Abstract
Candida albicans is a common cause of life-threatening fungal bloodstream infections. In the murine model of systemic candidiasis, the kidney is the primary target organ while the fungal load declines over time in liver and spleen. To better understand these organ-specific differences in host-pathogen interaction, we performed gene expression profiling of murine kidney, liver and spleen and determined the fungal transcriptome in liver and kidney. We observed a delayed transcriptional immune response accompanied by late induction of fungal stress response genes in the kidneys. In contrast, early upregulation of the proinflammatory response in the liver was associated with a fungal transcriptome resembling response to phagocytosis, suggesting that phagocytes contribute significantly to fungal control in the liver. Notably, C. albicans hypha-associated genes were upregulated in the absence of visible filamentation in the liver, indicating an uncoupling of gene expression and morphology and a morphology-independent effect by hypha-associated genes in this organ. Consistently, integration of host and pathogen transcriptional data in an inter-species gene regulatory network indicated connections of C. albicans cell wall remodelling and metabolism to the organ-specific immune responses.
Similar content being viewed by others
Introduction
Candida albicans is the most common cause of life-threatening fungal bloodstream and disseminated infections. The crude mortality of disseminated infections caused by Candida is >50%, higher than for bacterial blood stream infection1. The murine model of hematogenously disseminated candidiasis is most commonly used to study systemic candidiasis and to evaluate efficacy of antifungal therapy. Following intravenous infection, C. albicans initially infects almost all organs; however, while the fungal load increases over time in kidneys, it declines in liver and spleen2,3. Moreover, C. albicans is able to form filaments – a hallmark of pathogenicity - in the kidney within two hours after intravenous infection, whereas no hypha formation is observed in liver and spleen4. Thus, the kidney appears to be the prime target organ for disseminated candidiasis in mice. In contrast, disseminated candidiasis in humans affects kidneys but also commonly leads to infection of liver and spleen5.
The relative susceptibility of the kidneys in murine candidiasis has been linked to the specific immunological setup of this organ: Neutrophils and macrophages are present in higher numbers in liver and spleen than in the kidneys of naive animals2. In addition, in comparison to spleen and liver, leukocytes are recruited later to C. albicans-infected kidneys2. Early accumulation of neutrophils has a protective effect and mononuclear phagocytes can directly kill C. albicans in vitro and in vivo2,6,7. As kidney-resident macrophages and inflammatory monocytes are important for fungal clearance in the kidney, the delay in the early phagocytic response in the kidney seems to be critical for progression of infection6,7. However, at later stages of infection continued neutrophil recruitment induced by locally produced cytokines and chemokines in the kidney directly contributes to immunopathology3. Thus, the immune response, fungal clearance and pathological consequences vary significantly in different organs during systemic C. albicans infection in the murine model.
C. albicans expresses a variety of virulence factors that contribute to pathogenesis, including the morphological transition between yeast and hypha, the expression of hypha-associated virulence factors and metabolic flexibility6,8,9. In addition to morphogenesis, the C. albicans cell wall structure is strongly influenced by environmental factors, such as available carbon sources or environmental stresses, leading to rapid cell wall remodelling processes in vitro and in vivo that affect interaction with immune cells10,11,12. Furthermore, C. albicans rapidly adapts to the changing environments encountered during infection by changes in gene expression, translation, and post-translational modifications13,14.
Thus, both the host and the fungus respond rapidly to the changing conditions during systemic candidiasis. Such changes can be monitored on the transcriptional level by in vivo gene expression profiling to estimate the functional adjustments during invasive growth of C. albicans. Recently, Xu et al. analysed 479 C. albicans and 46 mouse genes from kidney samples using the NanoString technology15. Although less than 2% of the Candida genome was covered, a clear induction of C. albicans hypha-associated as well as zinc and iron responsive genes was observed. Similarly, RNASeq of C. albicans cells infecting the mouse kidney revealed regulation of iron metabolism, hypha formation, and metabolic adaptation during infection16.
Even though these studies significantly contributed to our understanding of disseminated candidiasis, they focused on the kidney as the main target organ. However, given the different immune responses in different organs, it is likely that organ-specific host-pathogen interactions occur and contribute to the different course of infection in murine organs. Understanding these differences may be crucial in understanding pathogenicity of life-threatening systemic C. albicans infections. Therefore, we performed a time-course transcriptional analysis of liver, spleen, and kidney samples from mice infected intravenously with C. albicans. Our results demonstrate not only a delayed immune response in the kidney compared to liver and spleen, but also qualitative differences in the responses of these organs. Microarray analyses of C. albicans cells likewise showed organ-specific adaptation processes. This includes an increased expression of hypha-associated genes in the liver in the absence of visible filamentation, suggesting that gene expression and morphogenesis is uncoupled under these specific conditions in vivo.
Results and Discussion
Murine spleen, liver, and kidney show different kinetics of transcriptional responses to systemic candidiasis
Transcriptional profiling is a powerful tool to elucidate host-pathogen interactions during infection17,18 that has been successfully applied to the murine model of disseminated candidiasis15,16,19,20,21. However, previous studies focused on the kidney as the main target organ. Given the differing progression of infection in kidney, liver, and spleen in mice and the frequent involvement of liver and spleen in human disseminated candidiasis, we performed a transcriptional profiling of these three organs over the time course of infection in mice.
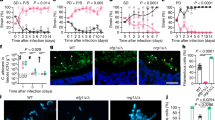
Principal component analysis (PCA) of the murine transcriptional data sets showed that 54% of the variance (PC1 and PC2) among all transcripts in the data sets could be attributed to organ-specific expression differences (Fig. 1A). Temporal changes also contributed to variance (7%) and biological replicates from individual organs clustered according to time point (Fig. 1A). PCA reflects general expression trends in the data which are due to alterations in the expression of the majority of genes represented on the microarray. When specifically analyzing differentially expressed genes (DEGs), organ-specific kinetics of transcriptional changes became evident: Most of the changes in gene expression in liver and spleen were observed already 24 h p.i., whereas in kidney the number of DEGs continuously increased over time (Fig. 1B). We thus performed cluster analysis to categorize the genes according to organ expression kinetics and analysed these clusters for enriched GO terms. All organs responded to infection by regulation of genes associated with immune processes and stress response (Suppl. Fig. 1A, Suppl. data 1 and 2) and, more specifically, upregulation of genes associated with the response to interferon β and cytokine biosynthesis (represented by cluster 5, Suppl. Fig. 1B). However, the expression kinetics differed between organs: As shown in Fig. 2A, several genes in TLR and NLR pathways were upregulated from 8 h to 24 h p.i. in the liver, while in the kidney sizable induction of these pathways was detected only after 24 h and further increased towards 72 h p.i. At 72 h p.i., complement activation was significantly induced in the liver. In the spleen, especially genes in T- and B-cell receptor signalling pathways were enriched; however, the involved genes were mainly downregulated compared to the control. The subset of genes displayed in Fig. 2B highlights the differences in the expression kinetics of genes associated with inflammation: Expression of most of these genes in the liver increased strongly early after infection with a tendency to return to control sample expression levels at later time points, whereas in kidney samples the largest changes were detected 72 h p.i.
Principle component (PC) analysis of mouse expression data (A) and numbers of differentially expressed genes (DEGs) (B).
(A) Left: Approximately 54% of the variation between data sets was captured by the first two PCs and coincided with the organ the sample was obtained from. Right: The third PC captured about 7% of the variance in the data and correlated with the temporal effects in the data set. (B) Numbers of genes differentially regulated (p < 0.05) in kidneys, liver and spleen during C. albicans infection as compared to the PBS control.
Gene expression changes in immune response pathways after systemic C. albicans infection.
(A) Number of differentially regulated genes in distinct immune pathways. The org.Mm.eg.db package for R was used to identify differentially expressed genes associated with the pathways “TLR signaling pathway” (GO:0002224), “NLR signaling pathway” (GO:0035872), “chemokine mediated signaling pathway” (GO:0070098), “complement activation” (GO:0006956), “T-cell receptor signaling pathway” (GO:0050852) and “B-cell receptor signaling pathway” (GO:0050853) including associated child-terms, respectively. (B) Gene expression of selected differentially expressed immune genes. For each organ, log2 fold-changes were gene-wise maximum scaled between −1 and 1 (scaled log2 FC). Measurements of probes mapping to the same gene were mean averaged.
Thus, while all organs responded to infection by downregulation of genes associated with organ-specific functions and upregulation of immune processes, the kinetics and type of immune responses differed.
Organ-specific transcriptional changes in systemic candidiasis reflect activation of local immune responses and organ damage
As discussed above, the transcriptional changes in the kidney were characterised by late upregulation of proinflammatory pathways, consistent with the reported delayed recruitment of immune cells in this organ2. Notable exceptions were the chemokine CXCL1 (also known as keratinocyte cell–derived chemokine [KC]) and the intercellular adhesion molecule ICAM-1, which is important for leukocyte transmigration. These genes were highly upregulated after 8 h in the kidney (Fig. 2B). As we analysed the transcriptome of whole organs, the precise cellular source of early immune transcripts is not clear. It has, however, been shown that CXCL1 is mainly produced by resident tissue macrophages via TLR4 signalling through MyD8822. In comparison to liver and spleen, the number of resident immune cells in the kidney is low2 and even though transcript abundance of CXCL1 increased in the kidney 8 h p.i., protein levels are known to increase more steeply only after 24 h3. This suggests that the potential of resident renal macrophages to produce this chemokine is not sufficient to drive early neutrophil recruitment at high levels. The progressive upregulation of proinflammatory pathways in the kidney over the course of infection can be explained partially by the increasing or persistent fungal burden which leads to ongoing immune stimulation. Fungal growth is furthermore associated with progressive renal damage3,23, reflected in our data set by (i) the upregulation of genes involved in wound healing and (ii) the downregulation of genes in “renal system processes”, “transport processes”, and “ion homeostasis” observed at later time points (Suppl. Fig. 1).
Organ-specific immune responses included the induction of the acute phase response in the liver (Suppl. Fig. 1A) and the TGF-β pathway and genes associated with lymphocyte activation and leukocyte proliferation in the kidney (Suppl. Fig. 1A). While the latter were upregulated at late time points in the kidney, their expression was continuously decreasing until 24 h in the spleen, with a tendency to return to basal levels towards 72 h p.i. As a secondary lymphoid organ, main functions of the spleen are associated with leukocyte activation and proliferation. Therefore, transiently reduced expression of these genes could be interpreted as reduced expression of genes with organ-specific functions. The transient nature of these alterations might thus reflect the successful control of fungal growth in the spleen. Downregulation was also observed in the liver for genes involved in liver function (“monocarboxylic acid metabolism” and “lipid homeostasis”, cluster 9, Suppl. Fig. 1). While these genes also showed a trend to return to basal levels at 72 h p.i., it was not as pronounced as for the spleen, indicating that the transcriptional response of this organ was continuing even though infection is controlled to a similar degree in liver and spleen. Furthermore, although the kinetics of fungal burden and immune cell recruitment in liver and spleen are similar during systemic candidiasis, the transcriptional immune response of these organs differed significantly: Components of the innate immune response, such as TLR and NLR signalling pathways, were induced to a higher extend in the liver. The lack of strong induction of genes associated with proinflammatory responses is consistent with the comparatively low and transient production of proinflammatory cytokines in the spleen during systemic candidiasis3. We find it noteworthy, however, that systemic responses were clearly induced in the liver, e.g. the acute phase response, and that genes involved in complement activation were upregulated especially after 72 h p.i. This might indicate the role of the liver for the systemic immune response during ongoing candidiasis independent of local pathogen control, and might explain why expression of genes involved in metabolic liver function (cluster 9, Suppl. Fig. 1) did not return to steady-state levels by 72 h p.
In summary, all organs showed downregulation of genes associated with organ function, likely as a result of organ impairment and/or as consequence of upregulation of genes dedicated to immune responses. Consistent with the development of fungal burden, these changes were transient in spleen and liver but increased over time in the kidney.
Transcriptional profiles of C. albicans indicate organ specific fungal adaptation in kidney and liver
One of the largest challenges for in vivo fungal transcription analysis is the relative abundance of host transcripts hampering the recovery of sufficient amounts of fungal RNA17. We used an enrichment protocol based on sequential lysis of host and fungal cells on ice. Determination of genome wide transcription levels by microarrays and of selected genes (TSP1, HSP90, HSP104) by quantitative RT-PCR (q RT-PCR) revealed approximately 10% variation between RNA isolated from spiked murine organs using our enrichment protocol and RNA isolated from the spiking culture by standard methods24. We were able to isolate sufficient amounts of high-quality fungal RNA to perform microarray analysis from C. albicans cells retrieved from the liver 8 h p.i., the kidneys 12 h p.i., and the kidneys 24 h p.i., enabling us to detect 6569 genes in the arrays. The amounts recovered at other time points were sufficient only for q RT-PCR.
The lower amounts of fungal RNA obtained from the kidney at 8 h p.i. or the liver at later time points are possibly due to lower amounts of total fungal biomass and RNA at these time points in the organs. We furthermore detected no cross-hybridization of uninfected kidney samples with the C. albicans microarrays, indicating that the detected transcripts originate from C. albicans. The microarray data were verified by qPCR analyses of selected genes (Suppl. Table 1).
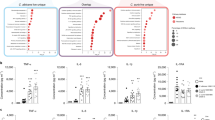
Compared to YPD culture (control), 917 C. albicans genes were differentially expressed in liver, 908 in kidney 12 h p.i., and 875 in kidney 24 h p.i., with 80 genes differentially expressed in all organs (Fig. 3, Suppl. data 3). These included infection-associated, highly upregulated individual genes, such as HWP1, ECE1, ALS1, ALS3, HYR1, and SOD5 (Table 1). Furthermore, GO categories related to cell wall and cell surface were significantly enriched in positively regulated genes of all samples (Fig. 4, Suppl. data 4), indicating remodelling of the cell wall and reflecting the upregulation of hypha-associated genes involved in adhesion such as ALS1, ALS3, DIF1, and HWP1 (Table 1).
C. albicans genes differentially expressed during systemic candidiasis in mice.
(A) Number of differentially expressed C. albicans genes in liver and kidney samples 8 h, 12 h and 24 h p.i. relative to common reference (log phase SC5314 grown in YPD at 37 °C). (B) Venn diagram representing the overlap of C. albicans DEGs in liver 8 h, kidney 12 h and kidney 24 h p.i. Within all three samples, 80 genes were differentially expressed in all samples, whereas most of the genes were differentially regulated at specific time points or organs.
GO terms enriched in positively regulated differentially expressed genes of C. albicans during systemic candidiasis.
The most significantly regulated ontologies were determined by the adjusted p-value (BP - biological process (orange); CC - cellular component (green), MF - molecular function (blue)). A full list of enriched GO terms is provided in Suppl. data 4.
While this demonstrates common regulation of a set of virulence-associated genes, most DEGs displayed organ- or time point-specific regulation (Fig. 3B). Some of the organ-specific differences might be due to the differences in the time points analysed (liver 8 h p.i. vs. kidney 12 h and 24 h p.i.), which is a limitation of this study; however, the higher similarity among gene expression profiles at two time points in the kidney compared to the liver (Suppl. Fig. 2A) still suggests organ-specific adaptations.
The transcriptional response of C. albicans during growth in the kidney is characterized by iron acquisition and metabolic adaptation
To elucidate which processes were affected by the DEGs, we performed GO term enrichment analysis and gene set enrichment analysis (GSEA) using published lists of in vivo and in vitro regulated C. albicans genes. Specific expression patterns deduced from GO term enrichment analysis included upregulation of “iron assimilation” in kidney samples at both time points, supported by GSEA results matching gene regulation in the kidney with iron homeostasis25 and consistent with two recently published in vivo transcriptional profiles of C. albicans cells infecting the mouse kidney15,16. This correlates well with local iron sequestration shown in kidney lesions26 and the upregulation of lactoferrin and haptoglobin by the host (Suppl. data 1), supporting the concept of nutritional immunity as a host defence mechanism27.
During early kidney infection, according to GO terms C. albicans furthermore upregulated translational processes (e.g. “ribosome biogenesis”, Fig. 4), which was supported by GSEA (ribosomal proteins in a set of putative targets of the transcription factor Fhl128). After 24 h in the kidney, the transcriptional profile of C. albicans indicated a response to starvation (Fig. 4), possibly as a result of rapid growth and hypha formation. In contrast, metabolic genes differentially expressed in the liver indicated catabolic processes and carbohydrate transport (Fig. 4). Some of the organ-specific metabolic adaptations of C. albicans could be a response to organ-specific nutrient supply. For example, while the glycerol biosynthetic gene RHR2 showed decreased expression in kidney, possibly reflecting the presence of the renal osmoprotectant glycero-phosphocholine15, RHR2 expression was increased in the liver (Suppl. data 3). Indeed, it has been shown that the ability of C. albicans to utilize external glycero-phosphocholine is important for virulence in murine systemic candidiasis29 and that RHR2 is essential for proliferation of C. albicans in the liver of intraperitoneally infected mice30.
Inference modelling identified three inter-species pathogen-host gene-regulatory networks linking fungal metabolism with the host immune response
Metabolic adaptation of C. albicans might also be influenced by immune reactions, especially restriction of nutrient availability upon phagocytosis13,31. To determine which fungal and host activities may directly influence each other, we inferred an inter-species pathogen-host gene-regulatory network32 using the ExTILAR algorithm33. This method integrates transcriptional data (DEGs of both species clustered by k-means; Suppl. Fig. 2B,C) with existing knowledge on transcription factor-dependent regulation of the genes to infer a regulatory network between DEGs of host (Suppl. data 5) and pathogen (Suppl. data 6). The resulting network showed three disconnected sub-networks each composed of mouse and Candida clusters.
The first network (Fig. 5A) was composed of a murine cluster containing downregulated genes involved in tissue homeostasis, remodelling and renal function, and thereby reflecting parts of the renal response to infection. This host cluster was both regulated by and regulating a cluster of upregulated fungal genes affecting iron homeostasis and biofilm formation (as observed in the kidney). This C. albicans cluster in turn was connected to the second C. albicans cluster, comprising downregulated genes involved in amino acid metabolism, similar to the observed changes in the liver. Thus, this sub-network suggests a link between host tissue function on the one side and fungal iron homeostasis and amino acid metabolism on the other, and reflects the organ-specific fungal transcription. Amino acid biosynthesis is affected by iron starvation in S. cerevisiae in a complex feedback loop involving transcription factors and biosynthesis gene containing iron-sulfur clusters34. Cysteine is important for the formation of iron-sulfur clusters and the C. albicans key iron regulator Sfu1 contains a cysteine-rich central domain that is typical for GATA-type transcription factors35. Cysteine might thus possibly link Sfu1 and amino acid biosynthesis via iron-sulfur clusters; this is however highly speculative. Alternatively, tissue damage caused by invasive fungal growth and the subsequent host response might significantly affect nutrient availability for and metabolism of the fungus.
Inferred inter-species pathogen-host gene-regulatory networks.
Nodes represent clusters of co-regulated genes for M. musculus (diamond) and C. albicans (triangle). Edges between two clusters represent directed regulatory interactions that are either positive (arrow) or negative (blunt end). A full list of genes and enriched GO terms of the clusters is provided in Suppl. Data 5 (mouse) and 6 (C. albicans).
In the second network (Fig. 5B) the central C. albicans cluster contained genes related to metabolic processes that were upregulated in the liver but downregulated in the kidney. This cluster was predicted to negatively affect the expression of genes in both, another fungal and a host gene cluster. In the host cluster, genes were associated with immune system processes and included genes such as IL6, CXCL10 and Interferon- and TNF-receptors, which were upregulated only in the kidney. The second C. albicans cluster also showed organ-specific regulation, with increased expression in the kidney and decreased expression in the liver. This cluster was significantly enriched for terms associated with the cell surface and included genes like CDC12, CHS3, and MYO2, that influence filamentation and cell wall composition and thereby could affect interaction with the immune system36. Moreover, binding sites for the transcription factor BCR1, which regulates biofilm formation and cell-surface associated genes37, was over-represented in the promoter region of genes in this cluster. Biofilm-associated genes were also found to be enriched in the kidney by GO term analysis (Fig. 4). Taken together, this sub-network indicates an organ-specific regulatory relationship between host immune responses, fungal metabolism and C. albicans morphology.
Similarly, the C. albicans clusters within the third network (Fig. 5C) also contained genes with organ-specific transcription patterns and functions for metabolic adaptation, while two of the four connected murine clusters (#1 and #3) were enriched in genes with functions for the immune response and organ-specific transcription.
Thus, all deduced networks support an organ-specific interplay between fungal metabolism and immune processes during infection. Immune reactions likely affect the availability of nutrients, e.g. upon phagocytosis and by iron sequestration13,27 but metabolic adaptations of C. albicans in turn can affect the interaction of the fungus with the immune system38.
Expression of hypha-associated genes in the absence of filamentation in the liver
This link between fungal metabolism and interaction with immune cells is likely to be mediated by changes of the fungal surface and cell wall38. Genes involved in biofilm formation were enriched at later time points in the kidney, consistent with observations by others15,16. GSEA furthermore identified a significant overlap of the in vivo transcriptome 24 h p.i. with genes differentially regulated in C. albicans ccr4Δ/Δ and sit4Δ/Δ39,40, both involved in cell wall maintenance and filamentation. The cell wall composition of C. albicans is also linked to morphogenesis, a central virulence attribute of C. albicans, and filamentation occurs early upon systemic infection of the murine kidney3. Not surprisingly, filamentation-associated genes were among the most strongly upregulated Candida genes in the kidney (Tables 1 and 2). In the liver, however, filamentation does not occur after intravenous infection2,41 (and own unpublished data). Thus, we were surprised to observe induction of hypha-associated genes (HAGs) and core filamentation response genes42 in the liver (Tables 1 and 2). To confirm the results obtained by microarrays, we performed qPCR on our RNA samples to analyse the expression levels of the six genes of the core filamentation response42. With the exception of DCK1, all core filamentation response genes showed a moderate to strong induction in both organs at all time-points under investigation and, with the exception of RBT1, similar expression levels in all samples (Table 2; Suppl. Fig. 3).
Expression of hypha-associated genes by C. albicans yeast cells in the liver could indicate cell wall remodelling
In addition to the core filamentation response genes, several genes encoding cell wall remodelling factors were specifically induced in C. albicans in the liver, e.g. CRH11 (cell wall transglycosylase), MP65 (cell surface mannoprotein), PHR1, BGL2 and PGA4 (glucanosyltransferases), and the chitinase gene CHT1.
It is known that the C. albicans cell wall is highly dynamic. In vitro, the exposure of C. albicans to different stresses that are encountered following phagocytosis can lead to rapid architectural changes and structural realignments of the cell wall43. Changes observed in vivo include altered exposure of chitin and β-glucan11,12. Both morphology and cell wall composition can influence the interaction with immune cells. For example, murine macrophages phagocytose C. albicans yeast cells more efficiently than hyphae44 and preferentially ingest O- and N-linked mannan-deficient mutants14. Cell wall glycosylation also affects phagosome maturation in macrophages45. We thus hypothesized that the induction of HAGs in yeast cells, as observed in the liver, could influence the interaction of the fungus with macrophages. To test this hypothesis, we inoculated log-phase yeast cells grown in YPD (0 min) into RPMI at 37 °C. In this condition, increased expression of ALS3, ECE1, HWP1, and IHD1 was detected by qPCR as early as 15 min after induction (data not shown), even though first germ tube formation only became visible after 45 min (Suppl. Fig. 4A). These C. albicans yeasts were then used in a phagocytosis assay with human monocyte-derived macrophages. We observed a significant increase in the phagocytosis rate of MDMs challenged with RPMI-induced yeast cells (Suppl. Fig. 4B,C) and alterations in the release of cytokines by macrophages stimulated with thimerosal-killed cells (Suppl. Fig. 4D). While this suggests that the induction of the filamentation program is indeed associated with changes that influence the interaction with immune cells, this hypothesis requires further investigation.
It has, however, been well established that both phagocytosis and cytokine production depend on the interaction of the fungal surface and components of the cell wall with host cell pattern recognition receptors. As the cell wall of C. albicans responds to numerous environmental cues45, it is plausible that the yeast cells found in the liver differ from both, cells grown in vitro under yeast-favouring conditions and the yeasts grown under hypha-inducing conditions in our in vitro experiment. Morphology-independent expression of HAGs has also been observed during growth in the intestinal tract46, suggesting that hypha-like modification of the yeast surface can occur in specific host niches, and GSEA revealed a significant overlap of DEGs in the liver with genes upregulated in the caecum46. It is however difficult to predict to which extent the interaction with immune cells in the liver might be influenced by the observed changes of the transcriptome. Given that HAGs were upregulated in the liver 8 h p.i., after prolonged exposure to a likely hypha-inducing environment, it nonetheless appears striking that visible hypha were absent in this organ – especially as they readily form in the kidney and in vitro within this time span. Possibly, phagocytosis of C. albicans yeasts in the liver exposes the fungus to an environment that prevents hyphal growth.
C. albicans likely faces phagocytosis early after infection of the liver
According to GSEA, the greatest overlap of analysed transcriptomes was observed between C. albicans DEGs specific for liver and the transcriptional response to phagocytosis by bone marrow-derived macrophages (Suppl. data 7)47. The upregulation of catabolic and transport processes by C. albicans in the liver (Fig. 4) furthermore resembled previously observed transcriptional changes upon phagocytosis13,31. Stress responsive genes that were upregulated following phagocytosis by PMNs, e.g. SOD5, YHB1, GPX2, ASR1, and CAT1, were also significantly upregulated by C. albicans cells in the liver (Suppl. data 8). In the kidney, in contrast, SOD5 and SSB1 were upregulated while other core stress response genes showed decreased expression (e.g. HSP70, HSP90 and CCP1; Suppl. data 8). All of the stress-responsive genes with increased expression in the kidney 24 h p.i. are also known to be induced upon oxidative stress48, possibly indicating interaction with immigrating phagocytes at later stages of infection in the kidney. These organ-specific differential expression kinetics of stress-associated fungal genes might reflect the higher abundance of immune cells in the liver. Residential immune cells likely induce an early profound stress response in C. albicans, which we were able to detect in the liver 8 h p.i. It has been shown that interaction between C. albicans and resident macrophages also occurs in the kidney within 2 h p.i.49; however, this interaction does not inhibit fungal filamentation and could not be detected on day 1 p.i. The low transcription of fungal stress genes in the kidney 12 h p.i. observed by us further supports that the interaction with resident phagocytes in the kidney is only transient and that additional phagocytes need to be recruited upon infection to this organ2,49. Thus, low transcription of fungal stress genes in the kidney at the 12 h time point, followed by delayed induction (compared to liver infection) could reflect a fatal later onset of interaction with recruited immune cells in the kidney, whereas early phagocytosis by resident or rapidly recruited immune cells may contribute significantly to the control of C. albicans in the liver.
As discussed above, phagocytosis of Candida yeasts in the liver possibly exposes the fungus to an environment that prevents hyphal growth. Even though C. albicans filaments within macrophages in vitro31, recent findings in a zebrafish model50 suggest that some macrophages can inhibit filamentation in vivo51 and C. albicans DEGs in the zebrafish model showed a significant overlap with the transcriptome in the liver. Furthermore, phagocytosis by PMNs prevents C. albicans filamentation13,52. It therefore appears plausible that C. albicans responds to the hepatic environment by induction of a hypha-associated program while physical formation of hypha is counteracted by the activity of phagocytes. This hypothesis is supported by the observation that combined neutropenia and C5 deficiency in mice led to the development of foci of fungal invasion in liver and spleen. This was not observed in neutropenic C5-sufficient mouse strains, indicating that in the absence of neutrophils other phagocytes control Candida in the liver53. However, further studies are needed to determine the type of immune cells and exact mechanisms that prevent hypha formation and facilitate fungal clearance in the liver.
Comparability of gene expression profiles across different studies
One very surprising result of the GSEA was that we did not observe a significant overlap with transcriptional data from C. albicans in the murine data published previously15,16,21. One explanation could be the approach used for our GSEA analysis, in which we combined data from both time points for the kidney to compare this data set to the DEGs in the liver. This initial comparison, designed to detect inter-organ differences, might partially explain why no significant enrichment was observed with other in vivo transcription data sets from individual organs referenced to pre-infection samples.
To analyse the comparability of our and published data sets in more detail, we performed a direct comparison (Suppl. Data 9), revealing a 31–44% overlap of DEGs identified in the different studies, with 73–90% of the overlapping DEGs displayed similar trends in expression (up- or downregulated, respectively). While this overlap may appear to be low, it should be interpreted considering that technical differences between the different studies influence the fold-change gene expression and thereby the DEGs identified depending on the cut off. For example, the NanoString technology used by Xu et al.15 is highly sensitive and might thus detect low-expression DEGs missed in our study, but is also limited to the set of genes included in the methodological set up. Similarly, RNASeq after specific enrichment procedures, as performed by Amorim-Vaz et al.16, is likely to be superior in sensitivity compared to microarrays. Amorim-Vaz et al. furthermore analysed time points different to our study, which will affect the direct comparison. Further technical differences hampering a meaningful comparison are the different control conditions chosen for C. albicans in vitro and the fungal strain used19.
While all these factors influence the DEGs identified in the different studies, it should be highlighted that distinct pathways (generally identified by GO analysis) were found to be induced in the kidney in all studies. This includes the upregulation of iron acquisition systems, filamentation and cell wall modification as well as specific virulence factors. This demonstrates that these pathways can be robustly identified in different settings and by different technologies and thus underlines their importance for the infection process. It also shows that pathway analyses (via GO-analysis or other bioinformatical approaches) are able to produce more robust and often more informative results than individual analysis of selected genes, for which expression levels might be strongly influenced by the chosen method.
In summary, our study clearly shows that at the transcriptional level both host responses and fungal adaptation during disseminated candidiasis are organ-specific. In the kidney, late onset of innate defence mechanisms, likely due to the comparatively low number of resident immune cells and slow recruitment of additional effector cells, facilitates fungal proliferation accompanied by filament formation, upregulation of iron acquisition mechanisms, and metabolic adaptation. Failure to control fungal growth likely drives the observed exacerbated induction of proinflammatory responses, thereby contributing to immunopathology. In contrast, innate immune factors are quickly upregulated in the liver and the fungal response indicates possible phagocytosis that might help to explain the noteworthy lack of filamentation in this organ. The distinct environments in the different organs likely drive the observed differential expression of cell wall and cell surface modifying enzymes. This in turn may lead to structural alterations that can affect interaction with immune cells and thus possibly contribute to the specific course of infections in different organs.
Material and Methods
Ethics statement
All animal experiments were performed in accordance with the German animal protection law and were approved by the responsible Federal State authority (Thüringer Landesamt für Lebensmittelsicherheit und Verbraucherschutz) ethics committee (beratende Komission nach §15 Abs. 1 Tierschutzgesetz; permit no. 03–009/13). The animals were cared for in accordance with the European Convention for the Protection of Vertebrate Animals Used for Experimental and Other Scientific Purposes.
Strains and culture condition
Candida albicans SC5314 and the GFP expressing strain M137 (Fradin et al., 2005) were maintained as glycerol stocks at −80 °C. For experiments, single colonies obtained from YPD agar (1% w/v peptone, 1% w/v yeast extract, 2% w/v glucose, 2% w/v agar) were grown overnight in liquid YPD (without agar) at 30 °C or in RPMI 1640 at 37 °C in a shaking incubator.
Murine infection model
The infection was performed as described previously54. Briefly, SC5314 cells from liquid YPD overnight cultures were washed twice with ice-cold PBS and adjusted to the desired concentration. The infection dose was confirmed by plating. On Day 0, three mice per time point were infected via the lateral tail vein with C. albicans ranging from 2.5 × 104 to 6.25 × 106 cfu/g body weight. Female BALB/c mice of 10–12 weeks were used for infection. The remaining infection solution was centrifuged and the pellet was snap frozen in liquid nitrogen and stored at −80 °C for later RNA extraction.
After infection, the health status of the mice was examined twice a day by a veterinarian, and surface temperature and body weight were recorded daily. Mice were humanely sacrificed 8 h, 12 h, 24 h, and 72 h post infection. Immediately after euthanasia, kidneys, spleen, and liver were removed aseptically, rinsed with sterile PBS and snap frozen in liquid nitrogen. PBS mock-infected animals served as controls.
RNA isolation
We designed an RNA isolation protocol for isolation of both fungal and murine RNA from the same sample based on step-wise isolation of host and fungal RNA (Suppl. Fig. 5). First, organs were aseptically homogenized in RLT buffer (Qiagen) with 1% β-Mercaptoethanol (β-ME) on ice water using an Ika T10 basic UltraTurrax homogenizer. Then, homogenates were centrifuged at 3,000 g at 4 °C for 3 min. The supernatant was used for mouse tissue RNA isolation with the RNeasy Mini Kit (Qiagen) as described by the manufacturer. The remaining pellets were vortexed on a mini-beadbeater (Precellys) for 5 sec at 5000 m/s in 1 ml RLT buffer (Qiagen) with 1% β-ME. After centrifugation at 4 °C, fungal RNA was isolated from the remaining pellets as previously described24. The RNA quality was determined using a Bioanalyzer (Agilent Inc.), the quantity was measured with a Nanodrop ND1000 (Peqlab).
Gene expression profiling
Genome-wide gene expression of C. albicans was analysed with C. albicans-specific microarrays (ClinEuroDiag). The red channel represents hybridization with Cy5-labeled C. albicans RNA from experimental mouse sepsis while the green channel always shows hybridization with Cy3-labeled RNA from C. albicans yeast cells growing logarithmically in standard YPD medium at 37 °C with shaking (common control). C. albicans RNA labelling, microarray hybridization, and scanning were performed as previously described24.
Genome-wide gene expression profiling of mouse samples was performed using the MouseRef-8 Expression and MouseWG6 v2.0 BeadChips (Illumina) according to the manufacturer’s instruction. From the RNA isolation, 200 ng total RNA with a Bioanalyzer RIN greater than seven were used for amplification prior to chip hybridization. Samples were analysed using the iScan platform (Illumina) measuring the variation of expression rate of > 42,000 transcripts. All microarray data are MIAME compliant and raw data have been deposited at GEO (GSE83682). The gene expression of liver, spleen, and kidney tissue 8 h, 12 h, 24 h, and 72 h after intravenous infection was compared to control samples of mock infected mice 24 h after PBS injection.
Expression data analyses
All analyses were performed in R using packages provided by Bioconductor 2.2655. C. albicans two-colour microarray data were pre-processed using the limma package56. “Printtiplowess” normalisation was used on each array separately to correct for spatial effects or cross-hybridization. Array spots corresponding to the same gene were summarized using the duplicated correlation function of limma. Normalization of the arrays was performed using between-array quantile normalization. Log2 fold-changes (log2-FC) were calculated between in vivo samples and the common reference using limma. Genes with Benjamini-Yekutieli corrected p-value < 0.05 were considered as differentially expressed.
Mouse data was annotated using appropriate Illumina manifest files. For each platform raw data were pre-processed independently using the lumi package57 for R, including background correction (bgAdjust) and subsequent data normalization using variance stabilization transformation and quantile normalization. Detection calls for each probe were performed at default parameters settings and probes detected as absent in all samples were removed. Subsequently, the illuminaMousev2.db package for R was used for re-annotation and probe filtering. We retained only probes with a quality grade of “good” or “perfect”. The normalized and pre-processed data set for the two different microarray platforms was integrated into a combined data set by probe ID matching and subsequent between array loess normalization (affy package 1.44; ref. 58) with the “span” parameter set to 0.1. Quality of the integration procedure was verified by analysis of the expression values of known housekeeping genes and identified stable reference genes before and after loess normalisation. Differential expression was assessed using limma with a Benjamini-Yekutieli corrected p-value below 0.05 and an absolute log2-FC of 0.5 for each organ and time-point. To compare expression values of the control samples between organs we included the within-donor correlation for samples derived from the same animal estimated using the duplicate Correlation function provided by limma.
GO-term analysis of C. albicans expression data was performed using FungiFun259. The results contain GO-categories with Fisher’s exact test determined, Benjamini-Hochberg corrected p-values below 0.05.
Cluster analysis
Clustering of differentially expressed genes was performed using the clValid package60 for R, allowing direct comparison of the results from various clustering methods. For our analysis, we compared the results of 5 clustering algorithms (Hierarchical clustering, k-means, SOTA61, Diana and Clara62) for 3 to 15 clusters. The resulting scores of each validation measure were scaled between 0 and 1 and transformed such that a score of 1 represents the best result. Subsequently, all scores were combined using the average of the mean internal score and the mean stability score. Finally, we selected the clustering method and the number of clusters according to the maximal overall score. Subsequent gene ontology enrichment analyses for the genes of each cluster were performed using the GOstats package63 excluding all terms with less than 10 or more than 500 genes.
Gene Set Enrichment Analysis (GSEA)
For gene set enrichment analysis (GSEA) a gene list was produced from (i) the combined DEGs of C. albicans in kidney 12 h and 24 h and (ii) liver 8 h p.i., which were imported as “phenotypes” to be compared into the GSEA software v2.2.0 (Broad Institute)64. Analysis parameters were as follows: norm, meandiv; scoring_scheme, weighted; Metric:Diff_of_Classes; set_min, 15; nperm, 1000; set_max, 5000. Gene sets with an FDR < 25% were considered as significant enriched. This set was used for a gene set enrichment analysis (GSEA) based on the recently published list of regulated genes compiled from the literature16 expanded with another recently performed in vivo transcriptional profile of C. albicans15 and the gene list “GSEA_Nantel_2012” kindly supplied to CGD by Andre Nantel (www.candidagenome.org/download/community/GSEA_Nantel_2012/).
Inference of pathogen-host gene-regulatory network
The ExTILAR algorithm33 was used for the inference of an inter-species pathogen-host gene-regulatory network (GRN) based on prior knowledge about transcription factor binding sites (TFBSs) and about genes known to affect the activity of transcription factors. oPOSSUM 3.065 was used to detect over-represented TFBSs for each cluster. For clusters formed by the genes of M. musculus we restricted the promoter region of the target genes between 2,000 base pairs (bp) upstream and 0 bp downstream of the transcription start site. TFs corresponding to TFBSs with a Z-score of 10 or more and a Fisher-score of 7 or more (default values) were considered as potential regulators of the respective cluster. For C. albicans, the S. cerevisiae tool required the mapping of ORF-IDs to gene symbols of the Saccharomyces Genome Database (SGD). oPOSSUM was then used at default values restricting the number of background genes to 5,000 randomly selected. TFs corresponding to TFBSs with a Z-score of 10 or more and a Fisher score of 5 or more were considered as potential regulators of the respective cluster.
Based on the temporal mean cluster expression profile and the respective standard deviation (SD) as well as the TF-to-gene information obtained from oPOSSUM we inferred 10,000 networks. For each inference, we sampled expression data from a normal distribution using the mean and SD of the respective cluster profile/time point. ExTILAR was applied at default parameters including stepwise forward selection for the optimization of the structure-template. The final stable consensus network was derived by selection of edges between clusters that were inferred in more than 50 percent of all networks. Using the stable consensus network structure as a template we applied an ordinary least squares approach for parameter estimation of the final network model.
Real time qPCR
For microarray validation and expression analysis of core filamentation response genes, qRT-PCR was performed using the my-Budget 5 × EvaGreen QPCR Mix II (Bio&Sell) in a C1000TM Thermal Cycler (BioRad) according to manufacture recommendations. Gene specific primers are listed in Suppl. Table 2. Relative gene expression levels were determined by the 2ΔΔCt method with ACT1 and EFB1 as housekeeping genes. RNA isolated from the C. albicans infection solution served as reference. Murine RNA from mock-infected mice served as negative control.
Additional Information
How to cite this article: Hebecker, B. et al. Dual-species transcriptional profiling during systemic candidiasis reveals organ-specific host-pathogen interactions. Sci. Rep. 6, 36055; doi: 10.1038/srep36055 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history
19 December 2016
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Perlroth, J., Choi, B. & Spellberg, B. Nosocomial fungal infections: epidemiology, diagnosis, and treatment. Med. Mycol. 45, 321–346 (2007).
Lionakis, M. S., Lim, J. K., Lee, C.-C. R. & Murphy, P. M. Organ-specific innate immune responses in a mouse model of invasive candidiasis. J. Innate Immun. 3, 180–199 (2011).
MacCallum, D. M., Castillo, L., Brown, A. J. P., Gow, N. A. R. & Odds, F. C. Early-expressed chemokines predict kidney immunopathology in experimental disseminated Candida albicans infections. PloS One 4, e6420 (2009).
Lionakis, M. S. & Netea, M. G. Candida and host determinants of susceptibility to invasive candidiasis. PLoS Pathog. 9, e1003079 (2013).
Lewis, R. E. et al. Epidemiology and sites of involvement of invasive fungal infections in patients with haematological malignancies: a 20-year autopsy study. Mycoses 56, 638–645 (2013).
Brown, G. D. Innate antifungal immunity: the key role of phagocytes. Annu. Rev. Immunol. 29, 1–21 (2011).
Ngo, L. Y. et al. Inflammatory monocytes mediate early and organ-specific innate defense during systemic candidiasis. J. Infect. Dis. 209, 109–119 (2014).
Moyes, D. L. et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 532, 64–68 (2016).
Sudbery, P. E. Growth of Candida albicans hyphae. Nat. Rev. Microbiol. 9, 737–748 (2011).
Ene, I. V. et al. Host carbon sources modulate cell wall architecture, drug resistance and virulence in a fungal pathogen. Cell. Microbiol. 14, 1319–1335 (2012).
Hopke, A. et al. Neutrophil Attack Triggers Extracellular Trap-Dependent Candida Cell Wall Remodeling and Altered Immune Recognition. PLoS Pathog. 12, e1005644 (2016).
Marakalala, M. J. et al. Differential adaptation of Candida albicans in vivo modulates immune recognition by dectin-1. PLoS Pathog. 9, e1003315 (2013).
Fradin, C. et al. Granulocytes govern the transcriptional response, morphology and proliferation of Candida albicans in human blood. Mol. Microbiol. 56, 397–415 (2005).
Lewis, L. E. et al. Stage specific assessment of Candida albicans phagocytosis by macrophages identifies cell wall composition and morphogenesis as key determinants. PLoS Pathog. 8, e1002578 (2012).
Xu, W. et al. Activation and alliance of regulatory pathways in C. albicans during mammalian infection. PLoS Biol. 13, e1002076 (2015).
Amorim-Vaz, S. et al. RNA Enrichment Method for Quantitative Transcriptional Analysis of Pathogens In Vivo Applied to the Fungus Candida albicans. mBio 6, e00942–915 (2015).
Allert, S., Brunke, S. & Hube, B. In Vivo Transcriptional Profiling of Human Pathogenic Fungi during Infection: Reflecting the Real Life? PLoS Pathog. 12, e1005471 (2016).
Weber, M. et al. Hepatic induction of cholesterol biosynthesis reflects a remote adaptive response to pneumococcal pneumonia. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 26, 2424–2436 (2012).
Walker, L. A. et al. Genome-wide analysis of Candida albicans gene expression patterns during infection of the mammalian kidney. Fungal Genet. Biol. FG B 46, 210–219 (2009).
MacCallum, D. M. Massive induction of innate immune response to Candida albicans in the kidney in a murine intravenous challenge model. Fems Yeast Res. 9, 1111–1122 (2009).
Andes, D., Lepak, A., Pitula, A., Marchillo, K. & Clark, J. A simple approach for estimating gene expression in Candida albicans directly from a systemic infection site. J. Infect. Dis. 192, 893–900 (2005).
De Filippo, K., Henderson, R. B., Laschinger, M. & Hogg, N. Neutrophil chemokines KC and macrophage-inflammatory protein-2 are newly synthesized by tissue macrophages using distinct TLR signaling pathways. J. Immunol. Baltim. Md 1950 180, 4308–4315 (2008).
Majer, O. et al. Type I interferons promote fatal immunopathology by regulating inflammatory monocytes and neutrophils during Candida infections. PLoS Pathog. 8, e1002811 (2012).
Wächtler, B., Wilson, D., Haedicke, K., Dalle, F. & Hube, B. From Attachment to Damage: Defined Genes of Candida albicans Mediate Adhesion, Invasion and Damage during Interaction with Oral Epithelial Cells. PLoS One 6, e17046 (2011).
Chen, C., Pande, K., French, S. D., Tuch, B. B. & Noble, S. M. An iron homeostasis regulatory circuit with reciprocal roles in Candida albicans commensalism and pathogenesis. Cell Host Microbe 10, 118–135 (2011).
Potrykus, J. et al. Fungal iron availability during deep seated candidiasis is defined by a complex interplay involving systemic and local events. PLoS Pathog. 9, e1003676 (2013).
Crawford, A. & Wilson, D. Essential metals at the host-pathogen interface: nutritional immunity and micronutrient assimilation by human fungal pathogens. FEMS Yeast Res. 15 (2015).
Lavoie, H. et al. Evolutionary tinkering with conserved components of a transcriptional regulatory network. PLoS Biol. 8, e1000329 (2010).
Bishop, A. C. et al. Glycerophosphocholine utilization by Candida albicans: role of the Git3 transporter in virulence. J. Biol. Chem. 288, 33939–33952 (2013).
Desai, J. V. et al. Coordination of Candida albicans Invasion and Infection Functions by Phosphoglycerol Phosphatase Rhr2. Pathog. Basel Switz. 4, 573–589 (2015).
Lorenz, M. C., Bender, J. A. & Fink, G. R. Transcriptional response of Candida albicans upon internalization by macrophages. Eukaryot. Cell 3, 1076–1087 (2004).
Schulze, S., Schleicher, J., Guthke, R. & Linde, J. How to Predict Molecular Interactions between Species? Front. Microbiol. 7, 442 (2016).
Vlaic, S. et al. The extended TILAR approach: a novel tool for dynamic modeling of the transcription factor network regulating the adaption to in vitro cultivation of murine hepatocytes. BMC Syst. Biol. 6, 147 (2012).
Philpott, C. C., Leidgens, S. & Frey, A. G. Metabolic remodeling in iron-deficient fungi. Biochim. Biophys. Acta 1823, 1509–1520 (2012).
Lan, C.-Y. et al. Regulatory networks affected by iron availability in Candida albicans. Mol. Microbiol. 53, 1451–1469 (2004).
Warenda, A. J. & Konopka, J. B. Septin function in Candida albicans morphogenesis. Mol. Biol. Cell 13, 2732–2746 (2002).
Nobile, C. J. et al. A recently evolved transcriptional network controls biofilm development in Candida albicans. Cell 148, 126–138 (2012).
Brown, A. J. P., Brown, G. D., Netea, M. G. & Gow, N. A. R. Metabolism impacts upon Candida immunogenicity and pathogenicity at multiple levels. Trends Microbiol. 22, 614–622 (2014).
Dagley, M. J. et al. Cell wall integrity is linked to mitochondria and phospholipid homeostasis in Candida albicans through the activity of the post-transcriptional regulator Ccr4-Pop2. Mol. Microbiol. 79, 968–989 (2011).
Lee, C.-M., Nantel, A., Jiang, L., Whiteway, M. & Shen, S.-H. The serine/threonine protein phosphatase SIT4 modulates yeast-to-hypha morphogenesis and virulence in Candida albicans. Mol. Microbiol. 51, 691–709 (2004).
Rozell, B., Ljungdahl, P. O. & Martínez, P. Host-pathogen interactions and the pathological consequences of acute systemic Candida albicans infections in mice. Curr. Drug Targets 7, 483–494 (2006).
Martin, R. et al. A core filamentation response network in Candida albicans is restricted to eight genes. PloS One 8, e58613 (2013).
Ene, I. V. et al. Cell Wall Remodeling Enzymes Modulate Fungal Cell Wall Elasticity and Osmotic Stress Resistance. mBio 6, e00986 (2015).
Keppler-Ross, S., Douglas, L., Konopka, J. B. & Dean, N. Recognition of yeast by murine macrophages requires mannan but not glucan. Eukaryot. Cell 9, 1776–1787 (2010).
Bain, J. M. et al. Candida albicans hypha formation and mannan masking of β-glucan inhibit macrophage phagosome maturation. mBio 5, e01874 (2014).
Rosenbach, A., Dignard, D., Pierce, J. V., Whiteway, M. & Kumamoto, C. A. Adaptations of Candida albicans for growth in the mammalian intestinal tract. Eukaryot. Cell 9, 1075–1086 (2010).
Marcil, A. et al. Analysis of PRA1 and its relationship to Candida albicans- macrophage interactions. Infect. Immun. 76, 4345–4358 (2008).
Enjalbert, B., MacCallum, D. M., Odds, F. C. & Brown, A. J. P. Niche-specific activation of the oxidative stress response by the pathogenic fungus Candida albicans. Infect. Immun. 75, 2143–2151 (2007).
Lionakis, M. S. et al. CX3CR1-dependent renal macrophage survival promotes Candida control and host survival. J. Clin. Invest. 123, 5035–5051 (2013).
Chen, Y. Y. et al. Dynamic transcript profiling of Candida albicans infection in zebrafish: a pathogen-host interaction study. PloS One 8, e72483 (2013).
Gilbert, A. S., Wheeler, R. T. & May, R. C. Fungal Pathogens: Survival and Replication within Macrophages. Cold Spring Harb. Perspect. Med. 5, a019661 (2015).
Rubin-Bejerano, I., Fraser, I., Grisafi, P. & Fink, G. R. Phagocytosis by neutrophils induces an amino acid deprivation response in Saccharomyces cerevisiae and Candida albicans. Proc. Natl. Acad. Sci. USA 100, 11007–11012 (2003).
Fulurija, A., Ashman, R. B. & Papadimitriou, J. M. Neutrophil depletion increases susceptibility to systemic and vaginal candidiasis in mice, and reveals differences between brain and kidney in mechanisms of host resistance. Microbiol. Read. Engl. 142 (Pt 12), 3487–3496 (1996).
Jacobsen, I. D., Lüttich, A., Kurzai, O., Hube, B. & Brock, M. In vivo imaging of disseminated murine Candida albicans infection reveals unexpected host sites of fungal persistence during antifungal therapy. J. Antimicrob. Chemother. 69, 2785–2796 (2014).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Du, P., Kibbe, W. A. & Lin, S. M. lumi: a pipeline for processing Illumina microarray. Bioinforma. Oxf. Engl. 24, 1547–1548 (2008).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. affy–analysis of Affymetrix GeneChip data at the probe level. Bioinforma. Oxf. Engl. 20, 307–315 (2004).
Priebe, S., Kreisel, C., Horn, F., Guthke, R. & Linde, J. FungiFun2: a comprehensive online resource for systematic analysis of gene lists from fungal species. Bioinforma. Oxf. Engl. 31, 445–446 (2015).
Hartigan, J. A. Clustering algorithms. (Wiley, 1975).
Herrero, J., Valencia, A. & Dopazo, J. A hierarchical unsupervised growing neural network for clustering gene expression patterns. Bioinforma. Oxf. Engl. 17, 126–136 (2001).
Kaufman, L. & Rousseeuw, P. J. Finding Groups in Data: An Introduction to Cluster Analysis. (Wiley, 2005).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinforma. Oxf. Engl. 23, 257–258 (2007).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Kwon, A. T., Arenillas, D. J., Worsley Hunt, R. & Wasserman, W. W. oPOSSUM-3: advanced analysis of regulatory motif over-representation across genes or ChIP-Seq datasets. G3 Bethesda Md 2, 987–1002 (2012).
Acknowledgements
We thank Christine Dunker for providing material to optimize the RNA isolation method and Volha Skrahina for qRT-PCR primers. For experimental assistance we thank Markus Bläss, Melanie Polke, Birgit Weber, Nadja Jablonowski, and Stephanie Wisgott. This work was supported by the German Federal Ministry of Education and Health (BMBF) Germany, (FKZ 01EO1002, Integrated Research and Treatment Center, Center for Sepsis Control and Care (CSCC), the Jena School for Microbial Communication (JSMC) and by the Hans Knöll Institute Jena. IDJ, MB and JL are supported by the Deutsche Forschungsgemeinschaft CRC/Transregio 124 ‘Pathogenic fungi and their human host: Networks of interaction’ subprojects C5 (IDJ, MB) and INF (JL). SV is supported by the German Federal Ministry of Education and Research (BMBF, ‘InfectControl,’ subproject FINAR).
Author information
Authors and Affiliations
Contributions
I.D.J. and B.H. designed the experiments. B.H. performed murine work, RNA isolation, and C. albicans microarrays. Cytokine and phagocytosis assays were performed by B.H. and M.K. S.V. performed differential murine gene expression analysis. Cluster analysis and inference network analysis were performed by S.V. and T.C. C. albicans gene expression analysis was performed by T.C., B.H. and J.L. S.B. and B.H. performed GSEA. B.H., B.H., M.B. and I.D.J. interpreted the data. B.H. and I.D.J. wrote the paper; all authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hebecker, B., Vlaic, S., Conrad, T. et al. Dual-species transcriptional profiling during systemic candidiasis reveals organ-specific host-pathogen interactions. Sci Rep 6, 36055 (2016). https://doi.org/10.1038/srep36055
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep36055
This article is cited by
-
Communication between immune system and mycobiota impacts health and disease
Proceedings of the Indian National Science Academy (2022)
-
Microbiota-driven interleukin-17 production provides immune protection against invasive candidiasis
Critical Care (2020)
-
Nucleic acids enrichment of fungal pathogens to study host-pathogen interactions
Scientific Reports (2019)
-
Coordinated host-pathogen transcriptional dynamics revealed using sorted subpopulations and single macrophages infected with Candida albicans
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.