Abstract
Similar to the SNP (single nucleotide polymorphism) data, there is non-random association of the DNA methylation level (we call it methylation disequilibrium, MD) between neighboring methylation loci. For the case-control study of complex diseases, it is important to identify the association between methylation levels combination types (we call it methylecomtype) and diseases/phenotypes. We extended the classical framework of SNP haplotype-based association study in population genetics to DNA methylation level data, and developed a software EWAS to identify the disease-related methylecomtypes. EWAS can provide the following basic functions: (1) calculating the DNA methylation disequilibrium coefficient between two CpG loci; (2) identifying the MD blocks across the whole genome; (3) carrying out case-control association study of methylecomtypes and identifying the disease-related methylecomtypes. For a DNA methylation level data set including 689 samples (354 cases and 335 controls) and 473864 CpG loci, it takes only about 25 min to complete the full scan. EWAS v1.0 can rapidly identify the association between combinations of methylation levels (methylecomtypes) and diseases. EWAS v1.0 is freely available at: http://www.ewas.org.cn or http://www.bioapp.org/ewas.
Similar content being viewed by others
Introduction
DNA methylation is an important epigenetic modification by adding a methyl group to the 5 position of cytosine, and forming 5-methylcytosine (5-mC)1,2. The DNA methylation associates with a number of key processes, such as the regulation of gene expression3,4, aging5,6, genomic imprinting7,8 and development9,10,11.
Recently years, with the development of chip and sequencing technology, such as Illumina Infinium HumanMethylation450 BeadChip (450 K methylation array) and bisulfite sequencing technology, it is possible for us to measure DNA methylation on a genome wide scale. Only for the GPL13534 platform 450k platform, including 485,577 probes) in NCBI GEO database12,13, at present, there are about 648 series including more than 42789 samples, and the number of samples is increasing rapidly. Under this background, the epigenome-wide association studies are developed. The epigenome-wide association studies aim to systematically identify epigenetic variants associated with complex diseases or phenotypes for case-control studies14,15. So far, many complex diseases have been successfully analyzed by using the epigenome-wide association studies, such as Parkinson’s disease15, Schizophrenia16, Psoriasis17.
Similar to linkage disequilibrium (LD) of single nucleotide polymorphisms (SNP), there is non-random association of the DNA methylation level between neighboring methylation loci. Here, we call the non-random association methylation disequilibrium (MD). Some previous studies also indicated that the decay distance of MD is about 1 kb ~10 kb18,19. For the case-control study of SNP data, a classical analysis is to identify complex disease-related haplotype (the combination of SNP alleles). The most popular software that can identify the disease-related haplotype is Haploview20. Here we extended the classic framework of haplotype-based association study in Haploview to DNA methylation data. Our aim is to develop a software EWAS that can identify the association between combinations of methylation levels (here, the combinations of methylation levels are called, methylecomtype) and diseases in case-control study. EWAS can be used to calculate the MD coefficient between any two DNA methylation loci, identify the MD blocks and perform the methylecomtypes-based case-control association study. We hope the classic framework of haplotype-based association study can be widely used in the epigenome-wide association studies.
Methods
DNA methylation level β value
The data used in the EWAS software is the methylation status β-values. β-value is usually defined as β = methylated signals/(methylated signals + unmethylated signals + 100). The β-values range from 0 to 1. The users can obtain the DNA methylation profile through the methylation chip (such as Illumina 450 K array) or the methylation sequencing (such as bisulfite sequencing).
The DNA methylation disequilibrium (MD)
The DNA methylation disequilibrium is derived from the concept of SNP linkage disequilibrium (LD)21,22,23,24. The LD means that the alleles of two SNPs were non-random association. Here, we changed the DNA methylation β value into two levels (H: high DNA methylation level and L: low DNA methylation level) based on some threshold T (e.g. T = 0.5), and defined the MD and related concepts as following:
(1) methylecomtype (methylation levels combination types): we defined the methylecomtype as the combinations of methylation levels (H or L). For example, we supposed that there are three CpG loci, and each locus had two levels: H and L. For all the samples, there may be total 23 = 8 types of methylecomtypes: HHH, HHL, HLH, HLL, LHH, LHL, LLH and LLL. Each sample is one of the eight types of methylecomtypes. If there is an association between loci, the types of methylecomtypes will less than eight.
(2) methylation equilibrium (ME): We supposed that there were two methylation loci M1 and M2. M1 had two levels H1 and L1, and M2 had two levels H2 and L2. If there is no association between the methylation levels of two loci M1 and M2, we believed that M1 and M2 were Methylation equilibrium (ME). In other words, the methylation level of M1 locus is independent of the methylation level of M2 locus. We then described the ME by using the principle of independence:

where pH1H2 is the frequency of methylecomtype H1H2, pL1H2 is the frequency of methylecomtype L1H2, pH1L2 is the frequency of methylecomtype H1L2, pL1L2 is the frequency of methylecomtype L1L2, pH1 is the frequency of high DNA methylation level H1 at M1 locus, pL1 is the frequency of low DNA methylation level L1 at M1 locus, pH2 is the frequency of high DNA methylation level H2 at M2 locus, and pL2 is the frequency of low DNA methylation level L2 at M2 locus. These frequencies can be calculated as follows:


(3) MD (methylation disequilibrium): We still assumed that there are two methylation loci M1 and M2. M1 had two levels: H1 and L1, and M2 had two levels H2 and L2. If there is non-random association between the methylation levels of two loci M1 and M2, we believed that M1 and M2 were Methylation disequilibrium (MD). We then gave the general methylation disequilibrium coefficient gd as follows25,26:

Standardization of gd
In this study, we provided two approaches, which are commonly used in LD analyses (LD coefficient D′ and r2), to standardize gd. The standardized MD coefficient gd′ and gr2 can be calculated as27:


The range of gD′ is from 0 to 125.
gr2 can be calculated as28:

The range of gr2 is also from 0 to 1.
MD block
In this study, we defined MD block as a chromosome region consisting of a group of closely related DNA methylation loci. That is, in a block region, DNA methylation loci were high MD with each other. Here, we used the Gabriel et al.’s algorithm to identify the MD blocks29.
Methylecomtype based association study
For each methylecomtype in a MD block, we carry out a case/control χ2 test to identify whether the methylecomtype is associated with a disease or phenotype. At the same time, we also calculate OR ratio and its 95% CI for each methylecomtype in every block.
Results
Overview of the EWAS
EWAS is written entirely using Java, and can be run on any platform which has installed Java 1.8 or later version. By using command line, EWAS is able to process whole genome DNA methylation data with hundreds of samples in acceptable time. EWAS has three basic functions: (1) calculating the MD coefficient gD′ and gr2 between any two DNA methylation loci in a genome region; (2) scanning the entire genome using an elastic sliding window to identify the MD blocks; (3) identifying the disease-related methylecomtypes. Users can easily complete the epigenome wide gMaplotype association by executing the command:

The input data
For case/control association study, the input file should include the information of sample status and DNA methylation levels. The statuses of samples were listed in the first row of the input file. In EWAS, we use 1,0 or −1 to represent case, control or unknown status. From the second row, each line in the input file is a DNA methylation locus and its detailed information including: name of the locus, chromosome number, physical position in chromosome, and DNA methylation level β-values for each of the samples. To facilitate subsequent calculation, we firstly sort the DNA methylation loci based on their chromosome number and physical position. Then all the DNA methylation level β-values were changed into two levels (H: high DNA methylation level and L: low DNA methylation level). All the sorted data were saved in a temporary file ‘out_sort.txt’. Users also can use the temporary file to carry out their personalized analysis.
If users only want to calculate the MD coefficients, identify the MD blocks, and calculate the frequencies of methylecomtypes, they only need to set the statuses of all samples to −1. The example data (example_coefficient.txt and example_block.txt) can be found in EWAS website.
Calculating the MD coefficient gD′ and gr2
EWAS can calculate the MD coefficients gD′ and gr2 (see method) between different DNA methylation loci. The gD′ and gr2 are used to measure whether there is association between DNA methylation levels at two methylation loci. The ranges of both gD′ and gr2 are from 0 to 1. Larger gD′ or gr2 represents the stronger associations of DNA methylation levels between two loci. For a given list of DNA methylation data (see ‘The input data’ section or example file ‘example_coefficient.txt’), EWAS can output MD information for each pair of DNA methylation loci. The output results include: the name of pair-wise DNA methylation loci, the chromosome number, the position, the MD coefficients gD′, the MD coefficients gr2. Users can use the following command to calculate and output MD coefficients:

Identifying MD blocks and calculating methylecomtype frequency
EWAS use the Gabriel et al.’s algorithm to identify the MD blocks29. For a chromosome region, we first calculated the MD coefficients gD′ for each pair-wise DNA methylation loci. If 95% of MD coefficients gD′ show strong MD, then this region is considered to be a MD block by EWAS. A part of Java codes and methods that identify the block can be found in Haploview20.
In this study, we use a sliding window to scan the entire genome. For the larger block, we design an elastic sliding window method to avoid breaking the real MD block structure. Firstly, we set a fixed length of sliding window (such as 20 loci) to identify the MD blocks in the window. If the last locus in the window is not located in a block, we will slide the window to the next locus. If the last locus in the window is located in an identified block, the block is likely to be broken by the border of the window. Then we increase the size of sliding window to two fold (such as 40 loci) size of fixed window. By using this strategy, we can scan the entire genome in a single run.
For each identified block, EWAS can also integrate the types of methylecomtype (methylation levels combination types) in the block region, and calculate the frequencies of methylecomtypes. For more details, see method.
For a given list of DNA methylation data (see ‘The input data’ section or example file ‘example_block.txt’), users can identify the MD blocks and calculated the frequency of methylecomtype using the following command:

The output results include: the name of a block, the chromosome number, the start and end position, and the list of DNA methylation loci in the block, the methylecomtype in the block and the frequency of methylecomtype.
Epigenome-wide association studies for methylecomtype
For a MD block region, EWAS will count the number of methylecomtype in case samples and control samples, and perform a chi-square test to identify the significant methylecomtypes associated with diseases or phenotypes. The association test is integrated into the framework of epigenome-wide methylecomtype association study.
Then we describe the complete process of epigenome-wide methylecomtype association study. There are five main steps: (1) sort the DNA methylation loci; (2) change DNA methylation β-values into two levels H and L; (3) scan the genome and identify the MD blocks; (4) calculate the frequency of methylecomtypes in MD block region; (5) carry out chi-square test to identify the significant methylecomtypes. Researchers can use the command ‘-ewas’ to automatically complete all the five steps. The output file includes the following information: the chromosome number, the name of a block, the start and end position, the list of DNA methylation loci in the block, the frequency of methylecomtype in case and control, the chi-square value, P-values for chi-square test, OR and its 95% CI.
Running time
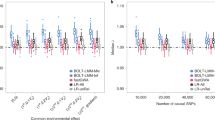
We tested the running time using a EWAS dataset (GSE42861) downloaded from GEO database (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE42861)30. The dataset includes 689 samples (354 Rheumatoid arthritis case and 335 normal controls). The DNA methylation levels of samples were tested by using Illumina 450 K array. There were total 473864 loci on the autosomal chromosomes, for each loci, the β-value was used to present the methylation status. All the tests were run on a personal computer with 2.5 GHz Intel(R) Core(TM) i3-3120M CPU and 4 G of RAM. We firstly sorted the data based on their chromosome physical location. This process takes about 12 minutes. Then we used the sorted data to carry out Epigenome-wide methylecomtype association studies (identify the MD blocks, calculate the frequency of methylecomtypes, and carry out chi-square test) for different window sizes. The range of window size is from 2 loci to 50 loci. From Fig. 1A we can see that the running time increased with the window size increased. Figure 1B showed the relationship between the window sizes and block numbers. We observed that the number of blocks reached a stable state and was almost not changed when the window size is larger than 20. Therefore, the default window size in EWAS software was set to 20. This process took about 13 minutes. This indicated that the EWAS software can process whole epigenome scale datasets in a feasible time on personal computers (PC).
Discussions
In this study, we developed a DNA methylation methylecomtype analysis tools EWAS v1.0. We extended the classic framework of SNP haplotype-based association study to DNA methylation methylecomtype-based association study. EWAS v1.0 can not only calculate the DNA methylation disequilibrium coefficient between two CpG loci, but also calculate the MD between methylation loci and identify the disease-related methylecomtypes.
In addition, EWAS v1.0 also can carry out epigenome-wide T test for beta-value of single DNA methylation locus. For a given list of DNA methylation data (see ‘The input data’ section or example file ‘example_ewas.txt’), users can calculate the T test value and p value for each methylation locus using the following command:

Due to technical limitations, we only consider the methylecomtype (methylation levels combination types). We change the DNA methylation β-values into two levels (H: high DNA methylation level and L: low DNA methylation level) based on some threshold T (e.g. T = 0.5) for analyzing the association of the DNA methylation level between neighboring methylation loci, but there is some useful information losing in this process. With the development of technology, the detection of methylaltion loci will be more accurate. We can develop some meplotype analysis modules for the further study of complex diseases.
Future
We will develop our software in the future from the following aspects:
-
1
epigenome-wide association study for SMP (single methylation polymorphism);
-
2
epigenome-wide association study for meplotype (methylation haplotype);
-
3
epigenome-wide association study for gene region;
-
4
epigenome-wide association study for KEGG pathway;
-
5
epigenome-wide association study for GO categories;
-
6
epigenome-wide association study for network;
-
7
epigenome-wide association study for interacting with genetic marker;
-
8
epigenome-wide association study for gene expresssion;
The data resource and software will be found on the EWAS (epigenome-wide association study) website: http://www.ewas.org.cn. or http://www.bioapp.org/ewas.
Additional Information
How to cite this article: Xu, J. et al. EWAS: epigenome-wide association studies software 1.0 – identifying the association between combinations of methylation levels and diseases. Sci. Rep. 6, 37951; doi: 10.1038/srep37951 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Michelotti, G. A. et al. Epigenetic regulation of human alpha1d-adrenergic receptor gene expression: a role for DNA methylation in Sp1-dependent regulation. FASEB J 21, 1979–1993, doi: fj.06-7118com 10.1096/fj.06-7118com (2007).
Jones, P. A. & Takai, D. The role of DNA methylation in mammalian epigenetics. Science 293, 1068–1070, doi: 10.1126/science.1063852 293/5532/1068 (2001).
Linn, F., Heidmann, I., Saedler, H. & Meyer, P. Epigenetic changes in the expression of the maize A1 gene in Petunia hybrida: role of numbers of integrated gene copies and state of methylation. Mol Gen Genet 222, 329–336 (1990).
Shirodkar, A. V. et al. A mechanistic role for DNA methylation in endothelial cell (EC)-enriched gene expression: relationship with DNA replication timing. Blood 121, 3531–3540, doi: 10.1182/blood-2013-01-479170 blood-2013-01-479170 (2013).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol 14, R115, doi: 10.1186/gb-2013-14-10-r115 (2013).
Jones, M. J., Goodman, S. J. & Kobor, M. S. DNA methylation and healthy human aging. Aging Cell 14, 924–932, doi: 10.1111/acel.12349 (2015).
Li, E., Beard, C. & Jaenisch, R. Role for DNA methylation in genomic imprinting. Nature 366, 362–365, doi: 10.1038/366362a0 (1993).
Reik, W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 447, 425–432, doi: nature05918 10.1038/nature05918 (2007).
Li, E., Bestor, T. H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926, doi: 0092-8674(92)90611-F (1992).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257, doi: S0092-8674(00)81656-6 (1999).
Illingworth, R. et al. A novel CpG island set identifies tissue-specific methylation at developmental gene loci. PLoS Biol 6, e22, doi: 10.1371/journal.pbio.0060022 07-PLBI-RA-3186 (2008).
Edgar, R. & Barrett, T. NCBI GEO standards and services for microarray data. Nat Biotechnol 24, 1471–1472, doi: nbt1206-1471 10.1038/nbt1206-1471 (2006).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets--update. Nucleic Acids Res 41, D991–995, doi: 10.1093/nar/gks1193 gks1193 (2013).
Verma, M. Epigenome-Wide Association Studies (EWAS) in Cancer. Curr Genomics 13, 308–313, doi: 10.2174/138920212800793294 (2012).
Moore, K., McKnight, A. J., Craig, D. & O’Neill, F. Epigenome-wide association study for Parkinson’s disease. Neuromolecular Med 16, 845–855, doi: 10.1007/s12017-014-8332-8 (2014).
Montano, C. et al. Association of DNA Methylation Differences With Schizophrenia in an Epigenome-Wide Association Study. JAMA psychiatry 73, 506–514, doi: 10.1001/jamapsychiatry.2016.0144 (2016).
Zhou, F. et al. Epigenome-Wide Association Analysis Identified Nine Skin DNA Methylation Loci for Psoriasis. J Invest Dermatol 136, 779–787, doi: 10.1016/j.jid.2015.12.029 (2016).
Schmitz, R. J. et al. Patterns of population epigenomic diversity. Nature 495, 193–198, doi: 10.1038/nature11968 (2013).
Liu, Y. et al. GeMes, clusters of DNA methylation under genetic control, can inform genetic and epigenetic analysis of disease. Am J Hum Genet 94, 485–495, doi: 10.1016/j.ajhg.2014.02.011 (2014).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265, doi: 10.1093/bioinformatics/bth457 bth457 (2005).
Balding, D. J. A tutorial on statistical methods for population association studies. Nat Rev Genet 7, 781–791, doi: nrg1916 10.1038/nrg1916 (2006).
Rosenberg, N. A. et al. Genome-wide association studies in diverse populations. Nat Rev Genet 11, 356–366, doi: nrg2760 10.1038/nrg2760 (2010).
Hirschhorn, J. N. & Gajdos, Z. K. Genome-wide association studies: results from the first few years and potential implications for clinical medicine. Annu Rev Med 62, 11–24, doi: 10.1146/annurev.med.091708.162036 (2011).
Stranger, B. E., Stahl, E. A. & Raj, T. Progress and promise of genome-wide association studies for human complex trait genetics. Genetics 187, 367–383, doi: genetics.110.120907 10.1534/genetics.110.120907 (2011).
Mueller, J. C. Linkage disequilibrium for different scales and applications. Brief Bioinform 5, 355–364 (2004).
Hill, W. G. & Robertson, A. Linkage Disequilibrium in Finite Populations. Theoretical and Applied Genetics 38, 266–231 (1968).
Lewontin, R. C. The Interaction of Selection and Linkage. I. General Considerations; Heterotic Models. Genetics 49, 49–67 (1964).
Hill, W. G. & Robertson, A. Linkage disequilibrium in finite populations. Theor. Appl. Genet 38, 226–231 (1968).
Gabriel, S. B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229, doi: 10.1126/science.1069424 (2002).
Liu, Y. et al. Epigenome-wide association data implicate DNA methylation as an intermediary of genetic risk in rheumatoid arthritis. Nat Biotechnol 31, 142–147, doi: 10.1038/nbt.2487 (2013).
Acknowledgements
This work was supported in part by the National Natural Science Foundation of China (Grant No. 31200934) and the Natural Science Foundation of Heilongjiang Province, China (Grant No. C2016036). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Jing Xu, Di Liu, Linna Zhao, Ying Li, Lin Gao and Yongshuai Jiang wrote the main manuscript text, Jing xu, Zhaoyang Wang and Changgui Lei developed the software, Zhaoyang Wang, Yang Chen analyzed the data, Yongshuai Jiang, LijunYuan and Fanwu Kong conceived and designed the experiments. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Xu, J., Liu, D., Zhao, L. et al. EWAS: epigenome-wide association studies software 1.0 – identifying the association between combinations of methylation levels and diseases. Sci Rep 6, 37951 (2016). https://doi.org/10.1038/srep37951
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37951
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.