Abstract
Brown planthopper (BPH) is a phloem sap-sucking insect pest of rice which causes severe yield loss. We cloned the BPH18 gene from the BPH-resistant introgression line derived from the wild rice species Oryza australiensis. Map-based cloning and complementation test revealed that the BPH18 encodes CC-NBS-NBS-LRR protein. BPH18 has two NBS domains, unlike the typical NBS-LRR proteins. The BPH18 promoter::GUS transgenic plants exhibited strong GUS expression in the vascular bundles of the leaf sheath, especially in phloem cells where the BPH attacks. The BPH18 proteins were widely localized to the endo-membranes in a cell, including the endoplasmic reticulum, Golgi apparatus, trans-Golgi network, and prevacuolar compartments, suggesting that BPH18 may recognize the BPH invasion at endo-membranes in phloem cells. Whole genome sequencing of the near-isogenic lines (NILs), NIL-BPH18 and NIL-BPH26, revealed that BPH18 located at the same locus of BPH26. However, these two genes have remarkable sequence differences and the independent NILs showed differential BPH resistance with different expression patterns of plant defense-related genes, indicating that BPH18 and BPH26 are functionally different alleles. These findings would facilitate elucidation of the molecular mechanism of BPH resistance and the identified novel alleles to fast track breeding BPH resistant rice cultivars.
Similar content being viewed by others
Introduction
The brown planthopper (BPH), Nilaparvata lugens Stål (Homoptera:Delphacidae) is the most destructive insect pest affecting rice plants in many rice growing countries besides rice stem borers that affect rice production in some regions under favorable climatic conditions. It is a phloem sap–sucking insect, making it a vector for the transmission of viral diseases such as ragged stunt and grassy stunt viruses1. Heavy BPH infestation causes serious damage to rice crop as shown by symptoms of complete drying and mortality known as “hopper burn”. In recent years, infestations of BPH have intensified in many countries as BPH developed the ability to attack resistant plants and gained resistance to widely used pesticides. Previous studies showed that host-plant resistance is an effective, environment-friendly approach to reducing BPH damage and increasing yield potential. To date, 30 BPH resistance loci have been reported from cultivated rice germplasm and as well from five wild Oryza species sources2,3. Among these, the Bph3, Bph14, BPH26 and BPH29 genes have been identified by map-based gene cloning. The Bph3 locus was revealed to be a cluster of three genes encoding plasma membrane-localized lectin receptor kinases (OsLecRK1-OsLecRK3)4. The Bph14 gene encodes a protein containing a coiled-coil nucleotide-binding site (CC-NBS) and a leucine-rich repeat (LRR) motif, and mediates a resistance mechanism similar to the defense mechanism against pathogens through the activation of the salicylic acid (SA)-dependent pathway5. The BPH26 gene also encodes a CC-NBS-LRR protein, and mediated sucking inhibition in the phloem sieve element6. The BPH29 encoding B3 DNA-binding domain confers BPH resistance through activation of SA pathway7.
The mechanism of host resistance to a broad range of BPH populations is still elusive. Innate immune response plays a critical role in the survival of plants against pathogens or insects. Plants have developed two strategies of immunity against attack of pathogens: pathogen-associated molecular patterns (PAMPs)-triggered immunity (PTI) and effector-triggered immunity (ETI)8. On the external face of the host cell, conserved microbial elicitors called PAMPs are recognized by receptor proteins, which trigger PTI. Pathogens evolve to suppress PTI by secreting virulence molecules called effectors into the host cell. The recognition of effector proteins by resistance (R) proteins induces ETI. Receptor kinases and a set of NBS-LRR proteins are involved in recognizing PAMPs or effectors and turning on the host-resistance pathways. In rice, most of the cloned R genes encode CC-NBS-LRR type proteins or receptor kinases4,5,6,8,9,10,11. Several effector genes in rice pathogens of blast and bacterial blight have been revealed12.
Plants also have developed elaborated protection systems against herbivore attack. The herbivore-associated molecular patterns (HAMPs) or the herbivore associated elicitors (HAEs) are recognized by plant cells, which triggers signal transduction pathways that connect herbivore-specific elicitors to the expression of suitable defense genes13. Various elicitors in the insects’ oral secretions have been discovered and have been well reviewed by Wu and Baldwin14. Recently, candidate effectors which appear to elicit plant defenses or promote plant-insect interactions have been reported15,16. It was proposed that HAMP-triggered immunity (HTI) and ETI are also applicable to plant-insect interactions15.
Phloem-feeding insects (PFIs) such as planthoppers, aphids, and whiteflies have specialized mouthparts and stylets that navigate through the apoplastic space of different cell layers, allowing them to reach phloem cells, puncture and ingest the sap. PFIs initially secrete sheath saliva, which is hypothesized to form a protective layer around stylets, and watery saliva during probing and feeding, which is thought to be involved in modulating the host-cell process16. Several genes conferring resistance to PFIs have been identified (tomato Mi-1 encoding CC-NBS-LRR protein17, melon Vat encoding CC-NBS-LRR protein18 as well as the above four rice BPH resistance genes4,5,6,7).
It is imperative to identify more BPH-resistance genes to elucidate their interactions for understanding the mechanism of resistance toward the development of durable broad-spectrum BPH-resistant varieties. In rice breeding programs, BPH18 was utilized to breed a BPH resistant variety in japonica background through marker-assisted selection. The variety, Anmi, harboring BPH18 showed BPH resistance at the seedling as well as at adult stages in Korea19. In this study, we report that the BPH18 gene, a unique resistance gene derived from a distantly related wild Oryza species (O. australiensis)1, encodes a novel type of CC-NBS-NBS-LRR protein and confers BPH resistance.
Results
Map-based cloning and complementation test revealed that BPH18 encodes a CC-NBS-NBS-LRR protein
In our previous study, BPH18 was identified in an introgression line IR65482-7-216-1-2 (designated as IR65482 hereinafter), inheriting the gene from the wild species O. australiensis1. BPH18 was mapped in an 843-kb interval between the markers R10289S and RM6869 and completely co-segregated with marker 7312.T4A on the long arm of chromosome 121. For the fine-mapping of BPH18, we planted 3,100 BC4F2 plants derived from the Junam/IR65482 cross and genotyped all the plants with two markers, BN45 and BN52, which were developed at about 450-kb distance forward and backward of the marker 7312.T4A. We identified 149 plants that had recombinant genotype in the BN45 - BN52 interval. The BC4F3 progenies from the selected 149 BC3F2 plants were genotyped again with BN45 and BN52 to select plants that have homozygous recombinant genotype in this interval. Thus, 130 BC4F3 plants were selected and their seeds were harvested. The BC4F4 plants from these selected homozygous recombinant 130 BC4F3 plants were subjected to BPH bioassay, and the selected BC4F3 lines were genotyped with additional markers in the target region. These analyses revealed that the BPH18 was delimited to a 27-kb region based on the Nipponbare genome sequence flanked by the markers BIM3 and BN162 (Fig. 1a). In this region, four genes were annotated in the Rice Genome Annotation Project database (http://rice.plantbiology.msu.edu/): LOC_Os12g37280 annotated as an LRR protein gene, LOC_Os12g37290 annotated as a resistance protein gene containing the NBS domain, and LOC_Os12g37300 and LOC_Os12g37310 annotated as retrotransposon genes. Sequencing the region from IR65482 and Junam revealed that a 14-kb region including the two retrotransposon genes were absent in IR65482. The remaining region contained the LOC_Os12g37280 and the LOC_Os12g37290 in IR65482 (Fig. 1a).
(a) Fine mapping of BPH18. Numbers under the linkage map indicate the number of recombinants detected between the markers at the BPH18 locus. Gene models annotated in the Rice Genome Annotation Project database (http://rice.plantbiology.msu.edu/, RGAP 7 version) were shown as black filled arrows and the actual BPH18 gene was shown with a red arrow. The grey parallelogram indicates the genomic region which is absent in the resistance donor line, IR65482, compared with Junam. NF and NR represent primers for testing a combined gene model. (b) Identification of the BPH18 full-length cDNA from the resistant line. RT-PCR was conducted with NF and NR primers. And RACE PCRs were done to determine 5′ and 3′ ends of cDNA. Finally 3,934 bp of the BPH18 full-length cDNA was obtained by PCR with NFCF and NFCR primers. M: DNA size marker, IR: IR65482. (c) The BPH18 genomic structure and the BPH18 protein structures of the resistance donor line. The filled-gray boxes indicate the protein coding region and the blank boxes indicate 5′ and 3′ un-translated regions (UTRs). The deduced BPH18 protein consists of 1,226 amino acids and has a CC domain, two NBS domains, and an LRR domain.
Between the two genes, we set the LOC_Os12g37290 encoding the NBS domain protein as the candidate gene for BPH18 and did complementation experiment. The 6.4-kb genomic region of LOC_Os12g37290, including promoter and terminator from the resistant line (Supplementary Fig. S1), was transferred into the susceptible japonica variety, Ilmi. The deduced protein of LOC_Os12g37290 had the NBS domain (Supplementary Fig. S1), but did not carry the LRR domain that is present in most NBS-LRR R proteins at the C-terminal regions20. However, the transgenic plants did not show enhanced resistance (Supplementary Fig. S2). Therefore, we speculated that LOC_Os12g37290 encoding the NBS domain and LOC_Os12g37280 encoding the LRR domain will form one gene to encode the usual NBS-LRR protein. To test this, reverse transcription PCR (RT-PCR) was conducted with a forward primer (NF) in LOC_Os12g37290 and a reverse primer (NR) in LOC_Os12g37280. This resulted in a clear PCR product (Fig. 1b), suggesting that two ORFs predicted is due to a false annotation. The full-length cDNA of the combined gene model was identified by 5′ RACE and 3′ RACE PCR (Fig. 1b). The gene consists of three exons encoding a protein of 1,226 amino acids with a CC motif, two NBS domains, and a LRR motif (Fig. 1c, Supplementary Fig. S3). We further examined the gene encoding CC-NBS-NBS-LRR, its ORF from IR65482 was isolated and placed between its own promoter and terminator regions (Supplementary Fig. S4a) and introduced into a susceptible cultivar Dongjin. Transgenic plants that expressed the introduced genes showed enhanced BPH resistance compared to the parental line (Fig. 2a,c,e), indicating that the gene encoding CC-NBS-NBS-LRR protein confers BPH resistance. This was confirmed by the introduction of the 14-kb genomic fragment covering the entire gene from IR65482 (Supplementary Fig. S4b). The transgenic lines also showed enhanced resistance to BPH (Fig. 2b,d,f). Additional functional evidence was provided by generating BPH18-RNAi transgenic plants in the resistant introgression line. Of the six RNAi lines, five lines showing a suppressed expression of BPH18 displayed significantly reduced BPH resistance (Supplementary Fig. S5). These data conclude that the gene encoding CC-NBS-NBS-LRR protein is BPH18 gene and is responsible for BPH resistance.
(a,b) BPH bioassay of the BPH18 transgenic lines (T1 generation). IR65482, resistant parental line; Dongjin, susceptible background variety; TC1-7, transgenic lines harboring the fusion gene of BPH18 promoter::BPH18 ORF::BPH18 terminator; TG1-7, transgenic lines harboring the full BPH18 genomic region (14 kb), including its promoter and terminator. (c,d) BPH resistance scores of the BPH18-transgenic lines at the seedling stage. Lower scores indicate a higher resistance to the insect. The BPH resistance score of the rice seedlings was evaluated according to a method described by Huang et al.53. Data are means ± standard deviation. (e,f) RT-PCR analysis of BPH18 in the transgenic T0 lines. The BPH18 primer pair flanked the whole ORF (3,681bp). Ubiquitin (Ubq) gene was used as an internal control.
The sequence comparison of BPH18 between the resistant donor line and the susceptible variety (Junam) revealed that the susceptible allele lacked the last part of the second NBS domain and the whole LRR domain due to premature stop codon in the beginning of the third exon (Supplementary Fig. S6). The absence of the conserved domain of BPH18 may make Junam susceptible to BPH.
BPH18 is close to the BPH26 and its first NBS domain is partial
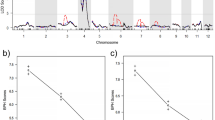
Phylogenetic analysis of BPH18 with previously identified rice NBS-LRR R proteins based on the NBS domain and LRR domain sequences showed that BPH18 is closest to BPH26 cloned from chromosome 12, and they exhibit highest similarity to the Pib protein conferring rice blast resistance located on chromosome 221 (Fig. 3a,b). These three proteins have two NBS domains unlike other typical NBS-LRR proteins. Bph14, which was the first identified BPH-resistance protein in rice5, was much farther related with BPH18 and BPH26 (Fig. 3a,b). Among these R proteins, LRR domain sequences are much more divergent than NBS domain sequences (The sum of branch length of the phylogenetic tree based on NBS domain sequences was 14.612 while that based on LRR domain sequences was 23.078).
(a) Phylogenetic tree based on NBS domain sequences. The sum of branch length was 14.612. (b) Phylogenetic tree based on LRR domain sequences. The sum of branch length was 23.078. Phylogenetic relationship was reconstructed using neighbor-joining distance method. Node supports are given in percentage of 1000 bootstrap replicates. Branch lengths are proportional to phylogenetic distances estimated from JTT amino acid substitution model.
A typical NBS domain of R proteins contains three sub-domains; a core NB (nucleotide binding), and two ARC sub-domains22,23. On the other hand, the second NBS regions of BPH18, BPH26 and Pib have all three conserved sub-domains, with the first NBS regions carrying only the NB sub-domain and lacking a portion of ARC1 and entire ARC2 (Supplementary Fig. S7). The first NBS domains of BPH18 and BPH26 have P-loop, RNBS-A, kinase-2, RNBS-B, and RNBS-C motifs but lacked GLPL, RNBS-D, and MHD motifs, which are in the ARC sub-domains24. The P-loop motif of the first NBS domain is much different from the consensus sequences. While the consensus sequence is GMGGIGKTT, that of BPH18 and BPH26 is GTSGDIREMS. Considering that the P-loop is the most-conserved motif in the NBS domains and that the lysine (K) and threonine (T) residues within the domain bind to ATP and a Mg2+ ion22, the first NBS domain is likely nonfunctional or has evolved to a diversified function.
The BPH18 expression pattern was consistent with BPH insect feeding site
We investigated the expression pattern of BPH18 using quantitative real-time PCR (qRT-PCR) and found that its transcript levels mainly found in leaf sheathes and weakly detected in leaf blades and roots (Fig. 4a). BPH18 was expressed before and after BPH infestation, indicating that it is expressed constitutively (Fig. 4b). To study a detailed expression pattern of the gene, we produced the BPH18 promoter-GUS transgenic plants. Histochemical analysis revealed that strong GUS activities were detected in the vascular bundles of the leaf sheath (Fig. 4c), where a BPH’s stylet targets.
(a) Expression of BPH18 in the leaf sheath, leaf blade and root. The mean was calculated based on the average of four biological repeats. The expression level in the samples was quantified relative to the first replicate of time 0. (b) Time-course expression of BPH18 in the leaf sheath before and after BPH infestation. The 0 means the time point just before BPH infestation. The mean was calculated based on the average of five biological repeats. The expression level in the samples was quantified relative to the first replicate of root. (c–e) GUS expression driven by the BPH18 promoter. Relatively strong GUS activity was detected in the leaf sheath (c). The leaf sheath was cross-sectioned (d). The rectangle region in Fig. 4d was magnified (e). Ph, phloem; Xy, xylem.
BPH18 localized to endo-membranes
The subcellular localization of BPH18 was investigated through its transient expression fused with Green fluorescent protein (GFP) or Red fluorescent protein (RFP) in rice protoplasts. BPH18:GFP and BPH18:RFP showed both reticular and punctate patterns in contrast to free GFP and RFP patterns which were localized to cytosol (Supplementary Fig. S8). To investigate the localization of BPH18 in detail, we co-expressed BPH18:GFP with an endoplasmic reticulum (ER) marker (Bip:RFP)25, cis-Golgi marker (ManI:RFP)26, and trans-Golgi network (TGN) marker (N-ST:RFP)27. We also co-expressed BPH18:RFP and prevacuolar compartments (PVC) marker (GFP:SYP21)28. The BPH18 fluorescence signals were co-localized partly with the ER, Golgi, TGN, and PVC markers (Fig. 5a–d), suggesting that the BPH18 protein is widely distributed to various endo-membranes, including ER, Golgi, TGN, and PVC.
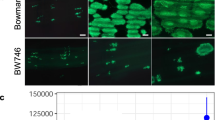
BPH18 proteins co-expressed with endoplasmic reticulum maker (Bip:RFP) (a), cis-Golgi marker (ManI:RFP) (b), trans-Golgi network marker (N-ST:RFP) (c), and prevacuolar compartment marker (GFP:SYP21) (d). Protoplasts were prepared from rice seedling shoots (a) and rice Oc cell line (b–d). Fluorescence signals were detected from the protoplasts under confocal microscopy. BF, bright field; Scale bar = 10 μm.
BPH18 involved in both antibiosis and antixenosis resistance mechanism
Of the three different mechanisms of resistance to BPH (antibiosis, antixenosis and tolerance)7, rice plants employ two major resistance strategies against herbivores5: antibiosis, which reduces insect feeding, growth rate, or survival, and antixenosis, which affects insect settling, colonization, or oviposition. To investigate the mechanism of resistance, we evaluated the BPH18 complementation transgenic lines, TC1 and TG7, along with the susceptible variety Dongjin and resistant donor line IR65482. The cultivar Dongjin showed a high BPH survival rate while the transgenic lines showed a significantly lower BPH survival rate (Fig. 6a). The antixenosis effect of BPH18 was assessed by the host choice test, which showed a significantly less number of BPH settling on rice seedlings in the transgenic plants compared to the susceptible Dongjin at 24–96 h after BPH infestation (Fig. 6b). These results suggest that BPH18 confers resistance via both antibiosis and antixenosis effects.
(a) Antibiosis effect of BPH18 was measured by BPH survival rate. Four-week-old seedlings were transferred in a glass tube with 7–11 nymphs of BPH per plant, and BPH survival rates were measured at seven days after BPH infestation when the susceptible variety began to die. The average values were obtained from twelve replications, and the error bar shows standard deviation. Statistical tests of difference among survival rates were done using Tukey’s honestly significant difference test. The letters on the bars represent groups, in which observations were not significantly different. The BPH resistance score of tested lines are shown on the graph at the right. The BPH resistance score of the rice seedlings was evaluated according to a method described by Huang et al.53. (b) Antixenosis effect of BPH18 measured by the host choice test. The number of BPH nymphs that settled on rice seedlings was shown at a time course of 3, 6, 24, 48, and 96 h after BPH were released into the pot covered with a light-transmitting mesh. Asterisks on bars represent significant difference between two lines by t-test (★α = 0.05, ★★α = 0.01).
BPH18 and BPH26 are in the same genomic location but they are functionally different alleles
In a recent study, another BPH resistance gene, BPH26, which is located at the BPH18 locus on chromosome 12 was cloned6. To reveal the relationship between these two genes, we developed NILs of BPH18 and BPH26 in the BPH-susceptible indica variety IR24. The whole genome sequencing of NILs (BPH18 and BPH26) and IR24 showed the integration of the donor segments including BPH18 or BPH26 gene into IR24 background (Supplementary Fig. S9). We analyzed the genomic structure of the surrounding regions of the BPH18 and BPH26 locus, respectively. The arrangement of the surrounding genes were quite similar between BPH18 and BPH26 loci (Fig. 7a), resulting that both genes located at the same locus. However, BPH18 and BPH26 showed remarkable differences at the genomic sequence level (Supplementary Fig. S10), despite both having three exons and two introns each. The length of the second intron and the third exon showed a 294-bp and 24-bp differences, respectively (Fig. 7b). The sequence difference was also confirmed by PCR in two NILs (Fig. 7c). In the protein coding sequence, BPH18 and BPH26 have 195 single nucleotide polymorphisms (SNPs) and four gaps (Supplementary Fig. S10). At the amino acid sequence level, we found 105 amino acids difference and five gaps, with 88 of the amino acids difference and all five gaps detected in the LRR domain (Supplementary Fig. S11). To compare the function of alleles, we performed the BPH bioassay in NIL-BPH18 and NIL-BPH26 plants with the BPH insects collected in Nueva Ecija Province, Philippines. The NIL-BPH26 was susceptible to this BPH biotype as IR24 but NIL-BPH18 showed clear resistance (Fig. 7c). These results indicate that BPH18 and BPH26 are functionally different alleles even though they are located at the same locus.
(a) Genomic structures near the BPH18 (top) and BPH26 (bottom) loci. The surrounding genes were annotated based on the whole genome sequence data of NIL-BPH18 and NIL-BPH26. Scale = 1 kb. (b) Gene structures of BPH18 and BPH26 from translation start codon to stop codon. Filled black boxes indicate protein coding region. The size (bp) of exon and intron were shown with number. (c) The BPH bioassay result of susceptible recurrent variety (IR24), NIL-BPH18, and NIL-BPH26. The percent of plant survival was observed after BPH infestation. The average was calculated from two seasons with replications. Error bar means standard deviation. Statistical difference shown as a or b was obtained through least significant difference test (α = 0.01). The sequence difference in the second intron among NIL-BPH18, NIL-BPH26, and IR24 was confirmed by PCR with the BPH18-ind2 primer set.
Plant defense responses to insects involve global changes in gene expression mediated by plant hormone SA and jasmonic acid (JA)/ethylene signaling pathways29. To investigate the pathway involved in the BPH resistance of BPH18, we observed the expression changes of the defense-related genes in IR24 and the two NILs after BPH infestation using qRT-PCR (Supplementary Fig. S12). Overall gene expression patterns of the plant defense-related genes were similar between the susceptible lines, IR24 and NIL-BPH26 and it differed with NIL-BPH18. In the susceptible lines, JA synthesis-related genes (LOX and AOS2), in SA synthesis-related gene (EDS1), ethylene receptor gene (EIN2), and a pathogen-related gene (PR1b) were strongly increased, especially at 72 h after BPH infestation. In contrast, in NIL-BPH18, none of the defense-related genes tested in this study was strongly activated by BPH insect, suggesting that unknown pathway may be involved in BPH resistance in NIL-BPH18. However, the gene expression patterns of the defense-related genes were significantly different between NIL-BPH18 and NIL-BPH26.
Discussion
As a PFI, the BPH probes intercellular plant tissues to establish feeding sites in the phloem sieve elements30. Remarkable similarities between plant responses to phloem feeders and pathogens have been found31. To date, six PFI-resistance genes (Mi-1, Vat, Bph3, Bph14, BPH26, and BPH29) have been isolated in plants. Four of these encode CC-NBS-LRR proteins which are typical among plant R proteins against pathogens. BPH18 and BPH26 proteins have a CC-NBS-NBS-LRR domain structure, which is basically similar to the typical CC-NBS-LRR R proteins. This commonality among the PFI-resistance proteins and R proteins against plant pathogens might indicate the similar molecular mechanism in resistance. As intracellular receptors, NBS-LRR R proteins sense pathogen effectors directly or host protein modifications induced by pathogen molecules while pathogens secrete into the plant cell to suppress immune response and trigger potent innate immune responses20,32,33,34. Additionally NBS-LRR proteins as helpers of defense signaling transduce signals downstream of some pathogen-activated NBS-LRR proteins35. PFIs puncture the phloem cell and secret watery saliva that contains complex mixtures of lipoproteins, phospholipids, and carbohydrates, as well as numerous enzymes with proteolytic, hydrolytic, oxidative, or cell wall-degrading activities30. These factors probably aid in stylet penetration and could detoxify defensive compounds in the host plant30. The PFI oral secretions are a potential source of effectors or avirulence (avr) factors. Elucidation of the molecular mechanism of BPH effectors corresponding with Bph14, BPH18, BPH26 and BPH29 is needed to get basic knowledge on resistance to BPH.
The BPH18/BPH26 and Pib encode R proteins of the unique domain structure of CC-NBS-NBS-LRR having two NBS domains with unconserved P-loops, respectively. Of the 480 NBS-LRR genes identified in the japonica rice genome36, only four genes encode proteins having two NBS domains in which NBS domains are partially duplicated. The first NBS domains of BPH18/BPH26 and Pib are partial, lacking most of the ARC sub-domains, and their P-loop motifs were much different from the consensus sequence. When NBS-LRR proteins function as a helper which regulates signal transduction following pathogen effector recognition by other NBS-LRR proteins, the P-loop is not essential for these functions. For example, rice Pb1 conferring durable resistance to neck blast disease37, its orthologue in Arabidopsis ADR1-L2 involved in bacterial pathogen resistance35 might be regarded as helper NBS-LRR proteins for defense signaling. They encode a CC-NBS-LRR protein having a degenerate P-loop in the NBS domain. These types of NBS-LRR proteins need to be studied to check whether they function as sensor, helper, or both.
Plants deploy intracellular immune receptors such as NBS-LRR proteins to perceive pathogen invasion. Through the subcellular localization experiments, it is revealed that BPH18 is localized widely to endo-membranes, including ER, Golgi, TGN, and PVC. Plant NBS-LRR R proteins reside in diverse subcellular locations, and each R protein will be in the place of its effector or effector target23. Membrane trafficking is emerging as a central theme in plant innate immunity and has been implicated in immune receptor activation, defense signaling, and targeting of cellular cargo to pathogen invasion sites38. Critical components of host-membrane trafficking are prime targets of pathogen effectors39. The Xanthomonas campestris pv. vesicatoria type III effector protein XopJ is localized in the Golgi body and plasma membrane, suppressing protein secretion and callose deposition, which leads to the weakening of cell wall-associated defense responses40. A Pseudomonas syringae virulence protein, HopM1, mediates the destruction of an immunity-associated protein, AtMIN7, which is involved in vesicle trafficking pathway and cell wall-associated defense; both HopM1 and AtMIN7 are localized to the trans-Golgi network/early endosome41,42. The ARFA1b/c, an ARF GTPase localized in the multi-vesicular body/PVC, is required for callose deposition for pre-invasive penetration resistance against powdery mildew in barley43. The Arabidopsis TIR-NB-LRR R protein, RPP1A, which confers resistance to the oomycete Hyalopernospora parasitica, resides in the ER/Golgi apparatus44. Based on these examples, we hypothesize that the BPH18 protein is possibly involved in recognizing the invasion of BPH and in detecting some effector proteins of BPH, which targets endo-membranes and interferes with the vesicle trafficking pathway and cell wall-associated defense, including callose deposition.
Even though the transgenic lines harboring BPH18 showed significant antibiosis effect, the effect was weaker than in the original resistant donor line. However, BPH18 showed their antixenosis effect in transgenic lines, which was similar with the resistant donor (Fig. 6a,b). Bph6 from the indica rice variety Swarnalata also had antibiosis and antixenosis effects45. Interestingly, Bph6 conferred a higher resistance when it was introgressed into an indica-susceptible genetic background than when it was introgressed into a japonica-susceptible genetic background. This might explain why BPH18 transferred into a japonica variety showed a weaker antibiosis effect than the original indica resistance donor line. One another possible explanation is that at least two more minor Quantitative Trait Loci (QTLs) were found on the short arm of chromosome 5 and on the end of chromosome 12 in the previous BPH18 mapping study1. Incorporating these QTLs would further enhance BPH resistance of BPH18 harboring rice lines.
BPH18 and BPH26 are on the same locus in the long arm of chromosome 12. Similarly the Pi2, Piz-t, and Pi9 genes are in the same genomic region of rice chromosome 6 including a cluster of nine NBS-LRR genes10,46. Pi2 and Piz-t are two different resistant alleles by eight amino-acid differences for the fourth NBS-LRR gene in this cluster, and Pi9 corresponds to the second NBS-LRR gene. Whole genome sequencing of the NIL-BPH18 and NIL-BPH26 revealed that BPH18 and BPH26 located at the same locus (Fig. 7a). This might imply that BPH18 and BPH26 are the same genes with different alleles. Unlike the Pi2 and Piz-t alleles, many DNA and amino acid sequence differences were found between the BPH18 and BPH26 genes. This may be derived from evolutionary divergence between the donor sources of these two genes. While BPH26 came from O. sativa indica variety ADR52, BPH18 was originated from O. australiensis.
The amino acid sequences of LRR domains of BPH18 and BPH26 are very much divergent while their NBS domains are well conserved (Supplementary Fig. S11), which suggests that LRR domains determine resistance specificities in the case of BPH18 and BPH26. The LRR domains of NBS-LRR proteins recognize pathogen effectors directly and determine resistance specificities. The LRR domain of Pi-ta binds directly with its cognate fungal effector Avr-Pita, and a single amino acid difference in the LRR domain of Pi-ta distinguishes resistant and susceptible alleles9,47. The eight amino-acid differences within the LRR domains between Pi2 and Piz-t determine resistance specificity46. In comparison of the L6 and L11 alleles of flax, polymorphisms in LRR domains account for the specific recognition of AvrL567 by L6 and AvrL11 by L1148,49. Also, it was revealed that six amino acid changes confined to LRR domains determine the difference between P and P2 rust resistance specificities in flax50. The in planta association of Arabidopsis RPP1 resistance protein and its cognate oomycete effector ATR1 was mediated by the LRR domain of RPP151. In addition to discovering effectors of BPH, molecular mechanism of their recognition by BPH resistance proteins should be elucidated, and their LRR domains might be strong candidates for effector recognition domains.
The NIL-BPH18 and NIL-BPH26 in the same susceptible background, IR24, showed different resistance reactions to the BPH strain collected in Nueva Ecija Province, Philippines. While, the NIL-BPH26 showed susceptibility like IR24, NIL-BPH18 showed higher resistance to BPH. Similar result was observed in the previous report. Neither BPH26 nor BPH25 in susceptible Taichung65 background showed resistance to the BPH strain, Japan-KG-06 which is BPH2-virulent biotype. When they coexist, the line showed resistance to that biotype6. However, BPH18 without BPH25 showed resistance to the Nueva Ecija BPH population, supporting that this BPH insect belongs to BPH2-virulent biotype, and BPH18 and BPH26 are functionally different alleles. Different resistance reactions to the same BPH population suggests that BPH18 and BPH26 recognizes the different effectors or the modifications of different plant proteins caused by effectors through the variable LRR domain probably. Gene expression patterns of the plant defense-related genes were quite different between the NIL-BPH26 and NIL-BPH18. This result indicates that the two BPH resistance genes utilize different resistance pathways after BPH attacks. It was observed that in Bph14 and BPH29 lines, JA synthesis-related genes are not upregulated but SA synthesis-related genes were strongly expressed by BPH infestation5,7. In NIL-BPH26, both SA and JA synthesis-related genes were strongly induced by BPH, suggesting that BPH26 may activate JA and SA-dependent resistance pathway. In NIL-BPH18, there was no significant transcription activation of the previously identified defense-related genes. Similarly, RNA sequencing experiment revealed that the transcript level of SA dependent pathway genes are not different between NIL-BPH15 and its susceptible recipient line52. But the expression level of many regulatory genes including hormone signaling genes, receptor kinases, and transcription factors were changed by BPH infestation. Like BPH15, unidentified resistance pathway may lead the BPH resistance in NIL-BPH18.
Transfer of the BPH18 gene conferring resistance could control planthopper infestation in rice. The BPH18 gene derived from wild rice should be utilized in breeding programs as a new source of resistance to increase rice production.
Methods
BPH insect materials
The BPH insects were collected from rice fields in South Korea in 2003. A pure BPH population was developed from a single colony of BPH and was grown on the susceptible japonica variety Taebaekbyeo in a glass house1. This Korean BPH biotype was used for fine-mapping and the experiments with transgenic plants in Suwon, South Korea. At IRRI, Philippines, we used the BPH population collected in 2011 from Nueva Ecija Province which is the major rice cultivation area in the Philippines. A pair of BPH insects was collected and cultured in the glass house on the susceptible variety, T(N)1. After culturing and maintaining several generations, this BPH population was used for evaluation of NIL-BPH18 and NIL-BPH26.
Fine-mapping of BPH18
The 3,100 BC4F2 plants derived from the cross between the susceptible cultivar Junam and IR65482-7-216-1-2 as a resistant donor were used as the fine-mapping population. We developed two cleaved amplified polymorphic sequences (CAPS) markers, BN45 and BN52, flanking the BPH18 region of about 1.1 Mb. We genotyped all the plants with these two markers and selected the plants that had a recombinant genotype in this interval. A total of 149 plants were selected. The BC4F3 lines from these selected plants were planted, each line comprising 13 plants. All the plants of this BC4F3 population were genotyped again with the two flanking markers from which 130 plants that had homozygous recombinant genotype were selected. Seeds from the selected plants were harvested and BC4F4 progenies were sown for BPH bioassay. The bioassay was done with the Korean BPH biotype using the modified bulk seedling test (MBST) method1. Seedlings at the three-leaf stage were infested with second- or third-instar nymphs at a density of 10–12 nymphs per seedling. When all the seedlings of the susceptible control died, the tested plants were evaluated as resistant or susceptible depending on survival or death of seedlings. Eight CAPS and InDel markers (Supplementary Table S1) in the BN52-BN45 region were used for genotyping the selected homozygous recombinant BC4F3 plants. The 27-kb genomic region, revealed to harbor BPH18 by fine-mapping in the BPH18 donor line, was sequenced.
Identification of the BPH18 full-length cDNA
To test whether the LOC_Os12g37280 and LOC_Os12g37290 genes make one gene encoding a CC-NBS-NBS-LRR protein, the NF and the NR primers were used for RT-PCR with cDNA synthesized from the RNA of the BPH18 donor line. To identify the full-length cDNA of this combined gene model, 5′ RACE and 3′ RACE PCR was conducted using the CapFishingTM Full-length cDNA Premix Kit (Seegene, Korea). The full-length cDNA of BPH18 was confirmed by PCR with NFCF and NFCR primers. All primers for molecular analysis of BPH18 gene are shown in Supplementary Table S2.
Complementation test and RNAi experiment
Firstly, a 6.4-kb genomic DNA fragment of the LOC_Os12g37290 gene in the BPH18 donor was amplified with RPL2F and RPL2R primers and finally it was inserted into the pCAMBIA1300 binary vector. The BPH-susceptible japonica variety, Ilmi, was transformed with this construct using the Agrobacterium-mediated method. The T1 plants were subjected to BPH bioassay. Based on the new gene model, combining LOC_Os12g37290 and LOC_Os12g37280, the promoter, ORF, and terminator part were amplified with BPH18-pro, BPH18-ORF, and BPH18-ter primer pairs, respectively, then inserted into pPZP vector consecutively. Alternatively the 8-kb genomic region, including the LOC_Os12g37280 gene in the BPH18 donor, was amplified with a pair of LRR-8.0 primers. The purified PCR product was inserted into the already constructed pCAMBIA1300 vector, harboring the LOC_Os12g37290 gene with In-Fusion HD Cloning Kit (Clontech, USA). Thus, the constructed vector included the whole 14-kb genomic region of the LOC_Os12g37290 gene and LOC_Os12g37280 gene of the BPH18 donor. The BPH-susceptible japonica variety, Dongjin, was transformed with this construct and the T1 plants having the transgene were selected by PCR or bar-strip test using the AgraStrip LL Rice Strip Test Kit (Romer Labs, USA). Then, the plants were subjected to BPH bioassay with the South Korea BPH strain. To generate the RNAi construct for the BPH18 gene, we amplified a 435-bp fragment of BPH18 donor cDNA using primers BPH18i-F and BPH18i-R. Finally the fragments were cloned into the destination vector, pANDAβ. The BPH-resistant NILs in Junam background, NIL-BPH18, was transformed with this construct and the T2 plants having the RNAi construct were subjected to BPH bioassay. The BPH resistance score of the rice seedlings was evaluated according to the method described by Huang et al.53.
Domain structure and phylogenetic analysis
The domain structure of BPH18 was analyzed using the CD SEARCH program of the NCBI website (http://www.ncbi.nlm.nih.gov/). The CC structure was predicted by Paircoil254 program (http://groups.csail.mit.edu/cb/paircoil2/). The amino acid sequences of the identified rice NBS-LRR R proteins and human APAF-1 protein were downloaded, and their NBS and LRR domain sequences were used for sequence alignment and phylogenetic analysis. The protein alignment was generated with ClustalW55. We used MEGA 6.056 to reconstruct neighbor-joining trees. For the tree analysis, we performed 1000 bootstrap replicates to assess the support for the nodes.
Gene expression analysis of BPH18
Five-leaf-stage plants of the resistance donor line (IR65482-7-216-1-2) were infested with the South Korea BPH strain and sampled after 0, 3, 6, 12, 24, and 48 h with five replications. Time 0 means the time point just before BPH infestation. Total RNA was extracted from the leaf sheaths and then converted into cDNA using a PrimeScript 1st cDNA synthesis kit (TaKaRa, Japan). The expression of BPH18 was evaluated by TaqMan qRT-PCR using ABI7900HT machine (Applied Biosystems, USA). The expression level in the samples was quantified relative to the first replicate of Time 0. To investigate BPH18 expression in different tissues, we extracted total RNAs from the leaf sheath, leaf, and root of five-leaf-stage plants. The expression level in the samples was quantified relative to the first replicate of root. A genomic DNA fragment (2,340 bp to 1 bp from the translation start site) containing the promoter region of BPH18 was amplified by PCR using primers attB-BPH18-pro-F and attB-BPH18-pro-R. This fragment was inserted into the upstream of the beta-glucuronidase (GUS) coding region in the pBGWFS7 binary vector. Transgenic plants carrying the above construct were generated in Dongjin cultivar background.
Subcellular localization
The full-length BPH18 coding region (3.7 kb) without a stop codon was amplified with primers BPH18-loc-F and BPH18-loc-R from the IR65482-7-216-1-2 cDNA. Finally, the fragment was cloned into the downstream of the ZmUbi1 promoter and in frame with GFP in the pGA3452 vector and with RFP in the pGA3574 vector, respectively. Protoplasts were prepared from leaves of rice seedlings and the rice Oc cell line (suspension culture) which was derived from roots of rice seedlings. The constructs were co-transformed into protoplast through electroporation with several markers, including the ER marker (Bip:RFP), cis-Golgi marker (ManI:RFP), trans-Golgi network marker (N-ST:RFP), and prevacuolar compartments (PVC) marker (GFP:SYP21). After incubation, the protoplasts were observed under confocal laser-scanning microscopy (LSM 510 META, Zeiss, Germany) and images were obtained using Zeiss LSM Image Browser. The experiments were done at least twice for each marker and the representative images were taken for publication.
BPH resistance mechanism analysis
To test the antibiosis effect, we grew T1 plants of the two BPH18 transgenic lines, TC1 and TG7, with their susceptible wild-type variety Dongjin and BPH18 donor. Twelve four-week-old seedlings of each line were transferred into test tubes with water. Around 7-11 BPH insects of second- or third-instar nymphs were put into each test tube. When the Dongjin seedlings began to die, both live and dead BPH insects were counted and the BPH survival rates were calculated for each test tube. The BPH resistance score of the rice seedlings in each test tube was evaluated according to a method described by Huang et al.53 as follows; 0 None of the leaves shrank and the plant was healthy, (1) One leaf was yellowing, (3) One to two leaves were yellowing or one leaf shrank, (5) One to two leaves shrank or one leaf shriveled, (7) Three to four leaves shrank or two to four leaves shriveled, the plant was still alive, (9) The plant died. To test the antixenosis effect of BPH18, TC1, TG7, and the BPH18 donor line (IR65482) were compared with the susceptible variety Dongjin. We transplanted five four-week-old seedlings of the test lines and Dongjin into a 20-cm pot, placing rice seedlings along the circumference of the pot, alternatively. A 5-cm Petri dish was placed at the center of the pot where BPH nymphs were released, and the pot was covered with a light-transmitting mesh.
Two replications were done for each experiment. The number of hoppers that had settled on each plant was recorded at 3, 6, 24, 48 and 96 h after infestation.
Development of NILs, its BPH bioassay, and qRT-PCR of defense-related genes
A BPH susceptible indica rice variety, IR24 as a recurrent parent, was crossed with IR65482-7-216-1-2 (BPH18 donor) and ADR52 (BPH26 donor), respectively. The F1 plants were backcrossed to the recurrent parent. The BC1F1 plants were screened with known linked markers to select those plants that contain the resistance gene from the donor parent. The selected plants carrying the target gene were used in the next backcrossing cycle. This procedure was repeated through the 3rd backcross. The BC3F1 plants having the target gene were selfed to produce the BC3F2 generation which were again selected and selfed until BC3F5. The bioassay was done by the MBST method1 during the dry and wet seasons of 2014 at the IRRI, Philippines. Seedlings at the three-leaf stage were infested with second- or third-instar nymphs at a density of 10–12 nymphs per seedling, done in two replications. The percent of plant survival was observed one week after infestation or once the susceptible check was dead. For qRT-PCR of defense-related genes, seven-day-old seedling plants were infested with the Nueva Ecija BPH population. Leaf sheaths from three plants per line were collected before (0 h) and after BPH infestation (8, 24, 48, 72 h). Total RNAs were extracted using TRIzol reagent (Life technologies, USA) and genomic DNAs were removed by treatment of DNase I (TURBO DNA-free kit, Life technologies), then first strand cDNAs were synthesized by ImProm-II Reverse Transcription system (Promega, USA). Real-time PCRs were performed using SYBR Select Master Mix (Life technologies) with ABI7500 machine (Applied Biosystems). The primer sequences for the target genes were same with the previous reports by Du et al.5 and Wang et al.7. OsAct1 gene was used as an internal control and the relative expression level was calculated based on the ΔΔCt method. Each data point represents the mean value of three biological replications.
Whole genome sequencing and data analysis
Whole genome sequencing of NIL-BPH18, NIL-BPH26, and IR24 were conducted by Illumina Hiseq2500 platform at 30X coverage depth (pair-end 125 bp sequencing with average 500 bp insert library). De novo assembly was done by SOAPdenovo2 software57 with k-mer 59 after filtering the raw reads. The BPH18/BPH26 gene with the surrounding regions was identified through pairwise sequence alignment between the newly created scaffolds and indica reference genome (93–11). The raw sequence reads were aligned against 93–11 reference genome using BWA software58, resulting in SAM files. Using SAMtools59, the SAM files were converted to BAM files and the consensus sequences were extracted. Finally the chromosomes were formed after comparing the reference based assembly and de novo assembly. The dot plot alignment of chromosome 12 was visualized using Mummer software60.
Additional Information
Accession codes: The BPH18 sequences from IR65482-7-216-1-2 (accession no. KF890252) and Junam (accession no. KJ850252) were deposited in the NCBI GenBank database.
How to cite this article: Ji, H. et al. Map-based Cloning and Characterization of the BPH18 Gene from Wild Rice Conferring Resistance to Brown Planthopper (BPH) Insect Pest. Sci. Rep. 6, 34376; doi: 10.1038/srep34376 (2016).
Change history
11 November 2016
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Jena, K. K., Jeung, J. U., Lee, J. H., Choi, H. C. & Brar, D. S. High-resolution mapping of a new brown planthopper (BPH) resistance gene, Bph18(t), and marker-assisted selection for BPH resistance in rice (Oryza sativa L.). Theor. Appl. Genet. 112, 288–297 (2006).
Cheng, X., Zhu, L. & He, G. Towards understanding of molecular interactions between rice and the brown planthopper. Mol. Plant 6, 621–634 (2013).
Wu, H. et al. Fine mapping of brown planthopper (Nilaparvata lugens Stal) resistance gene Bph28(t) in rice (Oryza sativa L.). Mol. Breed. 33, 909–918 (2014).
Liu, Y. et al. A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice. Nat. Biotechnol. 33, 301–305 (2015).
Du, B. et al. Identification and characterization of Bph14, a gene conferring resistance to brown planthopper in rice. Proc. Natl. Acad. Sci. USA. 106, 22163–22168 (2009).
Tamura, Y. et al. Map-based cloning and characterization of a brown planthopper resistance gene BPH26 from Oryza sativa L. ssp. indica cultivar ADR52. Sci. Rep. 4, 5872 (2014).
Wang, Y. et al. Map-based cloning and characterization of BPH29, a B3 domain-containing recessive gene conferring brown planthopper resistance in rice. J. Exp. Bot. 66, 6035–6045 (2015).
Dodds, P. N. & Rathjen, J. P. Plant immunity: towards an integrated view of plant-pathogen interactions. Nat. Rev. Genet. 11, 539–548 (2010).
Bryan, G. T. et al. A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta . Plant Cell 12, 2033–2046 (2000).
Qu, S. et al. The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics 172, 1901–1914 (2006).
Ling, Y. & Weilin, Z. Genetic and biochemical mechanism of rice resistance to planthopper. Plant Cell Rep. doi: 10.1007/s00299-016-1962-6 (2016).
Lo Presti, L. et al. Fungal effectors and plant susceptibility. Annu. Rev. Plant Biol. 66, 513–545 (2015).
Santamaria, M. E., Martinez, M., Cambra, I., Grbic, V. & Diaz, I. Understanding plant defence responses against herbivore attacks: an essential first step towards the development of sustainable resistance against pests. Transgenic Res. 22, 697–708 (2013).
Wu, J. & Baldwin, I. T. New insights into plant responses to the attack from insect herbivores. Annu. Rev. Genet. 44, 1–24 (2010).
Hogenhout, S. A. & Bos, J. I. Effector proteins that modulate plant–insect interactions. Curr. Opin. Plant Biol. 14, 422–428 (2011).
Rodriguez, P. A. & Bos, J. I. Toward understanding the role of aphid effectors in plant infestation. Mol. Plant Microbe Interact. 26, 25–30 (2013).
Rossi, M. et al. The nematode resistance gene Mi of tomato confers resistance against the potato aphid. Proc. Natl. Acad. Sci. USA. 95, 9750–9754 (1998).
Dogimont, C., Bendahmane, A., Chovelon, V. & Boissot, N. Host plant resistance to aphids in cultivated crops: genetic and molecular bases, and interactions with aphid populations. C. R. Biol. 333, 566–573 (2010).
Suh, J.-P. et al. Development of elite breeding lines conferring Bph18 gene-derived resistance to brown planthopper (BPH) by marker-assisted selection and genome-wide background analysis in japonica rice (Oryza sativa L.). Field Crops Research 120, 215–222 (2011).
McHale, L., Tan, X., Koehl, P. & Michelmore, R. W. Plant NBS-LRR proteins: adaptable guards. Genome Biol. 7, 212 (2006).
Wang, Z. X. et al. The Pib gene for rice blast resistance belongs to the nucleotide binding and leucine-rich repeat class of plant disease resistance genes. Plant J. 19, 55–64 (1999).
Takken, F. L., Albrecht, M. & Tameling, W. I. Resistance proteins: molecular switches of plant defence. Curr. Opin. Plant Biol. 9, 383–390 (2006).
Rafiqi, M., Bernoux, M., Ellis, J. G. & Dodds, P. N. In the trenches of plant pathogen recognition: Role of NB-LRR proteins. Semin. Cell Dev. Biol. 20, 1017–1024 (2009).
van Ooijen, G. et al. Structure-function analysis of the NB-ARC domain of plant disease resistance proteins. J. Exp. Bot. 59, 1383–1397 (2008).
Min, M. K. et al. Overexpression of Arabidopsis AGD7 causes relocation of Golgi-localized proteins to the endoplasmic reticulum and inhibits protein trafficking in plant cells. Plant Physiol. 143, 1601–1614 (2007).
Saint-Jore-Dupas, C. et al. Plant N-glycan processing enzymes employ different targeting mechanisms for their spatial arrangement along the secretory pathway. Plant Cell 18, 3182–3200 (2006).
Liu, T. Y. et al. PHO2-dependent degradation of PHO1 modulates phosphate homeostasis in Arabidopsis. Plant Cell 24, 2168–2183 (2012).
Foresti, O., daSilva, L. L. & Denecke, J. Overexpression of the Arabidopsis syntaxin PEP12/SYP21 inhibits transport from the prevacuolar compartment to the lytic vacuole in vivo . Plant Cell 18, 2275–2293 (2006).
Walling, L. L. The myriad plant responses to herbivores. J. Plant Growth Regul. 19, 195–216 (2000).
Thompson, G. A. & Goggin, F. L. Transcriptomics and functional genomics of plant defence induction by phloem-feeding insects. J. Exp. Bot. 57, 755–766 (2006).
Klingler, J. et al. Aphid resistance in Medicago truncatula involves antixenosis and phloem-specific, inducible antibiosis, and maps to a single locus flanked by NBS-LRR resistance gene analogs. Plant Physiol. 137, 1445–1455 (2005).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Lukasik, E. & Takken, F. L. STANDing strong, resistance proteins instigators of plant defence. Curr. Opin. Plant Biol. 12, 427–436 (2009).
Jacob, F., Vernaldi, S. & Maekawa, T. Evolution and Conservation of Plant NLR Functions. Front. Immunol. 4, 297 (2013).
Bonardi, V. et al. Expanded functions for a family of plant intracellular immune receptors beyond specific recognition of pathogen effectors. Proc. Natl. Acad. Sci. USA. 108, 6463–16468 (2011).
Zhou, T. et al. Genome-wide identification of NBS genes in japonica rice reveals significant expansion of divergent non-TIR NBS-LRR genes. Mol. Genet. Genomics 271, 402–415 (2004).
Hayashi, N. et al. Durable panicle blast-resistance gene Pb1 encodes an atypical CC-NBS-LRR protein and was generated by acquiring a promoter through local genome duplication. Plant J. 64, 498–510 (2010).
Teh, O. K. & Hofius, D. Membrane trafficking and autophagy in pathogen-triggered cell death and immunity. J. Exp. Bot. 65, 1297–1312 (2014).
Frei dit Frey, N. & Robatzek, S. Trafficking vesicles: pro or contra pathogens? Curr. Opin. Plant Biol. 12, 437–443 (2009).
Bartetzko, V. et al. The Xanthomonas campestris pv. vesicatoria type III effector protein XopJ inhibits protein secretion: evidence for interference with cell wall-associated defense responses. Mol. Plant Microbe Interact. 22, 655–664 (2009).
Nomura, K., Debroy, S., Lee, Y. H., Pumplin, N., Jones, J. & He, S. Y. A bacterial virulence protein suppresses host innate immunity to cause plant disease. Science 313, 220–223 (2006).
Nomura, K. et al. Effector-triggered immunity blocks pathogen degradation of an immunity-associated vesicle traffic regulator in Arabidopsis. Proc. Natl. Acad. Sci. USA. 108, 10774–10779 (2011).
Bohlenius, H., Morch, S. M., Godfrey, D., Nielsen, M. E. & Thordal-Christensen, H. The multivesicular body-localized GTPase ARFA1b/1c is important for callose deposition and ROR2 syntaxin-dependent preinvasive basal defense in barley. Plant Cell 22, 3831–3844 (2010).
Weaver, L. M., Swiderski, M. R., Li, Y. & Jones, J. D. The Arabidopsis thaliana TIR-NB-LRR R-protein, RPP1A; protein localization and constitutive activation of defence by truncated alleles in tobacco and Arabidopsis. Plant J. 47, 829–840 (2006).
Qiu, Y., Guo, J., Jing, S., Zhu, L. & He, G. High-resolution mapping of the brown planthopper resistance gene Bph6 in rice and characterizing its resistance in the 93–11 and Nipponbare near isogenic backgrounds. Theor. Appl. Genet. 121, 1601–1611 (2010).
Zhou, B. et al. The eight amino-acid differences within three leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea . Mol. Plant. Microbe Interact. 19, 1216–1228 (2006).
Jia, Y., McAdams, S. A., Bryan, G. T., Hershey, H. P. & Valent, B. Direct interaction of resistance gene and avirulence gene products confers rice blast resistance. EMBO. J. 19, 4004–4014 (2000).
Dodds, P. N. et al. Direct protein interaction underlies gene-for-gene specificity and coevolution of the flax resistance genes and flax rust avirulence genes. Proc. Natl. Acad. Sci. USA. 103, 8888–8893 (2006).
Ellis, J. G., Dodds, P. N. & Lawrence, G. J. Flax rust resistance gene specificity is based on direct resistance-avirulence protein interactions. Annu. Rev. Phytopathol. 45, 289–306 (2007).
Dodds, P. N., Lawrence, G. J. & Ellis, J. G. Six amino acid changes confined to the leucine-rich repeat beta-strand/beta-turn motif determine the difference between the P and P2 rust resistance specificities in flax. Plant Cell 13, 163–178 (2001).
Krasileva, K. V., Dahlbeck, D. & Staskawicz, B. J. Activation of an Arabidopsis resistance protein is specified by the in planta association of its leucine-rich repeat domain with the cognate oomycete effector. Plant Cell 22, 2444–2458 (2010).
Lv, W. et al. BAC and RNA sequencing reveal the brown planthopper resistance gene BPH15 in a recombination cold spot that mediates a unique defense mechanism. BMC Genomics 15, 674 (2014).
Huang, Z., He, G., Shu, L., Li, X. & Zhang, Q. Identification and mapping of two brown planthopper resistance genes in rice. Theor. Appl. Genet. 102, 929–934 (2001).
McDonnell, A. V., Jiang, T., Keating, A. E. & Berger, B. Paircoil2: improved prediction of coiled coils from sequence. Bioinformatics 22, 356–358 (2006).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Delcher, A. L., Phillippy, A., Carlton, J. & Salzberg S. L. Fast aligorithms for large-scale genome alignment and comparison. Nucleic Acids Res. 30, 2478–2483 (2002).
Acknowledgements
This research was supported by grants from the National Institute of Agricultural Sciences (No. PJ0068172013), Rural Development Administration (RDA), Korea; the Next-Generation BioGreen 21 Program (Plant Molecular Breeding Center, No. PJ008128), RDA, Korea; National Institute of Crop Science, RDA, Korea, and the International Rice Research Institute (Grant No. DRPC 2011-134), Metro Manila, Philippines. We thank Prof. Inwhan Hwang (POSTECH, Korea) for providing ER, Golgi, and PVC markers, Dr. Ajay Kohli (IRRI) for supporting qRT-PCR, and Prof. Rod Wing (Arizona Genomics Institute, University of Arizona, USA) for reviewing the manuscript. We are thankful to science editorial team of IRRI communication for carefully editing the manuscript.
Author information
Authors and Affiliations
Contributions
K.K.J. conceived the experiments. H.J., S.-R.K. and K.K.J. wrote the paper. H.J., J.-P.S., S.-M.K., H.K., G.-S.L., U.-H.Y., T.-H.K., H.L., S.-C.S. and K.K.J. performed mapping and BPH bioassay. Y.-H.K. and H.-M.P. generated transgenic plants. H.J., S.-R.K. and S.L.H. conducted gene expression analysis. S.-R.K., J.Y. and G.A. performed sub-cellular localization. K.K.J. and S.L.H. developed NILs in IR24 background. N.S. and G.M. analyzed NGS data. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ji, H., Kim, SR., Kim, YH. et al. Map-based Cloning and Characterization of the BPH18 Gene from Wild Rice Conferring Resistance to Brown Planthopper (BPH) Insect Pest. Sci Rep 6, 34376 (2016). https://doi.org/10.1038/srep34376
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34376
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.