Abstract
The receptor tyrosine kinase Tie-2 is involved in vessel remodeling and maturation, and has been regarded as a potential target for the treatment of various solid tumors. The absence of novel, potent and selective inhibitors severely hampers the understanding of the therapeutic potential of Tie-2. In the present work, we describe the discovery of novel type-I inhibitors of Tie-2 by structure-based virtual screening. Preliminary SAR was also performed based on one active compound, and several novel inhibitors with low micro-molar affinity were discovered. To directly compare the efficiency between different filtering strategies in selecting VS candidates, two methods were separately carried out to screen the same chemical library, and the selected VS candidates were then experimentally assessed by in vitro enzymatic assays. The results demonstrate that the hit rate is improved when stricter drug-likeness criteria and less number of molecules for clustering analysis are used, and meanwhile, the molecular diversity of the compounds still maintains. As a case study of TIE-2, the information presented in this work underscores the importance of selecting an appropriate selection strategy in VS campaign, and the novel inhibitors identified and the detailed binding modes of action provide a starting point for further hit-to-lead optimization process.
Similar content being viewed by others
Introduction
Angiogenesis is involved in the formation of new capillaries from existing vasculature, where a primitive vascular network is assembled due to the differentiation and proliferation of endothelial cells1. The activation of angiogenesis usually occurs in embryonic development but can also be found in normal physiological processes such as wound healing and certain stages of the menstrual cycle. Aberrant angiogenesis is demonstrated to be the cause of numerous life-threatening diseases including cancer, inflammatory disorders, ischemic diseases and various retinopathies2. The growth of tumors has been shown to rely on the progress of angiogenesis, and interference with the formation of vascular system is believed to be an effective strategy for the treatment of various solid tumors3.
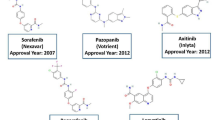
Great success has been made in the development of drugs targeting angiogenesis signaling pathways in the past years. The members of the vascular endothelial cell growth factor (VEGF) and tyrosine kinase with immunoglobulin and epidermal growth factor homology domains-2 (Tie-2) have been shown to be essential factors in vascular development. The VEGF family, such as Flt-1 (VEGF-R1), Flk-1/KDR (VEGF-R2), and Flt-4 (VEGF-R3), plays critical roles in the sprouting process of angiogenesis4,5. While, Tie-2 receptors have been implicated in further stabilization, maturation and remodeling of preliminary vessels6,7,8. It is currently well established that blockage of VEGF and/or Tie-2 signaling pathways can significantly attenuate tumor-induced angiogenesis and suppress tumor growth and metastasis in a variety of solid tumors. Numerous anti-angiogenesis inhibitors targeting VEGF signaling have been under clinical assessment, and several of them, such as sunitinib (Sutent) and sorafenib (Nexavar), have been approved by the Food and Drug Administration (FDA)4,9,10. Despite of the encouraging clinical outcomes from VEGF inhibitors, more patients emerge to become resistant to currently available drugs11,12,13,14,15.
The emergence of drug resistance and the increasing need for better therapeutic strategies lead to the development of second-generation anti-angiogenesis drugs targeting different stages of vessel growth. Among them, angiopoietins (ANGs) and their physiologic receptors, such as Tie-2/TEK receptor that is expressed principally on vascular endothelium, have become very promising therapeutic targets. A substantial body of evidence has been reported that combination of different anti-angiogenesis inhibitors can obtain superior therapeutic outcomes compared with treatment using either agent alone in a variety of xenograft models16,17. Several inhibitory antibodies of ANGs (ANG-1 and/or ANG-2), such as Trebananib, MEDI-3617, CVX-060, REGN-910 and AMG-78018,19,20, have entered clinical trials, but the development of selective small-molecule inhibitors of Tie-2 is still in urgent need with only two candidates with poor kinase selectivity in early clinical phase, i.e. CEP-11981 (Phase I, Cephalon, Inc.) and ARRY-614 (Phase I, Array Biopharma. Inc.)21,22. However, hundreds of molecules were found effect in inhibiting Tie-2 activity including some FDA-approved tyrosine kinase inhibitors, several of which showed quite good inhibitory activity in nM level. Though these inhibitors may not be developed specially towards Tie-2, they are still helpful in understanding the binding patterns of Tie-2 and offer clues for the design of selective Tie-2 inhibitors.
Structure-based virtual screening (VS) strategy has been successfully applied in identifying novel inhibitors of a specific protein target23,24,25,26,27,28,29,30. However, the prediction accuracy of molecular docking and the percentage rate of active compounds are always low. Except for the influence of docking simulations, the effect of applying different selection strategy in selecting VS candidates is also obvious. Though numerous attempts have been made to improve the efficiency of VS in in silico models, few approaches were experimentally validated27,31,32. In the present work, structure-based VS was performed to identify type-I inhibitors of Tie-2 using different drug-likeness filtering criteria (Fig. 1). In VS campaign, clustering analysis can be performed based on the top ranked compounds to maximize the molecular diversity of the candidates. But this practice may have a pronounced influence on the overall hit rates. So we also assessed the influence of different number of clustering compounds on the overall hit rates by using the same docking method (Glide docking) and chemical library (ChemBridge), and two groups of VS candidates obtained from different clustering strategies were then experimentally evaluated. Afterwards, substructure searching was carried out to optimize the active compound (TP-C2-9, IC50 = 1.86 μM) by introducing different substitutions on the core structure and to give a preliminary structure-activity relationships (SAR) analysis. The activities of the compounds discovered in this work are all in low μM range, which is much higher than some already known inhibitors, but these novel structures could be a starting point for medicinal chemists to optimize the activity and benefit to developing more potent and selective Tie-2 inhibitors. Molecular modeling of the active compounds proposed the binding mode to Tie-2. Our results demonstrate that the hit rates can be well improved when stricter drug-likeness criteria are used and top-200 rather than top-1000 compounds are included in clustering analysis, and meanwhile, the molecular diversity of the compounds still maintains. The information provided and the active compounds identified in this work can help to guide the design and discovery of novel and potent inhibitors of Tie-2 as well as the structure-based VS of other protein targets.
Results and Discussions
Structure-based virtual screening
At present, two DFG-in structures (Type-I) of Tie-2 complexes (PDB entries: 2WQB and 3L8P)33,34 are available in RCSB Protein Data Bank35. Both crystal structures were firstly assessed for their ability to distinguish the known inhibitors from non-inhibitors of Tie-2 using two different scoring modes of docking (Glide SP and XP). The distributions of the docking scores for both inhibitors and non-inhibitors were plotted in Fig. 2. The student’s t-test was used to assess the significance of the difference between two sets of docking scores with the associated probability (p value). Moreover, the AUC values (area under curve) of the ROC (receiver operating characteristic) curves for both crystals based on the docking scores were also plotted in Supplementary Fig. S3 to quantitatively evaluate the prediction accuracy. We can find that the p values for 3L8P are much lower than those for 2WQB and the AUC values from SP and XP docking of 3L8P crystal are both higher than those from 2WQB, indicating that the discrimination power of molecular docking based on 3L8P is much higher. Thus, the crystal structure of 3L8P was employed in our VS campaign. For both 2WQB and 3L8P crystals, the performance of Glide XP docking is slightly better than that of SP docking, but the computational cost of XP docking is relatively much higher. Therefore, the Glide SP docking was utilized in the first-round screening of the entire ChemBridge database (~1,020,000 compounds). Afterwards the top-ranked 100,000 compounds from the SP docking were submitted to the second-round screening using the Glide XP docking, and the top-ranked 2,000 compounds were then saved.
In order to assess the influence of different criteria in selecting VS candidates on the final hit percentage of the active compounds, two selection strategies were separately carried out to filter the same top-ranked 2,000 compounds from the Glide XP docking (Fig. 1). The ADMET properties of the 2,000 compounds were firstly predicted by ACD/ADME package36. As we can see in Fig. 1, two different drug-likeness criteria were used: (1) logP/logD (pH = 7.0) > 6.0; violation number of Lipinski’s rule of five ≥ 237; violation number of Oprea’s rules of drug-likeness ≥ 438. (2) logP/logD (pH = 7.0) > 5.5; violation number of Lipinski’s rule of five ≥ 237; violation number of Oprea’s rules of drug-likeness ≥ 338. All the molecules satisfying either criterion for each group were removed. The remaining compounds from both groups were then filtered by ROES rules to remove molecules containing toxic, reactive, and undesirable functional groups. For Strategy 1, the top-ranked 1,000 compounds were submitted to clustering analysis to maximize the structural diversity on the basis of the Tanimoto distance predicted from the FCFP_4 fingerprints. For Strategy 2, clustering analysis was performed with only the top-ranked 200 compounds. The Tanimoto Coefficient (tanimoto) metric was used for clustering based on the FCFP_4 fingerprints of the compounds in DS 2.5. The same number of 70 clusters was manually defined for each strategy prior to the clustering analysis, and the compounds were extracted using the method of selecting the cluster centers (a maximum dissimilarity method) to find diverse molecules39. Finally, almost the same number of molecules was obtained from each strategy. However some compounds were not available for purchasing from the ChemBridge vendor. So eventually 60 compounds were purchased from Strategy 1 and 56 compounds were purchased from Strategy 2. The inhibitory activities of these compounds were then validated by biological experimental assays. Based on the most potent compound, structural optimization was also carried out to give a preliminary structure activity relationship (SAR) analysis using substructure searching method by querying the entire ChemBridge library for compounds with the same core structure.
The performance of different filtering strategies in selecting VS candidates
In order to assess the biological inhibitory activity of the VS compounds, Z’-Lyte enzymatic activity assay based on FRET (Fluorescence Resonance Energy Transfer) technology was developed as described above. The Tie-2 inhibitor (CAS: 948557–43–5) was previously reported as a tool inhibitor that can potently inhibit Tie-2 activity with an IC50 value of 0.25 μM, and was used as a positive control inhibitor in our study. The tested inhibitory activity of this Tie-2 inhibitor is 0.31 μM, which is in good agreement with the ref. 40. At first, all the compounds obtained from both filtering strategies were preliminary screened for their ability to inhibit Tie-2 kinase activity at the concentration of 10 μg/mL. The results were summarized and plotted in Fig. 3 and Fig. 4. It is easy to find that the performance of the candidates chosen from the clustering of the top-ranked 200 compounds is better than those chosen from the clustering of the top-ranked 1,000 compounds. The probability p value of the two sets is 0.00018 indicating that they are statistically different. As we can see in Fig. 4, over 80 percentage of the purchased compounds from Strategy 2 were found to inhibit ≥50% Tie-2 activity at 10 μg/mL, and ~40% compounds can inhibit ≥80% Tie-2 activity. However, only ~20% compounds from Strategy 1 were observed to inhibit ≥50% Tie-2 activity, and <10% compounds were found to inhibit ≥80% activity. This indicates that the hit rate from Strategy 2 is much higher than that from Strategy 1. Different selection strategies in selecting the same 2,000 VS candidates have a pronounced influence on the overall hit rates of the active compounds. The results demonstrate that the hit rates can be well improved when stricter drug-likeness criteria and top-200 rather than top-1000 compounds for clustering analysis are used.
In fact, different drug-likeness criteria may affect the ADMET properties of the hit compounds, but may have little influence on the inhibitory activity. So, further analyses were carried out based on the VS result. We find that only six compounds (TP-C1-4, TP-C1-6, TP-C1-13, TP-C1-18, TP-C1-20, TP-C1-31) from Strategy 1 were filtered out using the criteria applied in Strategy 2. Among them, TP-C1-4 is active and the others are inactive. So the final hit rate of Strategy 1 is only slightly influenced when the criteria of Strategy 2 are applied. The likelihood of the difference based on the new data sets with the six compounds excluded from the list was also calculated, and the p value is 0.0002 implying that there is also a statistical difference between the two sets. Moreover, a power analysis to see how many compounds are needed to test to be confident was also performed and the results were summarized in Supplementary Table S2. A sample size of 50 is sufficient to get a power of 0.939, which says we have a 93% chance of detecting a difference if it collects 50 compounds. From this point of view, the high hit rates from Strategy 2 should be mainly contributed by using different clustering strategies. However stricter drug-likeness filtering criteria are also suggested since the potential druggability of the discovered hit compounds is somewhat better. In addition, the rankings of the tested compounds from both Strategy 1 and Strategy 2 were also summarized and plotted (see Supplementary Fig. S2). We can see that the rankings of all the compounds obtained from Strategy 2 are within 200, and only a limited number of compounds from Strategy 1 are within top-ranked 200, which is in part responsible for the observed difference in activity. Furthermore, the rankings of the compounds with >60% inhibitory activity @ 10 μg/ml from Strategy 1 are 132, 147, 189, 375, 38 and 252 respectively, which are all higher than 400. In other words, no compounds with rankings lower than 400 are found potent inhibitors. The rankings of two most potent inhibitors, TP-C1-17 and TP-C1-47 showing <10 μM activity, are both higher than 200, which in another perspective supports the idea that higher success rate of Strategy 2 could be the result of considering higher ranked compounds due to the using of a lower number of compounds for clustering.
Molecular diversity of VS candidates
The main purpose of clustering top-ranked 1,000 rather than fewer molecules in Strategy 1 is to maximize the molecular diversity of the screened compounds. So we also compared the molecular diversity of the compounds from Strategy 1 to that from Strategy 2. FCFP_6 fingerprint and tanimoto metric were used to calculate molecular diversity in DS 2.5. As is shown in Table 1, the molecular diversity of the candidates from both strategies was calculated based on the fingerprint distance metrics, which is defined as 1 - similarity for every pair of molecules. We can find that the average fingerprint distances for Strategy 1 and Strategy 2 are 0.881 and 0.859, respectively, which suggests that the molecular diversity of the candidates from Strategy 2 with fewer clustering compounds still maintains compared with that from Strategy 1. The distributions of the fingerprint distance for the molecules from Strategy 1 and Strategy 2 were also plotted in Supplementary Fig. S4. We can find that both sets are normal-distributed and well overlapped. The maximum fingerprint distances are almost identical (Strategy 1: 0.959; Strategy 2: 0.958), but the minimum fingerprint distances are obviously different (Strategy 1: 0.621; Strategy 2: 0.370). This means that several compounds from Strategy 2 are structurally similar. However these compounds have little influence on the average fingerprint distance, so most compounds from Strategy 2 are still structurally diverse. Therefore, using fewer molecules in clustering analysis is suggested to improve the hit rates, but the exact number of compounds should be assessed in advance based on the molecular diversity of the selected compounds. However, the outcome from this study can only be a welcome indication when no extensive testing of the other protein targets is carried out. So this work can only be regarded as a case study, which is possibly target dependent. Several VS projects against other protein targets are ongoing in our lab, and we hope the outcome could be generalized.
Biological activity and binding mechanisms of the active compounds
Based on the results of preliminary screening assays, ~10% compounds from Strategy 1 and ~40% compounds from Strategy 2 were found to inhibit ≥80% Tie-2 enzymatic activity. The dose-response effects of these candidate compounds were then evaluated to determine the half-maximal inhibitory concentrations (IC50). The compounds were tested in duplicate or more for every compound concentration. The dose-response curve was generated and plotted for each compound, and the IC50 value was calculated from each curve. The cutoff percentage inhibitory values of the compounds for IC50 determination were set to 60% and 80% for Strategy 1 and Strategy 2, respectively. 6 compounds from Strategy 1 and 18 compounds from Strategy 2 were finally tested, and the results were summarized in Table 2. The IC50 values of 13 compounds from Strategy 2 (TP-C2) were under 10 μM, while only 2 compounds from Strategy 1 (TP-C1) exhibited <10 μM activity.
The IC50 values of three most potent compounds from each strategy (TP-C1-17, 47, 52 and TP-C2-9, 33, 50) are 4.45 μM, 5.15 μM, 15.3 μM, 1.86 μM, 3.31 μM, and 1.73 μM, respectively (Figs 5 and 6). The novelty of these inhibitors was also assessed by evaluating the structural similarity to the reported type-I inhibitors of Tie-2 that were retrieved from the BindingDB database. The molecular similarity was computed based on the pairwise Tanimoto similarity using FCFP_6 fingerprints41. According to our calculations, the maximum similarities of compounds TP-C1-17, 47, 52 and TP-C2-9, 33, 50 are 0.125, 0.141, 0.147, 0.268, 0.204, and 0.252, respectively, where all the obtained known inhibitors were compared to each compound and the similarity value of the most similar inhibitor was kept. All the 6 compounds show low structural similarity to the reported inhibitors, indicating that the candidate compounds discovered are novel Tie-2 inhibitors.
The binding mechanisms of three most potent inhibitors from each strategy were further investigated by the Glide XP docking simulations (Figs 5 and 6). The dominating residues for the binding of these 6 compounds are Val838, Lys855, Ala905, Ala981, Asp982 and Phe983. However, the interaction pattern in each inhibitor/Tie-2 system is different. In the binding of compound TP-C1-17 to Tie-2, the nitrogen atom of pyrazine group in TP-C1-17 can form a hydrogen bond with the backbone of Asp982. The pyrazine ring and the substituted methyl group can form arene-H contacts with the residues Lys855 and Phe983. In addition, arene-H interaction can also be found between the benzene ring and the residue Val838. The inhibitory activity of the compound TP-C1-47 is similar to that of TP-C1-17, but the interaction patterns are relatively different. Three hydrogen bonds interacting with the residues Lys855 and Ala905 are observed. The side chains of Asp982 can form arene-H interactions with the benzene ring in TP-C1-47 rather than a hydrogen bond. The role of Lys855, which interacts with the benzene ring also via arene-H contacts, is similar to the binding of TP-C1-17. The activity of compound TP-C1-52 is ~3 fold decreased compared to those of TP-C1-17 and TP-C1-47. This may result from the lack of strong hydrophobic interactions in the surrounding region of Asp982, Phe983 and Lys855, where only a hydrogen bond is observed between Lys855 and the nitrogen atom of the thiadiazole group. Another hydrogen bond can also be found between the carbonyl of isoindole and the backbone of Ala905. In addition, the isoindole ring in TP-C1-52 can also form arene-H interactions with Val838.
In the binding of compound TP-C2-9, the benzene rings in both sides of the molecules, which can form π interactions with Lys855, Phe983, Ile830 and Leu971, play major roles in ligand binding. Different from the other 5 compounds, TP-C2-9 can form a hydrogen bond with Asn909 showing a unique binding mode of action. For compound TP-C2-33, hydrophobic interactions also dominate the binding to Tie-2, where Ala905, Ala981 and Phe983 form arene-H interactions with TP-C2-33. A hydrogen bond is also formed between the nitrogen atom of pyridine group in TP-C2-33 and the backbone of Asp982, which is similar to the hydrogen binding of TP-C1-17. In the structure of TP-C2-50, the biphenyl group and naphthalene ring are quite hydrophobic, which form strong arene-H interactions with Val838, Lys855 and Ala981. Besides, the substituted hydroxyl group in naphthalene ring can interact with the backbone of Ala905 via hydrogen bonding.
Preliminary structure−activity relationship (SAR) analysis
The candidate compound (TP-C2-9), which represents novel chemotypes for Tie-2 inhibitors, was selected for further optimization and preliminary SAR analysis. Hit optimization was performed based on the technique of substructure search to retrieve the compounds that contain the query substructure in ChemBridge database. The core scaffold of compound TP-C2-9, which is colored red in Tables 3 and 4, was taken as the template structure for searching. 60 analogues of TP-C2-9 were finally selected and purchased for the second-round screening. These compounds were firstly assessed for their ability to inhibit Tie-2 at the concentration of 5 μg/mL. The results of kinase assay clearly revealed that different substitutes on R1, R2 and R3 phenyl ring have obvious effects on the inhibitory activity. The IC50 values of six compounds (TP-C3-1, TP-C3-3, TP-C3-21, TP-C3-43, TP-C3-45 and TP-C3-55) are lower than 5 μM, wherein the most potent one (TP-C3-43) can reach 1.64 μM. However, we can also observe that most analogues exhibit weak inhibitory activity (<50% at 5 μg/mL). The low hit rates of the compounds from substructure search, on the contrary, demonstrated the effectiveness of VS that selected TP-C2-9 (IC50 = 1.86 μM) rather than other analogues as the preferential compound.
The inhibitory activity of compound TP-C3-3 (IC50 = 2.82 μM) is only slightly decreased compared to that of TP-C2-9, where two methoxy groups are removed from the R2 phenyl ring. This indicates that the removal of methoxy groups has little influence on ligand binding. However, replacement of 3-methyl group by 4-tert-butyl in R1 ring (compound TP-C3-2) leads to~10 fold decreases in activity compared with compound TP-C3-3. To gain rational insights into the difference of the binding modes, MM/GBSA binding energy calculations and decomposition analysis based on MD simulations were performed by examining the contribution of each residue to binding. Figure 7 illustrated the energetic contributions of the dominating residues from the MM/GBSA decomposition analysis. We can find that the contributions of most residues for both compounds TP-C2-9 and TP-C3-3 are similar, implying that removal of methoxy groups from R2 ring does not affect the main interactions between inhibitors and Tie-2. For the binding of compound TP-C3-2 (see Supplementary Fig. S1), several critical interaction patterns are influenced, especially the arene-H contacts with the residues Ile902 and Leu971. As illustrated in Fig. 7, the contributions of Ile902 and Leu971 for TP-C3-2, which are dominated by the non-polar interactions, are obviously smaller than those for TP-C3-3. In addition, the impaired interactions with Tyr904 also account for the inferior binding affinity of TP-C3-2. Interestingly, the binding to Asp982, where arene-H interactions can be observed, is found stronger than both compounds TP-C2-9 and TP-C3-3, however, this cannot make up for the decreased binding affinity to other amino acids.
The methyl group on R1 was found crucial for inhibitor binding since the inhibitory activities of all the compounds (TP-C3-4~TP-C3-15 and TP-C3-26~TP-C3-30) were greatly reduced when R1 methyl is removed. The IC50 value of the most potent one within this series (TP-C3-15) is only 13.79 μM with ~10 fold decline in activity compared to TP-C2-9. This phenomenon is mainly caused by the interruption of arene-H contacts between Phe983 and methyl group. Substitutions of methyl groups on different positions of R2 phenyl can significantly influence the potency. Compound TP-C3-21 with methyl groups introduced on 2, 4-positions of R2 ring displays strong inhibitory activity (71.19% inhibition, 5 μg/mL), while almost no activity was observed for TP-C3-22 with methyl groups substituted on 3, 5-positions. Detailed energy decomposition analysis revealed that the impaired interactions with the residues Val838, Lys855 and Asn909 determinate the decreased binding affinity of TP-C3-22 to Tie-2. The residues Val838 and Lys855 can form arene-H contacts with compound TP-C3-21, but these interactions are not observed in the binding of TP-C3-22 (see Supplementary Fig. S1), which directly leads to the reduction of the binding energy. Except for the impaired van der Waals interactions with Val838 and Lys855, the electrostatic interactions with Asn909 (~1 kcal/mol drop in energy) are also responsible for the decreased binding affinity. Thus, introducing methyl groups on different positions of R2 phenyl ring can cause structural rearrangement of the inhibitor as well as the side chains of surrounding amino acids, which leads to different binding potency. Compound TP-C3-43 is the most potent inhibitor with an IC50 value of 1.64 μM, but replacement of chlorine by methyl on the 3-position of R1 ring (TP-C3-47) is unfavorable. Structural analysis (see Supplementary Fig. S1) revealed that most critical interactions are not observed in the binding of TP-C3-47 to Tie-2. Even small variations in structure may cause significant change of inhibitor-protein interaction patterns and finally lead to the change of activity. Compounds listed in Table 5 mainly focus on modifications with saturated or unsaturated six-member rings introduced on R1 or R2 phenyl, but no compounds displayed strong inhibitory activity at the concentration of 5 μg/mL, implying that no sufficient space around R1 or R2 ring can accommodate bulk groups.
Based on the discussions above, most of the dominating amino acids are hydrophobic, so the non-polar interactions, especially the bindings to the residues Ile830, Val838, Ile902, Ala981 and Leu971 via π/arene-H contacts, play key roles in inhibitor binding. In addition, the hydrophilic regions involving the residues Lys855, Asn909 and Asp982 may benefit to form H-bond interactions and should be paid special attention in further optimization. The inhibitors discovered and the detailed binding modes of action facilitate the subsequent focused medicinal chemistry study to expand the SAR of potential drug candidates.
Conclusions
Targeted intervention of ANG/Tie-2 signaling pathway in preliminary vessels represents a promising strategy for the treatment of various life-threatening diseases. In the present work, a class of compounds was identified as potential inhibitors of Tie-2 through structure-based VS and substructure searching. By comparing the performance of two different filtering strategies employed in VS, we found that the hit rate can be well improved when stricter drug-likeness criteria and top-200 rather than top-1000 compounds for clustering analysis were used, and meanwhile, the molecular diversity of the compounds still maintained. Further hit-based optimization was also performed to give a preliminary SAR using substructure searching technology. The information presented and the novel inhibitors discovered in this work will benefit to developing new anti-angiogenesis drug candidates.
Materials and Methods
Preparations for Protein Structures and Small Molecule Datasets
All structure preparations for molecular docking were carried out using the Protein Preparation wizard in Maestro 9.0 program42 starting from previously reported crystal structures of human Tie-2 in complex with CEP11207 (PDB code: 3L8P)21 and a thiazolopyrimidine inhibitor (PDB code: 2OO8)43. All water molecules in the original crystal structures were removed from the coordinate set. Hydrogen atoms and partial charges were added to the protein, followed by a restrained partial minimization which is terminated when the root-mean-square deviation reached 0.3 Å.
The reported inhibitors of Tie-2 were retrieved from the BindingDB database44. In order to evaluate the prediction accuracy of docking simulation in distinguishing type-I inhibitors from non-inhibitors, the type-II inhibitors and those with high IC50 values (IC50 > 100 μM) were removed from the dataset of known inhibitors45. The eventual inhibitor dataset was composed of 200 structurally diverse molecules by extracting from the above cleared dataset using the Find Diverse Molecule module in Discovery Studio 2.5 (DS 2.5)46. The 20,000 non-inhibitors of Tie-2 was randomly selected from the ChemBridge chemical database (~1,020,000 screening compounds) using DS 2.546. All the selected inhibitors and non-inhibitors were then minimized using the OPLS-2005 force field47. Preparations of all the compounds were subsequently performed using the LigPrep module in Schrödinger 9.048, where the protonated states were predicted at pH = 7.0 ± 2.0.
Molecular Docking Simulations
The Glide module in Schrödinger was utilized to perform all the docking simulations. The protein grid box for molecular docking was produced by centering on the ligand in the active site using the Receptor gird generation module. The scaling factor for van der Waals radii was set to 0.8, and the partial atomic charge cutoff value was set to 0.15. Eventually, all the prepared compounds were docked into the corresponding crystal structures of Tie-2. Two different precision modes of docking were utilized, including the standard precision (SP) and extra precision (XP). Though the computational speed of the high throughput virtual screening (HTVS) mode is the fastest among the three docking modes, the prediction accuracy is quite low49. So the HTVS docking mode was not employed in our VS workflow.
Molecular Dynamics (MD) Simulations and Binding energy Calculations
MD simulations were carried out to elucidate the binding patterns of the studied inhibitors. The Tie-2/inhibitor complexes predicted by the docking simulations were employed as the starting structures for the MD simulations. All the MD simulations were carried out in Amber1450. The electrostatic potential of each inhibitor was calculated at the HF/6-31 G* level after optimization by semi-empirical AM1 method in Gaussian 09 program51. Then, the atomic partial charges of the inhibitors were generated by fitting the electrostatic potentials using the RESP fitting method52.
The ff99SB force field53 and the general AMBER force field (gaff)54 were used for the proteins and inhibitors, respectively. Each system was inserted into a cubic TIP3P water box with at least 8 Å away from any solute atom. The long-range electrostatics was handled by the Particle Mesh Ewald (PME) method55.
Each system was firstly minimized by the sander program in three steps: 1) 500 cycles of steepest descent and 500 cycles of conjugate gradient minimizations with 50 kcal/mol/Å2 restraint on protein backbone; 2) 500 cycles of steepest descent and 500 cycles of conjugate gradient minimizations with 10 kcal/mol/Å2 restraint; 3) 1000 cycles of steepest descent and 4000 cycles of conjugate gradient minimizations without any restraint. Afterwards the systems were gradually heated from 0 to 300 K in 50 ps with 2.0 kcal/mol/Å2 restraint, and 5 ns NPT (P = 1 atm and T = 300 K) MD simulations were then carried out with a time step of 2 fs. All hydrogen bonded atoms were constrained using the SHAKE algorithm56. The interval time of saving the coordinates was set to 10 ps.
The Molecular Mechanics/Generalized Born Solvent Area (MM/GBSA) method implemented in the mm_pbsa module in Amber11 was employed to calculate the binding energies of the studied inhibitors57,58,59,60,61. The dielectric constants for solvent and solute were set to 80 and 1, respectively. The modified GB model (igb = 2)62 developed by Onufriev and coworkers was used to calculate the polar part of desolvation energy, and the LCPO algorithm63 was employed to compute the non-polar part of desolvation based on the solvent accessible surface area (SASA): ΔGSA = 0.0072 × ΔSASA. Due to the low prediction accuracy and relatively high computational cost, the entropic contributions to the total binding free energy were not calculated57,64. Eventually, 200 structures evenly extracted from 3 to 5 ns MD trajectories were used to predict binding energies.
The MM/GBSA decomposition analysis was used to highlight the important residues to ligand binding59,65. Except for the non-polar part of the solvation free energy (∆GSA), which was calculated based on ΔSASA using the ICOSA algorithm65, the other terms were calculated based on the same parameters used in the MM/GBSA calculations.
Reagents and Materials for Experiments
All anhydrous solvents and reagents used in this work were obtained from commercial sources. The Tie-2 kinase inhibitor (CAS#: 948557-43-5, purity ≥ 97%) was purchased from Merck Calbiochem (Darmstadt, Germany) and used as a positive control inhibitor in bioassays40. Both the purified recombinant human Tie-2 protein (Catalog#: PV3628) and Z′-Lyte® kinase assay kit (Catalog#: PV3178) were purchased from Thermo Fisher Scientific Inc. The candidate compounds from VS were purchased from ChemBridge and were tested by LC-MS and/or NMR to confirm sample identity and purity (compound information are listed in Supplementary Table S1). All the purchased screening compounds were firstly dissolved in dimethyl sulfoxide (DMSO) at a stock concentration of 5 mg/ml. The final working concentration of DMSO is less than 1%.
Enzymatic Activity Assay
Compounds were tested for their ability to inhibit Tie-2 kinase activity using Z′-Lyte® kinase assay kit (ser/thr 5 peptide) with NUNC 384-well plate. In the first step, the test compounds and ATP solution were firstly prepared at 4 × desired concentration. The mixture of recombinant human Tie-2 protein and non-phosphorylated peptide at 2 × desired concentration was prepared in kinase reaction buffer A consisting of 50 mM HEPES (pH 7.5), 0.01% BRIJ-35, 10 mM MgCl2, 4 mM MnCl2, 1 mM EGTA and 2 mM DTT. Reactions were then carried out in a solution containing 2.5 μL test compounds, 5 μL Tie-2/Tyr 05 peptide mixture (10 ng Tie-2 and 2 μM Tyr 05 peptide) and 2.5 μL ATP (100 μM). Then, the assay plate was shaken for 30 seconds on a plate shaker, and incubated for 1 hour at room temperature (20~25 °C). After sufficient kinase reactions, 5 μL development reagent A was added to each well, and the plate was incubated for another hour. Finally, 5 μL stop reagent was added to terminate the reactions, and the coumarin and fluorescein emission signals (excitation: 400 nm; emission: 445 and 520 nm, respectively) were measured on a fluorescence plate reader (BioTek Synergy 4).
Additional Information
How to cite this article: Pan, P. et al. In Silico Exploration for Novel Type-I Inhibitors of Tie-2/TEK: The Performance of Different selection strategy in Selecting Virtual Screening Candidates. Sci. Rep. 6, 37628; doi: 10.1038/srep37628 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Risau, W. Mechanisms of angiogenesis. Nature 386, 671–674 (1997).
Siemann, D. W., Chaplin, D. J. & Horsman, M. R. Vascular‐targeting therapies for treatment of malignant disease. Cancer 100, 2491–2499 (2004).
Villanueva, M. T. Angiogenesis: A sudden rush of blood to the tumour. Nat. Rev. Cancer 15, 135–135 (2015).
Musumeci, F., Radi, M., Brullo, C. & Schenone, S. Vascular Endothelial Growth Factor (VEGF) Receptors: Drugs and New Inhibitors. J. Med. Chem. 55, 10797–10822 (2012).
Ishak, R. S., Aad, S. A., Kyei, A. & Farhat, F. S. Cutaneous manifestations of anti-angiogenic therapy in oncology: review with focus on VEGF inhibitors. Crit. Rev. Oncol./Hematol. 90, 152–164 (2014).
Augustin, H. G., Koh, G. Y., Thurston, G. & Alitalo, K. Control of vascular morphogenesis and homeostasis through the angiopoietin–Tie system. Nat. Rev. Mol. Cell Biol. 10, 165–177 (2009).
Hu, J. et al. Endothelial cell-derived angiopoietin-2 controls liver regeneration as a spatiotemporal rheostat. Science 343, 416–419 (2014).
Biel, N. M. & Siemann, D. W. Targeting the Angiopoietin-2/Tie-2 axis in conjunction with VEGF signal interference. Cancer Lett. In press (2014).
Demetri, G. D. et al. Efficacy and safety of sunitinib in patients with advanced gastrointestinal stromal tumour after failure of imatinib: a randomised controlled trial. The Lancet 368, 1329–1338 (2006).
Escudier, B. et al. Sorafenib for treatment of renal cell carcinoma: final efficacy and safety results of the phase III treatment approaches in renal cancer global evaluation trial. J. Clin. Oncol. 27, 3312–3318 (2009).
Bergers, G. & Hanahan, D. Modes of resistance to anti-angiogenic therapy. Nat. Rev. Cancer 8, 592–603 (2008).
Casanovas, O., Salazar, R. & Tabernero, J. Multi-target angiokinase inhibitors to fight resistance. Cell Cycle 13, 2649–2650 (2014).
Vasudev, N. S. & Reynolds, A. R. Anti-angiogenic therapy for cancer: current progress, unresolved questions and future directions. Angiogenesis 17, 471–494 (2014).
Tozer, G. et al. Tumour vascular disrupting agents: combating treatment resistance. Br. J. Radiol. (2014).
Rivera, L. B. et al. Intratumoral Myeloid Cells Regulate Responsiveness and Resistance to Antiangiogenic Therapy. Cell reports 11, 577–591 (2015).
Gerald, D., Chintharlapalli, S., Augustin, H. G. & Benjamin, L. E. Angiopoietin-2: an attractive target for improved antiangiogenic tumor therapy. Cancer Res. 73, 1649–1657 (2013).
Brown, J. L. et al. A Human Monoclonal Anti-ANG2 Antibody Leads to Broad Antitumor Activity in Combination with VEGF Inhibitors and Chemotherapy Agents in Preclinical Models. Mol. Cancer Ther. 9, 145–156 (2010).
Leow, C. C. et al. MEDI3617, a human anti-angiopoietin 2 monoclonal antibody, inhibits angiogenesis and tumor growth in human tumor xenograft models. Int. J. Oncol. 40, 1321–1330 (2012).
Huang, H. et al. Specifically targeting angiopoietin-2 inhibits angiogenesis, Tie2-expressing monocyte infiltration, and tumor growth. Clin. Cancer. Res. 17, 1001–1011 (2011).
Papadopoulos, K. et al. In Patients with Advanced Solid Tumor Malignancies. 2011 ASCO Annual Meeting Abstracts Part 1. 29, TPS159 (2011).
Hudkins, R. L. et al. Synthesis and Biological Profile of the pan-Vascular Endothelial Growth Factor Receptor/Tyrosine Kinase with Immunoglobulin and Epidermal Growth Factor-Like Homology Domains 2 (VEGF-R/TIE-2) Inhibitor 11-(2-Methylpropyl)-12, 13-dihydro-2-methyl-8-(pyrimidin-2-ylamino)-4 H-indazolo [5, 4-a] pyrrolo [3, 4-c] carbazol-4-one (CEP-11981): A Novel Oncology Therapeutic Agent. J. Med. Chem. 55, 903–913 (2012).
Gross, S. et al. In Kinases Proceedings of the Cancer Meeting: Lessons from the Clinic. Abstract nr 14 (2006).
Saha, I. et al. Binding Activity Prediction of Cyclin-Dependent Inhibitors. J. Chem. Inf. Model. 55, 1469–1482 (2015).
Xu, L. et al. Discovery of novel inhibitors targeting the macrophage migration inhibitory factor via structure-based virtual screening and bioassays. J. Med. Chem. 57, 3737–3745 (2014).
Shen, M. et al. Discovery of Rho-kinase inhibitors by docking-based virtual screening. Mol. BioSyst. 9, 1511–1521 (2013).
Singh, N., Tiwari, S., Srivastava, K. K. & Siddiqi, M. I. Identification of Novel Inhibitors of Mycobacterium tuberculosis PknG Using Pharmacophore Based Virtual Screening, Docking, Molecular Dynamics Simulation, and Their Biological Evaluation. J. Chem. Inf. Model. 55, 1120–1129 (2015).
Tian, S. et al. Assessing an Ensemble Docking-Based Virtual Screening Strategy for Kinase Targets by Considering Protein Flexibility. J. Chem. Inf. Model. 54, 2664–2679 (2014).
Lindh, M. et al. Toward a Benchmarking Data Set Able to Evaluate Ligand-and Structure-based Virtual Screening Using Public HTS Data. J. Chem. Inf. Model. 55, 343–353 (2015).
Middendorp, S. J., Puthenkalam, R., Baur, R., Ernst, M. & Sigel, E. Accelerated Discovery of Novel Benzodiazepine Ligands by Experiment-Guided Virtual Screening. ACS Chem. Biol. 9, 1854–1859 (2014).
Kincaid, V. A. et al. Virtual Screening for UDP-Galactopyranose Mutase Ligands Identifies a New Class of Antimycobacterial Agents. ACS Chem. Biol. 10, 2209–2218 (2015).
Steindl, T. M., Schuster, D., Laggner, C. & Langer, T. Parallel Screening: A Novel Concept in Pharmacophore Modeling and Virtual Screening†. J. Chem. Inf. Model. 46, 2146–2157 (2006).
Therrien, E. et al. Docking Ligands into Flexible and Solvated Macromolecules. 7. Impact of Protein Flexibility and Water Molecules on Docking-Based Virtual Screening Accuracy. J. Chem. Inf. Model. 54, 3198–3210 (2014).
Luke, R. W. A. et al. Novel thienopyrimidine and thiazolopyrimidine kinase inhibitors with activity against Tie-2 in vitro and in vivo. Bioorg. Med. Chem. Lett. 19, 6670–6674 (2009).
Hudkins, R. L. et al. Synthesis and Biological Profile of the pan-Vascular Endothelial Growth Factor Receptor/Tyrosine Kinase with Immunoglobulin and Epidermal Growth Factor-Like Homology Domains 2 (VEGF-R/TIE-2) Inhibitor 11-(2-Methylpropyl)-12,13-dihydro-2-methyl-8-(pyrimidin-2-ylamino)-4H-indazolo[5,4-a]pyrrolo[3,4-c]carbazol-4-one (CEP-11981): A Novel Oncology Therapeutic Agent. J. Med. Chem. 55, 903–913 (2012).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
ACD/ADME Suite 5.0, Advanced Chemistry Development Inc., Toronto, Canada (2011).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Del. Rev. 64 Supplement, 4–17 (2012).
Oprea, T. I. Property distribution of drug-related chemical databases. J. Comput.-Aided Mol. Des. 14, 251–264 (2000).
Hassan, M., Bielawski, J. P., Hempel, J. C. & Waldman, M. Optimization and visualization of molecular diversity of combinatorial libraries. Mol. Divers. 2, 64–74 (1996).
Semones, M. et al. Pyridinylimidazole inhibitors of Tie2 kinase. Bioorg. Med. Chem. Lett. 17, 4756–4760 (2007).
Willett, P. Similarity-based virtual screening using 2D fingerprints. Drug Discov. Today 11, 1046–1053 (2006).
Schrödinger. Maestro, Version 9.0 Schrödinger LLC, New York, NY. (2009).
Hodous, B. L. et al. Synthesis, structural analysis, and SAR studies of triazine derivatives as potent, selective Tie-2 inhibitors. Bioorg. Med. Chem. Lett. 17, 2886–2889 (2007).
Nicola, G., Liu, T., Hwang, L. & Gilson, M. BindingDB: a protein-ligand database for drug discovery. Biophys. J. 102, 61a (2012).
Liu, T., Lin, Y., Wen, X., Jorissen, R. N. & Gilson, M. K. BindingDB: a web-accessible database of experimentally determined protein–ligand binding affinities. Nucleic Acids Res. 35, D198–D201 (2007).
Discovery Studio 2.5, Accelrys Inc., San Diego, CA (2009).
Kaminski, G. A., Friesner, R. A., Tirado-Rives, J. & Jorgensen, W. L. Evaluation and Reparametrization of the OPLS-AA Force Field for Proteins via Comparison with Accurate Quantum Chemical Calculations on Peptides†. J. Phys. Chem. B 105, 6474–6487 (2001).
Schrödinger, M. Version 9.0. Schrödinger LLC, New York, NY (2009).
Pan, P. et al. Identification and Preliminary SAR Analysis of Novel Type-I Inhibitors of TIE-2 via Structure-Based Virtual Screening and Biological Evaluation in in vitro Models. J. Chem. Inf. Model. 55, 2693–2704 (2015).
Case, D. et al. AMBER 14 (University of California), San Francisco (2014).
Frisch, M. J. et al. Gaussian 09, Gaussian. Inc., Wallingford, CT (2009).
Bayly, C. I., Cieplak, P., Cornell, W. & Kollman, P. A. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model. J. Phys. Chem. 97, 10269–10280 (1993).
Duan, Y. et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24, 1999–2012 (2003).
Wang, J. M., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: An W log (N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Ryckaert, J. P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Wang, J., Wang, W., Kollman, P. A. & Case, D. A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graphics Model. 25, 247–260 (2006).
Kollman, P. A. et al. Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Accounts Chem. Res. 33, 889–897 (2000).
Hou, T., Li, N., Li, Y. & Wang, W. Characterization of Domain-peptide Interaction Interface: Prediction of SH3 Domain-Mediated Protein-protein Interaction Network in Yeast by Generic Structure-Based Models. J. Proteome Res. 11, 2982–2995 (2012).
Hou, T., Zhang, W., Case, D. A. & Wang, W. Characterization of domain-peptide interaction interface: A case study on the amphiphysin-1 SH3 domain. J. Mol. Biol. 376, 1201–1214 (2008).
Hou, T. J. et al. Characterization of Domain-Peptide Interaction Interface. Mol. Cell. Proteomics 8, 639–649 (2009).
Onufriev, A., Bashford, D. & Case, D. A. Exploring protein native states and large-scale conformational changes with a modified generalized born model. Proteins: Struct., Funct., Bioinf. 55, 383–394 (2004).
Weiser, J., Shenkin, P. S. & Still, W. C. Approximate atomic surfaces from linear combinations of pairwise overlaps (LCPO). J. Comput. Chem. 20, 217–230 (1999).
Hou, T. J., Li, Y. Y. & Wang, W. Prediction of peptides binding to the PKA RII alpha subunit using a hierarchical strategy. Bioinformatics 27, 1814–1821 (2011).
Gohlke, H., Kiel, C. & Case, D. A. Insights into protein-protein binding by binding free energy calculation and free energy decomposition for the Ras-Raf and Ras-RalGDS complexes. J. Mol. Biol. 330, 891–914 (2003).
Acknowledgements
This study was supported by the National Science Foundation of China (21575128), the National Major Basic Research Program of China (2016YFB0201700 and 2016YFA0501701), and the Innovation fund for Technology based Firms (14C26214402675, 2013J4400175).
Author information
Authors and Affiliations
Contributions
P.P. and T.H. conceived the projects, P.P., H.S. and H.L. conducted the experiments of structure-based virtual screening, P.P., D.L. and W.Z. conducted the biological inhibitory activity experiments of the compounds, X.K. and Y.L. analyzed the results and studied the binding mechanism of the active compounds. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Pan, P., Sun, H., Liu, H. et al. In Silico Exploration for Novel Type-I Inhibitors of Tie-2/TEK: The Performance of Different Selection Strategy in Selecting Virtual Screening Candidates. Sci Rep 6, 37628 (2016). https://doi.org/10.1038/srep37628
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep37628
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.