Abstract
Leucine rich repeat kinase 2 is a complex enzyme with both kinase and GTPase activities, closely linked to the pathogenesis of several human disorders including Parkinson’s disease, Crohn’s disease, leprosy and cancer. LRRK2 has been implicated in numerous cellular processes; however its physiological function remains unclear. Recent reports suggest that LRRK2 can act to regulate the cellular catabolic process of macroautophagy, although the precise mechanism whereby this occurs has not been identified. To investigate the signalling events through which LRRK2 acts to influence macroautophagy, the mammalian target of rapamycin (mTOR)/Unc-51-like kinase 1 (ULK1) and Beclin-1/phosphatidylinositol 3-kinase (PI3K) pathways were evaluated in astrocytic cell models in the presence and absence of LRRK2 kinase inhibitors. Chemical inhibition of LRRK2 kinase activity resulted in the stimulation of macroautophagy in a non-canonical fashion, independent of mTOR and ULK1, but dependent upon the activation of Beclin 1-containing class III PI3-kinase.
Similar content being viewed by others
Introduction
Leucine rich repeat kinase 2 is one of the key genetic factors contributing to the risk of developing Parkinson’s disease (PD), an irreversible, progressive neurodegenerative movement disorder primarily associated with neuronal cell loss in the Substantia nigra pars compacta. Coding mutations in the LRRK2 gene are the most frequent genetic cause of familial PD, with polymorphisms in LRRK2 associated with an increased risk of idiopathic PD1,2,3,4. In addition to this, genome wide association (GWA) studies recently identified the LRRK2 locus as being involved in the risk for PD5, Crohn’s disease6 and multibacillary leprosy7,8. Mutations in LRRK2 have also been linked to cancer9, and the LRRK2 region was identified as being subject to frequent carcinogenic alterations10. The LRRK2 gene is therefore related to the etiopathogenesis of at least four human diseases, making it the focus of increasing attention as a putative drug target.
The physiological function of LRRK2 is as yet unclear. It is a complex enzyme, with active kinase and GTPase domains that are thought to reciprocally regulate one another’s activity11,12. As detailed in the following section, several studies have indicated a putative role for LRRK2 in the control of macroautophagy, a process used by the cell to maintain a healthy microenvironment by removing misfolded proteins and damaged organelles13. The molecular mechanism underlying this association remains to be fully understood. While LRRK2 over-expression was associated with a macroautophagy-dependent induction of toxicity coupled with neurite atrophy14, LRRK2 knock down was shown to both reduce and potentiate the autophagic flux15,16. Moreover, the overexpression of full-length LRRK2, or its kinase domain, as well as inhibition of LRRK2 kinase activity induced alterations of the macroautophagy-lysosomal pathway17,18,19. Macroautophagy was shown to be altered in human fibroblasts carrying LRRK2 pathogenic mutations associated with PD20,21, in neurons derived from those human fibroblasts22 and in transgenic or LRRK2 knock-out mouse models23. Finally, pathogenic mutations in LRRK2 have been linked to deregulation of chaperone mediated autophagy (CMA)24. More generally, LRRK2 was associated with vesicle trafficking and synaptic functionality25,26, and with endocytosis and trans-Golgi network homeostasis27,28. A hypothetical function for LRRK2 in the regulation of macroautophagy, and in general in vesicle homeostasis, is compelling considering that the macroautophagy/lysosomal system has an increasingly appreciated link to the etiology of PD29, while it has long been considered a central player in the pathogenesis of Crohn’s, leprosy and cancer.
The data presented herein demonstrate that the kinase activity of LRRK2 acts as a negative regulator of macroautophagy in astrocyte cell models. Our results suggest that LRRK2 may act to control a non-canonical pathway alternative and parallel to that regulated by the mammalian target of rapamycin (mTOR) and Unc-51 Like Kinase 1 (ULK1), but dependent on the presence of an active Beclin-1 complex. These data have important implications for the study of the physiological and pathological functions of LRRK2, in particular for any pharmacological intervention based upon LRRK2 inhibition.
Results
Inhibition of LRRK2 kinase activity increases LC3-II levels
LRRK2 is expressed at high levels in astrocytes within the brain30,31. Human H4 neuroglioma cells, originally derived from an astrocytoma, were previously used as a model to study LRRK2 function in macroautophagy18,30. Based on a previous work by our group18, we here replicate and expand our previous analysis confirming that treatment of H4 cells for 150 minutes (acute treatment) or 18 hours (chronic treatment) with LRRK2 kinase inhibitors, either LRRK2in132 or GSK2578215A33 result in a concentration dependent increase of LC3-II (Fig. 1a,b); no concomitant toxicity was recorded for the LRRK2in1 while a decrease in cell survival was detected for GSK2578215A starting at 30 μM (Supplementary Fig. S1a,b)34. A major confounding factor when using chemical inhibitors of enzymes is the possibility of off target effects. Although the inhibitors used are structurally distinct, it is critical to demonstrate that the cellular phenotypes measured are specific to the protein of interest. To achieve this, and as already previously proposed by our group, endogenous LRRK2 protein levels in H4 cells were decreased (~50%) by stable expression of LRRK2 shRNA (Fig.1c,d). 150 minutes (Fig. 1e,f) or 18 hours (Supplementary Fig. S1c) inhibition of LRRK2 kinase activity by LRRK2in1 increased LC3-II in scrambled controls cells but not in LRRK2 knocked-down cells, strongly suggesting that this is a LRRK2 dependent phenomenon. Interestingly, in this model system we consistently see no increase in basal LC3-II when knockin-down LRRK2. Further investigations are needed, however we suggest this may happen either because the knock-down is never complete, or alternative splicing isoforms may originate. Moreover, it is worth noticing that, with the knock-down strategy, we remove the kinase as well as the GTPase activities and the protein-protein interaction domains of LRRK2. Therefore the chemical kinase inhibition and the protein knock-down approaches may not represent a perfect phenocopy of each other. Primary astrocytes from LRRK2 knock-out mice were prepared and assessed for response to LRRK2 inhibition. An elevated, basal level of LC3-II was detected in the knock-out cells that was not significantly increased following treatment with LRRK2in1 (Fig. 1g,h, see Supplementary Fig. S1e for astrocytes evaluation by GFAP staining).
Inhibition of LRRK2 kinase leads to a dose response and time dependent increase in LC3-II.
150 minutes (a) or 18 hours (b) LRRK2in1 and GSK2578215A dose-response. The gels shown are representative of 3 independent experiments, LC3-II was quantified against β-actin, quantification was done for each single experiment; after normalization against the control in DMSO, data were pooled together in the dose-response curve (mean and SEM). (c) H4 cells with stable LRRK2 knock-down (~50% LRRK2 knock-down for KD#644167.03b transfection) as quantified in (d); LRRK2 was quantified (sum of the upper and lower bands) against β-actin, statistical analysis was performed by unpaired student t-test (mean and SD, p value = 0.0236). (e) 150 minutes LRRK2in1 treatment in scramble controls and in LRRK2 knock-down cells; the image shown (reporting 3 samples for each condition) is representative of 3 independent experiments and is quantified in (f); LC3-II was quantified against β-actin; statistical analysis was performed by 1way Anova followed by Tukey post-hoc test (mean and SD, **p < 0.01; *p < 0.05). (g) primary astrocytes from wild-type and LRRK2 knock-out mice treated with LRRK2in1 for 18 hours, the image shown is representative of 3 independent cell preparations, each replicated with 3 samples. (h) LC3-II was quantified against β-actin for each single experiment; after normalization against the internal control in DMSO, data were pulled together and statistical analysis was performed by 1way Anova followed by Tukey post-hoc test. (mean and SD, *p < 0.05).
LRRK2 dependent increase of LC3-II is not due to decreased autophagosome-lysosome fusion
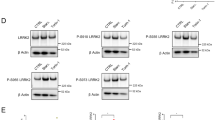
We have previously suggested that the increased levels of LC3-II after LRRK2 kinase inhibition were consequence of an increase in autophagosome production rather than a decrease in degradation. We here corroborate this data by improving the experimental setting including Torin-1 control, evaluating two distinct time-points of treatment, assessing the drug Chloroquine alongside with Bafilomycin (BafA1) and performing co-localization analysis of LAMP1 and p62. As first, H4 cells were co-treated with LRRK2 kinase inhibitor and BafA1 or Chloroquine. Results showed an additive effect of LRRK2in1 over BafA1 (or Chloroquine) treatment alone (Fig. 2a,b, Supplementary Fig. S2a) that appeared at 150 minutes and became significant at 18 hours (Fig. 2c–e), similarly to that observed with Torin-1, an mTOR inhibitor. This confirmed, as previously reported18, that the LC3-II increase following inhibition of LRRK2 kinase activity, is different from the effect of BafA1, thus suggesting an increase in macroautophagy flux34. Since BafA1, which inhibits autophagosome-lysosome fusion, has an additional, distinct action in preventing lysosomal acidification35, the pH of cellular vesicles was assessed in H4 cells treated for 150 minutes or 18 hours with LRRK2in1 or Torin-1, in the presence or absence of BafA1. Using neutral red accumulation no disruption of vesicle acidification was detected either with LRRK2in1 or Torin-1 (Supplementary Fig. S2b), further confirming that the inhibition of LRRK2 kinase activity does not affect lysosomal function. p62 is a cargo protein used to target substrates for degradation through autophagy. We evaluated the co-localization between p62 and LAMP1 (a lysosomal marker) to assess the fusion between lysosomes and autophagosomes34,36 (Supplementary Fig. S3) showing that, whereas BafA1 was able to reduce the co-localization between p62 and LAMP1 as consequence of inhibition of autophagosome-lysosome fusion, cells treated with Torin-1 or LRRK2in1 were characterized by a higher proportion of co-localized vesicles suggesting that no impairment in autophagy degradation was occurring under such treatments.
Increase in LC3-II after inhibition of LRRK2 kinase is due to an induction of the macroautophagy flux.
(a) 18 hours treatment with LRRK2in1 in the presence and absence of BafA1 to block the autophagy flux. The gel shown is representative of 3 independent experiments, each performed in triplicate and it is quantified in (b). (b) LC3-II is quantified against β-actin; statistical analysis was performed by 1way Anova followed by Tukey post-hoc test (mean and SD, **p < 0.01). 150 minutes (c) and 18 hours (d) treatment with LRRK2in1 or Torin-1 (to induce macroautophagy) in the presence and absence of BafA1. The gels shown are representative of at least 4 independent experiments summarized in (e). (e) LC3-II is quantified against β-actin for 5 replicates (150 minutes) and 4 replicates (18 hours); quantification was done for each single experiment; after normalization against the internal control in DMSO, data were pulled together and statistical analysis was performed by 1way Anova followed by Tukey post-hoc test (mean and SD, ***p < 0.001, *p < 0.05).
Inhibition of LRRK2 kinase activity induces macroautophagy independently of mTOR/ULK1
In our previous work18 we did not record any alteration of P70S6K and S6 phosphorylation when macroautophagy was induced by LRRK2 kinase inhibition. This suggested that, in this specific case, induction of macroautophagy may not follow the canonical mTOR pathway. With a new set of tailored experiments we here properly assess that previous suggestion and corroborate the idea that inhibition of LRRK2 kinase activity induces macroautophagy independently of mTOR/ULK1. In canonical macroautophagy, ULK1 is the key downstream effector of mTOR and AMP-activated protein kinase (AMPK) for phagophore generation37. Inactivation of mTOR (i.e. following Torin-1 treatment) results in de-phosphorylation of ULK1, and subsequent macroautophagy induction. Canonical macroautophagy induced by Torin-1 treatment and involving mTOR inhibition (as indicated by reduction in the phosphorylation level of p70S6 kinase at Thr389) was strongly associated with decreased mTOR-dependent phosphorylation of ULK1 at Ser758 (Fig. 3a). 150 minutes of LRRK2-in1 treatment was sufficient to increase the levels of the macroautophagy marker LC3-II, however no reduction of ULK1 phosphorylation on Ser758, nor alteration of mTOR activity was observed, as indicated by phosphorylation levels of p70S6K at Thr389 (Fig. 3a).
Increased macroautophagy following inhibition of LRRK2 kinase is mTOR independent.
(a) 150 minutes treatment with LRRK2in1 or Torin-1 (to induce macroautophagy through mTOR). Phosphorylation of P70S6K and ULK1 were used as readout for mTOR inhibition. (b) 150 minutes treatment with LRRK2in1 or Torin-1 in scramble controls and in H4 cells with stable ULK1 knock-down (~70% ULK1 knock-down). The gel shown is representative of 4 independent experiments that are quantified in (c). (c) Quantification was done for each single experiment and LC3-II was normalized against β-actin. After normalization against the control in DMSO, data were pulled together and statistical analysis was performed by 1way Anova followed by Tukey post-hoc test (mean and SD, *p < 0.05). (d) 18 hours treatment with LRRK2in1 in the presence and absence of Torin-1. Phosphorylation on P70S6K was used as control for mTOR inhibition. The gel shown is representative of 3 independent experiments each performed in triplicate and quantified in (e). (e) LC3-II was quantified against β-actin; statistical analysis was performed by 1way Anova followed by Tukey post-hoc test (mean and SD, **p < 0.01, ***p < 0.001).
In order to assess whether ULK1 is required for LRRK2-induced macroautophagy, ULK1 protein levels were stably reduced by 70% in H4 cells transfected with ULK1 shRNA. 150 minute treatment of LRRK2in1 led to a comparable increase of LC3-II production in stable ULK1 knock-down as compared to scrambled control cells. These data indicate that ULK1 is dispensable for macroautophagy induced by LRRK2 kinase activity inhibition (Fig. 3b,c).
Co-treatment with LRRK2in1 and Torin-1 resulted in a stronger increase of LC3-II levels, as compared to single treatments with either LRRK2in1 or Torin-1 (Fig. 3d,e). This data supports an additive effect of the combined inhibition of LRRK2 and mTOR over the macroautophagy flux, further suggesting that these kinases act in parallel pathways to control macroautophagy induction and that the regulation of autophagy by LRRK2 is independent of mTOR/ULK1. Similar results were obtained when m-TOR was alternatively inhibited by aminoacid starvation (Supplementary Fig. S4a,b) or following Rapamycin treatment (Supplementary Fig. S4c), further demonstrating that LRRK2 control over autophagy is independent of mTOR.
Activation of macroautophagy after inhibition of LRRK2 kinase requires PI3P/Beclin-1
In addition to the ULK-kinase complex, canonical macroautophagy induction requires the class III PI3-kinase complex containing Beclin-138. WIPI-2 is directly recruited to the nascent autophagosome by the presence of PI3P in the autophagosome membrane, following the activity of the Beclin-1 complex. We have previously shown that LRRK2in1 increases the number of WIPI2 positive punctae in cells18. To confirm that this effect can be reproduced, we repeated the prior experiment using a different quantification algorithm (Supplementary Fig. S5). This result suggests that, in contrast to mTOR inhibition and ULK1 activation, PI3P and WIPI-2 may be required for macroautophagy induced by LRRK2 inhibition. To further test this hypothesis, we performed a new and tailored set of experiments. LC3-II levels after LRRK2in1 treatment were assessed in H4 cells treated with wortmannin (or VPS34in1) to inhibit PI3K (or specifically VPS34) and consequently the production of PI3P. Co-treatment with wortmannin (Fig. 4a,b) or VPS34in1 (Fig. 4c,d) and either LRRK2in1 or Torin-1 blocked LRRK2-induced macroautophagy and mTOR-induced macroautophagy respectively further confirming that PI3P is required for autophagosome production under LRRK2 inhibition.
Increase in macroautophagy after inhibition of LRRK2 kinase and Beclin-1.
(a) 150 minutes treatment with LRRK2in1 or Torin-1 (to induce macroautophagy) in the presence and absence of wortmannin. The gel shown is representative of 3 independent experiments as quantified in (b). (b) LC3-II was quantified against β-actin; quantification was done for each single experiment; after normalization against the control in DMSO, data were pulled together and statistical analysis was performed by 1way Anova followed by Tukey post-hoc test (mean and SD, ***p < 0.001, **p < 0.01). (c) 150 minutes treatment with LRRK2in1 or Torin-1 (to induce macroautophagy) in the presence and absence of VPS34in1. The gel shown is representative of 4 independent experiments as quantified in (d). (d) LC3-II was quantified against β-actin; quantification was done for each single experiment; after normalization against the control in DMSO, data were pulled together and statistical analysis was performed by 1way Anova followed by Tukey post-hoc test (mean and SD, ***p < 0.001, **p < 0.01). (e) 150 minutes treatment with LRRK2in1 in H4 cells stably expressing scrambled shRNA controls or shRNA for Beclin-1. The gel shown is representative of 3 independent experiments, each performed in triplicate and it is quantified in (f). (f) LC3-II was normalized against β-actin and statistical analysis was performed by Anova followed by Tukey post-hoc test (mean and SD, *p < 0.05).
Finally, in order to assess whether Beclin-1 is required for LRRK2-induced macroautophagy, Beclin-1 protein levels were stably reduced in H4 cells transfected with Beclin-1 shRNA(s). Knock-down of Beclin-1 increased the basal level of LC3-II and this was not further increased after 150 minutes treatment with LRRK2-in 1, in contrast to scrambled shRNA controls which showed an enhancement of LC3-II compared to basal levels (Fig. 4e,f). This data further suggested that the macroautophagy pathway controlled by Beclin-1, but not ULK1, is required for the negative regulation by LRRK2 kinase.
Discussion
Despite its key role in the etiology of a number of human diseases, there is as yet no consensus regarding the physiological function of LRRK2; leading to the suggestion that LRRK2 may have contrasting roles depending on the cell type and condition under investigation39. A number of reports have implicated LRRK2 in the regulation of macroautophagy, in endocytosis and metabolism, however the precise molecular function of LRRK2 in these processes is still not defined. The data reported here demonstrate that, at least one of the functions supported by the endogenous LRRK2 kinase activity in a H4 glioma cell line and in primary astrocytes is involved in the non-canonical control of macroautophagy, working parallel to the mTOR/ULK1 pathway and dependent on PI3P and Beclin-1 activity.
The complex events regulating macroautophagy are yet to be fully characterised40. Two of the key regulatory systems have, however, been identified: the mTOR/ULK1 and the Beclin-1 pathways. While ULK1 is phosphorylated under basal conditions to repress macroautophagy, ULK1 phosphorylation is lost when macroautophagy flux is induced, via the orchestrated inhibition of mTOR in concert with as yet unidentified phosphatases41,42. Beclin-1 works in two distinct complexes, both necessary for the production of PI3P that is required first for the formation and later for the maturation of the nascent autophagosome. Non-canonical pathways have been described which do not require proteins crucial for canonical autophagy such as ULK1, Beclin-1 and LC343,44,45. As an example, the phenotype of the double knock-out for both ULK1 and ULK2 is embryonic lethal, but mice embryonic fibroblasts on this background are able to activate a residual autophagy pathway that is, by definition, independent of ULK1/2 proteins46. Moreover, the knock-down of both ULK1 and ULK2 in a B-cell line (DT40) did not alter autophagy47, thus confirming that an ULK1/2 alternative route does exist and suggesting different cell types have different ways to control and sustain macroautophagy. Finally, small molecules enhancers of rapamycin (SMERs) have been isolated to induce mTOR-independent autophagy48.
In the current study, pharmacological inhibition of LRRK2 kinase activity resulted in an increase of LC3 processing consistent with induction of macroautophagy. Inhibition of mTOR using Torin-1 causes a complete loss of ULK1 phosphorylation on Ser758 and this loss of phosphorylation on ULK1 is required for translocation of the ULK1/2 complex to the pre-autophagosomal structure37. In contrast, the lack of alteration in ULK1 phosphorylation following LRRK2 kinase inhibition, suggests the possible activation of a non-canonical pathway independent of ULK1. Autophagosomes produced due to loss of LRRK2 kinase activity were positive for WIPI-2 thus suggesting the activity of Beclin-1 and the generation of PI3P was conserved. Indeed, the inhibition of PI3P production by wortmannin or VPS34in1 was able to inhibit the induction of LRRK2-regulated macroautophagy. Finally, Beclin-1 knock-down cells were not responsive to LRRK2in1 treatment, thus confirming that Beclin-1/PI3P are involved either upstream or downstream of the induction of macroautophagy via LRRK2 inhibition. A possible link between Beclin-1 and LRRK2 is intriguing given the existing literature. The G2019S-LRRK2 mutant (which shows increased kinase activity), is able to bind to and phosphorylate Bcl-2 resulting in dysregulation of mitophagy49. Beclin-1 contains Bcl-2 binding sites that are important for its macroautophagy regulatory activities50. Beclin-1 has been found to be phosphorylated and regulated by the stress responsive kinases MAPKAPK2 and 351; LRRK2 has been suggested to bind and phosphorylate MAP2K3 (MKK3) upstream of MAPKAPK2/3 in the p38-MAPK pathway52. It is also of note that the Beclin complex has a complex role in the regulation of macroautophagy, dependent upon the binding partners Beclin-1 is associated with53,54,55. In the presence of ATG14L or UVRAG, Beclin-1 acts as a positive regulator of macroautophagy – consistent with the data presented herein from H4 cells. In contrast, when bound to RUBICON Beclin-1 acts as a repressor of autophagy. Clarifying how LRRK2 relates to these distinct Beclin complexes will provide further insight into the precise mechanisms whereby LRRK2 can impact on autophagy, and may provide an explanation for the divergence in experimental data across the research literature. Strikingly, regulation of macroautophagy by LRRK2 is echoed by studies on Death Associated Protein kinase 1 (DAPK1), another member of the ROCO protein family to which LRRK2 belongs, and that has been proposed to be involved in the control of macroautophagy by both direct and indirect phosphorylation of Beclin-156,57. Members of the ROCO family share a high homology in their GTPase and COR domains, while the kinase domains of LRRK2 and DAPK1 belong to different families. There is accumulating evidence, however, that the kinase and the GTPase activities present in LRRK2 are able to regulate one another58. These observations suggest that an understanding of both the kinase and GTPase activities of LRRK2 may be required to fully illuminate LRRK2’s role in the regulation of macroautophagy. Within the current study we make use of chemical inhibition of LRRK2 kinase activity to infer how the endogenous protein regulates macroautophagy by physiological signaling in cells. However, we cannot currently determine if the effects here are solely due to kinase activity or are partially mediated through, for example, the GTPase function of LRRK2. Further experiments addressing the GTPase activity of LRRK2, either through genetic manipulations or small molecules that target this region of the protein, are therefore important in the future.The close genetic ties between LRRK2 and PD have resulted in extensive efforts to understand LRRK2 in the context of this disorder. The recent discovery of significant links between LRRK2 and a range of other, apparently unrelated, human disorders has further emphasised the importance of this protein to human health. A major challenge for LRRK2 research is, therefore, to reconcile the involvement of LRRK2 in the pathological pathways underlying these disparate disorders. Several complementary strands of evidence suggest that one key physiological function of LRRK2 is with regard to the control of macroautophagy and vesicle dynamics, raising the possibility that this may be the common theme behind the different disorders in the pathogenesis of which LRRK2 has been implicated. Another major problem in the study of LRRK2 patho-physiology is represented by the fact that it has been associated with a multiple array of cellular functions59, and has many protein interactors60, suggesting that LRRK2 acts as a hub protein able to work with a wide range of partners. This has led to the suggestion that LRRK2 may have different roles in different cell types and its function may be different depending on the situation/stimuli. The data presented in this study provides new insight into the dissection of the mechanism of LRRK2 function, information that will be critical for understanding the connection between LRRK2 and disease pathogenesis, to provide the basis for therapeutic intervention directed at LRRK2s activities and to consider the side effects and safety issues of chronic LRRK2 kinase inhibition in a human context.
Materials and Methods
Reagents
The LRRK2in1 compound and the VPS34in1 were purchased from the Division of Signal Transduction Therapy, School of Life Sciences, and University of Dundee, UK. The GSK2578215A compound was purchased from Tocris. Bafilomycin A1 (B1793-2UG), Chloroquine (C6628) and Wortmannin (W3144-250UL) were purchased from Sigma-Aldrich. Torin-1 (CAY10997) was purchased from Cayman Chemicals.
Antibodies used were as follows: LC3 antibody (NB100-2220, Novus Biologicals); LRRK2 antibodies (MJFF#2, 3514-1/ab133474, Epitomics); total S6 antibody (2317, Cell Signalling); phospho Ser235/236 S6 antibody (2211S, Cell Signalling); total P70S6K antibody (sc-8418, Santa Cruz); phospho Thr389 P70S6K (sc-11759, Santa Cruz); total ULK1 antibody (8054 and 4773, Cell Signalling); phospho Ser757 ULK1 antibody (6888, Cell Signalling); Beclin-1 antibody (3738, Cell Signalling); β-actin antibody (A1978, Sigma Aldrich); β-tubulin (T6199, Sigma Aldrich).
H4 cell culture and treatment
H4 cells (ATCC number HTB-148) were grown in DMEM containing 10% FCS. After 24 hours from plating, when at 80% confluence, H4 cells were treated with LRRK2 inhibitors LRRK2in1 or GSK2578215A, with BafA1 or Torin-1 or wortmannin at the concentrations and for the time reported in each experiment. For each experiment, DMSO vehicle controls were added. After washing in Dulbecco’s phosphate buffered saline (DPBS) cells were collected in a lysis buffer containing: 0.5% Triton X-100, 2 mM ethylen di-ammonium tetra acetic acid (EDTA), 150 mM NaCl, 0.5% sodium deoxycholate, 0.1% sodium dodecyl sulphate (SDS), protease inhibitors (cOmplete, protease inhibitor cocktail, Roche) and phosphatase inhibitors (Halt phosphatase inhibitor cocktail, Pierce) in 50 mM TRIS-HCl pH 7.5.
Primary astrocytes culture
Primary astrocyte cultures were prepared from P3-4 mouse forebrain following a protocol adapted from Schildge et al.61. Briefly, brains were isolated in cold HBSS (Sigma-Aldrich). The forebrains were dissected and meninges removed. Forebrains were dissociated before incubation in papain (Worthington) at 37 °C for 30 min. The samples were triturated by pipetting and subsequently plated in DMEM supplemented with 10% FCS and 1% penicillin/streptomycin. At 14 divisions, microglia were dissociated by agitation for 1 hr at room temperature and astrocytes were re-plated following brief trypsinization to expand the culture. Astrocytes were aged to 45/47 divisions, plated in 6 well plates (1 × 106 cells/well), and at 80% confluency the cells were treated with LRRK2in1 or DMSO (control) as reported in figures. Following treatment, the cells were washed in DPBS, collected by scraping and cell pellets were frozen until Western blot analysis.
Western blotting
Cell lysates were frozen immediately upon collection or kept at 4 °C for 30 minutes; following thawing, they were clarified by centrifugation at 13000 rpm for 5 minutes at 4 °C, protein concentration was assessed by BCA assay (BCA Protein Assay Kit, Pierce) and 10–15 μg aliquots were added of NuPAGE sample buffer containing 2-mercaptoethanol (Invitrogen), denatured for 10 minutes at 70 °C and analysed using NuPAGE, Novex precasted Bis-Tris 4–12% (Invitrogen), in MES running buffer (Invitrogen). After electrophoresis, gels were blotted onto 0.45 μm cut-off, PVDF through conventional blotting. Membranes were blocked and processed with peroxidase-conjugated antibodies for enhanced chemiluminescence (ECL) detection. Films were acquired as images in jpg format using an EPSON Perfection 4870 photo scanner and processed by the ImageJ software ( http://rsbweb.nih.gov/ij/).
Statistics
All the results have been repeated in at least 3 independent experiments (details are given in each figure legend). In the case of Western blot analysis for phospho-proteins, samples were divided in half and run in two parallel gels; one was processed for the phospho-antibody and one for the total-antibody (to avoid membrane stripping and re-probing). The total and the phospho bands were normalized against the respective β-actin loading control; then, the ratio of (normalized)phospho over (normalized)total protein was calculated. Statistical analyses were performed by the use of the Prism software (GraphPad) as described in each figure legend.
Generation of stable knock-down
H4 cells were transfected with 2 to 10 μg LRRK2 shRNA, ULK1 shRNA, Beclin-1 shRNA or scramble Open Biosystems GIPZ shRNAmir (V3LHS-644167, V2LHS-33057, V3LHS-349512 and -349514, Thermo Fisher Scientific) using PEI (Sigma-Aldrich) transfection reagent according to the manufacturer’s instructions. ShRNA vectors contain a puromycin resistance gene. Cells were treated with 2 μg/ml puromycin supplemented DMEM 48 hrs after transfection and kept under selection for expansion. Puromycin selection was removed 24 hours before the experiment to avoid interference of the antibiotic with the treatment.
Additional Information
How to cite this article: Manzoni, C. et al. mTOR independent regulation of macroautophagy by Leucine Rich Repeat Kinase 2 via Beclin-1. Sci. Rep. 6, 35106; doi: 10.1038/srep35106 (2016).
References
Hernandez, D. G., Reed, X. & Singleton, A. B. Genetics in Parkinson disease: Mendelian versus non-Mendelian inheritance. J Neurochem, doi: 10.1111/jnc.13593 (2016).
Paisan-Ruiz, C. et al. Cloning of the gene containing mutations that cause PARK8-linked Parkinson’s disease. Neuron 44, 595–600, doi: 10.1016/j.neuron.2004.10.023 (2004).
Singleton, A. B., Farrer, M. J. & Bonifati, V. The genetics of Parkinson’s disease: progress and therapeutic implications. Mov Disord 28, 14–23, doi: 10.1002/mds.25249 (2013).
Zimprich, A. et al. Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron 44, 601–607, doi: 10.1016/j.neuron.2004.11.005 (2004).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat Genet 46, 989–993, doi: 10.1038/ng.3043 (2014).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124, doi: 10.1038/nature11582 (2012).
Fava, V. M. et al. A Missense LRRK2 Variant Is a Risk Factor for Excessive Inflammatory Responses in Leprosy. Plos Negl Trop Dis 10, e0004412, doi: 10.1371/journal.pntd.0004412 (2016).
Zhang, F. R. et al. Genomewide association study of leprosy. N Engl J Med 361, 2609–2618, doi: 10.1056/NEJMoa0903753 (2009).
Inzelberg, R. et al. The LRRK2 G2019S mutation is associated with Parkinson disease and concomitant non-skin cancers. Neurology 78, 781–786, doi: 10.1212/WNL.0b013e318249f673 (2012).
Agalliu, I. et al. Higher frequency of certain cancers in LRRK2 G2019S mutation carriers with Parkinson disease: a pooled analysis. JAMA Neurol 72, 58–65, doi: 10.1001/jamaneurol.2014.1973 (2015).
Biosa, A. et al. GTPase activity regulates kinase activity and cellular phenotypes of Parkinson’s disease-associated LRRK2. Hum Mol Genet 22, 1140–1156, doi: 10.1093/hmg/dds522 (2013).
Webber, P. J. et al. Autophosphorylation in the leucine-rich repeat kinase 2 (LRRK2) GTPase domain modifies kinase and GTP-binding activities. J Mol Biol 412, 94–110, doi: 10.1016/j.jmb.2011.07.033 (2011).
Wirth, M., Joachim, J. & Tooze, S. A. Autophagosome formation–the role of ULK1 and Beclin1-PI3KC3 complexes in setting the stage. Semin Cancer Biol 23, 301–309, doi: 10.1016/j.semcancer.2013.05.007 (2013).
Plowey, E. D., Cherra, S. J. 3rd., Liu, Y. J. & Chu, C. T. Role of autophagy in G2019S-LRRK2-associated neurite shortening in differentiated SH-SY5Y cells. J Neurochem 105, 1048–1056, doi: 10.1111/j.1471-4159.2008.05217.x (2008).
Alegre-Abarrategui, J. et al. LRRK2 regulates autophagic activity and localizes to specific membrane microdomains in a novel human genomic reporter cellular model. Hum Mol Genet 18, 4022–4034, doi: 10.1093/hmg/ddp346 (2009).
Schapansky, J., Nardozzi, J. D., Felizia, F. & LaVoie, M. J. Membrane recruitment of endogenous LRRK2 precedes its potent regulation of autophagy. Hum Mol Genet 23, 4201–4214, doi: 10.1093/hmg/ddu138 (2014).
Gomez-Suaga, P. et al. Leucine-rich repeat kinase 2 regulates autophagy through a calcium-dependent pathway involving NAADP. Hum Mol Genet 21, 511–525, doi: 10.1093/hmg/ddr481 (2012).
Manzoni, C. et al. Inhibition of LRRK2 kinase activity stimulates macroautophagy. Biochim Biophys Acta 1833, 2900–2910, doi: 10.1016/j.bbamcr.2013.07.020 (2013).
Saez-Atienzar, S. et al. The LRRK2 inhibitor GSK2578215A induces protective autophagy in SH-SY5Y cells: involvement of Drp-1-mediated mitochondrial fission and mitochondrial-derived ROS signaling. Cell Death Dis 5, e1368, doi: 10.1038/cddis.2014.320 (2014).
Bravo-San Pedro, J. M. et al. The LRRK2 G2019S mutant exacerbates basal autophagy through activation of the MEK/ERK pathway. Cell Mol Life Sci 70, 121–136, doi: 10.1007/s00018-012-1061-y (2013).
Manzoni, C. et al. Pathogenic Parkinson’s disease mutations across the functional domains of LRRK2 alter the autophagic/lysosomal response to starvation. Biochem Biophys Res Commun 441, 862–866, doi: 10.1016/j.bbrc.2013.10.159 (2013).
Sanchez-Danes, A. et al. Disease-specific phenotypes in dopamine neurons from human iPS-based models of genetic and sporadic Parkinson’s disease. EMBO Mol Med 4, 380–395, doi: 10.1002/emmm.201200215 (2012).
Tong, Y. et al. Loss of leucine-rich repeat kinase 2 causes age-dependent bi-phasic alterations of the autophagy pathway. Mol Neurodegener 7, 2, doi: 10.1186/1750-1326-7-2 (2012).
Orenstein, S. J. et al. Interplay of LRRK2 with chaperone-mediated autophagy. Nat Neurosci 16, 394–406, doi: 10.1038/nn.3350 (2013).
Arranz, A. M. et al. LRRK2 functions in synaptic vesicle endocytosis through a kinase-dependent mechanism. J Cell Sci 128, 541–552, doi: 10.1242/jcs.158196 (2015).
Belluzzi, E. et al. LRRK2 phosphorylates pre-synaptic N-ethylmaleimide sensitive fusion (NSF) protein enhancing its ATPase activity and SNARE complex disassembling rate. Mol Neurodegener 11, 1, doi: 10.1186/s13024-015-0066-z (2016).
Chia, R. et al. Phosphorylation of LRRK2 by casein kinase 1alpha regulates trans-Golgi clustering via differential interaction with ARHGEF7. Nat Commun 5, 5827, doi: 10.1038/ncomms6827 (2014).
Matta, S. et al. LRRK2 controls an EndoA phosphorylation cycle in synaptic endocytosis. Neuron 75, 1008–1021, doi: 10.1016/j.neuron.2012.08.022 (2012).
Xilouri, M., Brekk, O. R. & Stefanis, L. Autophagy and Alpha-Synuclein: Relevance to Parkinson’s Disease and Related Synucleopathies. Mov Disord 31, 178–192, doi: 10.1002/mds.26477 (2016).
Henry, A. G. et al. Pathogenic LRRK2 mutations, through increased kinase activity, produce enlarged lysosomes with reduced degradative capacity and increase ATP13A2 expression. Hum Mol Genet 24, 6013–6028, doi: 10.1093/hmg/ddv314 (2015).
Moehle, M. S. et al. LRRK2 inhibition attenuates microglial inflammatory responses. J Neurosci 32, 1602–1611, doi: 10.1523/JNEUROSCI.5601-11.2012 (2012).
Deng, X. et al. Characterization of a selective inhibitor of the Parkinson’s disease kinase LRRK2. Nat Chem Biol 7, 203–205, doi: 10.1038/nchembio.538 (2011).
Reith, A. D. et al. GSK2578215A; a potent and highly selective 2-arylmethyloxy-5-substitutent-N-arylbenzamide LRRK2 kinase inhibitor. Bioorg Med Chem Lett 22, 5625–5629, doi: 10.1016/j.bmcl.2012.06.104 (2012).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222, doi: 10.1080/15548627.2015.1100356 (2016).
Mauvezin, C., Nagy, P., Juhasz, G. & Neufeld, T. P. Autophagosome-lysosome fusion is independent of V-ATPase-mediated acidification. Nat Commun 6, 7007, doi: 10.1038/ncomms8007 (2015).
Tooze, S. A. et al. Assessing mammalian autophagy. Methods Mol Biol 1270, 155–165, doi: 10.1007/978-1-4939-2309-0_12 (2015).
Papinski, D. & Kraft, C. Regulation of Autophagy By Signaling Through the Atg1/ULK1 Complex. J Mol Biol, doi: 10.1016/j.jmb.2016.03.030 (2016).
Johnson, C. W., Melia, T. J. & Yamamoto, A. Modulating macroautophagy: a neuronal perspective. Future Med Chem 4, 1715–1731, doi: 10.4155/fmc.12.112 (2012).
Lewis, P. A. & Manzoni, C. LRRK2 and human disease: a complicated question or a question of complexes? Sci Signal 5, pe2, doi: 10.1126/scisignal.2002680 (2012).
Ktistakis, N. T. & Tooze, S. A. Digesting the Expanding Mechanisms of Autophagy. Trends Cell Biol, doi: 10.1016/j.tcb.2016.03.006 (2016).
Alers, S., Loffler, A. S., Wesselborg, S. & Stork, B. Role of AMPK-mTOR-Ulk1/2 in the regulation of autophagy: cross talk, shortcuts, and feedbacks. Mol Cell Biol 32, 2–11, doi: 10.1128/MCB.06159-11 (2012).
McAlpine, F., Williamson, L. E., Tooze, S. A. & Chan, E. Y. Regulation of nutrient-sensitive autophagy by uncoordinated 51-like kinases 1 and 2. Autophagy 9, 361–373, doi: 10.4161/auto.23066 (2013).
Codogno, P., Mehrpour, M. & Proikas-Cezanne, T. Canonical and non-canonical autophagy: variations on a common theme of self-eating? Nat Rev Mol Cell Biol 13, 7–12, doi: 10.1038/nrm3249 (2012).
Mauthe, M. et al. Resveratrol-mediated autophagy requires WIPI-1-regulated LC3 lipidation in the absence of induced phagophore formation. Autophagy 7, 1448–1461 (2011).
Wang, J. et al. A non-canonical MEK/ERK signaling pathway regulates autophagy via regulating Beclin 1. J Biol Chem 284, 21412–21424, doi: 10.1074/jbc.M109.026013 (2009).
Cheong, H., Lindsten, T., Wu, J., Lu, C. & Thompson, C. B. Ammonia-induced autophagy is independent of ULK1/ULK2 kinases. Proc Natl Acad Sci USA 108, 11121–11126, doi: 10.1073/pnas.1107969108 (2011).
Alers, S. et al. Atg13 and FIP200 act independently of Ulk1 and Ulk2 in autophagy induction. Autophagy 7, 1423–1433 (2011).
Sarkar, S. et al. Small molecules enhance autophagy and reduce toxicity in Huntington’s disease models. Nat Chem Biol 3, 331–338, doi: 10.1038/nchembio883 (2007).
Su, Y. C., Guo, X. & Qi, X. Threonine 56 phosphorylation of Bcl-2 is required for LRRK2 G2019S-induced mitochondrial depolarization and autophagy. Biochim Biophys Acta 1852, 12–21, doi: 10.1016/j.bbadis.2014.11.009 (2015).
Decuypere, J. P., Parys, J. B. & Bultynck, G. Regulation of the autophagic bcl-2/beclin 1 interaction. Cells 1, 284–312, doi: 10.3390/cells1030284 (2012).
Wei, Y. et al. The stress-responsive kinases MAPKAPK2/MAPKAPK3 activate starvation-induced autophagy through Beclin 1 phosphorylation. Elife 4, doi: 10.7554/eLife.05289 (2015).
Gloeckner, C. J., Schumacher, A., Boldt, K. & Ueffing, M. The Parkinson disease-associated protein kinase LRRK2 exhibits MAPKKK activity and phosphorylates MKK3/6 and MKK4/7, in vitro. J Neurochem 109, 959–968, doi: 10.1111/j.1471-4159.2009.06024.x (2009).
Funderburk, S. F., Wang, Q. J. & Yue, Z. The Beclin 1-VPS34 complex–at the crossroads of autophagy and beyond. Trends Cell Biol 20, 355–362, doi: 10.1016/j.tcb.2010.03.002 (2010).
Matsunaga, K. et al. Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nat Cell Biol 11, 385–396, doi: 10.1038/ncb1846 (2009).
Zhong, Y. et al. Distinct regulation of autophagic activity by Atg14L and Rubicon associated with Beclin 1-phosphatidylinositol-3-kinase complex. Nat Cell Biol 11, 468–476, doi: 10.1038/ncb1854 (2009).
Zalckvar, E., Berissi, H., Eisenstein, M. & Kimchi, A. Phosphorylation of Beclin 1 by DAP-kinase promotes autophagy by weakening its interactions with Bcl-2 and Bcl-XL. Autophagy 5, 720–722 (2009).
Zalckvar, E. et al. DAP-kinase-mediated phosphorylation on the BH3 domain of beclin 1 promotes dissociation of beclin 1 from Bcl-XL and induction of autophagy. EMBO Rep 10, 285–292, doi: 10.1038/embor.2008.246 (2009).
Liu, Z., Mobley, J. A., DeLucas, L. J., Kahn, R. A. & West, A. B. LRRK2 autophosphorylation enhances its GTPase activity. FASEB J 30, 336–347, doi: 10.1096/fj.15-277095 (2016).
Wallings, R., Manzoni, C. & Bandopadhyay, R. Cellular processes associated with LRRK2 function and dysfunction. FEBS J 282, 2806–2826, doi: 10.1111/febs.13305 (2015).
Manzoni, C., Denny, P., Lovering, R. C. & Lewis, P. A. Computational analysis of the LRRK2 interactome. PeerJ 3, e778, doi: 10.7717/peerj.778 (2015).
Schildge, S., Bohrer, C., Beck, K. & Schachtrup, C. Isolation and culture of mouse cortical astrocytes. J Vis Exp, doi: 10.3791/50079 (2013).
Acknowledgements
The authors would like to acknowledge generous research support from the Michael J. Fox Foundation, Parkinson’s UK and the Rosetrees Trust. PAL is a Parkinson’s UK research fellow (grant F1002). This work was supported in part by the Wellcome Trust/MRC Joint Call in Neurodegeneration award (WT089698) to the UK Parkinson’s Disease Consortium (UKPDC) whose members are from the UCL Institute of Neurology, the University of Sheffield and the MRC Protein Phosphorylation Unit at the University of Dundee, by a MRC New Investigator Research Grant (MR/L010933/1) to PAL, and by MRC programme grant MR/N026004/1 to JH, HPF and PAL. This research was supported in part by the Intramural Research Program of the NIH, National Institute on Aging. SAT was supported by the Francis Crick Institute which receives its core funding from Cancer Research UK (FC001187), the UK Medical Research Council (FC001187), and the Wellcome Trust (FC001187).We thank Dr. Claudio Galbiati and Dr. Miriam Manzoni at the Università degli Studi di Bergamo (IT) for the development of the image quantification script for Supplementary Figure S5. We thank the UCL Cancer Institute CAGE Facility, which provided the Open Biosystems GIPZ shRNAmir. The CAGE Facility is supported by the Welton Foundation, BRC and in part by the Cancer Research UK – UCL Centre.
Author information
Authors and Affiliations
Contributions
P.A.L. directed the research. P.A.L. and C.M. designed the experiments. C.M. performed most of the experiments. M.S. supervised the generation of knock-down cells. A.M. and D.R. prepared and were involved in the experiments with primary astrocytes. S.D. helped in the setup of experimental procedures. S.A.T., J.H., R.B., H.P.-F. and M.R.C. supported the work with guidance, advice and reagents. C.M. and P.A.L. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Manzoni, C., Mamais, A., Roosen, D. et al. mTOR independent regulation of macroautophagy by Leucine Rich Repeat Kinase 2 via Beclin-1. Sci Rep 6, 35106 (2016). https://doi.org/10.1038/srep35106
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep35106
This article is cited by
-
Macroautophagy in CNS health and disease
Nature Reviews Neuroscience (2022)
-
Exploring the focal role of LRRK2 kinase in Parkinson’s disease
Environmental Science and Pollution Research (2022)
-
Mammalian/mechanistic target of rapamycin (mTOR) complexes in neurodegeneration
Molecular Neurodegeneration (2021)
-
The Role of iPSC Modeling Toward Projection of Autophagy Pathway in Disease Pathogenesis: Leader or Follower
Stem Cell Reviews and Reports (2021)
-
LRRK2 Biology from structure to dysfunction: research progresses, but the themes remain the same
Molecular Neurodegeneration (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.