the probability is higher at both ends. The even end shows a similar vertically elongated landscape as the center region with high probability distributed throughout the wide-angle area of  . This landscape is broadly distributed over
. This landscape is broadly distributed over  . At the odd end, the free-energy landscape is widely distributed both vertically and horizontally. The Cα-Cα distance between A21 and V36 can exceed 20 Å only at the odd end.
. At the odd end, the free-energy landscape is widely distributed both vertically and horizontally. The Cα-Cα distance between A21 and V36 can exceed 20 Å only at the odd end.
Two-dimensional free-energy landscape as a function of the distance  between the two Cα atoms of A21 and V36 and the angle
between the two Cα atoms of A21 and V36 and the angle  formed by the three Cα atoms of K28, A30, and I32 for the (a–c) Aβ42 and (a’–c’) Aβ40 amyloid fibrils (unit = kcal/mol) (a and a’) at the even end, (b and b’) in the center region, and (c and c’) at the odd end.
formed by the three Cα atoms of K28, A30, and I32 for the (a–c) Aβ42 and (a’–c’) Aβ40 amyloid fibrils (unit = kcal/mol) (a and a’) at the even end, (b and b’) in the center region, and (c and c’) at the odd end.
Because the odd end took both closed and open form, we calculated the the fractions of the open and closed forms, as listed in Table 1. Here, when the Cα-Cα distance  in Fig. 4 was longer than 20 Å, we regarded it as the open form. Free energy difference ΔFclosed→open from the closed forms to the open is also shown. Free energy difference ΔFclosed→open is the same order of room temperature (kBT = 0.59 kcal/mol), and the open form is often taken at the odd end.
in Fig. 4 was longer than 20 Å, we regarded it as the open form. Free energy difference ΔFclosed→open from the closed forms to the open is also shown. Free energy difference ΔFclosed→open is the same order of room temperature (kBT = 0.59 kcal/mol), and the open form is often taken at the odd end.
Figure 5 presents typical conformations at points A–D in Fig. 4(a–c). In conformation B, which is common in the bulk region, the N- and C-termini are closely spaced, and the side chains are closely packed between the two β-sheets, similar to the PDB conformation in Fig. 2(a). At the even end, conformations A and B are both observed. The loop region of conformation A is swollen, and some of its side chains (I32 and L34) protrude from between the two β-sheets. However, the N- and C-termini remain close, as in conformation B. The Cα-Cα distance  was almost identical at the even end and in the center region (Fig. 3(c)), despite the slightly larger RMSD at the even end than in the center (see Fig. 2(c)). This difference is attributed to fluctuations in the loop region. The even end fluctuates as conformation A while maintaining a short Cα-Cα distance. Conformations C and D are found only at the odd end. The N- and C-termini are slightly opened in conformation C, and decidedly opened in conformation D.
was almost identical at the even end and in the center region (Fig. 3(c)), despite the slightly larger RMSD at the even end than in the center (see Fig. 2(c)). This difference is attributed to fluctuations in the loop region. The even end fluctuates as conformation A while maintaining a short Cα-Cα distance. Conformations C and D are found only at the odd end. The N- and C-termini are slightly opened in conformation C, and decidedly opened in conformation D.
Typical conformations of the Aβ42 peptide.
The figures were created using PyMOL60.
Reason why only the odd end opens
The larger fluctuations at both ends than in the bulk have been already reported in previous MD studies28,29. This finding is relatively trivial because both ends (unlike the bulk) are exposed to the solvent. However, the differences between the two ends are nontrivial. To understand these differences, we calculated the probability that each amino acid residue forms an intermolecular parallel β-sheet by using Define Secondary Structure of Proteins (DSSP)30. The probability distributions of the intermolecular parallel β-sheets are shown in Fig. 6. Because DSSP was used, formation of intermolecular β-sheet indicates that intermolecular hydrogen bonds were formed. In both Aβ42 and Aβ40 amyloid fibrils, residues V18 to D23 and I31 to V36 (delineated by the black dotted rectangles) form intermolecular parallel β-sheets with high probability. The former group is β1; the latter is β2. Because the glycine residue G33 in β2 takes a wide range of dihedral angles ϕ and ψ, the β1 region forms a β-sheet with higher probability than β2. According to the PDB conformation of Fig. 1(a), Aβ peptides are slightly twisted, and β2 does not lie directly below β1. Thus, β1 is more exposed than β2 at the even end, whereas β2 is more exposed than β1 at the odd end (indicated by the blue dotted ellipses in Fig. 1(a)). Both of these β-sheets, β1 at the even end and β2 at the odd end, might be easily broken in their environment. However, the amino acid residues in β1 form intermolecular hydrogen bonds with higher probability than those in β2. In general, a secondary structure with more hydrogen bonds fluctuates less than that with less hydrogen bonds31,32. In other words, the secondary structure with more hydrogen bonds is firmer than that with less hydrogen bonds. In this case, β1 fluctuates less than β2. β1 is like hard board, whereas β2 is like fluttering paper. β1 and β2 stick together by the hydrophobic interaction of their side chains8 except for β1 at the even end and β2 at the odd end because they are exposed to water. At the even end, β1 does not fluctuate much even if it does not sick with β2, because β1 forms firmer β-sheet by hydrogen bonds with the neighboring peptides. Figure 6 shows that β2 at the even end forms less intermolecular hydrogen bonds than β2 at the odd end. However, because it sticks with β1 of the neighboring Aβ peptide (i.e. 2nd peptide) by the hydrophobic interaction, β2 does not fluctuate much at the even end, either. On the other hand, because β2 at the odd end does not stick with β1 and forms less intermolecular hydrogen bonds than β1 at the even end, β2 at the odd end fluctuates more than β1 and β2 at the even end. Consequently, at the even end, where β1 is exposed, the Aβ peptide retains its closed forms (conformations A and B in Fig. 5) and constrains its fluctuations. At the odd end, where β2 is exposed, the Aβ peptide fluctuates comparatively widely and adopt many conformations, including the open conformations C and D.
We remark that the stagger33,34 of 2BEG, the model used here, is −1. That is, the N-terminus of Aβ peptide i interacts with the C-terminus of peptide i − 1 in the peptide numbering of Figs 2 and 3. There are other Aβ amyloid fibril models, such as 2LMN, 2LMO, 2LMP, etc., which have either more negative stagger or positive stagger. Depending on the sign of the stagger, β1 is exposed at the different end. Revealing which end opens in these models is a future research project.
Although the 3D coordinates of the Aβ peptides at the two fibril ends have not been experimentally determined, their different characteristics have been reported. Ban et al. observed that the Aβ fibrils extend only in one direction35,36. This unidirectionality of fibril extension implies different conformations of the odd and even ends, although the growing end has not been experimentally identified. Recently, Uchihashi and Konno observed a single amyloid fibril of yeast prion-protein sup35 by high-speed atomic force microscopy (AFM)37. Consistent with our MD simulations, they observed fluctuations at one end of the fibril; the other end remained steady. Furthermore, they observed that one sup35 molecule binds to the stable end, initiating elongation at that end. If the stable end also extends in the Aβ fibril, we can surmise that elongation proceeds from the even end, which fluctuated less than the odd end in our MD simulations.
In previous MD studies of Aβ fibril extension38,39, an Aβ peptide was added to either end of the fibril, while restraining the positions of the Aβ atoms at the end. According to Han and Schulten, the extension speed is 40 times faster at the even end than at the odd end because the additional Aβ peptide is locked in by the exposed hydrophobic residues at the even end38. However, Schwierz et al.39 reported a much smaller difference in the odd- and even-end extension speeds than that reported by Han and Schulten. In other simulations, residues already assembled into a β-strand easily form another strand with another peptide19,21,22. Our MD simulations revealed much less fluctuation of the even end than of the odd end, and stronger β-sheet formation by the β1 exposed at the even end. From these results, we infer that if an Aβ peptide is added to either end of the Aβ amyloid fibril without any restriction in the MD simulation, the extension speed would be much faster at the even end than at the odd end. We expect that the even end maintains a closed form in the existing intermolecular β-sheet and can readily form a new intermolecular β-sheet with the additional Aβ peptide. In contrast, the fluctuating odd end will less easily form a β-sheet with the additional Aβ peptide. The structural and fluctuational differences between the odd and even ends of the Aβ amyloid fibril may provide important insights into fibril extension.
Conclusion
We revealed the structural and fluctuational differences between the even and odd ends of the Aβ fibrils in all-atom MD simulations. The even end always takes the closed form and fluctuates much less than the odd end. The fluctuating odd end can adopt both closed and open forms. The conformational flexibility of the odd end is attributed to the exposed β2, which forms weak hydrogen bonds. Meanwhile, the β1 exposed at the even end forms stronger hydrogen bonds and is more spatially constrained than the β2 exposed at the odd end. These results can explain the unidirectionality of fibril extension: The even end easily forms new β-sheets with another Aβ peptide, because it has already formed a stable β-sheet in the amyloid fibril. Our findings well agree with the results of recent high-speed AFM experiments.
By revealing the atomic structures at the fibril ends, we can design drugs that prevent Aβ amyloid fibril extension. Methods to inhibit the amyloid fibril formation have been attempted by many researchers40,41,42,43. From our MD simulations, the Aβ amyloid fibril is expected to elongate at the even end, where β1 is exposed. It may be a good strategy to design a molecule that binds to β1 for an inhibitor. The knowledge obtained from our simulations will be useful for understanding the amyloidogenesis mechanism and for overcoming amyloid diseases.
Methods
Modeling the initial conditions
The initial conformations of the amyloid fibril with 20 Aβ42 peptides, the amino-acid sequence of which was DAEFRHDSGYEVHHQKLVFFAEDVGSNKGAIIGLMVGGVVIA, were prepared as follows: model 1 of the 2BEG PDB file8 was used. Five Aβ (17–42) peptides formed intermolecular β-sheet structures between neighboring peptides in this PDB. The two edge peptides were removed because they were slightly distorted and unsuitable for a longer amyloid fibril. Ten copies of the other three peptides were aligned by the rigid translation so that intermolecular β-sheet structures between the trimers could be formed. By minimizing the potential energy with the conjugate gradient method in vacuum, an amyloid fibril with 30 Aβ (17–42) peptides was obtained. Ten of the 30 Aβ (17–42) peptides were then removed to obtain an amyloid fibril with 20 Aβ (17–42) peptides. Removing different ten peptides, three different conformations of 20 Aβ (17–42) peptides were obtained. The amino-acid residues 1–16, the conformations of which were not determined via the NMR experiments, were added with the dihedral angles of ϕ = ψ = 180°. However, these dihedral angles were not fixed at 180°, but flexibly fluctuated during the MD simulations. The residues 1–16 took random conformations and rarely formed secondary structures, as shown in Fig. 6. N- and C-termini of the peptide were left uncapped. The initial conformations of the amyloid fibril with 20 Aβ40 peptides were prepared in a similar manner. They were also obtained from model 1 of the 2BEG PDB file, but I41 and A42 were removed; the amino-acid sequence was DAEFRHDSGYEVHHQKLVFFAEDVGSNKGAIIGLMVGGVV. Employing three different initial velocities for each initial conformation, MD simulations were performed from nine different initial conditions for both Aβ42 and Aβ40 systems (18 initial conditions in total).
The Aβ42 amyloid fibril system consisted of 20 Aβ42 peptides, 57,876 water molecules, and 60 sodium counter ions. The Aβ40 amyloid fibril system consisted of 20 Aβ40 peptides, 58,015 water molecules, and 60 sodium ions. The total number of atoms was 186,228 and 186,065 for the Aβ42 and Aβ40 systems, respectively. A cubic simulation box was employed with periodic boundary conditions. The side length of the initial simulation box was L = 124.29 Å for both systems.
Molecular dynamics simulations
MD simulations were performed by the Generalized-Ensemble Molecular Biophysics program developed by one of the authors (H.O.). This program has been applied to several biomolecules44,45,46,47. For the Aβ peptides and water models, we applied the AMBER parm99SB force field48 and the TIP3P rigid-body model49, respectively. The electrostatic potential was calculated using the particle-mesh Ewald (PME) method50. The cut-off distance was rc = 12 Å for the Lennard-Jone potential. Temperature was controlled at 298 K using the Nosé-Hoover thermostat51,52,53, and pressure was controlled at 0.1 MPa using the Andersen barostat54. The symplectic55 quaternion scheme was used for the rigid-body water molecules56,57. Reversible multiple time-step MD techniques were also applied58. The time step was taken to be Δt = 0.5 fs for the bonding interactions of the peptide atoms, Δt = 2.0 fs for the non-bonding interactions of the peptide atoms and those between the peptide atoms and solvent molecules, and Δt = 4.0 fs for the interaction between the solvent molecules. Because the symplectic rigid-body algorithm was used for the water molecules here, Δt can be taken to be as long as 4.0 fs57. We performed an MD simulation for 200 ns from each initial condition. Averages of all physical quantities were taken over the last 100 ns and nine initial conditions.
Analysis of the simulation results
Root mean square deviations (RMSD) in Fig. 2 was calculated for the backbone N, Cα, and C atoms with respect to the reference conformation. Chain C of model 1 of the NMR structure (PDB ID: 2BEG) was used for the reference conformation8. Error bars of RMSDs and Cα-Cα distances represent standard errors calculated using the bootstrap method59 for the nine MD simulations from different initial conditions. The number of bootstrap cycles was 1 × 107.
Two-dimensional free-energy landscape  in Fig. 4 was calculated from the probability distribution
in Fig. 4 was calculated from the probability distribution  as a function of the Cα-Cα distance
as a function of the Cα-Cα distance  between A21 and V36 and angle
between A21 and V36 and angle  formed by three Cα atoms of K28, A30, and I32. It is given by
formed by three Cα atoms of K28, A30, and I32. It is given by

where kB is the Boltzmann constant, T is a temperature of 298 K, and F0 is the minimum value of  . The landscapes at the even and odd ends were calculated from the probability distribution of one Aβ peptide at the respective ends. The landscape in the center region of the fibril was calculated by taking an average of six Aβ peptides in the center region.
. The landscapes at the even and odd ends were calculated from the probability distribution of one Aβ peptide at the respective ends. The landscape in the center region of the fibril was calculated by taking an average of six Aβ peptides in the center region.
Free energy difference ΔFclosed→open from the closed form to the open form in Table 1 was calculated from the fraction of these forms as

Additional Information
How to cite this article: Okumura, H. and Itoh, S. G. Structural and fluctuational difference between two ends of Aβ amyloid fibril: MD simulations predict only one end has open conformations. Sci. Rep. 6, 38422; doi: 10.1038/srep38422 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Sipe, J. D. & Cohen, A. S. Review: History of the amyloid fibril. J. Struct. Biol. 130, 88–98 (2000).
Chiti, F. & Dobson, C. M. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 75, 333–366 (2006).
Chiti, F. & Dobson, C. M. Amyloid formation by globular proteins under native conditions. Nat. Chem. Bio. 5, 15–22 (2009).
Knowles, T. P. J., Vendruscolo, M. & Dobson, C. M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 15, 384–396 (2014).
Sunde, M. et al. Common core structure of amyloid fibrils by synchrotron X-ray diffraction. J. Mol. Biol. 273, 729–739 (1997).
Petkova, A. T. et al. A structural model for Alzheimer’s β-amyloid fibrils based on experimental constraints from solid state NMR. Proc. Natl. Acad. Sci. USA 99, 16742–16747 (2002).
Yagi-Utsumi, M., Kato, K. & Nishimura, K. Conformation of Amyloid β with the Disordered N-Terminal Segment Followed by the Stable C-Terminal β Structure. PLoS ONE 11, e0146405 (2016).
Lührs, T. et al. 3D structure of Alzheimer’s amyloid-β (1–42) fibrils. Proc. Natl. Acad. Sci. USA 102, 17342–17347 (2005).
Hasegawa, K., Ono, K., Yamada, M. & Naiki, H. Kinetic modeling and determination of reaction constants of Alzheimer’s β-amyloid fibril extension and dissociation using surface plasmon resonance. Biochemistry 41, 13489–13498 (2002).
Cohen, S. I. A. et al. Proliferation of amyloid-β42 aggregates occurs through a secondary nucleation mechanism. Proc. Natl. Acad. Sci. USA 110, 9758–9763 (2013).
Kittel, C. chap. 17, 487-514 (John Wiley and Sons, Inc., New York, 2004).
Buch, V., Milet, A., Vacha, R., Jungwirth, P. & Devlin, J. P. Water surface is acidic. Proc. Natl. Acad. Sci. USA 104, 7342–7347 (2007).
Nguyen, H. D. & Hall, C. K. Molecular dynamics simulations of spontaneous fibril formation by random-coil peptides. Proc. Natl. Acad. Sci. USA 101, 16180–16185 (2004).
Nguyen, P. H., Li, M. S., Stock, G., Straub, J. E. & Thirumalai, D. Monomer adds to preformed structured oligomers of A β-peptides by a two-stage dock-lock mechanism. Proc. Natl. Acad. Sci. USA 104, 111–116 (2007).
Itoh, S. G. & Okamoto, Y. Amyloid-β (29–42) dimer formations studied by a multicanonical-multioverlap molecular dynamics simulation. J. Phys. Chem. B 112, 2767–2770 (2008).
O’Brien, E. P., Okamoto, Y., Straub, J. E., Brooks, B. R. & Thirumalai, D. Thermodynamic Perspective on the Dock-Lock Growth Mechanism of Amyloid Fibrils. J. Phys. Chem. B 113, 14421–14430 (2009).
Reddy, G., Straub, J. E. & Thirumalai, D. Dry amyloid fibril assembly in a yeast prion peptide is mediated by long-lived structures containing water wires. Proc. Natl. Acad. Sci. USA 107, 21459–21464 (2010).
Urbanc, B., Betnel, M., Cruz, L., Bitan, G. & Teplow, D. B. Elucidation of Amyloid β-Protein Oligomerization Mechanisms: Discrete Molecular Dynamics Study. J. Am. Chem. Soc. 132, 4266–4280 (2010).
Larini, L. & Shea, J.-E. Role of β-Hairpin Formation in Aggregation: The Self-Assembly of the Amyloid-β (25–35) Peptide. Biophys. J. 103, 576–586 (2012).
Itoh, S. G. & Okumura, H. Hamiltonian Replica-Permutation Method and Its Applications to an Alanine Dipeptide and Amyloid-β (29–42) Peptides. J. Comput. Chem. 34, 2493–2497 (2013).
Itoh, S. G. & Okumura, H. Dimerization Process of Amyloid-β (29–42) Studied by the Hamiltonian Replica-Permutation Molecular Dynamics Simulations. J. Phys. Chem. B 118, 11428–11436 (2014).
Chiang, H.-L., Chen, C.-J., Okumura, H. & Hu, C.-K. Transformation Between α-Helix and β-Sheet Structures of One and Two Polyglutamine Peptides in Explicit Water Molecules by Replica-Exchange Molecular Dynamics Simulations. J. Comput. Chem. 35, 1430–1437 (2014).
Gurry, T. & Stultz, C. M. Mechanism of Amyloid-β Fibril Elongation. Biochemistry 53, 6981–6991 (2014).
Vacha, R., Linse, S. & Lund, M. Surface Effects on Aggregation Kinetics of Amyloidogenic Peptides. J. Am. Chem. Soc. 136, 11776–11782 (2014).
Lemkul, J. A. & Bevan, D. R. Assessing the Stability of Alzheimer’s Amyloid Protofibrils Using Molecular Dynamics. J. Phys. Chem. B 114, 1652–1660 (2010).
Okumura, H. & Itoh, S. G. Amyloid Fibril Disruption by Ultrasonic Cavitation: Nonequilibrium Molecular Dynamics Simulations. J. Am. Chem. Soc. 136, 10549–10552 (2014).
Viet, M. H. et al. Picosecond dissociation of amyloid fibrils with infrared laser: A nonequilibrium simulation study. J. Chem. Phys. 143, 155101 (2015).
Buchete, N.-V., Tycko, R. & Hummer, G. Molecular dynamics simulations of Alzheimer’s β-amyloid protofilaments. J. Mol. Biol. 353, 804–821 (2005).
Buchete, N.-V. & Hummer, G. Structure and dynamics of parallel β-sheets, hydrophobic core, and loops in Alzheimer’s Aβ fibrils. Biophys. J. 92, 3032–3039 (2007).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 12, 2577–2637 (1983).
Oroguchi, T., Hashimoto, H., Shimizu, T., Sato, M. & Ikeguchi, M. Intrinsic dynamics of restriction endonuclease EcoO109I studied by molecular dynamics simulations and X-ray scattering data analysis. Biophys. J. 96, 2808–2822 (2009).
Inagaki, K., Satoh, T., Itoh, S. G., Okumura, H. & Kato, K. Redox-dependent conformational transition of catalytic domain of protein disulfide isomerase indicated by crystal structure-based molecular dynamics simulation. Chem. Phys. Lett. 618, 203–207 (2015).
Petkova, A. T., Yau, W. M. & Tycko, R. Experimental Constraints on Quaternary Structure in Alzheimer’s β-Amyloid Fibrils. BioChem. 45, 498–512 (2006).
GhattyVenkataKrishna, P. K., Uberbacher, E. C. & Cheng, X. Effect of the amyloid β hairpin’s structure on the handedness of helices formed by its aggregates. FEBS Lett. 587, 2649–2655 (2013).
Ban, T., Hamada, D., Hasegawa, K., Naiki, H. & Goto, Y. Direct observation of amyloid fibril growth monitored by thioflavin T fluorescence. J. Biol. Chem. 278, 16462–16465 (2003).
Ban, T. et al. Direct observation of Aβ amyloid fibril growth and inhibition. J. Mol. Biol. 344, 757–767 (2004).
Uchihashi, T. & Konno, H. The 96th Annual Meeting of the Chemical Society of Japan Kyotanabe, 1S5–13 (2016).
Han, W. & Schulten, K. Fibril Elongation by Aβ (17–42): Kinetic Network Analysis of Hybrid-Resolution Molecular Dynamics Simulations. J. Am. Chem. Soc. 136, 12450–12460 (2014).
Schwierz, N., Frost, C. V., Geissler, P. L. & Martin, Z. Dynamics of Seeded Aβ40-Fibril Growth from Atomistic Molecular Dynamics Simulations: Kinetic Trapping and Reduced Water Mobility in the Locking Step. J. Am. Chem. Soc. 138, 527–539 (2016).
Hayashi, H. et al. A Seed for Alzheimer Amyloid in the Brain. J. Neurosci. 24, 4894–4902 (2004).
Milojevic, J., Esposito, V., Das, R. & Melacini, G. Understanding the Molecular Basis for the Inhibition of the Alzheimer’s Aβ-Peptide Oligomerization by Human Serum Albumin Using Saturation Transfer Difference and Off-Resonance Relaxation NMR Spectroscopy. J. Am. Chem. Soc. 129, 4282–4290 (2007).
Yoo, S. I. et al. Inhibition of Amyloid Peptide Fibrillation by Inorganic Nanoparticles: Functional Similarities with Proteins. Angew. Chem. Int. Ed. 50, 5110–5115 (2011).
Luo, J., Wärmländer, S. K. T. S., Gräslund, A. & Abrahams, J. P. Non-chaperone Proteins Can Inhibit Aggregation and Cytotoxicity of Alzheimer Amyloid β Peptide. J. Biol. Chem. 289, 27766–27775 (2014).
Okumura, H. Partial multicanonical algorithm for molecular dynamics and Monte Carlo simulations. J. Chem. Phys. 129, 124116 (2008).
Okumura, H. & Okamoto, Y. Temperature and pressure dependence of alanine dipeptide studied by multibaric-multithermal molecular dynamics simulations. J. Phys. Chem. B 112, 12038–12049 (2008).
Okumura, H. Temperature and pressure denaturation of chignolin: Folding and unfolding simulation by multibaric-multithermal molecular dynamics method. Proteins 80, 2397–2416 (2012).
Okumura, H. & Itoh, S. G. Transformation of a design peptide between the α-helix and β-hairpin structures using a helix-strand replica-exchange molecular dynamics simulation. Phys. Chem. Chem. Phys. 15, 13852–13861 (2013).
Hornak, V. et al. Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins 65, 712–725 (2006).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Essmann, U. et al. A smooth particle mesh ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Nosé, S. A molecular dynamics method for simulations in the canonical ensemble. Mol. Phys. 52, 255–268 (1984).
Nosé, S. A unified formulation of the constant temperature molecular dynamics methods. J. Chem. Phys. 81, 511–519 (1984).
Hoover, W. G. Canonical dynamics - equilibrium phase space distributions. Phys. Rev. A 31, 1695–1697 (1985).
Andersen, H. C. Molecular dynamics simulations at constant pressure and/or temperature. J. Chem. Phys. 72, 2384–2393 (1980).
Yoshida, H. Construction of higher-order symplectic integrators. Phys. Lett. A 150, 262–268 (1990).
Miller, T. F. et al. Symplectic quaternion scheme for biophysical molecular dynamics. J. Chem. Phys. 116, 8649–8659 (2002).
Okumura, H., Itoh, S. G. & Okamoto, Y. Explicit symplectic integrators of molecular dynamics algorithms for rigid-body molecules in the canonical, isobaric-isothermal, and related ensembles. J. Chem. Phys. 126, 084103 (2007).
Tuckerman, M., Berne, B. J. & Martyna, G. J. Reversible multiple time scale molecular dynamics. J. Chem. Phys. 97, 1990–2001 (1992).
Efron, B. 1977 Rietz Lecture - Bootstrap Methods: Another Look at the Jackknife. Ann. Stat. 7, 1–26 (1979).
Schrödinger, L. L. C. The PyMOL Molecular Graphics System, Version 1.8 (2015).
Acknowledgements
This work was supported by JSPS KAKENHI (26102550), the Okazaki Orion project, and the NINS program for cross-disciplinary study. We used super-computers at the Research Center for Computational Science, Okazaki Research Facilities, National Institutes of Natural Sciences in Japan.
Author information
Authors and Affiliations
Contributions
S.G.I. modeled the initial conformations. H.O. carried out simulations, analysed the results, and wrote the paper. All authors designed the research, discussed the results, and reviewed the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Okumura, H., Itoh, S. Structural and fluctuational difference between two ends of Aβ amyloid fibril: MD simulations predict only one end has open conformations. Sci Rep 6, 38422 (2016). https://doi.org/10.1038/srep38422
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep38422
This article is cited by
-
Identification of Bile Acid-Derived Chemical Chaperone(s) Targeting E46K-Mutated Alpha-Synuclein Protein to Treat Parkinson’s Disease: Molecular Modelling, Docking, ADME, and Simulation Studies
Applied Biochemistry and Biotechnology (2023)
-
Conformational Change of Amyloid-β 40 in Association with Binding to GM1-Glycan Cluster
Scientific Reports (2019)
-
Alkali ion influence on structure and stability of fibrillar amyloid-β oligomers
Journal of Molecular Modeling (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.