Abstract
To better understand genomic changes in the early generations after polyploidisation, we examined the chromosomal consequences of genomic merger in allotetraploid hybrids (4 nF1) (AABB, 4n = 148) of Carassius auratus red var. (RCC) (AA, 2n = 100) (♀) × Megalobrama amblycephala (BSB) (BB, 2n = 48) (♂). Complete loss of the paternal 5S rDNA sequence and the expected number of maternal chromosomal loci were found in 4 nF1, suggesting directional genomic changes occurred in the first generations after polyploidisation. Recent studies have reported instability of newly established allotetraploid genomes. To assess this in the newly formed 4 nF1 genome, we performed fluorescence in situ hybridisation on an allotetraploid gynogenetic hybrid (4 nG) (AABB, 4n = 148) and an allopentaploid hybrid (5 nH) (AABBB, 5n = 172) from 4 nF1 (♀) × BSB (♂) with 5S rDNA gene and centromere probes from RCC, the original diploid parent. The expected numbers of maternal chromosomal loci were found in 4 nG, while chromosomal locus deletions and chromosome recombinations were detected in 5 nH. These observations suggest that abnormal meiosis did not lead to obvious genomic changes in the newly established allotetraploid genomes, but hybridisation with the original diploid parent resulted in obvious genomic changes in the newly established allotetraploid genomes, as was found for the maternal genome.
Similar content being viewed by others
Introduction
Polyploidisation is a significant evolutionary process that results in rapid speciation1,2,3,4,5,6. Many diploids species are actually ancient polyploids that have undergone diploidisation. Recent studies, mostly in plants, suggested that allopolyploid formation could induce various types of genomic changes, including directional sequence elimination, random structural changes and chromosome structure7,8,9,10,11,12,13. Importantly, a variety of genomic changes has been shown to result in diploid-like chromosome pairing14,15,16,17. Although the potential contribution of genomic changes to the evolutionary success of polyploidy has been widely recognised, virtually no information is available on how newly established genomes have evolved after polyploidisation.
Because sex determination and development are disrupted by polyploidisation, polyploid species are common in plants, but rarely successful in animals18,19,20. In our previous study, fertile allotetraploid hybrids (4 nF1) (AABB, 4n = 148) were successfully obtained in the first generation of Carassius auratus red var. (RCC) (AA, 2n = 100, ♀) × Megalobrama amblycephala (BSB) (BB, 2n = 48, ♂) as a result of chromosome doubling of diploid hybrid embryos (AB, 2n = 74) by inhibition of the first cleavage21. Unexpectedly, abnormal chromosome behaviour during meiosis, but not bivalent pairing, occurred in 4 nF1 and resulted in a generation of gametes with different genetic compositions, including allotetraploid (AABB), autotriploid (AAA) and autodiploid (AA) gametes22,23. Thus, we obtained autotetraploids (AAAA, 4n = 200) in the self-cross progenies of 4 nF1 and allopentaploid hybrids (5 nH) (AABBB, 5n = 172) in backcross progenies of 4 nF1 (♀) × BSB (♂)21,24. In addition, gynogenetic allotetraploids (4 nG) (AABB, 4n = 148) were obtained by artificial gynogenesis from eggs of the 4 nF1 that were activated with UV-treated sperm of BSB, but not subject to treatment for doubling the chromosome number22. Because the progenitors for these allopolyploids are known, we can precisely determine the timing and processes of genomic changes after polyploidisation. Thus, these allopolyploids provide a model system to study early chromosomal evolution after polyploidisation.
Cytogenetics studies using fluorescence in situ hybridisation (FISH) have reported chromosomal changes in many polyploid species25,26,27,28,29. To further understand allopolyploid genome evolution in a broad context, we used the first generation 4 nF1 hybrids and their backcross progenies to explore potential genomic changes on polyploidisation. We determined the response of two dispersed chromosomal loci (5S rDNA and centromere) that may be particularly important for de novo allopolyploidy, because they are likely to be most vulnerable to genetic changes after polyploidisation and (or) hybridisation29,30,31. Our results revealed rapid genomic changes occurred in the first generations after polyploidisation and indicated instability of the newly established allotetraploid genome. The findings of this study provide new insights into chromosomal evolution in vertebrates.
Results
Organisation of 5S rDNA unit
Formation and genetic compositionof experimental fish is showed in Fig. 1. DNA fragments were amplified from BSB, RCC, 4 nF1, 4 nG and 5 nH using the 5SP1 and 5SP2R primers. Agarose gel electrophoresis identified three PCR fragments (approximately 200, 340 and 500 bp) from RCC, two PCR fragments (approximately 200 and 370 bp) from BSB and four PCR fragments (approximately 200, 340, 400 and 500 bp) from each of 4 nF1, 4 nG and 5 nH (Fig. 2).
DNA fragments amplified from RCC, BSB, 4 nF1, 4 nG and 5 nH.
M, DNA ladder markers (200-bp increments); lane 1, DNA from BSB; lane 2, DNA from RCC; lane 3, DNA from 4 nF1; lane 4, DNA from 4 nG; lane 5, DNA from 5 nH. RCC, Carassius auratus red var.; BSB, Megalobrama amblycephala; 4 nF1, allotetraploid hybrid; 4 nG, allotetraploid gynogenetic hybrid; 5 nH, allopentaploid hybrid.
A total of 340 clones were sequenced to examine the different patterns of the 5S rDNA sequences; 60 clones were from RCC, 40 clones were from BSB and 80 clones were from each of the 4 nF1, 4 nG and 5 nH hybrids (Table 1). Based on the BLASTn analysis, all the sequences from all five hybrids were confirmed as 5S rDNA repeat units. The 5S rDNA sequences from RCC fell into three distinct families (designated class I: 203 bp; class II: 340 bp; and class III: 477 bp), while the 5S rDNA sequences from BSB formed one family (designated class IV: 188 bp). All three RCC-derived families (class I, class II and class III) were detected in 4 nF1 and 4 nG, while the BSB-derived family (class IV) was not found in 4 nF1 and 4 nG. All four classes were detected in 5 nH (Table 1).
Southern blot hybridisation
Genomic DNA from RCC, BSB and 4 nF1 was digested with HindIII and ScaI. Southern hybridisation was performed using the 5S rDNA sequence from BSB as a probe. This probe hybridised with the genomic DNA from BSB, but not with the genomic DNA from RCC and 4 nF1 (Fig. 3). This result implies that the paternal 5S rDNA cluster is completely deleted in 4 nF1.
Southern blot hybridisation with the 5S rDNA sequence from BSB as a probe.
HindIII and ScaI were used to digest the genomic DNA of BSB, RCC and 4 nF1. Positive hybridisation was detected in the genomic DNA of BSB, but not in the genomic DNA of RCC and the 4 nF1 hybrid. Molecular weight makers (kb) are shown on the left. BSB, Megalobrama amblycephala; RCC, Carassius auratus red var.; 4 nF1, allotetraploid hybrid.
Fluorescence in situ hybridisation
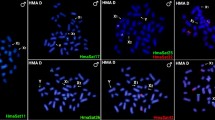
FISH hybridisation of the RCC-derived class I (203 bp, GenBank: GQ485555) 5S rDNA gene probe to the RCC and BSB metaphase chromosomes yielded eight 5S rDNA gene loci in RCC (Fig. 4A; Table 2), but none in BSB (Table 2). It was expected that the eight 5S rDNA loci will also be present in the 4 nF1, 4 nG and 5 nH hybrids because they were derived from RCC. However, we found that, while all eight 5S rDNA gene loci detected in the metaphase chromosomes of 4 nF1 and 4 nG and were similar to the eight RCC loci (Fig. 4B,C; Table 2), only five of the 5S rDNA gene loci were detected in the 5 nH metaphase chromosomes (Fig. 4D; Table 2).
FISH hybridisation signals in the metaphase chromosomes of RCC, 4 nF1, 4 nG and 5 nH with class I (203 bp) 5S rDNA as a probe.
The white arrows indicate the 5S rDNA gene loci. The eight 5S rDNA gene loci in RCC (A), 4 nF1 (B) and 4 nG (C) and the five 5S rDNA gene loci in 5 nH (D) are shown. Bars in (A–D): 3 μm. RCC, Carassius auratus red var.; 4 nF1, allotetraploid hybrid; 4 nG, allotetraploid gynogenetic hybrid; 5 nH, allopentaploid hybrid.
FISH hybridisation of the class II (340 bp, GenBank: GQ485556) 5S rDNA gene probe to the RCC and BSB metaphase chromosomes yield four 5S rDNA gene loci in RCC (Fig. 5A; Table 2), but none in BSB (Table 2). The chromosomal locus map for RCC revealed two large 5S rDNA gene loci on homologous submetacentric chromosomes and two small 5S rDNA gene loci on homologous subtelocentric chromosomes (Fig. 5A). Similarly, as expected, two large and two small 5S rDNA gene loci were found on homologous submetacentric chromosomes and homologous subtelocentric chromosomes, respectively, in 4 nF1 (Fig. 5B) and 4 nG (Fig. 5C). Unexpectedly, in 5 nH, one large 5S rDNA gene locus was located on a submetacentric chromosome and another was located on a metacentric chromosome (Fig. 5D), suggesting that these two large 5S rDNA gene loci were not located on homologous chromosomes.
FISH hybridisation signals in the metaphase chromosomes of RCC, 4 nF1, 4 nG and 5 nH with class II (340 bp) 5S rDNA as a probe.
The red arrows indicate the large 5S rDNA gene loci and the green arrows indicate the small 5S rDNA gene loci. The two big and two small 5S rDNA gene loci in RCC (A), 4nF1 (B) and 4 nG (C) are shown. A1 in (A) B1 in (B) and C1 in (C) indicate that the large 5S rDNA gene loci were located on homologous submetacentric chromosomes. A2 in (A) B2 in (B) and C2 in (C) indicate that the small 5S rDNA gene loci were located on homologous subtelocentric chromosomes. (D) The two large 5S rDNA gene loci (red and yellow arrows) and two small 5S rDNA gene loci (green arrows) in 5 nH are shown. D1 indicates that the large 5S rDNA gene loci were located on a submetacentric chromosome (red) and a metacentric chromosome (yellow). D2 indicates that the small 5S gene loci were located on homologous subtelocentric chromosomes; Bars in (A–D) 3 μm. RCC, Carassius auratus red var.; 4 nF1, allotetraploid hybrid; 4 nG, allotetraploid gynogenetic hybrid; 5 nH, allopentaploid hybrid.
FISH hybridisation of the class III (477 bp, GenBank: GQ485557) 5S rDNA gene probe to the RCC and BSB metaphase chromosomes yield eight 5S gene loci in RCC (Fig. 6A; Table 2), but none in BSB (Table 2). As expected, the eight RCC-derived 5S rDNA gene loci were detected in the metaphase chromosomes of 4 nF1, 4 nG and 5 nH and were similar to the loci in RCC (Fig. 6B–D; Table 2).
FISH hybridisation signals in the metaphase chromosomes of RCC, 4 nF1, 4 nG and 5 nH with class III (477 bp) 5S rDNA as a probe.
The white arrows indicate the 5S rDNA gene loci. The eight 5S gene loci in RCC (A), 4 nF1 (B), 4 nG (C) and 5 nH (D) are shown. Bars in (A–D): 3 μm. RCC, Carassius auratus red var.; 4 nF1, allotetraploid hybrid; 4 nG, allotetraploid gynogenetic hybrid; 5 nH, allopentaploid hybrid.
The RCC-derived centromere probe (GenBank: JQ086761) hybridised to 100 chromosomes in RCC (Fig. 7D), but none in BSB (Table 2). This species-specific centromere probe also hybridised to 100 metaphase chromosomes in 4 nF1 and 4 nG, as expected, but only about 65 to 70 RCC-derived centromere loci were detected in 5 nH metaphase chromosomes, rather than the expected 100 loci.
FISH hybridisation signals in the metaphase chromosomes of RCC, 4 nF1, 4 nG and 5 nH with the centromere probe.
The centromere probe hybridised to 100 chromosomes in RCC (A), 4 nF1 (B) and 4 nG (C) and to 69 chromosomes in 5 nH (D). Bars in (A–D): 3 μm. RCC, Carassius auratus red var.; 4 nF1, allotetraploid hybrid; 4 nG, allotetraploid gynogenetic hybrid; 5 nH, allopentaploid hybrid.
Discussion
Polyploidisation may increase genomic variation rates and is important for the formation in new polyploid species4. Evidence for genomic variations, including fragment loss, chromosomal rearrangement and rDNA loci changes have been reported in both synthesised polyploid7,27,32 and natural polyploid species25,26. Genomic variations usually occur in the early generations after polyploidisation, possibly reflecting instability in newly established polyploid genomes26,27. The results of the present study support previous observations that genomic changes occur in newly established polyploid genomes and reveal that these changes can begin as early as the first generation after polyploidisation.
Because of incompatibility between homeologous chromosomes, hybridization can boost genomic change33,34. The frequency of genomic change has been associated with divergence of the diploid parental genomes7. The 4 nF1 hybrid was formed by combining the two diploid genomes from RCC and BSB, two fish species in the family Cyprinidae, that belonged to different subfamilies (Cyprininae and Cultrinae)21, implying RCC and BSB are genetically distinct. In 4 nF1, not only was the paternal 5S rDNA unit deleted entirely, but so were the paternal sox21,35 and hox (unpublished data) gene families, suggesting that a large number of genomic changes had occurred in the newly established allotetraploid genome. A variety of genomic changes can result in diploid-like chromosome pairing, which has been reported to prevent meiotic irregularities and improve the efficiency of gamete production in polyploid species16,36. However, there is still no direct evidence that large numbers of genomic variations or unstable individuals are selected for during the establishment of polyploid species27,37,38,39. In previous study, we found that diploid-like chromosome pairing was not restored in 4 nF122,23. We speculated that mass deletion of paternal genetic material gave rise to excessive genomic modification in 4 nF1, which prevented diploid-like chromosome pairing and resulted in weak fertility and the generation of gametes with a different genetic composition. To avoid extinction, the unstable 4 nF1 individuals may have entered a novel evolutionary trajectory by abnormal meiosis and produced diploid gamete with two sets of RCC-derived chromosomes. Thus, unexpectedly, we obtained better fertile autotetraploids among the progenies of 4 nF1 and successfully established an autotetraploid fish line (F2–F9)24.
In some cases, it has been shown that hybridization had more effect on the change in genomic and gene expression than polyploidization40,41. In our study, 4nG result from genome doubling of germ cell, 5 nH was obtained by hybridization of 4nF1 (♀) × BSB. Thus, the 5S rDNA units and chromosomal loci (5S rDNA and centromere) remained intact in the 4nG genome, but obvious variations were found in 5 nH. In addition, our data also revealed the elimination of the entire paternal 5S rDNA unit and stabilisation of the maternal 5S rDNA units and chromosomal loci in the allotetraploid hybrids, implying that the paternal genome underwent greater polyploidisation-associated modifications than the maternal genome. Similar findings have been reported in polyploid plants30,42,43. The nucleo-cytoplasmic hypothesis might be an explanation for the apparent paternal genome lability. This hypothesis predicts that the paternal genome of a newly formed allopolyploid evolves most rapidly because the maternal cytoplasmic background leads to paternal genome instabilities7. However in 5 nH, the newly established maternal allotetraploid genome showed obvious variations in chromosomal loci, while the parental 5S rDNA units remained intact, suggesting that the maternal genome was more unstable than the parental genome. These results are opposite to those predicted by the nucleo-cytoplasmic hypothesis. We speculate that the genetic variations in the maternal chromosomal loci may be attributed to instability of the newly established allotetraploid genome. Further, the newly established allotetraploid genome consists of the BSB-derived genome, which may hinder or reduce the influence of the cytoplasmic background on the instability of the BSB-derived paternal genome.
Methods
Animals and crosses
All experiments, performed from 2012–2015, were approved by the Animal Care Committee of Hunan Normal University. The Administration of Affairs Concerning Animal Experimentation guidelines stated approval from the Science and Technology Bureau of China. The methods were carried out in accordance with the approved guidelines. Experimental individuals were fed in a pool with suitable illumination, water temperature, dissolved oxygen content and adequate forage in the Engineering Center of Polyploidy Fish Breeding of the National Education Ministry located at Hunan Normal University, China. Approval from the Department of Wildlife Administration is not required for the experiments conducted in this paper. Fish were deeply anesthetized with 100 mg/L MS-222 (Sigma-Aldrich) before dissection.
The 4nF1 hybrids (AABB, 4n = 148) of RCC (AA, 2n = 100, ♀) × BSB (BB, 2n = 48, ♂) were produced during the reproductive seasons (April to June) in 2012, 2013 and 2014. During the reproductive season of 2014, the gynogenetic allotetraploid hybrids (4 nG) (AABB, 4n = 148) were obtained by artificial gynogenesis from the eggs of the 4 nF1 that were activated with UV-treated sperm of BSB without treatment for doubling the chromosomes. During the reproductive season of 2015, 5 nH (AABBB, 5n = 172) was obtained in backcross progenies of 4nF1 (♀) × BSB (♂).
PCR amplification and sequencing
One pair of primers (5SP1: 5′-GCTATGCCCGATCTCGT CTGA-3′ and 5SP2R: 5′-CAGGTTGGTATGGCCGTAAGC-3′) was designed and synthesised to amplify the 5S rDNA repeats directly from genomic DNA by PCR. The PCR reactions and sequencing were performed as described by Qin et al.44. Sequences were analysed using ClustalW software (http://www.ebi.ac.uk/Tools/msa/clustalw2/).
Southern blot hybridisation
Genomic DNA (10 mg) from all the samples from RCC, BSB and 4nF1 was completely digested with the restriction endonucleases HindIII and ScaI, submitted to 0.8% agarose gel electrophoresis and transferred onto Hybond-N1 membrane45. The 5S rDNA sequences were labelled with Dig-11-dUTP (Roche), which was used as a probe and hybridised with the filter-immobilised DNA. Hybrid signal detection was performed with a DIG detection kit II (Innogent, China).
Fluorescence in situ hybridisation
Chromosome preparation was carried out on the kidney tissues of all samples, according to the procedures reported by Liu et al.21 The FISH probes for the 5S gene and species-specific centromere were amplified by PCR using 5SP1 and 5SP2R primer and the primer 5′-TTCGAAAAGAGAGAATAATCTA-3′ and 5′-AACTCGTCTAAACCCGAACTA-3′, respectively. The FISH probes were produced by Dig-11-dUTP labelling (using a nick translation kit; Roche, Germany) of the purified PCR products. FISH was performed according to the method described by He et al.46 For each type of fish hybrid, 200 metaphase spreads (20 metaphase spreads in each sample) of the chromosomes were analysed.
Additional Information
How to cite this article: Qin, Q. et al. Rapid genomic changes in allopolyploids of Carassius auratus red var. (♀) × Megalobrama amblycephala (♂). Sci. Rep. 6, 34417; doi: 10.1038/srep34417 (2016).
References
Masterson, J. Stomatal size in fossil plants: evidence for polyploidy in majority of angiosperms. Science 264, 421–424 (1994).
Comai, L. The advantages and disadvantages of being polyploid. Nature reviews Genetics 6, 836–846 (2005).
Rieseberg, L. H. & Willis, J. H. Plant speciation. Science 317, 910–914 (2007).
Otto, S. P. The evolutionary consequences of polyploidy. Cell 131, 452–462 (2007).
Mallet, J. Hybrid speciation. Nature 446, 279–283 (2007).
Wood, T. E. et al. The frequency of polyploid speciation in vascular plants. Proc Natl Acad Sci USA 106, 13875–13879 (2009).
Song, K. M. et al. Rapid genome change in synthetic polyploids of Brassica and its implications for polyploid evolution. Proc Natl Acad Sci USA 92, 7719–7723 (1995).
Chen, Z. J. & Pikaard, C. S. Transcriptional analysis of nucleolar dominance in polyploid plants: Biased expression/silencing of progenitor rRNA genes is developmentally regulated in Brassica. Proc Natl Acad Sci USA 94, 3442–3447 (1997).
Ozkan, H. et al. Allopolyploidy-induced rapid genome evolution in the wheat (Aegilops-Triticum) group. Plant Cell 13, 1735–1747 (2001).
Kashkush, K. et al. Transcriptional activation of retrotransposons alters the expression of adjacent genes in wheat. Nat Genet 33, 102–106 (2003).
Tate, J. A. et al. Evolution and expression of homeologous loci in Tragopogon miscellus (Asteraceae), a recent and reciprocally formed allopolyploid. Genetics 173, 1599–1611 (2006).
Leitch, A. R. & Leitch, I. J. Genomic plasticity and the diversity of polyploid plants. Science 320, 481–483 (2008).
Ha, M. et al. Small RNAs serve as a genetic buffer against genomic shock in Arabidopsis interspecific hybrids and allopolyploids. Proc Natl Acad Sci USA 106, 17835–17840 (2009).
Wendel, J. F. Genome evolution in polyploids. Plant Mol Biol 42, 225–249 (2000).
Wolfe, K. H. Yesterday’s polyploids and the mystery of diploidization. Nat Rev Genet 2, 333–341 (2001).
Paterson, A. et al. Ancient polyploidization predating divergence of the cereals and its consequences for comparative genomics. Proc Natl Acad Sci USA 101, 9903–9908 (2004).
Soltis, D. E. et al. Polyploidy and angiosperm diversification. Am J Bot 96, 336–348 (2009).
Luo, J. et al. Tempo and mode of recurrent polyploidization in the Carassius auratus species complex (Cypriniformes, Cyprinidae). Heredity 112, 415–427 (2014).
Li, X. Y. et al. Extra Microchromosomes Play Male Determination Role in Polyploid Gibel Carp. Genetics 203, 1415–1424 (2016).
Mei, J. & Gui, J. F. Genetic basis and biotechnological manipulation of sexual dimorphism and sex determination in fish. Sci China Life Sci. 58, 124–136 (2015).
Liu, S. J. et al. The formation of the polyploid hybrids from different subfamily fish crossing and its evolutionary significance. Genetics 176, 1023–1034 (2007).
Qin, Q. B. et al. The abnormal chromosome behavior during meiosis was revealed in allotetraploid of Carassius auratus red var. (♀) × Megalobrama amblycephala (♂). BMC genetics 15, 95 (2014).
Qin, Q. B. et al. Induced All-Female Autotriploidy in the Allotetraploids of Carassius auratus red var. (♀) × Megalobrama amblycephala (♂). Marine Biotechnology 17, 604–612 (2015).
Qin, Q. B. et al. The autotetraploid fish derived from hybridization of Carassius auratus red var. (female) × Megalobrama amblycephala (male). Biology of reproduction 91, 93, 1–11 (2014).
Pontes, O. et al. Chromosomal locus rearrangements are a rapid response to formation of the allotetraploid Arabidopsis suecica genome. Proc Natl Acad Sci USA 101, 18240–18245 (2004).
Lim, K. Y. et al. Rapid chromosome evolution in recently formed polyploids in Tragopogon (Asteraceae). PLoS ONE 3, e3353 (2008).
Xiong, Z. Y. et al. Homoeologous shuffling and chromosome compensation maintain genome balance in resynthesized allopolyploid Brassica napus. Proc Natl Acad Sci USA 108, 7908–7913 (2011).
Wang, Z. W. et al. A novel nucleo-cytoplasmic hybrid clone formed via androgenesis in polyploid gibel carp. BMC Res Notes. 4(82), 1–13 (2011).
Zhang, J. et al. Meiosis completion and various sperm responses lead to unisexual and sexual reproduction modes in one clone of polyploid Carassius gibelio. Scientific reports. 5, 10898 (2015).
Skalická, K. et al. Preferential elimination of repeated DNA sequences from the paternal, Nicotiana tomentosiformis genome donor of a synthetic, allotetraploid tobacco. The New Phytologist 166, 291–303 (2005).
Qin, Q. B. et al. Organization and Variation Analysis of 5S rDNA in gynogenetic offspring of Carassius auratus red var. (female) ×Megalobrama amblycephala (male). BMC Genetics 16, 26 (2015b).
Gaeta, R. T. et al. Genomic changes in resynthesized Brassica napus and their effect on gene expression and phenotype. Plant Cell 19, 3403–3417 (2007).
Buerkle, C. A. et al. The likelihood of homoploid hybrid speciation. Heredity 84, 441–451 (2000).
Rieseberg, L. H. et al. Major ecological transitions in wild sunflowers facilitated by hybridization. Science 301, 1211–1216 (2003).
Chen, L. et al. Novel genetic markers derived from the DNA fragments of Sox genes. Molecular and Cellular Probes 23, 157–165 (2009).
Comai, L. et al. Phenotypic instability and rapid gene silencing in newly formed Arabidopsis allotetraploids. Plant Cell 12, 1551–1567 (2000).
Howell, E. C. et al. A and C genome distinction and chromosome identification in Brassica napus by sequential fluorescence in situ hybridization and genomic in situ hybridization. Genetics 180, 1849–1857 (2008).
Cheung, F. et al. Comparative analysis between homoeologous genome segments of Brassica napus and its progenitor species reveals extensive sequencelevel divergence. Plant Cell 21, 1912–1928 (2009).
Xiong, Z. Y. & Pires, J. C. Karyotype and identification of all homoeologous chromosomes of allopolyploid Brassica napus and its diploid progenitors. Genetics 187, 37–49 (2011).
Albertin, W. et al. Numerous and rapid nonstochastic modifications of gene products in newly synthesized Brassica napus allotetraploids. Genetics 173, 1101icsti (2006).
Salmon, A. et al. Genetic and epigenetic consequences of recent hybridization and polyploidy in Spartina (Poaceae). Mol. Ecol. 14, 1163 (2005).
Lim, K. Y. et al. Molecular cytogenetic analyses and phylogenetic studies in the Nicotiana section Tomentosae. Chromosoma 109, 245–258 (2000).
Koukalova, B. et al. Fall and rise of satellite repeats in allopolyploids of Nicotiana over c. 5 million years. New Phytologist 186, 148–160 (2010).
Qin, Q. B. et al. Analysis of 5S rDNA organization and variation in polyploid hybrids from crosses of different fish subfamilies. Journal of Experimental Zoology Part B: Molecular and Developmental Evolution 314, 403–411 (2010).
Southern, E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98, 503–517 (1975).
He, W. G. et al. Organization and variation analysis of 5S rDNA in different ploidy-level hybrids of red crucian carp × topmouth culter. PLoS ONE 7, e38976 (2012).
Acknowledgements
This work was supported by the Major international cooperation projects of the National Natural Science Foundation of China (Grant No. 31210103918), Key Item of National Natural Science Foundation of China (Grant No. 31430088), Training Program of the Major Research Plan of the National Natural Science Foundation of China (Grant No. 91331105), the National Key Basic Research Program of China (Grant No. 2012CB722305), the Doctoral Fund of Ministry of Education of China (Grant No. 20114306130001), the National High Technology Research and Development Program of China (Grant No. 2011 AA100403), the Cooperative Innovation Center of Engineering and New Products for Developmental Biology of Hunan Province (20134486) and the construct program of the key discipline in Hunan province and China.
Author information
Authors and Affiliations
Contributions
Q.Q. carried out analyses and wrote the manuscript. S.L. contributed to the conception and design of the study. Z.L., L.C., Q.X. and Y.D.W. prepared figures. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Qin, Q., Lai, Z., Cao, L. et al. Rapid genomic changes in allopolyploids of Carassius auratus red var. (♀) × Megalobrama amblycephala (♂). Sci Rep 6, 34417 (2016). https://doi.org/10.1038/srep34417
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34417
This article is cited by
-
Comparative analyses of the Sox9a-Amh-Cyp19a1a regulatory Cascade in Autotetraploid fish and its diploid parent
BMC Genetics (2020)
-
Rapid Genomic and Epigenetic Alterations in Gynogenetic Carassius auratus Red Var. Derived from Distant Hybridization
Marine Biotechnology (2020)
-
Rapid Genomic and Genetic Changes in the First Generation of Autotetraploid Lineages Derived from Distant Hybridization of Carassius auratus Red Var. (♀) × Megalobrama amblycephala (♂)
Marine Biotechnology (2019)
-
FISH-based mitotic and meiotic diakinesis karyotypes of Morus notabilis reveal a chromosomal fusion-fission cycle between mitotic and meiotic phases
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.