Abstract
Little is known about the interplay between Ca2+ and H2O2 signaling in stressed cool-season turfgrass. To understand better how Ca2+ and H2O2 signals are integrated to enhance grass acclimation to stress conditions, we analyzed the rearrangements of endogenous ion accumulation, antioxidant systems and secondary metabolism in roots, stems and leaves of perennial ryegrass (Lolium perenne L.) treated with exogenous Ca2+ and H2O2 under salinity. Ca2+ signaling remarkably enhanced the physiological response to salt conditions. Ca2+ signaling could maintain ROS homeostasis in stressed grass by increasing the responses of antioxidant genes, proteins and enzymes. H2O2 signaling could activate ROS homeostasis by inducing antioxidant genes but weakened Ca2+ signaling in leaves. Furthermore, the metabolic profiles revealed that sugars and sugar alcohol accounted for 49.5–88.2% of all metabolites accumulation in all treated leaves and roots. However, the accumulation of these sugars and sugar alcohols displayed opposing trends between Ca2+ and H2O2 application in salt-stressed plants, which suggests that these metabolites are the common regulatory factor for Ca2+ and H2O2 signals. These findings assist in understanding better the integrated network in Ca2+ and H2O2 of cool-season turfgrass’ response to salinity.
Similar content being viewed by others
Introduction
Calcium (Ca2+) and hydrogen peroxide (H2O2) are vital secondary messengers in plant signaling networks, and they could trigger different physiological and molecular responses to various environmental stresses in plant1,2,3. Ca2+ and H2O2 never work alone as the secondary messengers in cell signal transduction, however, most of previous studies focused on the single Ca2+ or H2O2 4,5. Ca2+ signaling could readily regulate ion transport behavior and cell wall enzyme activities in plant cells6. H2O2 was mainly generated at the plant plasma membrane, and could enhance protein tyrosine phosphorylation or hyperoxidation in cytoplasm7. Cellular H2O2 level is controlled by a complex set of antioxidant enzymes such as catalase (CAT)8. Hu et al.9 reported that there was one crosstalk between Ca2+ and H2O2 signaling in the regulation of plant defenses. Some studies demonstrated that Ca2+ could activate CAT enzyme activity to minimize cellular H2O2 level, and function as an upstream component in H2O2 stress signals in plants10. However, Sun et al.2 suggested that H2O2 may function as an upstream signal to regulate Ca2+ in NaCl-stressed Populus euphratica cells. Despite their relevance in many cell biology aspects, integrated changes in spatio-temporal regulation of H2O2 and Ca2+ signaling remains elusive in stressed plants7,10,11.
Reactive oxygen species (ROS) accumulation, such as H2O2, HO and O2−, could be triggered by environmental stresses12. In plant cells, the reaction or conversion summarized by the expression: O2− → H2O2 → HO → H2O is catalyzed by different antioxidant enzymes such as superoxide dismutase (SOD, EC 1.15.1.1), peroxidase (POD, EC 1.11.1.7), ascorbate peroxidase (APX, EC 1.11.1.11), catalase (CAT, EC 1.11.1.16)13,14,15. Generally, these enzymes have multiple molecular forms (isoenzymes) in different plant cell organelles. For example, SOD can be divided into ChlCu/ZnSOD, CytCu/ZnSOD, MnSOD, NiSOD or FeSOD isoenzymes depending on their metal cofactor, which are localized in the chloroplasts, cytosol and mitochondria. All these isoforms play cooperative role in protecting each organelle and minimizing tissue injury caused by ROS16,17. In addition, the specific isoenzymes in many plants can be recognized as the biomarkers in response to various environmental stresses, and regulated by distinct genes12. APX, POD and CAT are the most abundant antioxidant enzymes in detoxifying H2O2, which catalyze the reduction of H2O2 to water in plant organelles18. APX reduces H2O2 in chloroplasts, mitochondria, peroxisomes and apoplastic space. CAT on the other hand acts in glyoxysomes and peroxisomes, which minimize cellular H2O2 level19. On the contrary, high H2O2 accumulation may increase CAT activity or CAT gene expression20. Consistently, H2O2 treatment could activate Ca2+ channels and simultaneously mediate Ca2+ influx in protoplasts and increase of [Ca2+]cyt in intact guard cells21. However, Yang and Poovaiah22 reported that Ca2+/CaM could down-regulate H2O2 levels in plants by stimulating the catalytic activity of plant CAT. Jiang and Huang23 showed that Ca2+ could increase CAT activity in heat stressed grasses. The regulatory mechanisms among Ca2+ and H2O2 signaling, and antioxidant enzymes also remain unclear, while several lines of evidence indicate one interaction between their multiple signal transduction.
Many studies have revealed that regulation of Ca2+ and H2O2 signaling involves many genes and metabolites in stressed plant5,22,24. Using gas chromatography-mass spectrometry (GC-MS) analysis of polar compounds, Shi et al.5 found that 42 metabolites including amino acids, organic acids, sugars, and sugar alcohols were regulated by exogenous Ca2+ treatment in bermudagrass [Cynodon dactylon (L.) Pers.] in response to cold stress. Dittami et al.24 showed that the accumulation level of mannitol, n-3 to n-6 ratio of polyunsaturated fatty acids and 22 amino acids changed after H2O2 treatment in brown alga Ectocarpus siliculosus. Furthermore, previous studies have demonstrated that the majority of the expressed genes could be activated by these metabolites, which may be the gene regulation end products25. Furuichi et al.26 reported that the increase of [Ca2+]cyt might initiate the expression of genes, such as AtSUC1 and AtSUC2, involved in biosynthesis of storage carbohydrate. On the contrary, exogenous application of hexoses also might mediate by monosaccharide-H+ co-transporters to increase [Ca2+]cyt level. In addition, H2O2 signaling activating gene expression also involved metabolites such as proline and sucrose8. Therefore, we speculated that the changes in metabolites abundance involved in the crosstalk between Ca2+ and H2O2 signaling may be regarded as a major regulation character in plants in response to stress environment.
Salinity is a major abiotic stress and increasingly threatens agriculture and food production worldwide27. Perennial ryegrass (Lolium perenne L.) is an extensively utilized cool-season grass globally used for forage, turfgrass and soil stabilization in United States, Australia, New Zealand, Japan and most of European countries because of its rapid establishment rate28. Our previous study confirmed that salt stress could cause oxidative injury and damage to cell membrane integrity in perennial ryegrass, and antioxidant enzymes decreased the stress toxicity12,17. In the present study, we further investigated the impact of exogenous Ca2+ or H2O2 on the secondary metabolism and oxidative signaling in perennial ryegrass under salt stress. The objective of this study was to further understand the orchestrated response of antioxidant system and metabolisms induced by H2O2/Ca2+ signaling in cool-season turfgrass in response to salt stress.
Results
The impact of exogenous Ca2+ on physiological features of salt-stressed perennial ryegrass
As shown in Fig. 1, 300 mM NaCl treatment enhanced the levels of MDA, EL and H2O2, but minimized the turf quality score in perennial ryegrass leaves. However, 7 mM Ca(NO3)2·4H2O added in 300 mM NaCl treatment solution reduced the levels of MDA, EL and H2O2, and increased the turf quality score compared with 300 mM NaCl treated plants (Fig. 1). These alleviated injury features in salt-treated plants induced by Ca2+ could be observed from the fourth and eighth day treatment (Supplementary Figure S1).
The impact of exogenous Ca2+ on endogenous ion accumulation in salt-stressed perennial ryegrass
Treatment with Ca2+ under salinity enhanced higher Ca2+ content in the leaves than that treated with NaCl alone (Fig. 2a). In the roots, NaCl treatment decreased the Ca2+ content compared with the controls, but exogenous application of Ca2+ relieved the decrease (Fig. 2c). In the leaves, external application of Ca2+ declined Mg2+ content at 4, 8 and 12 DAT, but treatment with NaCl alone stimulated the content decline at 12 DAT (Fig. 2a). In the stem, Mg2+ content increased under both salt treatment and salt+ Ca2+ treatment (Fig. 2b). The increased Mg2+ content in root was just found in treatment line with NaCl alone (Fig. 2c). For K+, NaCl treatment decreased the content in leaf, stem and root, but exogenous application of Ca2+ relieved the decrease in leaf. In the NaCl treated leaf, stem and root of perennial ryegrass, endogenous Na+ content showed remarkable increase but the increase was inhibited under the exogenous application of Ca2+. In addition, the NaCl treatment enhanced Ca2+/Mg2+ ratio in leaf but reduced it in stem and root. Exogenous application of Ca2+ under salinity caused a higher Ca2+/Mg2+ ratio in leaf and root compared with NaCl treatment alone. The K+/Na+ ratio decreased in leaf, stem and root in response to NaCl treatment. Exogenous application of Ca2+ relieved the decrease in leaf at 8 DAT and in NaCl-stressed root.
The impact of exogenous Ca2+ on the antioxidant systems of salt-stressed perennial ryegrass
To investigate the salinity response induced by Ca2+ signal, the SOD, POD, CAT and APX activities, as well as oxidative stress-responsive isoenzymes and genes were analyzed (Fig. 3). During the 12 days treatment, salinity stress induced greater SOD activity in leaves compared with controls, however, Ca2+ addition to NaCl-treated plants induced more pronounced increase at 8 and 12 DAT (Fig. 3a). An increase in POD and CAT activities was observed at 4 DAT, while the activities were depressed at 8 and 12 DAT. Application of Ca2+ upon salinity imposition enhanced POD activity at 4 and 12 DAT, and CAT activity at 4, 8 and 12 DAT respectively in comparison with NaCl only treatment (Fig. 3b). Salinity increased leaf APX activity at 4 DAT but decreased the activity at 12 DAT. Exogenous application of Ca2+ in salt-treated leaves induced higher APX activity compared with NaCl treatment alone at 4, 8 and 12 DAT. The changes of antioxidant isoenzymes for SOD, POD, CAT and APX confirmed these discovery on enzyme activity (Supplementary Figure S2).
Exogenous Ca2+ modulates antioxidant-related enzymatic activities in perennial ryegrass.
Effect of exogenous Ca2+ in the activity of SOD (A), POD (B), CAT (C) and APX (D). The plants were treated with 300 mM NaCl + 7 mM Ca(NO3)2·4H2O (T1) or 300 mM NaCl (T2) at 0, 4, 8 and 12 d. Vertical bars at the top indicate LSD values (P < 0.05) for the comparison of different treatments on a given day of treatment.
The impact of H2O2 and Ca2+ signaling on the oxidative status in salt-stressed perennial ryegrass
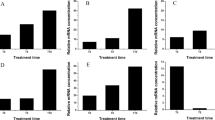
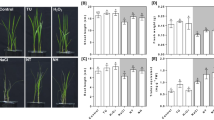
To elucidate further the integrated connection between exogenous H2O2/Ca2+ and oxidative protection against salinity stress, endogenous H2O2 and Ca2+ steady state levels were monitored in perennial ryegrass roots at 8 DAT (Fig. 4). Treatment with NaCl alone decreased Ca2+ accumulation, but stimulated the H2O2 and O2− production. H2O2 and O2− content also increased remarkably, and Ca2+ accumulation declined in H2O2 alone treated plants. In addition, treatment with both NaCl and H2O2 induced higher H2O2 and O2− accumulation and lower Ca2+ content. However, in the presence of NaCl, greater Ca2+ accumulation and lower H2O2 and O2− production were observed because of exogenous Ca2+ application. In contrast, depriving Ca2+ from the 300 mM NaCl treatment solution induced higher H2O2 and O2− accumulation and lower Ca2+ production compared with NaCl treatment alone (Fig. 4).
Meanwhile, to further assess the role of H2O2 and Ca2+ signaling in the ROS metabolism, expression profiles of calmodulin binding protein genes (CaM1 and CaM2) and ROS metabolism genes (ChlCu/ZnSOD, CytCu/ZnSOD, MnSOD, CAT, APX, GPX and GR) were analyzed in perennial ryegrass leaves and roots. When exposed to all treatments, the transcripts levels of APX remained down-regulated in leaves (Fig. 5). Under NaCl treatment, CytCu/ZnSOD, MnSOD, CAT, GPX and GR increased transcript abundant but ChlCu/ZnSOD was down-regulated in the leaves. All ROS genes were up-regulated in the salt-stressed roots Upon Ca2+ addition, ChlCu/ZnSOD, CAT, APX and GPX were down-regulated compared with control. However, application of H2O2 or depriving Ca2+ from NaCl treatment induced all ROS genes expression in leaves, except for CytCu/ZnSOD and APX. In roots, most of ROS genes also showed increased transcripts levels compared with control. Upon Ca2+ addition, CaM1 was induced but CaM2 inhibited in both leaves and roots. However, CaM2 showed higher transcripts than CaM1 in roots after application of H2O2. For other treatment lines, CaM1 and CaM2 were induced in both leaves and roots.
The impact of H2O2 and Ca2+ signaling on the metabolite profiles in salt-stressed perennial ryegrass
To identify candidate metabolites targets in salt-stressed plants, the H2O2 and Ca2+ treated samples were extracted and derivatized, and then analyzed by GC-MS. A total of 160 total peaks with fairly consistent retention times (RT) and excellent resolution were resolved from each polar extract. Based on the internal consistency of RT and retention indices (RI), 41 metabolites could be identified across all samples in this study (Supplementary Table S2). The identified metabolites included 15 organic acids, 12 sugars, 8 amino acids, 4 fatty acids and 2 polyols (Fig. 6; Supplementary Table S3). Each metabolite in each treatment was given in Table S3.
Under NaCl treatment, the total content of 41 metabolites increased in perennial ryegrass leaves. Upon Ca2+ addition, more metabolites accumulated at 8 DAT (Fig. 6d). However, H2O2 stress decreased the 41 metabolites total content after 2 days treatment (Fig. 6c,d). Both NaCl and H2O2 chemical treatments stimulated greater metabolites accumulation compared with NaCl or H2O2 application alone (Fig. 6c,d). Removing Ca2+ from NaCl treatment induced lower accumulation at 2 DAT (Fig. 6c) but higher metabolites content at 4 and 8 DAT (Fig. 6b–d) compared with NaCl treatment in leaves. However, treatments without Ca2+ in NaCl solution decreased the metabolites content in roots (Fig. 6e). In addition, treatments with NaCl or H2O2 increased the content of metobolites, but upon Ca2+ addition, metabolite accumulation declined in roots (Fig. 6e).
The impact of H2O2 and Ca2+ signaling on sugars and sugar alcohol content in salt-stressed perennial ryegrass
The increase of fatty acid, sugars and organic acids content was the major contributor of total metabolites content increase in leaves (Fig. 6). Eleven sugars and one sugar alcohol accounted for more than 66.0% of total metabolites content except H2O2 chemical treatments only 49.5% at 8 DAT. By further evaluating the effects of H2O2 and Ca2+ signaling on sugars and sugar alcohol content in detail, we found that all the twelve metabolites were highly accumulated after Ca2+ application compared with control and NaCl treatment lines (Fig. 7). With NaCl treatment, tagatose, idose, sucrose, talose, allose (5TMS) BP and psicose showed higher content compared with controls. However, NaCl treatment induced lower glucose and inositol content compared with controls. Tagatose, idose, glucose, talose and psicose content had higher content by depriving Ca2+ from NaCl treatment than by NaCl treatment alone in leaves (Fig. 7). H2O2 treatment induced lower accumulation of tagatose, idose, glucose, sucrose, cellobiose, psicose and allose (5TMS) BP, but greater content of talose, glucoheptose and allose (5TMS) MP in comparison with controls. In contrast, both NaCl and H2O2 chemical treatments stimulated higher abundance of tagatose, idose, sucrose, psicose and allose (5TMS) BP, but lower glucoheptose abundance compared with controls (Fig. 7).
H2O2 and Ca2+ signaling associated content of sugars and sugar alcohol in plant leaves aftert 8 days streatment.
Vertical bars indicate standard error of each mean for 11 sugars and one sugar alcohol (Inositol) at a given day of treatment. Experiments were repeated at least three times, each one with three replicates, and P-value was calculated by LSD’ t-test.
In the roots, we found that glucose abundance decreased significantly in all treatment lines (Fig. 8). NaCl stress increased tagatose, psicose, idose, allose (5TMS) BP, sucrose and cellobiose content. Interestingly, adding and removing Ca2+ from NaCl solution induced similar effect. Tagatose, psicose, idose, allose (5TMS) BP, sucrose, cellobiose and inositol abundance had lower accumulation compared with NaCl stress alone, but eventually declined. In addition, plant roots accumulated lower tagatose, psicose, idose, allose (5TMS) BP, glucose, sucrose, cellobiose and inositol under NaCl treatment without Ca2+ compared with controls. However, the treatment after Ca2+ addition increased sucrose content (Fig. 8). H2O2 treatments decreased glucose and sucrose, but increased cellobiose abundance. Both NaCl and H2O2 chemical treatments stimulated higher idose and allose (5TMS) BP abundance, but lower glucose, sucrose and cellobiose accumulation. Idose, allose (5TMS) BP, sucrose showed greater abundance under both NaCl and H2O2 chemical treatments than H2O2 alone. In contrast, both NaCl and H2O2 chemical treatments induced lower cellobiose content than H2O2 alone (Fig. 8).
H2O2 and Ca2+ signaling associated content of sugars and sugar alcohol in plant roots aftert 8 days streatment.
Vertical bars indicate standard error of each mean for 11 sugars and one sugar alcohol (Inositol) at a given day of treatment. Experiments were repeated at least three times, each one with three replicates, and P-value was calculated by LSD’ t-test.
Discussion
In this study, we initially characterized the impact of external Ca2+ or H2O2 on the physiological status of salt-treated cool-season turfgrass. Exogenous administration with Ca2+ alleviated the physiological damage induced by salt tress, as shown by the higher turf quality and lower EL, MDA and H2O2 content. This confirmed previous observations that Ca2+ treatment significantly improved the physiological response of stressed plants5. Similar results in the Ca2+ induced physiological behavior in stressed plants were previously found in tall fescue23, which suggest that Ca2+ acts on key signaling in the stress response pathways of cool-season turfgrass29. Furthermore, we found that the application of external H2O2 could trigger Ca2+ signals positively involved in the rearrangements of ROS accumulation and metabolite profiles.
To elucidate the mechanism of salt tolerance induced by external Ca2+ application and transport, we monitored internal levels of Ca2+, Mg2+, K+ and Na+ in NaCl stressed root, stem and leaf. We found that Ca2+ pool in leaves and roots is modulated in response to NaCl stress. NaCl treatment decreased Ca2+ accumulation in root, but exogenous application of Ca2+ relieved the decline. In the leaves, no significant difference was obtained between controls and NaCl treated line, however, a higher Ca2+ content was found after treatment with Ca2+ under salinity. These results suggest that Ca2+ metabolism is regulated by salinity, which was similar with heat23,30 and cold stresses5. In addition, the Ca2+ pool influx into leaves under salt stress may be a crucial salinity acclimation mechanism for perennial ryegrass. Our previous study had confirmed that the changes of macronutrient cations content, such as Ca2+, Mg2+, Na+, and K+, are necessary for salinity adaptation of perennial ryegrass12. In this study, NaCl treatment decreased K+ but increased Na+ concentration in leaves, stems and roots, which is consistent with our previous studies12,31. In roots, external Ca2+ application significantly increased K+ concentration but decreased Na+ concentration. Even though Ca2+ treatment did not increase K+ concentration in leaves, Na+ concentration was still lower compared with NaCl treatment alone. Moreover, external Ca2+ application significantly induced higher K+/Na+ ratio in roots at 8 days treatment leaves. Evidence has shown that regulation of K+ uptake and prevention of Na+ influx to maintain desirable K+/Na+ ratios in the cytosol are vital strategies of improving plant salt tolerance32. These results suggest that increasing the K+/Na+ ratio could be one regulation mechanism induced by Ca2+ signaling in salt stressed perennial ryegrass. Meanwhile, Mg2+ concentration decreased at 12 DAT in leaves, but increased in stems and roots. Interestingly, feeding salt-treated leaves with Ca2+ induced lower accumulation of Mg2+ and Na+ compared with the NaCl treatment alone. A higher Ca2+/Mg2+ ratio were observed in leaf and root than NaCl-stressed tissues. Low Ca2+/Mg2+ quotient in culture media is often thought as a restriction factor to restrict species from growing in these culture conditions33. We could deduce that the increase of Ca2+/Mg2+ ratio could be one adaption mechanism induced by Ca2+ signaling in salt stressed perennial ryegrass.
Some protective antioxidant enzymes/genes and metabolites are the backbone of the oxidative status in plants34,35. Although salinity conditions produced excessive H2O2 and O2− in plants, the higher antioxidant enzymes activity including SOD, POD, CAT and APX inhibited the excessive ROS accumulation to maintain ROS homeostasis12,36, and our enzymes activity data support this. Ca2+ signals perform functions upstream or downstream in the ROS signal transduction network in plants10. Notably, Ca2+ application under salinity inhibited the decrease of enzyme activity, and greater SOD, POD, CAT and APX activity could be obtained when exogenous application of Ca2+ in NaCl solution. These data are in agreement with that Ca2+ signaling could improve the stress adaptation by stimulating higher antioxidant enzymes activity in plants23, which were further confirmed by greater isoenzymes intensity of SOD (3–5), POD (2–5), CAT (1–2), APX (1–5) (Supplementary Figure S2). The increased antioxidant enzymes activity might be attributed to up/down regulated expression of the candidate genes17,34. Consistently, this study also observed that the expression of ChlCu/ZnSOD, CytCu/ZnSOD, MnSOD, CAT, APX, GPX and GR changed under salt stress. These were involved in the regulation of antioxidant enzyme activities. NaCl direct treatment increased transcript abundance of CytCu/ZnSOD, MnSOD, CAT, GPX and GR in leaves. In contrast, Ca2+ addition treatment induced the up-regulation of only three genes CytCu/ZnSOD, MnSOD and GR in the leaves. However, in roots, ChlCu/ZnSOD, CytCu/ZnSOD, MnSOD, CAT, APX and GR were up-regulated under exogenous Ca2+ application in NaCl treatment, indicating that the positive role in the Ca2+ dependent regulation of antioxidant enzyme gene was greater in root than in leaf under salt stress. These results were confirmed by the opposite changes detected in roots and leaves after removing Ca2+ from NaCl treatment.
Both Na+ and Ca2+ were added into half-strength Hoagland’s solution and first affected the normal functioning of the root in our experiment. H2O2 forms the basis of the ROS signals mediated priming phenomenon in stressed plants37. To understand further the mechanistic connection between H2O2 and Ca2+ signaling in response to salt stress, we investigated the endogenous steady state of Ca2+, H2O2 and O2− levels in roots. Results obtained further proved that exogenous application of Ca2+ in NaCl treatment had higher intensity of Ca2+ signaling in plant tissues than NaCl treatment alone38. Compared with direct NaCl stress, the weaker DAB and NBT staining for H2O2 and O2− after exogenous application of Ca2+ under stress conditions was observed. Moreover, removing all Ca2+ from the NaCl treatment solution induced higher H2O2 and O2− accumulation and lower Ca2+ production compared with NaCl treatment alone, indicating exogenous Ca2+ modulates salinity-induced oxidative damage by directly scavenging H2O2 and O2−. Consistently, H2O2 could function as an upstream or downstream component in the Ca2+ signaling network2. The paper reported that H2O2 induced [Ca2+]cyt increased in leaf cells39. However, in our study, H2O2 treatment alone decreased Ca2+ signaling and increased DAB and NBT staining for H2O2 and O2− in roots. In addition, treatment with both NaCl and H2O2 induced lower Ca2+ content but higher accumulation of H2O2 and O2−, indicating H2O2 in roots may be one negative regulator of Ca2+ signaling pathway in response to salinity stress. Calmodulin (CaM) genes, such as CaM1 and CaM2, could regulate different target intermediate in Ca2+ mediated signal transduction40. Data from the current experiments showed that H2O2 stress induced CaM1 and CaM2 expression in leaves. However, the H2O2 stressed roots exhibited the same CaM1 transcript level as the controls. Consistently, the strong induction of CaM1 and CaM2 in NaCl stressed leaves and roots reinforces the recent suggestion that H2O2 may function as an upstream signal activating Ca2+ signals in NaCl stressed perennial ryegrass2.
H2O2 is one of the major active oxygen species and could be synthesized by the catalyzed reaction of SOD34. Our results demonstrated that salt stress increased the H2O2 level while SOD activity and CytCu/ZnSOD and MnSOD expression level increased in leaves of perennial ryegrass. In salt-stressed roots, the H2O2 level significantly increased with an increasing expression of ChlCu/ZnSOD, CytCu/ZnSOD and MnSOD. Excessive H2O2 could be detoxified into H2O by the catalyzed reaction of CAT, GPX, APX or GR in plants cells41. Our previous study showed that NaCl stress increased the transcript abundance of CAT, GPX, APX or GR12. Achieving ROS homeostasis is necessary self-defense mechanism for plant in response to stress conditions42. Here, qPCR analysis showed that CAT and APX had higher transcript abundance than SOD genes such as ChlCu/ZnSOD, CytCu/ZnSOD and MnSOD in NaCl-stressed root. The same result was obtained under H2O2 treatment and H2O2+ NaCl treatment, suggesting that self-regulation for ROS homeostasis could be activated due to the excessive accumulation of H2O2 induced by NaCl stress. Together with exogenous Ca2+ signal could increase SOD, POD, CAT and APX activity, as well as may function as a downstream signal activated by H2O2, we speculated that Ca2+ and H2O2 signals may exist one common regulation orientation that maintains ROS homeostasis in salt-stressed cool-season turfgrass.
Since the accumulation of various metabolites changed when plants were exposed to salinity conditions43, metabolite profile analysis was performed in leaf and root samples to characterize how Ca2+ and H2O2 signals affect metabolite patterns in salt-stressed cool-season turfgrass. Here, 41 metabolites exhibited different response patterns in plants exposed to NaCl, Ca2+ or H2O2. Consistent with the previous research in barley (Hordeum vulgare L.)43 and Arabidopsis thaliana44, the accumulation of amino acids, organic acids and sugars changed in salt-stressed perennial ryegrass. Metabolites may therefore play vital roles in salinity response pathway. In addition, the present study also characterized a number of modified metabolites that were not observed in previous salinity metabolic response studies, such as polyol and fatty acids. Compared with NaCl treatment alone, the total content of 41 metabolites showed upward trend upon Ca2+ addition during the first 4 days exposure. In contrast, removing Ca2+ or adding H2O2 accelerated the upward trend. At 8 DAT, upon Ca2+ addition treatment obtained the highest accumulation compared with other treatment, but H2O2 alone treatment had the lowest metabolic accumulation during the 8 days treatment. The present data combined with previous findings involved in metabolites response under salt stress, may help to understand better the molecular basis of Ca2+ and ROS signaling in salt-treated plants.
Sugars not only directly provide energy and solutes for osmotic adjustment45,46, but also act as sugar-sensing signal to regulate key candidate gene expression47. Our metabolomic data demonstrated that the sugar and sugar alcohol contents accounted for 49.5–88.2% of all 41 metabolites accumulation in treated leaves, which lead us to hypothesize that sugars profiles is involved in sensing Ca2+ and ROS signaling in salt-treated leaves. First, NaCl treatment induced the production of tagatose, idose, sucrose, talose, allose (5TMS) BP and psicose but inhibited glucose and inositol content in leaves. However, we found that all eleven sugars and one sugar alcohol showed higher accumulation after Ca2+ application compared with control or NaCl treatment lines in leaves. On the contrary, NaCl treatment induced higher sugar than that in Ca2+ application lines in root. These results suggest that up-regulated sugars induced by Ca2+ signaling in leaf could contribute to the stronger response to salt stress. Secondly, the lowest contribution rate (49.5%) for sugars in total metabolites content was just obtained at H2O2 treatment alone at 8 DAT. Correspondingly, in leaves, we found that H2O2 treatment alone induced lower accumulation of tagatose, idose, glucose, sucrose, cellobiose, psicose and allose (5TMS) BP compared with controls or NaCl treatment lines, and just the talose, glucoheptose and allose (5TMS) MP content increased, which indicated that the positive role induced by Ca2+ signaling in leaf may be inhibited by H2O2 signaling. However, converse results were observed in stressed roots, which may be in relation to the different carbohydrate allocation in source and sink tissue under salt stress46.
In conclusion, the present study reported here reinforces the understanding of salt tolerance involved in H2O2 and Ca2+-based signaling response through coupling ion accumulation, antioxidant systems and secondary metabolism analysis in cool-season turfgrass. Increased Ca2+/Mg2+ ratio but decreased Na+/K+ ratio were observed following external Ca2+ application in salt-stressed grass. Exogenous Ca2+ signaling could induce higher antioxidant enzymes activity and the expression level of antioxidant gene and protein in plants to adapt salinity environment. However, H2O2 application decreased Ca2+ signaling but increased the transcript abundance of CAT and APX. In order to maintain ROS homeostasis, Ca2+ and H2O2 signals had one common regulation pattern in salt-stressed cool-season turfgrass. In addition, H2O2/Ca2+-mediated metabolites detected in this study could provide a dataset of common regulatory factors for signaling transduction and salinity acclimation in perennial ryegrass. These findings involved in overlapping roles of H2O2/Ca2+ signaling in salinity response mechanisms supply one novel strategy for a cool-season turfgrass’s adaptation under salt stress.
Methods
Plant materials and growth conditions
Perennial ryegrass ‘Quick start II’ seeds were planted into plastic pots (10 cm diameter, 15 cm deep) with sand as substrate. After seedlings had grown to 8 cm tall, they were mowed to a height of 5 cm and were irrigated twice weekly with half-strength Hoagland’s solution48. Forty-day-old seedlings were rinsed thoroughly using distilled water, and transferred into 300 mL Erlenmeyer flasks filled with approximately 290 mL half-strength Hoagland’s solution. The bottlenecks were closed with appropriate amount of absorbent paper twined using plastic food wrap. To prevent potential algal growth, the flasks were wrapped with aluminum foil. The nutrient solutions were completely replaced each week. All plants were grown under greenhouse conditions (daily temperature of 2471/20711C (day/night), photosynthetic active radiation (PAR) at 300 μmol m−2s−1, and a 14 h photoperiod) throughout the experiment.
Treatments
Experiment 1
Four weeks after canopy and root system establishment, plants were separated into three groups (Group I, II, III) according to the similar transpiration rate calculated based on the weight difference at three-day intervals before salt treatment initiation described by Hu et al.12. Group I plants were supplied with half-strength Hoagland’s solution throughout the whole experimental procedure as control line. Group II plants, as salt treatment line, were treated with 300 mM NaCl for 12 d. Group III plants, as Ca2+ treatment line, were treated with 300 mM NaCl + 7 mM Ca(NO3)2·4H2O for 12 d. All treatment solutions were prepared in half-strength Hoagland’s solution. Shoots samples for physiological, gene expression and metabolic analysis were harvested at 0, 4 d, 8 d and 12 d after treatment (DAT), respectively. Roots samples were harvested at the end of experiment. Treatments were arranged as a completely randomized design and each treatment having four plant-pot systems as four replicates.
Experiment 2
The growing conditions were similar with those of Experiment 1. Plants were separated into six groups with the same method as shown in Experiment 1. The six treatments arranged in a completely randomized block design with four replicates were as follows: (1) nutrient solution (control); (2) 300 mM NaCl + 7 mM Ca(NO3)2·4H2O; (3) 300 mM NaCl; (4) H2O2; (5) H2O2 + 300 mM NaCl; (6) 300 mM NaCl- Ca2+ (prepared in half-strength Hoagland’s nutrient solution without Ca ion). All reagents were dissolved in half-strength Hoagland’s nutrient solution. Shoot samples were harvested at 0, 12 h, 2 d, 4 d, 8 d and 12 d after treatment, respectively. Roots samples were harvested at the end of experiment.
Measurements
Cell membrane stability and turf quality
Cell membrane stability was determined by the electrolyte leakage (EL) level measured based on the method described by Hu et al.17. Fresh 0.1 g of uniform leaf segments were washed with deionized water, incubated in 15 mL deionized water, and then shaken for 24 h at room temperature. The initial conductance (Ci) was measured using a conductance meter (JENCO-3173, Jenco Instruments, Inc., San Diego, CA, USA). The samples were then autoclaved at 120 °C for 20 min to completely disrupt the tissues and to release all electrolytes. After cooling to room temperature, the conductance of the incubation solution with killed tissues (Cmax) was determined. Relative EL was calculated using the formula: EL (%) = (Ci/Cmax) × 100.
Turf quality was rated visually based on turfgrass color (percentage green leaves), plant density and degree of leaf wilting on a scale of 0 to 9, where 0 score indicated withered, yellow, thin or dead grass, while 6 indicated minimum acceptable level. A 9 score indicated green, dense and uniform grass49.
Antioxidant enzymes activities, cell lipid peroxidation and H2O2 level
Extract preparation and assays for SOD (EC 1.15.1.1), CAT (EC 1.11.1.6), POD (EC. 1.11.1.7), APX (EC 1.11.1.11), soluble protein, MDA and H2O2 level were conducted according to methods reported previously17. The SOD activity was measured according to the method of Jiang and Huang23. A 3 mL reaction mixture, composed of 50 mM phosphate buffer solution (PBS) (pH 7.8), 60 mM Riboflavin, 195 mM Met, 3mmM EDTA, 1.125 mM NBT and 0.1 mL enzyme extract, was placed under light at 3000 lux for 10 min, and was recorded at 560 nm absorbance by spectrophotometer (UV-2600, UNICO Instruments Co., Ltd., Shanghai, China). One unit of SOD activity was defined as the amount of enzyme that inhibited 50% photochemical reduction of NBT. POD activity was measured as an increase in absorbance at 470 nm for 1 min following the oxidation of guaiacol50. The reaction mixture contained 0.1 M HAC–NaAC buffer (pH 5.0), 20 mM guaiacol, 10 mM PBS (pH 7.0), 40 mM H2O2 and 0.1 mL enzyme extract. CAT activity was determined according to Chance and Maehly51. A reaction mixture of 50 mM PBS (pH 7.4), 45 mM H2O2 and 0.1 mL enzyme extract was measured at 240 nm absorbance for 1 min. CAT activity was determined through the decomposition of H2O2. APX activity was detected using the method described by Mittler and Zilinskas52.
Cell lipid peroxidation was determined by the MDA content measured as follows. One milliliter of enzyme extract was mixed with 2 ml of reaction solution containing 20% (v/v) trichloroacetic acid and 0.5% (v/v) TBA. The mixture was heated in a water bath at 95 °C for 30 min, then cooled quickly in ice-water bath to room temperature, and centrifuged at 14,000 rpm for 20 min. Absorbance of the supernatant was measured at 532 and 600 nm. MDA content was calculated based on subtracting the absorption at 600 nm from the absorption at 532 nm and calibrated with the extinction coefficient of 155 mM−1cm−1.
H2O2 level was determined based on the method described by Lin and Kao53. One milliliter of supernatant was mixed thoroughly with 1 ml of 0.1% titanium sulphate in 20% H2SO4 (v/v), and then centrifuged at 6000 × g for 15 min at room temperature. The supernatant was measured at 410 nm absorbance. The H2O2 level was calculated according the standard curve generated with known concentrations of H2O2 and calibrated by extinction coefficient of 0.28 μmol−1cm−1.
RNA isolation, cDNA synthesis and real-time PCR analysis
Total RNA extraction was performed from the leaves and roots using Trizol reagent (Invitrogen, Paisley, UK) according to the manufacturer’s instructions and then was incubated with RNase-free DNase (RQ1; Promega, Madison, WI, USA). RNA integrity was examined at 260 and 280 nm by spectrophotometer (UV-2600; UNICO Instruments, Shanghai, China) and checked on a gel electrophoresis in 1.5% agarose gels with 1 μL RNA (=0.5 μg μL−1). Subsequently, cDNA synthesis was performed from 2 μg purified RNA using cDNA synthesis kit according to the manufacturer’s protocol (Fermentas, Burlington, ON, Canada). The resultant cDNA was diluted six-fold and kept at −20 °C for RT-PCR analysis.
The transcript levels of the target genes were analyzed using ABI StepOne Plus Real-Time PCR system (Applied Biosystems, Foster City, CA) and SYBR Green Real-Time PCR Master Mix (Toyobo, Japan) in 20 mL reactions. Each reaction mix contained 2 ng of total RNA, 0.5 μL of each primer (Supplementary Table S1) and 10 μL master mix. Reactions were carried out as follows: initial denaturation at 95 °C for 3 min, 38 cycles of 10 s at 94 °C, 20 s at 50–55 °C, and 20 s at 72 °C, followed by 5 min at 72 °C. For specific product verification, a melting curve was performed from 82 °C at 0.2 °C increments with a 10 s hold between observations. YT521-B gene was used as a standard control in the RT-PCR reactions. The relative expression of specific genes was quantitated with comparative Ct method as described earlier54. The experiments were repeated twice with three replicates.
Histochemical staining for cytosolic calciumdetection, H2O2 and O2−
Labeling for cytosolic Ca2+ was performed by incubating cross-sections of the main root tissue with Fluo 3-AM (Molecular Probes, CAS#:121714-22-5) according to earlier reports55. Root tip was placed in 4 μM Fluo 3-AM dissolved in Hanks balanced salt solution (HBSS) containing 10 mM HEPES, 1mM Na2HPO4, 137 mM NaCl, 5 mM KCl, 1 mM CaCl2, 0.5 mM MgCl2, 5 mM glucose, 0.1% BSA, pH 7.4, and incubated at 37 °C for 60 min. And then the preincubated roots were washed with HBSS and fluorescence examination was performed using an IX70 inverted fluorescence microscope (Olympus, Japan), with the following settings: Ex 488 nm, Em 526 nm.
For histochemical detection of H2O2 and O2−, perennial ryegrass roots were stained with 3,3′-diaminobenzidine (DAB) and nitroblue tetrazolium (NBT), respectively, based on the method described by Zhang et al.56. The histochemical detection of H2O2 in perennial ryegrass roots was performed by directly immersion and infiltration of root quarters under vacuum with a freshly prepared DAB solution (1 mg mL−1, pH 3.8) at 25 °C in the dark for 30 min. For NBT staining to detect O2− in roots, perennial ryegrass roots were immersed and infiltrated with 3.5 mg mL−1 NBT staining solution in 25 mM KOH-HEPES buffer (pH 7.8) and incubated in the dark for 30 min. After staining, roots were bleached in acetic acid-glycerol-ethanol (1/1/3, v/v/v) solution at 100 °C for 5 min, and then stored in glycerol-ethanol (1/4, v/v) until photographs were taken. H2O2 as a brown color and O2− as a blue color were visualized.
Metabolite extraction and derivatization
Metabolites from controls, Ca2+, H2O2, NaCl treated leaves and roots were extracted based on the procedure reported by Roessner et al.57 with some modifications. Frozen leaves and roots were ground to a fine powder in liquid nitrogen, transferred into 2 mL Eppendorf tubes, and then extracted in 1.4 mL of 80% (v/v) aqueous methanol under intensive oscillation (200 rmp) at 25 °C for 2 h. Then, a 50 μL methyl nonadecanoate (2 mg mL−1 in chloroform) with 20 μL internal standard (2 mg mL−1 ribitol in water) was added. Extraction was carried out at 70 °C in a metal bath for 15 min (each 5 min vortex 5 s). The tube was centrifuged for 5 min at 12 000 g. The supernatant was decanted to new 10-mL tubes, and 1.5 mL of water and 0.75 mL of chloroform were added. The mixture was vortexed thoroughly and subsequently centrifuged for 15 min at 8000 g. 300 μL polar phase (methanol/water) was decanted into 1.5 mL HPLC vials and dried in a benchtop centrifugal concentrator (Labconco Corporation, Kansas City, MI) overnight. The dried residue was redissolved and derivatized with 80 μL of 20 mg mL−1 methoxyamine hydrochloride in pyridine for 2 h under intensive oscillation (200 rmp) at 37 °C, and followed by a 2 h treatment at 37 °C with 50 μL N-methyl-N- trimethylsilyltrifluoroacetamide (MSTFA).
Gas chromatography mass spectrometry (GC-MS) analysis
For GC-MS analysis, a 1 μL of the derivatization solutions was determined with a GC–MS (DSQII, Agilent 7890A/5975C, Hemel Hempstead, USA) system based on the method described by Hancock et al.58. Samples were injected into the DB5-MSTM column (15 m × 0.25 mm × 0.25 μm; J&W, Folsom, CA, USA) with a split ratio of 1:25. Injection temperature was set to 280 °C, the interface temperature was set at 290 °C, and the ion source temperature was adjusted to 200 °C. The column temperature was 5 min isothermal at 70 °C, then the GC oven temperature was raised to 260 °C with 5 °C min−1 in 2 min after injection, and finally held at 260 °C for 10 min. Helium gas was used as carrier set with a constant flow set at 1 mL min −1. The MS measurement were set at electron impact (EI) source, electron energy 70 eV, solvent delay 4 min and the scan range 30–650 m/z at 0.6 scan s−1. Retention time (RT), retention indices (RI, http://gmd.mpimp-golm.mpg.de/search.aspx) and NIST Mass Spectral Database (version 11) were implemented to identify the target metabolites. Only metabolite detected at least three in five samples was considered true. Each metabolite was finally identified based on the internal consistency of RT and RI. For each sample in controls and Ca2+, H2O2, NaCl treated leaves and roots, we performed four biological replicate and two technical replicates per biological experiment.
Statistical analysis
Statistical significance was performed by ANOVA using the Statistics Analysis System (SAS) (version 9.0 for Windows; SAS Institute, Cary, NC). Means for physiological measurements, gene expression and metabolites between treatments were separated using the Fisher’s least significant difference test (LSD) at P < 0.05 level.
Additional Information
How to cite this article: Hu, T. et al. H2O2 and Ca2+-based signaling and associated ion accumulation, antioxidant systems and secondary metabolism orchestrate the response to NaCl stress in perennial ryegrass. Sci. Rep. 6, 36396; doi: 10.1038/srep36396 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Zhu, X., Feng, Y., Liang, G., Liu, N. & Zhu, J. K. Aequorin-based luminescence imaging reveals stimulus-and tissue-specific Ca2+ dynamics in Arabidopsis plants. Mol. Plant 6, 444–455 (2013).
Sun, J. et al. H2O2 and cytosolic Ca2+ signals triggered by the PM H+‐coupled transport system mediate K+/Na+ homeostasis in NaCl‐stressed Populus euphratica cells. Plant Cell Environ. 33, 943–958 (2010).
Ma, Y. et al. COLD1 confers chilling tolerance in rice. Cell 160, 1209–1221 (2015).
Hung, S. H., Yu, C. W. & Lin, C. H. Hydrogen peroxide functions as a stress signal in plants. Bot. Bull. Acad. Sinica 46, 1–10 (2005).
Shi, H., Ye, T., Zhong, B., Liu, X. & Chan, Z. Comparative proteomic and metabolomic analyses reveal mechanisms of improved cold stress tolerance in bermudagrass (Cynodon dactylon (L.) Pers.) by exogenous calcium. J. Integr. Plant Biol. 56, 1064–1079, doi: 10.1111/jipb.12167 (2014).
Tuna, A. L. et al. The effects of calcium sulphate on growth, membrane stability and nutrient uptake of tomato plants grown under salt stress. Environ. Exp. Bot. 59, 173–178 (2007).
Rhee, S. G. H2O2, a Necessary Evil for Cell Signaling. Science 312, 1882–1883, doi: 10.1126/science.1130481 (2006).
Verslues, P. E. et al. Interaction of SOS2 with Nucleoside Diphosphate Kinase 2 and Catalases Reveals a Point of Connection between salt stress and H2O2 signaling in Arabidopsis thaliana. Mol. Cell Biol. 27, 7771–7780, doi: 10.1128/mcb.00429-07 (2007).
Hu, X. et al. Cross-talks between Ca2+/CaM and H2O2 in abscisic acid-induced antioxidant defense in leaves of maize plants exposed to water stress. Plant Growth Regul. 55, 183–198 (2008).
Jiang, M. & Zhang, J. Cross‐talk between calcium and reactive oxygen species originated from NADPH oxidase in abscisic acid‐induced antioxidant defence in leaves of maize seedlings. Plant Cell Environ. 26, 929–939 (2003).
Sun, J. et al. Hydrogen peroxide and nitric oxide mediate K+/Na+ homeostasis and antioxidant defense in NaCl-stressed callus cells of two contrasting poplars. Plant Cell Tiss. Org. 103, 205–215 (2010).
Hu, T., Li, H., Zhang, X., Luo, H. & Fu, J. Toxic effect of NaCl on ion metabolism, antioxidative enzymes and gene expression of perennial ryegrass. Ecotox. Environ. Safe. 74, 2050–2056 (2011).
Scandalios, J. G. Oxygen stress and superoxide dismutases. Plant Physiol. 101(1), 7–12 (1993).
Hernández, J. A., Ferrer, M. A., Jiménez, A., Barceló, A. R. & Sevilla, F. Antioxidant Systems and O2−/H2O2 Production in the Apoplast of Pea Leaves. Its Relation with Salt-Induced Necrotic Lesions in Minor Veins. Plant Physiol. 127, 817–831, doi: 10.1104/pp.010188 (2001).
Hernandez, M., Fernandez-Garcia, N., Diaz-Vivancos, P. & Olmos, E. A different role for hydrogen peroxide and the antioxidative system under short and long salt stress in Brassica oleracea roots. J. Exp. Bot. 61, 521–535, doi: 10.1093/jxb/erp321 (2010).
Scandalios, J. G. Oxidative stress: molecular perception and transduction of signals triggering antioxidant gene defenses. Braz. J. Med. Biol. Res. 38, 995–1014 (2005).
Hu, L., Li, H., Pang, H. & Fu, J. Responses of antioxidant gene, protein and enzymes to salinity stress in two genotypes of perennial ryegrass (Lolium perenne) differing in salt tolerance. J. Plant Physiol. 169, 146–156 (2012).
Abogadallah, G. M. Insights into the significance of antioxidative defense under salt stress. Plant Sign. Behav. 5, 369–374 (2010).
JG, S. Oxidative stress: molecular perception and transduction of signals triggering antioxidant gene defenses. Braz. J. Med. Biol. Res. 38, 995–1014 (2005).
Luna, C. M. et al. Drought controls on H2O2 accumulation, catalase (CAT) activity and CAT gene expression in wheat. J Exp. Bot. 56, 417–423, doi: 10.1093/jxb/eri039 (2005).
Pei, Z. M. et al. Calcium channels activated by hydrogen peroxide mediate abscisic acid signalling in guard cells. Nature 406, 731–734 (2000).
Yang, T. & Poovaiah, B. W. Hydrogen peroxide homeostasis: activation of plant catalase by calcium/calmodulin. PNAS 99, 4097–4102, doi: 10.1073/pnas.052564899 (2002).
Jiang, Y. & Huang, B. Effects of calcium on antioxidant activities and water relations associated with heat tolerance in two cool‐season grasses. J. Exp. Bot. 52, 341–349 (2001).
Dittami, S. M. et al. Integrative analysis of metabolite and transcript abundance during the short-term response to saline and oxidative stress in the brown alga Ectocarpus siliculosus. Plant Cell Environ. 34, 629–642, doi: 10.1111/j.1365-3040.2010.02268.x (2011).
Zhu, J., Dong, C. H. & Zhu, J. K. Interplay between cold-responsive gene regulation, metabolism and RNA processing during plant cold acclimation. Curr. Opin. Plant Biol. 10, 290–295 (2007).
Furuichi, T., Mori, I. C., Takahashi, K. & Muto, S. Sugar-induced increase in cytosolic Ca2+ in Arabidopsis thaliana whole plants. Plant Cell Physiol. 42, 1149–1155 (2001).
Shabala, S. & Cuin, T. A. Potassium transport and plant salt tolerance. Physiol. Plantarum 133, 651–669 (2008).
Xiong, Y. et al. Identification of quantitative trait loci controlling winter hardiness in an annual × perennial ryegrass interspecific hybrid population. Mol. Breeding 19, 125–136 (2007).
Sheen, J. Ca2+ -dependent protein kinases and stress signal transduction in plants. Science 274, 1900–1902 (1996).
Wu, H. C. & Jinn, T. L. Heat shock-triggered Ca2+ mobilization accompanied by pectin methylesterase activity and cytosolic Ca2+ oscillation are crucial for plant thermotolerance. Plant Sign. Behav. 5, 1252–1256 (2010).
Hu, L., Hu, T., Zhang, X., Pang, H. & Fu, J. Exogenous glycine betaine ameliorates the adverse effect of salt stress on perennial ryegrass. J. Am. Soc. Hortic. Sci. 137, 38–46 (2012).
Rus, A. et al. AtHKT1 is a salt tolerance determinant that controls Na+ entry into plant roots. PNAS 98, 14150–14155 (2001).
Rajakaruna, N., Siddiqi, M. Y., Whitton, J., Bohm, B. A. & Glass, A. D. Differential responses to Na+/K+ and Ca2+/Mg2+ in two edaphic races of the Lasthenia californica (Asteraceae) complex: a case for parallel evolution of physiological traits. New Phytol. 157, 93–103 (2003).
Mates, J. Effects of antioxidant enzymes in the molecular control of reactive oxygen species toxicology. Toxicology 153, 83–104 (2000).
Tanou, G. et al. Oxidative and nitrosative-based signaling and associated post-translational modifications orchestrate the acclimation of citrus plants to salinity stress. Plant J. 72, 585–599, doi: 10.1111/j.1365-313X.2012.05100.x (2012).
Zhu, J. K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 53, 247–273 (2002).
Tanou, G. et al. Proteomics reveals the overlapping roles of hydrogen peroxide and nitric oxide in the acclimation of citrus plants to salinity. Plant J. 60, 795–804 (2009).
Saijo, Y. et al. A Ca2+-dependent protein kinase that endows rice plants with cold-and salt-stress tolerance functions in vascular bundles. Plant Cell Physiol. 42, 1228–1233 (2001).
Hu, X. et al. Calcium-calmodulin is required for abscisic acid‐induced antioxidant defense and functions both upstream and downstream of H2O2 production in leaves of maize (Zea mays) plants. New Phytol. 173, 27–38 (2007).
Snedden, W. A. & Fromm, H. Calmodulin as a versatile calcium signal transducer in plants. New Phytol. 151, 35–66 (2001).
Choudhury, S., Panda, P., Sahoo, L. & Panda, S. K. Reactive oxygen species signaling in plants under abiotic stress. Plant Sign. Behav. 8, e23681 (2013).
Takahashi, F. et al. Calmodulin-dependent activation of MAP kinase for ROS homeostasis in Arabidopsis. Mol. Cell 41, 649–660 (2011).
Widodo Patterson, J. H., Newbigin, E., Tester, M. & Bacic, A. Roessner U. Metabolic responses to salt stress of barley (Hordeum vulgare L.) cultivars, Sahara and Clipper, which differ in salinity tolerance. J. Exp. Bot. 1–15, doi: 10.1093/jxb/erp243 (2009).
Kim, J. K., Bamba, T., Harada, K., Fukusaki, E. & Kobayashi, A. Time-course metabolic profiling in Arabidopsis thaliana cell cultures after salt stress treatment. J. Exp. Bot. 58, 415–424 (2007).
Hoekstra, F. A., Golovina, E. A. & Buitink, J. Mechanisms of plant desiccation tolerance. Trends Plant Sci. 6, 431–438 (2001).
Hu, T. et al. Differential responses of CO2 assimilation, carbohydrate allocation and gene expression to NaCl stress in perennial ryegrass with different salt tolerance. Plos One 8, e66090 (2013).
Ruan, Y. L. Sucrose metabolism: gateway to diverse carbon use and sugar signaling. Annu. Rev. Plant Biol. 65, 33–67 (2014).
Hoagland, D. R. & Arnon, D. I. The water-culture method for growing plants without soil. (2nd edit) 347 (Circular California Agricultural Experiment Station, 1950).
Turgeon, A. J. Turfgrass management. (ed Eighth) Prentice Hall, Englewood Cliffs, NJ. (1991).
Rao, M. V., Paliyath, G. & Ormrod, D. P. Ultraviolet-B-and ozone-induced biochemical changes in antioxidant enzymes of Arabidopsis thaliana. Plant Physiol. 110, 125–136 (1996).
Chance, B. & Maehly, A. Assay of catalases and peroxidases. Method Enzymol. 2, 764–775 (1955).
Mittler, R. & Zilinskas, B. A. Detection of ascorbate peroxidase activity in native gels by inhibition of the ascorbate-dependent reduction of nitroblue tetrazolium. Anal. Biochem. 212, 540–546 (1993).
Lin, J. N. & Kao, C. H. Effect of oxidative stress caused by hydrogen peroxide on senescence of rice leaves. Bot. Bull. Acad. Sinica 39, 161–165 (1998).
Li, X. B., Fan, X. P., Wang, X. L., Cai, L. & Yang, W. C. The cotton ACTIN1 gene is functionally expressed in fibers and participates in fiber elongation. Plant Cell. 17, 859–875 (2005).
Zhang, W., Fan, L. M. & Wu, W. H. Osmo-sensitive and stretch-activated calcium-permeable channels in Vicia faba guard cells are regulated by actin dynamics. Plant Physiol. 143, 1140–1151, doi: 10.1104/pp.106.091405 (2007).
Zhang, M. et al. ZmMKK3, a novel maize group B mitogen-activated protein kinase kinase gene, mediates osmotic stress and ABA signal responses. J. Plant Physiol. 169, 1501–1510 (2012).
Roessner, U., Wagner, C., Kopka, J., Trethewey, R. N. & Willmitzer, L. Simultaneous analysis of metabolites in potato tuber by gas chromatography–mass spectrometry. Plant J. 23, 131–142 (2000).
Hancock, R. D. et al. Physiological, biochemical and molecular responses of the potato (Solanum tuberosum L.) plant to moderately elevated temperature. Plant Cell Environ. 37, 439–450 (2014).
Acknowledgements
This research was supported by the China-Africa Center for Research and Education, CAS (No. SAJC201325) and National Natural Science Foundation of China (No. 31272194 and 31201653).
Author information
Authors and Affiliations
Contributions
J.M.F. and T.H. conceived and designed the experiments. T.H. performed the experiments and wrote the manuscript. K.C. and L.X.H. helped to analyze the data. E.A. revise the manuscript. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hu, T., Chen, K., Hu, L. et al. H2O2 and Ca2+-based signaling and associated ion accumulation, antioxidant systems and secondary metabolism orchestrate the response to NaCl stress in perennial ryegrass. Sci Rep 6, 36396 (2016). https://doi.org/10.1038/srep36396
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36396
This article is cited by
-
Lipid metabolism and antioxidant system contribute to salinity tolerance in halophytic grass seashore paspalum in a tissue-specific manner
BMC Plant Biology (2023)
-
Homocysteine-Thiolactone Modulates Gating of Mitochondrial Voltage-Dependent Anion Channel (VDAC) and Protects It from Induced Oxidative Stress
The Journal of Membrane Biology (2022)
-
Hydrogen peroxide-induced stress acclimation in plants
Cellular and Molecular Life Sciences (2022)
-
Antioxidant regulation of iron as a repressor for salt-induced leaf senescence in perennial grass species
Plant Growth Regulation (2021)
-
Screening of diverse tall fescue population for salinity tolerance based on SSR marker-physiological trait association
Euphytica (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.