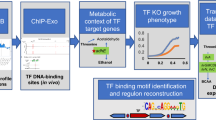
Abstract
A wide variety of Salmonella enterica serovars cause intestinal and systemic infections to humans and animals. Salmonella Patogenicity Island 1 (SPI-1) is a chromosomal region containing 39 genes that have crucial virulence roles. The AraC-like transcriptional regulator HilD, encoded in SPI-1, positively controls the expression of the SPI-1 genes, as well as of several other virulence genes located outside SPI-1. In this study, we applied a clustering method to the global gene expression data of S. enterica serovar Typhimurium from the COLOMBOS database; thus genes that show an expression pattern similar to that of SPI-1 genes were selected. This analysis revealed nine novel genes that are co-expressed with SPI-1, which are located in different chromosomal regions. Expression analyses and protein-DNA interaction assays showed regulation by HilD for six of these genes: gtgE, phoH, sinR, SL1263 (lpxR) and SL4247 were regulated directly, whereas SL1896 was regulated indirectly. Interestingly, phoH is an ancestral gene conserved in most of bacteria, whereas the other genes show characteristics of genes acquired by Salmonella. A role in virulence has been previously demonstrated for gtgE, lpxR and sinR. Our results further expand the regulon of HilD and thus identify novel possible Salmonella virulence genes.
Similar content being viewed by others
Introduction
The genus Salmonella groups Gram-negative bacteria that can infect humans and a great variety of animals, causing self-limiting enteritis or a systemic disease1,2. Salmonella comprises only two species, bongori and enterica, the latter is further divided into six subspecies and around 2500 serotypes or serovars3. S. enterica serovar Typhimurium (S. Typhimurium) can cause intestinal or systemic infections in different hosts; thus, it is widely used as a model to study the molecular virulence mechanisms of Salmonella4.
Around one-third of the Salmonella genome has been shaped by horizontal events; most of the acquired genes are clustered in regions denominated islands5,6. Salmonella pathogenicity island 1 (SPI-1) is a chromosomal region conserved in the two Salmonella species, which contains 39 genes that code for a type 3 secretion system (T3SS-1), different effector proteins and their chaperones, as well as for transcriptional regulators that control the expression of the genes within this island4,7. The T3SS and effector proteins encoded in SPI-1 are required for Salmonella invasion into intestinal epithelial cells and thus for the intestinal colonization leading to enteritis1,4,8. In vivo, the SPI-1 genes are expressed when Salmonella is in the intestinal lumen, associated with the epithelium or with extruding enterocytes9, and also in a subpopulation of Salmonella hyperreplicating in the cytosol of epithelial cells10. In vitro, the SPI-1 genes are expressed during the early stationary phase when Salmonella is grown in nutrient-rich media, such as the Luria-Bertani (LB) medium, and their expression is regulated by growth-phase, temperature, osmolarity, oxygen tension, long- and short-chain fatty acids concentration, pH, iron level and bile11,12,13,14,15,16,17,18,19.
Expression of the SPI-1 genes is controlled through a regulatory cascade formed by the transcriptional regulators HilD, HilA and InvF, encoded within this island4,20. HilD, a member of the AraC/XylS family of transcriptional regulators, directly induces the expression of HilA21,22,23, a regulator with an OmpR/ToxR-like DNA binding domain, which in turn, activates the expression of InvF24,25, another AraC/XylS-like regulator; HilA and InvF activate the expression of the SPI-1 genes encoding the T3SS-1 components and effector proteins, respectively4,20. Although HilD has a dominant role, the expression of HilA, and thus the SPI-1 genes, depends on a complex feed-forward positive regulatory loop formed by HilD, HilC and RtsA23,26; both HilC and RtsA are AraC-like regulators and bind the DNA sequence recognized by HilD27,28. HilD also induces directly, or indirectly through HilA, InvF or other regulators, the expression of several other virulence genes, including genes located in SPI-2, SPI-4, SPI-5 and other islands4,14,29,30,31,32,33,34. Furthermore, HilD directly controls the expression of the flhDC operon encoding FlhDC, the master transcriptional complex required for the expression of flagellar and chemotaxis genes30,35. Interestingly, FlhDC induces the expression of FliZ that somehow increases activity of HilD and thus the expression of the SPI-1 genes36. Therefore, HilD and FlhDC form a positive regulatory loop that co-regulates the expression of the SPI-1 and flagellar/chemotaxis genes. Important to note, like the SPI-1 genes, the flagellar/chemotaxis genes are also necessary for the invasion of Salmonella into host cells4,37,38. Thus, the HilD regulon comprises genes encoding different cellular functions that make to Salmonella an intestinal pathogen.
Gene expression is coordinated to carry out the different cellular activities; therefore, co-expression analyses can be used to infer biological functions of uncharacterized genes, as well as to identify the regulatory pathways governing the co-expression relationships. The high amount of global gene expression data that is currently available offers opportunities to investigate the gene co-expression networks in many organisms. COLOMBOS is a database that integrates a collection of transcriptomics results from both microarray and RNA-seq experiments for several prokaryotic species, including Salmonella39.
In this work, we identified genes co-expressed with the SPI-1 genes, by applying a clustering method to the S. Typhimurium SL1344 global gene expression data from the COLOMBOS database. This analysis indicates that most genes known to be directly or indirectly regulated by HilD are indeed co-expressed with the SPI-1 genes, including acquired genes located outside SPI-1 and the flagellar/chemotaxis ancestral genes. Interestingly, our results revealed nine novel genes that are co-expressed with SPI-1: gtgE, phoH, sinR, SL1344_1028 (SL1028), SL1344_1263 (SL1263 or lpxR), SL1344_1896 (SL1896), SL1344_3812 (SL3812), SL1344_4247 (SL4247) and SL1344_4433 (SL4433). Expression analyses and protein-DNA interaction assays show that HilD regulates six of these genes: gtgE, phoH, sinR, lpxR and SL4247 were regulated directly, whereas SL1896 was regulated indirectly. Sequence and genome context analyses indicate that gtgE, sinR, lpxR, SL1896 and SL4247 are genes that were acquired by Salmonella, whereas phoH is an ancestral gene. A role in virulence has been previously determined for gtgE, encoding an effector protein secreted by the two T3SSs of Salmonella; lpxR, that codes for an outer membrane protein that removes the 3´-acyloxyacyl group of lipid A; and sinR, encoding a putative LysR-family transcriptional regulator. The phoH gene encodes a protein that has an ATP-binding activity, which is conserved in most of bacteria, whereas the SL4247 and SL1896 genes encode hypothetical proteins. Therefore, our results further expand the virulence regulon of HilD and identify novel genes possibly involved in the pathogenesis of Salmonella.
Results
Identification of genes co-expressed with SPI-1
To identify genes co-expressed with SPI-1, we applied a clustering method to the genome-wide expression data of S. Typhimurium SL1344 from COLOMBOS, which generated a variety of clusters containing genes with a similar expression pattern. Then, SPI-1 genes were used independently as a bait to select those clusters that should include genes expected to be co-expressed with SPI-1. This analysis generated scores indicating the frequency with which a gene is clustered with the corresponding SPI-1 gene used as the bait (Table S1 in Supplementary File 1). For a better visualization, these data are represented in a heat map (Fig. 1). The co-expression pattern for each SPI-1 gene used as the bait show some differences; there are genes that were clustered with some SPI-1 genes but not with others (Fig. 1), which could be due to subtle differences in regulation between the SPI-1 genes or variations in the results from the global expression experiments analyzed. Therefore, the use as the bait of several SPI-1 genes increased the possibility to find genes co-expressed with SPI-1.
Genes co-expressed with SPI-1.
The heat map represents the frequency with which each gene, shown at the left side of this Fig. (co-expressed genes), was clustered with the corresponding SPI-1 gene used as the bait, shown above this Fig. The intensity of the green color in the heat map indicates the score obtained for each co-expressed gene, based in the color bar shown below the figure, ranging from Measure Values of 0 to 25. The frequency scores for all the co-expressed genes are displayed in Table S1 in Supplementary File 1. The co-expressed genes are classified in four groups: genes located in SPI-1, genes known to be co-regulated with SPI-1 that are located in other genomic islands, flagellar/chemotaxis genes and novel genes that are co-expressed with SPI-1, which are indicated with dark blue, orange, pink and brown color bars, respectively. For a better visualization, the novel genes that are co-expressed with SPI-1 are boxed. The left side of this Fig. also show whether the genes co-expressed with SPI-1 are located in any SPI. Null, indicates not located in any SPI.
Taken together, the results from our clustering analysis show that many genes known to be regulated by HilD, directly or through HilA, InvF or FlhDC, including the SPI-1 genes themselves and genes located in different genomic islands, as well as flagellar/chemotaxis genes4,34,40,41, are indeed co-expressed with SPI-1 (Fig. 1; Table S1 in Supplementary File 1). Interestingly, the gtgE, phoH, sinR, lpxR, SL1028, SL1896, SL3812, SL4247 and SL4433 genes were also found to be co-expressed with SPI-1 (Fig. 1; Table S1 in Supplementary File 1). Another clustering analysis, now using these nine genes as the bait, showed groups of co-expressed genes very similar to those obtained by using the SPI-1 genes as the bait (Fig. S1 in Supplementary File 2 and Table S2 in Supplementary File 1), further supporting the link in expression of all these genes.
Thus, these results indicate that the clustering method that we used was successful to find genes co-expressed with SPI-1, identifying gtgE, phoH, sinR, lpxR, SL1028, SL1896, SL3812, SL4247 and SL4433 as novel genes co-expressed with SPI-1.
HilD, but not HilA or InvF, regulates the expression of gtgE, phoH, sinR, lpxR, SL1896 and SL4247
To determine whether the expression of the novel genes found to be co-expressed with the SPI-1 genes is controlled by the major transcriptional regulators encoded in SPI-1, HilD, HilA and InvF, transcriptional fusions of these genes to the cat gene were constructed in plasmid pKK232–8, an expression reporter system that we have successfully used in S. Typhimurium14,42,43,44. These transcriptional fusions carry the intergenic region upstream of the respective gene tested. The chloramphenicol acetyl transferase (CAT)-specific activity directed by plasmids carrying the transcriptional fusions gtgE-cat, phoH-cat, sinR-cat, lpxR-cat, SL1028-cat, SL1896-cat, SL3812-cat, SL4247-cat or SL4433-cat, was determined in wild type (WT) S. Typhimurium strain SL1344 and its isogenic ΔhilD, ΔhilA and ΔinvF mutants, grown in LB medium a 37 °C, conditions that favor the expression of the SPI-1 genes14,42,44. As a control, the expression of a cat transcriptional fusion of invF, which is positively regulated by HilD through HilA, was also assessed. Expression of the gtgE-cat, phoH-cat, sinR-cat, lpxR-cat, SL1896-cat and SL4247-cat fusions was decreased in the ΔhilD mutant, but not in the ΔhilA and ΔinvF mutants, with respect to their expression levels shown in the WT strain (Table 1). Furthermore, the plasmid pK6-HilD, expressing HilD from an arabinose-inducible promoter, was able to increase the expression of these fusions in the ΔhilD mutant to WT levels or even higher (Fig. 2A–F). In contrast, the expression of the SL1028-cat, SL3812-cat and SL4433-cat fusions was not significantly reduced in the ΔhilD, ΔhilA or ΔinvF mutants and, as expected, the expression of the invF-cat fusion was decreased in the ΔhilD and ΔhilA mutants, but not in the ΔinvF mutant (Table 1). The SL1028-cat fusion showed a very low level of CAT activity (Table 1), which is consistent with the very low level of expression previously detected for SL1028 by RNA-seq-based transcriptomic analyses17.
HilD positively regulates gtgE, phoH, sinR, lpxR, SL1896 and SL4247 in S. Typhimurium.
Expression of the gtgE-cat (A), phoH-cat (B), sinR-cat (C), lpxR-cat (D), SL1896-cat (E) and SL4247-cat (F) transcriptional fusions contained in plasmids pgtgE-cat, pphoH-cat, psinR-cat, plpxR-cat, pSL1896-cat and pSL4247-cat, respectively, was tested in the WT S. Typhimurium strain SL1344 and its isogenic ΔhilD mutant carrying the vector pMPM-K6 or the plasmid pK6-HilD, which expresses HilD under an arabinose-inducible promoter. CAT-specific activity was determined from samples collected of bacterial cultures grown for 9 h in LB medium at 37 °C. Expression of HilD from pK6-HilD was induced by adding 0.001% L-arabinose to the medium at the beginning of the bacterial cultures. The data are the average of three independent experiments performed in duplicate. Bars represent the standard deviations. *Expression statistically different with respect to that shown by the same transcriptional fusions in the ΔhilD mutant carrying the vector pMPM-K6, P-value < 0.001.
These results show that HilD positively controls the expression of the gtgE, phoH, sinR, lpxR, SL1896 and SL4247 genes, independently of HilA and InvF.
HilD induces the expression of gtgE, phoH, sinR, lpxR and SL4247, but not that of SL1896, in the absence of other Salmonella-specific regulators
E. coli K-12 lacks around 1,400 genes present in S. Typhimurium, including several encoding transcriptional regulators, such as HilD, located in the SPIs and other Salmonella islands. Therefore, to determine whether the HilD-mediated expression of the gtgE, phoH, sinR, lpxR, SL1896 and SL4247 genes requires additional Salmonella-specific regulators, we determined the expression of the transcriptional fusions of these genes in the E. coli MC4100 strain carrying the plasmid pK6-HilD or the vector pMPM-K6Ω. As controls, the expression of cat transcriptional fusions of the hilA and ssaG genes, which are controlled by HilD directly and indirectly, respectively, were also assessed. It is important to note that the E. coli MC4100 strain carries a frameshift mutation in the flhDC operon and thus it does not express the flagellar transcriptional regulator FlhDC45,46; HilD positively regulates the expression of FlhDC in S. Typhimurium30,35. The expression of all these fusions tested was reduced in the E. coli MC4100 strain carrying the vector pMPM-K6Ω, with respect to their respective expression level shown in the WT S. Typhimurium strain (Fig. 3A–F), which is consistent with their positive regulation by HilD. The expression of HilD from plasmid pK6-HilD increased the activity of the fusions gtgE-cat, phoH-cat, sinR-cat, lpxR-cat, and SL4247-cat, similar at their respective levels reached in the WT S. Typhimurium strain or even higher (Fig. 3A–E), showing that the HilD-mediated expression of gtgE, phoH, sinR, lpxR and SL4247 does not need any other Salmonella-regulator nor FlhDC. In contrast, the expression of HilD from plasmid pK6-HilD did not increase the activity of the SL1896-cat fusion (Fig. 3F), indicating that HilD induces the expression of SL1896 through a factor not present in E. coli MC4100. As expected, the presence of HilD also induced the activity of the hilA-cat fusion, but not that of the ssaG-cat fusion (data not shown). To investigate whether HilD induces the expression of SL1896 through FlhDC, a ΔflhDC mutant derivative of S. Typhimurium SL1344 was constructed and the activity of the SL1896-cat fusion was determined in this strain. Additionally, a cat transcriptional fusion of trg, a gene regulated by FlhDC41,47, was constructed and analyzed as a control in these assays. The SL1896-cat fusion showed similar expression levels in the WT and ΔflhDC mutant strains (72 ± 9 and 82 ± 7, respectively), whereas the activity of the trg-cat fusion was drastically decreased in the ΔflhDC mutant (19 ± 8), with respect to its expression levels shown in the WT strain (154 ± 6), indicating that FlhDC is not required for the expression of SL1896 in the growth conditions tested.
HilD induces the expression of gtgE, phoH, sinR, lpxR and SL4247, but not SL1896, in E. coli MC4100. Expression of the gtgE-cat (A), phoH-cat (B), sinR-cat (C), lpxR-cat (D), SL4247-cat (E) and SL1896-cat (F) transcriptional fusions contained in plasmids pgtgE-cat, pphoH-cat, psinR-cat, plpxR-cat, pSL4247-cat and pSL1896-cat, respectively, was tested in the E. coli MC4100 strain carrying the vector pMPM-K6 or the plasmid pK6-HilD, which expresses HilD under an arabinose-inducible promoter. CAT-specific activity was determined from samples collected of bacterial cultures grown for 9 h in LB medium at 37 °C. Expression of HilD from pK6-HilD was induced by adding 0.001% L-arabinose to the medium at the beginning of the bacterial cultures. The data are the average of three independent experiments performed in duplicate. Bars represent the standard deviations. *Expression statistically different with respect to that shown by the same transcriptional fusions in the presence of the vector pMPM-K6, P-value < 0.001.
Thus, these data strongly support that HilD directly controls the expression of the gtgE, phoH, sinR, lpxR and SL4247 genes, and indirectly, through a regulator found in S. Typhimurium but not in E. coli MC4100, that of the SL1896 gene.
HilD binds to the regulatory regions of gtgE, phoH, sinR, lpxR and SL4247, but not to that of SL1896
To further define whether the HilD-mediated regulation of gtgE, phoH, sinR, lpxR, SL1896 and SL4247 is direct or indirect, we analyzed the interaction of HilD with the regulatory region of these genes. Affinity-purified maltose-binding protein (MBP)-HilD, which is active in vivo and specifically binds to HilD-target genes in vitro14,29, and the DNA fragments contained in the respective transcriptional fusion of each gene, were used to perform electrophoretic mobility shift assays (EMSAs). As a positive control, a DNA fragment containing the regulatory region of hilA was also assessed. Additionally, a DNA fragment containing the intergenic region upstream of ppk, a gene not regulated by HilD, or sigD, a gene not directly regulated by HilD, was included in the binding reactions as an internal negative control. MBP-HilD specifically bound the DNA fragments of gtgE, phoH, sinR, lpxR, SL4247 and, as expected, that of hilA, at concentrations of 0.1 to 1.0 μM; in contrast, at the same concentrations it did not bind the DNA fragment of SL1896, or those of the negative controls, ppk and sigD (Fig. 4A–G). At concentrations higher than 1.5 μM, MBP-HilD bound most of the DNA fragments tested, including the negative controls, indicating that it binds non-specifically at these concentrations (data not shown). In order to identify putative HilD-binding sites, we scanned the regulatory regions of the gtgE, phoH, sinR, lpxR and SL4247 genes with PSSMs representing the two HilD-binding consensus sequences reported previously27,30. Some HilD-binding sites were predicted in these genes (Fig. S2 in Supplementary File 2), which is consistent with their direct regulation by HilD.
HilD binds to the regulatory region of gtgE, phoH, sinR, lpxR and SL4247, but not to that of SL1896.
MBP-HilD binding to the DNA fragments contained in the gtgE-cat (A), phoH-cat (B), sinR-cat (C), lpxR-cat (D), SL4247-cat (E) and SL1896-cat (F) transcriptional fusions was analyzed by competitive nonradioactive EMSAs. As a positive control, the regulatory region of hilA was also assessed (G), and as a negative internal control, a DNA fragment containing the regulatory region of ppk or sigD was included in each DNA-binding reaction. PCR-amplified and purified DNA fragments were incubated with increasing concentrations (0 to 1 μM) of purified MBP-HilD fusion protein. The DNA-protein complexes (indicated by an asterisk) were resolved in a nondenaturing 5% polyacrylamide gel and stained with ethidium bromide.
Together with the expression analyses, these binding assays demonstrate that HilD directly regulates the expression of the gtgE, phoH, sinR, lpxR and SL4247 genes, and indirectly that of the SL1896 gene.
Discussion
Co-regulation can ensure the coordinated expression of genes located in different chromosomal regions whose products are required for specific cellular functions. For instance, the transcriptional regulator HilD, encoded in SPI-1, positively controls the expression of the genes within this island, as well as several other genes located outside SPI-1, which mediate Salmonella invasion of host cells4,30. Additionally, HilD also positively regulates several genes necessary for Salmonella replication inside host cells, including the SPI-2 genes4,14,29.
In this work, by applying a clustering method to S. Typhimurium SL1344 global expression data from the COLOMBOS database, we show that most of the known genes regulated by HilD, including the flagellar/chemotaxis genes, are indeed co-expressed with SPI-1; moreover, nine novel genes that are co-expressed with SPI-1 were identified: gtgE, phoH, sinR, SL1028, lpxR, SL1896, SL3812, SL4247 and SL4433. Furthermore, we demonstrate that HilD is required for the expression of the gtgE, phoH, sinR, lpxR, SL1896 and SL4247 genes, but not for the SL1028, SL3812 and SL4433 genes, when S. Typhimurium SL1344 is grown in conditions that favor the expression of the SPI-1 genes. FlhDC, the master regulator of the flagellar/chemotaxis genes, was not required either for the expression of the SL1028, SL3812 and SL4433 genes in the growth conditions tested (data not shown), indicating that other regulators could link the expression of these genes with SPI-1. Additionally, we show that HilD can induce the expression of gtgE-cat, phoH-cat, sinR-cat, lpxR-cat, SL3812-cat and SL4247-cat, but not SL1896-cat, transcriptional fusions, in the E. coli MC4100 strain, and thus in the absence of other Salmonella-specific regulators or FlhDC; consistently, HilD bound to the regulatory regions of gtgE, phoH, sinR, lpxR and SL4247, but not to that of SL1896. Previously, by using chromatin immunoprecipitation-sequencing (ChIP-seq) and ChIP-qPCR, it was found that HilD binds in vivo to DNA regions associated with the sinR, lpxR and SL4247 (STM14_5184) S. Typhimurium 14028s genes; furthermore, it was shown that HilD can induce the expression of a lpxR-lacZ translational fusion in the E. coli AMD054 strain (flhDC+)31. Thus, HilD positively and directly regulates the expression of the gtgE, phoH, sinR, lpxR and SL4247 genes, and positively but indirectly controls the expression of the SL1896 gene.
The gtgE, phoH, sinR, lpxR, SL4247 and SL1896 genes are located in different S. Typhimurium chromosomal regions (Table S3 in Supplementary File 1). The phoH gene has a G + C content (50.7%) similar to the average G + C content of the Salmonella genome (52%), indicating that this is an ancestral gene; whereas the gtgE, sinR, lpxR, SL4247 and SL1896 genes have low G + C contents (34.4%, 39.7%, 47.6%, 46.6% and 40.8%, respectively), supporting that these genes were acquired for S. Typhimurium by horizontal transfer. Consistently, the phoH gene is highly conserved in most of bacteria and some archaea (Supplementary File 3)48 and the gtgE, sinR and SL4247 (STM4310) genes are located in S. Typhimurium genomic islands6,49,50,51,52. Genome context and BLAST analyses revealed that the lpxR and SL1896 genes are also located in S. Typhimurium genomic islands (Fig. 5). The lpxR (SL1263) gene is located in a S. Typhimurium region that is absent in E. coli K-12, which is flanked by the ydiY and thrS genes, encoding a conserved putative protein and the enzyme threonyl-tRNA synthetase, respectively. This region also carries the SL1264, SL1265, SL1330A and rfc (SL1266) genes, which have low G + C content, with the exception of SL1265; thus, we denominated this region as island SL1263-66 (Fig. 5A). SL1264 and SL1330A are genes of unknown function, whereas the SL1265 and rfc genes are predicted to code for a DNA/RNA non-specific endonuclease and an O-antigen polymerase, respectively. In E. coli K-12, instead the SL1263-66 island, the ydiY and thrS genes flank the arpB_1 and arpB-2 pseudogenes (Fig. 5A). Interestingly, S. bongori contains the region spanning the SL1263 and SL1265 genes, but not the SL1330A and rfc genes, suggesting that S. Typhimurium acquired the SL1263-66 island by distinct horizontal transfer events; in agreement, the genes within this island show different G + C contents (Fig. 5A). On the other hand, the SL1896 gene is present in S. Typhimurium and S. bongori, but not in E. coli K-12; it is located between the yedF and fliE genes, encoding a conserved putative protein and the flagellar basal-body protein FliE, respectively (Fig. 5B). In E. coli K-12, the yedF and fliE genes flank a region carrying the yedK, yedL and yedM genes, as well as the yedN_1, yedN_2 and intG pseudogenes (Fig. 5B).
The SL1263 and SL1896 genes are located in Salmonella genomic islands that are absent in E. coli K-12. Schematic view of the regions between the ydiY and thrS ancestral genes (A), and the yedF and fliE ancestral genes (B), in E. coli K-12 MG1655, S. Typhimurium SL1344 and S. bongori NCTC 12419. The regions different in S. Typhimurium SL1344 with respect to those of E. coli K-12 MG1655, as well as the region present in S. Typhimurium SL1344 but not in S. bongori NCTC 12419, are indicated (see text for description). Pseudogenes are marked with an X. Broken arrows represent long genes. The G + C content for each of the S. Typhimurium SL1344 genes is shown.
The products of the gtgE, sinR, lpxR, SL4247 and SL1896 genes are highly conserved in S. enterica, and in some cases also in S. bongori; furthermore, orthologs for SinR, LpxR, SL4247 and SL1896 are present in some other bacteria (Supplementary File S3).
A role in Salmonella virulence has been previously determined for gtgE, lpxR and sinR. The gtgE gene, located in the Gifsy-2 bacteriophage49, encodes a T3SS effector protein (GtgE) that is translocated into host cells, where it cleaves the Rab29, Rab-32 and Rab-38 GTPases; depletion of Rab-32 prevents activation of a pathway for Salmonella killing inside macrophages53,54,55. GtgE is present in S. Typhimurium but not in S. Typhi (Supplementary File 3); interestingly, the ectopic expression of GtgE allows S. Typhi to survive and replicate within macrophages and tissues from mice, a nonpermissive host54. Consistently, GtgE is required for the systemic disease caused by S. Typhimurium in mice49,55,56. GtgE can be secreted through both the T3SS-1 and the T3SS encoded in SPI-2 (T3SS-2)53,56; furthermore, its expression is induced in growth conditions favoring the expression of SPI-1 and also in those for SPI-256,57. Therefore, the expression of gtgE is controlled by HilD in SPI-1-inducing conditions and probably by another regulator in SPI-2-inducing conditions, which would coordinate the secretion of GtgE through the T3SS-1 and T3SS-2, respectively. Similarly, the expression of slrP, also encoding an effector protein secreted through both T3SS-1 and T3SS-2, is controlled by HilD in SPI-1-inducing conditions and by the response regulator PhoP in SPI-2-inducing conditions32. In addition to gtgE, the Gifsy-2 bacteriophage carries other Salmonella virulence genes, such as sodCI and sseI49,58. The lpxR gene, located in the 1263–66 island, encodes a Ca2+-dependent outer membrane enzyme (LpxR) that removes the 3´-acyloxyacyl residue of lipid A, the hydrophobic anchor of lipopolysaccharide (LPS)59. LpxR is required for S. Typhimurium growth inside macrophages, probably by its activity on lipid A that could be beneficial to evade host immune surveillance, as well as by its negative effect on the amount of the inducible nitric oxide synthase, which would reduce the nitric oxide-mediated antibacterial cellular response60,61,62. In addition to HilD, the expression of lpxR in SPI-1-inducing conditions is positively regulated by SlyA, a MarR-like regulator63. HilD and SlyA cooperate to directly control the expression of ssrAB in SPI-1-inducing conditions (our unpublished results), which could also apply for lpxR. The other genes located in the SL1263-66 island (Fig. 5A) remain uncharacterized. The sinR gene, located in SPI-6 (also know as Salmonella enterica centisome 7 genomic island)51, encodes a putative LysR-family transcriptional regulator50. No targets of SinR have been determined; however, a S. Typhimurium sinR insertion mutant is attenuated in replication within macrophages, which supports its regulatory role64. SPI-6 also carries other Salmonella virulence genes, such as pagN, sfaCD, sciG, rhs1 and those encoding a type 6 secretion system64,65,66. Whether the regulation by HilD implies that the gtgE, lpxR and sinR genes also have a role in the Salmonella invasion of host cells and thus in the intestinal infection, as for most other genes regulated by HilD, needs to be investigated.
The phoH gene was firstly characterized in E. coli K-12, as encoding a protein (PhoH) that has an ATP-binding activity, and as positively controlled by the transcriptional regulator PhoB in response to phosphate limitation67. It was found that PhoH is homologous to the N-terminal ATPase domain of superfamily I helicases68 and that phoH is not an essential gene in E. coli K-1269; however, even when PhoH orthologs are present in most of bacteria and some archaea (Supplementary File 3), its function remains unknown. Moreover, to our knowledge, there are not previous studies involving any other regulator, in addition to the PhoR/B system, in the expression of phoH. It is tempting to speculate that the regulation by HilD recruited the PhoH activity as a factor that can contribute to the Salmonella pathogenesis.
The SL4247 and SL1896 genes encode hypothetical proteins (Table S3 in Supplementary File 1). Interestingly, the SL4747 (STM4310) gene is upstream of the putative rtsA-rtsB-SL4249(STM4313)-SL4248(STM4312) operon52, which is directly regulated by HilD23,27. The rtsA and rtsB genes code for transcriptional regulators involved in the expression of the SPI-1 and flagellar genes23,70. On the other hand, the SL1896 gene is located in a region containing a large cluster of flagellar genes (data not shown). However, our results show that HilD does not control the expression of SL1896 through FlhDC, the master regulator of the flagellar/chemotaxis genes, neither through the SPI-1 regulators HilA and InvF (Table 1). It is known that HilD can also control gene expression through HilC, SprB, RtsA and SsrA/B4,14, and possibly, as suggested by our results, through SinR and SlyA; thus, HilD could act on SL1896 through any of these regulators.
Whether or not the phoH, SL4247 and SL1896 genes, or even the SL1028, SL3812 and SL4433 genes that are co-expressed with SPI-1 but not regulated by HilD, have a role in Salmonella virulence is a matter of our current investigation.
Several studies support the notion that HilD induces the expression of its target genes mainly by counteracting the repression exerted by the histone-like nucleoid structuring protein (H-NS) on the respective promoters21,22,27,29,71,72. Genome-wide transcriptional and/or binding analyses support that H-NS represses gtgE, sinR, lpxR, SL1896 and SL424773,74,75; however, whether HilD induces the expression of these genes by acting as an anti-repressor of H-NS, and how it positively controls the expression of phoH, remains to be determined.
The method and parameters that we initially used in this study for clustering the S. Typhimurium SL1344 global gene expression results from the COLOMBOS database, were successful to identify gtgE, phoH, sinR, lpxR, SL1896 and SL4247, as novel genes regulated by HilD. By using less-stringent clustering parameters, we found 34 additional genes whose pattern of expression can be linked to that of the SPI-1 genes (Table S4 in Supplementary File 1). Interestingly, between these genes are SL1265 and SL4248, located in the S. Typhimurium genomic islands carrying lpxR (SL1263) and SL4247, respectively, as well as slyA that is positively regulated by HilD (our unpublished results). This strongly supports that more targets of HilD can be found among these 34 genes.
Our findings further expand the virulence regulon of HilD and reveal novel factors possibly involved in the pathogenesis of Salmonella.
Methods
Bioinformatics analyses
The S. Typhimurium SL1344 compendium in the COLOMBOS database (www.colombos.net) contains transcriptional expression values for 4655 genes, from 213 condition contrasts. We used this dataset and a clustering method to find genes co-expressed with SPI-1. Firstly, the k-means algorithm76 was applied 100 times to generate clusters of genes by their expression profiles, using K values of 1024 and 466. Then, consensuses of these clusters were obtained with the consensus clustering method77, using grouping frequencies of 40% and 60%, for the K values of 1024 and 466, respectively, to assign a gene in a consensus cluster. Finally, consensus clusters containing a particular gene, used as bait, were selected and the frequency with which a determined gene is present in these consensus clusters was obtained. Specifically, SPI-1 genes were used as the bait; then, the genes present in at least one selected consensus cluster were considered as genes co-expressed with SPI-1. The same procedure was followed when the novel genes found to be co-expressed with SPI-1 were used as the bait. For less-stringent clustering conditions, grouping frequencies of 30% and 60%, and of 40% and 50%, for the K values of 1024 and 466, respectively, were used to assign a gene in a consensus cluster. This clustering method with similar parameters has been successful to group genes of Escherichia coli K-12 with a related biological function (Sánchez, M., unpublished data).
Position-specific scoring matrices (PSSMs) 1 and 2, representing HilD-binding consensus sequences, were generated by using the consensus program78 and the HilD-binding sites reported by Oleknovich and Kadner and by Singer et al., respectively27,30. Scanning of the regulatory regions of tested genes with these PSSMs was performed with the matrix-scan program78 using a P value of 1e-3. The resulting hits are reported with a corresponding significance score, which is a log-transformation of the E-value.
The orthologs for the products of the gtgE, phoH, sinR, lpxR, SL1028, SL1896, SL3812, SL4247 and SL4433 genes were downloaded from a database of orthologs detected as Reciprocal Best Hits using soft masking and Smith-Waterman alignments (http://popolvuh.wlu.ca/Orthologs)79.
Media and culture conditions
Bacterial cultures were grown at 37 °C in LB medium containing 1% tryptone, 0.5% yeast agar and 1% NaCl, pH 7.5. When necessary, media were supplemented with ampicillin (200 μg ml−1), streptomycin (100 μg ml−1) or kanamycin (30 μg ml−1). Cultures for CAT assays were performed as described previously14,42.
Construction of the mutant strain lacking flhDC
Bacterial strains used in this work are listed in Table S5 in Supplementary File 2. Non-polar deletion of the flhDC operon in the S. Typhimurium SL1344 strain was generated by the λRed recombinase system, as reported previously80, using the respective primers described in Table S6 in Supplementary File 2, generating the strain DTM90. This mutant strain was verified by PCR amplification and sequencing.
Construction of plasmids
Plasmids and primers used in this work are listed in Tables S5 and S6, respectively, in Supplementary File 2. To construct the plasmids containing the transcriptional fusions gtgE-cat, phoH-cat, sinR-cat, trg-cat, SL1028-cat, lpxR-cat, SL1896-cat, SL3812-cat, SL4247-cat and SL4433-cat, DNA fragments containing the intergenic region upstream of gtgE, phoH, sinR, trg, SL1028, lpxR, SL1896, SL3812, SL4247 or SL4433 were amplified by PCR with the primer pairs gtgE-Rv1/gtgE-Fw2, phoH-Rv/phoH-Fw, sinRH3-Rv33/sinRB2-Fw44, trg-Rv1/trg-Fw2, SL1028-Rv1/SL1028-Fw2, SL1263-Rv55/SL1263-Fw77, SL1896-Rv1/SL1896-Fw2, SL3812-Rv1/SL3812-Fw2, SL4247-Rv1/SL4247-Fw2 and SL4433-Rv1/SL4433-Fw2, respectively. The PCR products were digested with BamHI and HindIII restriction enzymes and then cloned into the BamHI and HindIII sites of the vector pKK232-8, which carries a promotorless cat gene (Amersham Pharmacia LKB Biotechnology), generating plasmids pgtgE-cat, pphoH-cat, psinR-cat, ptrg-cat, pSL1028-cat, plpxR-cat, pSL1896-cat, pSL3812-cat, pSL4247-cat and pSL4433-cat. To construct the plasmid pK6-HilD, the hilD structural gene was amplified by PCR using the primer pair HilDK6-F/HilDexR-PstI and chromosomal DNA from WT S. Typhimurium SL1344 as template. This PCR product was digested with NcoI and PstI restriction enzymes and then cloned into the vector pMPM-K6Ω81 digested with the same restriction enzymes. pK6-HilD expresses HilD from an arabinose-inducible promoter.
CAT assays
The chloramphenicol acetyl transferase (CAT) assays and protein quantification to calculate CAT specific activities were performed as previously described82.
Statistical analysis
Results from CAT assays were analyzed using One-Way analysis of variance (ANOVA) with the Dunnett multiple comparison test for Table 1, or unpaired two-tailed Student’s t test for Figs 2 and 3. This statistical analysis was performed using Prism 5 program version 5.04 (GraphPad Software, San Diego, CA).
Expression and purification of MBP-HilD
Maltose binding protein (MBP)-HilD was expressed in E. coli BL21/DE3 containing pMAL-HilD1 and purified by using an amylose column, as described previously14.
EMSAs
DNA fragments containing the intergenic region upstream of gtgE, phoH, sinR, lpxR, SL1896 and SL4247 were obtained by PCR amplification with the same primer pairs used to construct the respective transcriptional fusion to the cat reporter gene. DNA fragments containing the intergenic region upstream of hilA, used as a positive control, and sigD or ppk, used as internal negative controls, were obtained by PCR amplification with the primer pairs hilA2R-HindIII/hilA1F-BamHI, sigD-H3R/sigD-BHIF and PPK-Rv1/PPK-Fw1, respectively. PCR products were purified using the QIAquick PCR purification kit (Qiagen). Each PCR product (≈100 ng) was mixed with an equal amount of the PCR product of sigD or ppk and increasing concentrations of purified MBP-HilD in a binding buffer containing 10 mM Tris (pH 8.0), 50 mM KCl, 1 mM dithiothreitol (DTT), 0.5 mM EDTA, 5% glycerol and 10 μg ml−1 bovine serum albumin (BSA), in a final volume of 20 μl. Protein-DNA binding reactions were incubated at room temperature for 20 min and then separated by electrophoresis in 5% non-denaturing acrylamide gels in 0.5 X Tris-borate-EDTA buffer at room temperature. The DNA fragments were stained with ethidium bromide and visualized with an Alpha-Imager UV transilluminator (Alpha Innotech Corp.).
Additional Information
How to cite this article: Martínez-Flores, I. et al. In silico clustering of Salmonella global gene expression data reveals novel genes co-regulated with the SPI-1 virulence genes through HilD. Sci. Rep. 6, 37858; doi: 10.1038/srep37858 (2016).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Haraga, A., Ohlson, M. B. & Miller, S. I. Salmonellae interplay with host cells. Nat Rev Microbiol 6, 53–66 (2008).
Sánchez-Vargas, F. M., Abu-El-Haija, M. A. & Gómez-Duarte, O. G. Salmonella infections: an update on epidemiology, management, and prevention. Travel Med Infect Dis 9, 263–277, doi: 10.1016/j.tmaid.2011.11.001 (2011).
Brenner, F. W., Villar, R. G., Angulo, F. J., Tauxe, R. & Swaminathan, B. Salmonella nomenclature. J Clin Microbiol 38, 2465–2467 (2000).
Fàbrega, A. & Vila, J. Salmonella enterica serovar Typhimurium skills to succeed in the host: virulence and regulation. Clin Microbiol Rev 26, 308–341, doi: 10.1128/CMR.00066-12 (2013).
Fookes, M. et al. Salmonella bongori provides insights into the evolution of the Salmonellae. PLoS Pathog 7, e1002191, doi: 10.1371/journal.ppat.1002191 (2011).
Porwollik, S. & McClelland, M. Lateral gene transfer in Salmonella. Microbes Infect 5, 977–989 (2003).
Hansen-Wester, I. & Hensel, M. Salmonella pathogenicity islands encoding type III secretion systems. Microbes Infect 3, 549–559 (2001).
Moest, T. P. & Meresse, S. Salmonella T3SSs: successful mission of the secret(ion) agents. Curr Opin Microbiol 16, 38–44, doi: 10.1016/j.mib.2012.11.006 (2013).
Laughlin, R. C. et al. Spatial segregation of virulence gene expression during acute enteric infection with Salmonella enterica serovar Typhimurium. MBio 5, e00946–00913, doi: 10.1128/mBio.00946-13 (2014).
Knodler, L. A. et al. Dissemination of invasive Salmonella via bacterial-induced extrusion of mucosal epithelia. Proc Natl Acad Sci USA 107, 17733–17738, doi: 10.1073/pnas.1006098107 (2010).
Lawhon, S. D., Maurer, R., Suyemoto, M. & Altier, C. Intestinal short-chain fatty acids alter Salmonella typhimurium invasion gene expression and virulence through BarA/SirA. Mol Microbiol 46, 1451–1464 (2002).
Altier, C. Genetic and environmental control of Salmonella invasion. J Microbiol 43 Spec No, 85–92 (2005).
Jones, B. D. Salmonella invasion gene regulation: a story of environmental awareness. J Microbiol 43 Spec No, 110–117 (2005).
Bustamante, V. H. et al. HilD-mediated transcriptional cross-talk between SPI-1 and SPI-2. Proc Natl Acad Sci USA 105, 14591–14596 (2008).
Ellermeier, J. R. & Slauch, J. M. Fur regulates expression of the Salmonella pathogenicity island 1 type III secretion system through HilD. J Bacteriol 190, 476–486, doi: 10.1128/JB.00926-07 (2008).
Hung, C. C. et al. The intestinal fatty acid propionate inhibits Salmonella invasion through the post-translational control of HilD. Mol Microbiol 87, 1045–1060, doi: 10.1111/mmi.12149 (2013).
Kroger, C. et al. An infection-relevant transcriptomic compendium for Salmonella enterica Serovar Typhimurium. Cell Host Microbe 14, 683–695, doi: 10.1016/j.chom.2013.11.010 (2013).
Golubeva, Y. A., Ellermeier, J. R., Cott Chubiz, J. E. & Slauch, J. M. Intestinal long-chain fatty acids act as a direct signal to modulate expression of the Salmonella pathogenicity island 1 type III secretion system. MBio 7, e02170–02115, doi: 10.1128/mBio.02170-15 (2016).
Eade, C. R. et al. Bile acids function synergistically to repress invasion gene expression in Salmonella by destabilizing the invasion regulator HilD. Infect Immun, doi: 10.1128/IAI.00177-16 (2016).
Golubeva, Y. A., Sadik, A. Y., Ellermeier, J. R. & Slauch, J. M. Integrating global regulatory input into the Salmonella pathogenicity island 1 type III secretion system. Genetics 190, 79–90, doi: 10.1534/genetics.111.132779 (2012).
Schechter, L. M., Damrauer, S. M. & Lee, C. A. Two AraC/XylS family members can independently counteract the effect of repressing sequences upstream of the hilA promoter. Mol Microbiol 32, 629–642 (1999).
Schechter, L. M. & Lee, C. A. AraC/XylS family members, HilC and HilD, directly bind and derepress the Salmonella typhimurium hilA promoter. Mol Microbiol 40, 1289–1299 (2001).
Ellermeier, C. D., Ellermeier, J. R. & Slauch, J. M. HilD, HilC and RtsA constitute a feed forward loop that controls expression of the SPI1 type three secretion system regulator hilA in Salmonella enterica serovar Typhimurium. Mol Microbiol 57, 691–705 (2005).
Lostroh, C. P., Bajaj, V. & Lee, C. A. The cis requirements for transcriptional activation by HilA, a virulence determinant encoded on SPI-1. Mol Microbiol 37, 300–315 (2000).
Lostroh, C. P. & Lee, C. A. The HilA box and sequences outside it determine the magnitude of HilA-dependent activation of P(prgH) from Salmonella pathogenicity island 1. J Bacteriol 183, 4876–4885 (2001).
Saini, S., Ellermeier, J. R., Slauch, J. M. & Rao, C. V. The role of coupled positive feedback in the expression of the SPI1 type three secretion system in Salmonella. PLoS Pathog 6, e1001025, doi: 10.1371/journal.ppat.1001025 (2010).
Olekhnovich, I. N. & Kadner, R. J. Role of nucleoid-associated proteins Hha and H-NS in expression of Salmonella enterica activators HilD, HilC, and RtsA required for cell invasion. J Bacteriol 189, 6882–6890 (2007).
Olekhnovich, I. N. & Kadner, R. J. DNA-binding activities of the HilC and HilD virulence regulatory proteins of Salmonella enterica serovar Typhimurium. J Bacteriol 184, 4148–4160 (2002).
Martínez, L. C., Banda, M. M., Fernández-Mora, M., Santana, F. J. & Bustamante, V. H. HilD induces expression of Salmonella pathogenicity island 2 genes by displacing the global negative regulator H-NS from ssrAB. J Bacteriol 196, 3746–3755, doi: 10.1128/JB.01799-14 (2014).
Singer, H. M., Kuhne, C., Deditius, J. A., Hughes, K. T. & Erhardt, M. The Salmonella Spi1 virulence regulatory protein HilD directly activates transcription of the flagellar master operon flhDC. J Bacteriol 196, 1448–1457, doi: 10.1128/JB.01438-13 (2014).
Petrone, B. L., Stringer, A. M. & Wade, J. T. Identification of HilD-regulated genes in Salmonella enterica serovar Typhimurium. J Bacteriol 196, 1094–1101, doi: 10.1128/JB.01449-13 (2014).
Cordero-Alba, M. & Ramos-Morales, F. Patterns of expression and translocation of the ubiquitin ligase SlrP in Salmonella enterica serovar Typhimurium. J Bacteriol 196, 3912–3922, doi: 10.1128/JB.02158-14 (2014).
Main-Hester, K. L., Colpitts, K. M., Thomas, G. A., Fang, F. C. & Libby, S. J. Coordinate regulation of Salmonella pathogenicity island 1 (SPI1) and SPI4 in Salmonella enterica serovar Typhimurium. Infect Immun 76, 1024–1035, doi: 10.1128/IAI.01224-07 (2008).
Thijs, I. M. et al. Delineation of the Salmonella enterica serovar Typhimurium HilA regulon through genome-wide location and transcript analysis. J Bacteriol 189, 4587–4596, doi: 10.1128/JB.00178-07 (2007).
Mouslim, C. & Hughes, K. T. The effect of cell growth phase on the regulatory cross-talk between flagellar and Spi1 virulence gene expression. PLoS pathogens 10, e1003987, doi: 10.1371/journal.ppat.1003987 (2014).
Chubiz, J. E., Golubeva, Y. A., Lin, D., Miller, L. D. & Slauch, J. M. FliZ regulates expression of the Salmonella pathogenicity island 1 invasion locus by controlling HilD protein activity in Salmonella enterica serovar typhimurium. J Bacteriol 192, 6261–6270, doi: 10.1128/JB.00635-10 (2010).
Ibarra, J. A. et al. Induction of Salmonella pathogenicity island 1 under different growth conditions can affect Salmonella-host cell interactions in vitro. Microbiology 156, 1120–1133, doi: 10.1099/mic.0.032896-0 (2010).
Eichelberg, K. & Galan, J. E. The flagellar sigma factor FliA (σ28) regulates the expression of Salmonella genes associated with the centisome 63 type III secretion system. Infect Immun 68, 2735–2743 (2000).
Moretto, M. et al. COLOMBOS v3.0: leveraging gene expression compendia for cross-species analyses. Nucleic Acids Res 44, D620–623, doi: 10.1093/nar/gkv1251 (2016).
Wang, Q., Mariconda, S., Suzuki, A., McClelland, M. & Harshey, R. M. Uncovering a large set of genes that affect surface motility in Salmonella enterica serovar Typhimurium. J Bacteriol 188, 7981–7984, doi: 10.1128/JB.00852-06 (2006).
Frye, J. et al. Identification of new flagellar genes of Salmonella enterica serovar Typhimurium. J Bacteriol 188, 2233–2243, doi: 10.1128/JB.188.6.2233-2243.2006 (2006).
Martínez, L. C. et al. Integration of a complex regulatory cascade involving the SirA/BarA and Csr global regulatory systems that controls expression of the Salmonella SPI-1 and SPI-2 virulence regulons through HilD. Mol Microbiol 80, 1637–1656, doi: 10.1111/j.1365-2958.2011.07674.x (2011).
Martínez, L. C. et al. In silico identification and experimental characterization of regulatory elements controlling the expression of the Salmonella csrB and csrC genes. J Bacteriol 196, 325–336, doi: 10.1128/JB.00806-13 (2014).
De la Cruz, M. A. et al. The two-component system CpxR/A represses the expression of Salmonella virulence genes by affecting the stability of the transcriptional regulator HilD. Front Microbiol 6, 807, doi: 10.3389/fmicb.2015.00807 (2015).
Park, K., Choi, S., Ko, M. & Park, C. Novel sF-dependent genes of Escherichia coli found using a specified promoter consensus. FEMS Microbiol Lett 202, 243–250 (2001).
Ferenci, T. et al. Genomic sequencing reveals regulatory mutations and recombinational events in the widely used MC4100 lineage of Escherichia coli K-12. J Bacteriol 191, 4025–4029, doi: 10.1128/JB.00118-09 (2009).
Zhao, K., Liu, M. & Burgess, R. R. Adaptation in bacterial flagellar and motility systems: from regulon members to ‘foraging’-like behavior in E. coli. Nucleic Acids Res 35, 4441–4452, doi: 10.1093/nar/gkm456 (2007).
Kazakov, A. E., Vassieva, O., Gelfand, M. S., Osterman, A. & Overbeek, R. Bioinformatics classification and functional analysis of PhoH homologs. In Silico Biol 3, 3–15 (2003).
Ho, T. D. et al. Identification of GtgE, a novel virulence factor encoded on the Gifsy-2 bacteriophage of Salmonella enterica serovar Typhimurium. J Bacteriol 184, 5234–5239 (2002).
Groisman, E. A., Sturmoski, M. A., Solomon, F. R., Lin, R. & Ochman, H. Molecular, functional, and evolutionary analysis of sequences specific to Salmonella. Proc Natl Acad Sci USA 90, 1033–1037 (1993).
Folkesson, A., Lofdahl, S. & Normark, S. The Salmonella enterica subspecies I specific centisome 7 genomic island encodes novel protein families present in bacteria living in close contact with eukaryotic cells. Res Microbiol 153, 537–545 (2002).
Wilson, J. W. & Nickerson, C. A. A new experimental approach for studying bacterial genomic island evolution identifies island genes with bacterial host-specific expression patterns. BMC Evol Biol 6, 2, doi: 10.1186/1471-2148-6-2 (2006).
Spano, S., Liu, X. & Galan, J. E. Proteolytic targeting of Rab29 by an effector protein distinguishes the intracellular compartments of human-adapted and broad-host Salmonella. Proc Natl Acad Sci USA 108, 18418–18423, doi: 10.1073/pnas.1111959108 (2011).
Spano, S. & Galan, J. E. A Rab32-dependent pathway contributes to Salmonella typhi host restriction. Science 338, 960–963, doi: 10.1126/science.1229224 (2012).
Spano, S., Gao, X., Hannemann, S., Lara-Tejero, M. & Galan, J. E. A Bacterial Pathogen Targets a Host Rab-Family GTPase Defense Pathway with a GAP. Cell Host Microbe 19, 216–226, doi: 10.1016/j.chom.2016.01.004 (2016).
Niemann, G. S. et al. Discovery of novel secreted virulence factors from Salmonella enterica serovar Typhimurium by proteomic analysis of culture supernatants. Infect Immun 79, 33–43, doi: 10.1128/IAI.00771-10 (2011).
Uzzau, S., Figueroa-Bossi, N., Rubino, S. & Bossi, L. Epitope tagging of chromosomal genes in Salmonella. Proc Natl Acad Sci USA 98, 15264–15269 (2001).
McLaughlin, L. M. et al. The Salmonella SPI2 effector SseI mediates long-term systemic infection by modulating host cell migration. PLoS Pathog 5, e1000671, doi: 10.1371/journal.ppat.1000671 (2009).
Reynolds, C. M. et al. An outer membrane enzyme encoded by Salmonella typhimurium lpxR that removes the 3′-acyloxyacyl moiety of lipid A. J Biol Chem 281, 21974–21987, doi: 10.1074/jbc.M603527200 (2006).
Eriksson, S. et al. Salmonella typhimurium mutants that downregulate phagocyte nitric oxide production. Cell Microbiol 2, 239–250 (2000).
Kawano, M., Manabe, T. & Kawasaki, K. Salmonella enterica serovar Typhimurium lipopolysaccharide deacylation enhances its intracellular growth within macrophages. FEBS Lett 584, 207–212, doi: 10.1016/j.febslet.2009.11.062 (2010).
Kawasaki, K., Teramoto, M., Tatsui, R. & Amamoto, S. Lipid A 3′-O-deacylation by Salmonella outer membrane enzyme LpxR modulates the ability of lipid A to stimulate Toll-like receptor 4. Biochem Biophys Res Commun 428, 343–347, doi: 10.1016/j.bbrc.2012.10.054 (2012).
Navarre, W. W. et al. Co-regulation of Salmonella enterica genes required for virulence and resistance to antimicrobial peptides by SlyA and PhoP/PhoQ. Mol Microbiol 56, 492–508, doi: 10.1111/j.1365-2958.2005.04553.x (2005).
Klumpp, J. & Fuchs, T. M. Identification of novel genes in genomic islands that contribute to Salmonella typhimurium replication in macrophages. Microbiology 153, 1207–1220, doi: 10.1099/mic.0. 2006/004747-0 (2007).
Mulder, D. T., Cooper, C. A. & Coombes, B. K. Type VI secretion system-associated gene clusters contribute to pathogenesis of Salmonella enterica serovar Typhimurium. Infect Immun 80, 1996–2007, doi: 10.1128/IAI.06205-11 (2012).
Lambert, M. A. & Smith, S. G. The PagN protein of Salmonella enterica serovar Typhimurium is an adhesin and invasin. BMC Microbiol 8, 142, doi: 10.1186/1471-2180-8-142 (2008).
Kim, S. K., Makino, K., Amemura, M., Shinagawa, H. & Nakata, A. Molecular analysis of the phoH gene, belonging to the phosphate regulon in Escherichia coli. J Bacteriol 175, 1316–1324 (1993).
Koonin, E. V. & Rudd, K. E. Two domains of superfamily I helicases may exist as separate proteins. Protein Sci 5, 178–180, doi: 10.1002/pro.5560050124 (1996).
Zhou, L., Lei, X. H., Bochner, B. R. & Wanner, B. L. Phenotype microarray analysis of Escherichia coli K-12 mutants with deletions of all two-component systems. J Bacteriol 185, 4956–4972 (2003).
Ellermeier, C. D. & Slauch, J. M. RtsA and RtsB coordinately regulate expression of the invasion and flagellar genes in Salmonella enterica serovar Typhimurium. J Bacteriol 185, 5096–5108 (2003).
Schechter, L. M., Jain, S., Akbar, S. & Lee, C. A. The small nucleoid-binding proteins H-NS, HU, and Fis affect hilA expression in Salmonella enterica serovar Typhimurium. Infect Immun 71, 5432–5435 (2003).
Olekhnovich, I. N. & Kadner, R. J. Crucial roles of both flanking sequences in silencing of the hilA promoter in Salmonella enterica. J Mol Biol 357, 373–386 (2006).
Ono, S. et al. H-NS is a part of a thermally controlled mechanism for bacterial gene regulation. Biochem J 391, 203–213, doi: 10.1042/BJ20050453 (2005).
Navarre, W. W. et al. Selective silencing of foreign DNA with low GC content by the H-NS protein in Salmonella. Science 313, 236–238 (2006).
Lucchini, S. et al. H-NS mediates the silencing of laterally acquired genes in bacteria. PLoS Pathog 2, e81 (2006).
Hastie, T., Tibshirani, R. & Friedman, J. The elements of statistical learning. (Springer, NY, 2001).
Monti, S., Tamayo, P., Mesirov, J. & Golub, T. Consensus clustering: a resampling-based method for class discovery and visualization of gene expression microarray data. Mach Learn 52, 91–118 (2003).
Medina-Rivera, A. et al. RSAT 2015: Regulatory Sequence Analysis Tools. Nucleic Acids Res 43, W50–56, doi: 10.1093/nar/gkv362 (2015).
Moreno-Hagelsieb, G. & Latimer, K. Choosing BLAST options for better detection of orthologs as reciprocal best hits. Bioinformatics 24, 319–324, doi: 10.1093/bioinformatics/btm585 (2008).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97, 6640–6645 (2000).
Mayer, M. P. A new set of useful cloning and expression vectors derived from pBlueScript. Gene 163, 41–46 (1995).
Puente, J. L., Bieber, D., Ramer, S. W., Murray, W. & Schoolnik, G. K. The bundle-forming pili of enteropathogenic Escherichia coli: transcriptional regulation by environmental signals. Mol Microbiol 20, 87–100 (1996).
Acknowledgements
This work was supported by the Dirección General de Asuntos del Personal Académico de la UNAM/México (IN203415 to V.H.B. and IN207813 to J.C.-V.), and the Consejo Nacional de Ciencia y Tecnología (CONACYT)/México (254531 to V.H.B.). C.C.P. was supported by a pre-doctoral fellowship from CONACYT (290934). We thank M.M. Banda for constructing the DTM90 strain, and F.J. Santana and M. Fernández-Mora for technical assistance.
Author information
Authors and Affiliations
Contributions
V.H.B. designed the research. I.M.-F., M.S.-P., H.S., J.C.-V. and V.H.B. conceived and designed the bioinformatics analysis. M.S.-P. designed the clustering method. I.M.-F., M.S.-P. and H.S. performed the bioinformatics analysis. D.P.-M. and V.H.B. conceived and designed the biological experiments. D.P.-M. and C.C.P. performed the biological experiments. I.M.-F., D.P.-M., H.S. and V.H.B. analyzed the data. I.M.-F. and V.H.B. drafted the manuscript. All authors read and approved the final version of the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Martínez-Flores, I., Pérez-Morales, D., Sánchez-Pérez, M. et al. In silico clustering of Salmonella global gene expression data reveals novel genes co-regulated with the SPI-1 virulence genes through HilD. Sci Rep 6, 37858 (2016). https://doi.org/10.1038/srep37858
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37858
This article is cited by
-
HilD induces expression of a novel Salmonella Typhimurium invasion factor, YobH, through a regulatory cascade involving SprB
Scientific Reports (2019)
-
HilD and PhoP independently regulate the expression of grhD1, a novel gene required for Salmonella Typhimurium invasion of host cells
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.