Abstract
The Pacific cupped oyster, Crassostrea gigas, is one of the major aquacultural shellfish species that has been introduced to Europe and America from its native source in the West Pacific. In Taiwan, the cultivated cupped oysters along the west coast have been identified as C. gigas for over centuries; however, several molecular phylogenetic studies have cast doubt upon the existence of this species in Taiwan and adjacent waters. Indeed, our analyses of mitochondrial cytochrome oxidase I (COI) sequences from 313 Crassostrea collected from 12 locations along Taiwanese and southern Chinese coastlines confirm that all samples were the Portuguese oyster, C. angulata, rather than C. gigas. Multiple lines of evidence, including haplotypic and nucleotide diversity of the COI gene, demographic history and population genetics, suggest that Taiwanese C. angulata is unique, probably experienced a sudden population expansion after the Last Glacial Maxima around 20,000 years ago and has a significantly limited genetic connectivity across the Taiwan Strait. Our study applies an extended sampling and DNA barcoding to confirm the absence of C. gigas in natural and cultivated populations in Taiwan and southern China, where we only found C. angulata. We highlight the importance of conserving the gene pool of the C. angulata population in Taiwan, particularly considering the current threats by large-scale environmental disturbances such as marine pollution, habitat destruction and climate change.
Similar content being viewed by others
Introduction
Cupped oysters belonging to the genus Crassostrea are among the most important cultivated shellfish species in the world1. Among the three species of Crassostrea listed by the FAO, the Pacific cupped oyster, C. gigas, contributed an estimated 555k tons to global aquaculture production in 20131. Taiwan’s cultivated cupped oyster species has been identified as C. gigas for over a century and has produced 27,793 tons in 2013 having an economic value of over US$20 million2. However, molecular phylogenetic analyses of Crassostrea species have raised concern over the taxonomic status of C. gigas in Taiwan3,4,5,6,7,8,9. Molecular phylogenetic studies on the origin of the Portuguese oyster, C. angulata, using small numbers of Taiwanese cupped oysters as the Asian reference, unexpectedly showed that those from Taiwan were grouped with C. angulata from the Atlantic3,4 instead of being clustered with C. gigas from Japan. C. angulata is a species assumed to be native to the northeastern Atlantic but is morphologically and enzymatically indistinguishable from C. gigas5,6,10. In contrast, karyotype analysis highlighted the close genetic similarity of these two taxa in comparison with other cupped oyster species11, but distinct differences between their respective karyotypes were also observed on certain chromosome pair and restriction enzyme ideograms12,13,14. Divergence time estimation based on the completed mitochondrial genomes of Crassostrea oysters confirmed that C. gigas and C. angulata are closely-related and probably diverged about 2.7 million years ago (mya)15,16. Subsequent sampling along the Chinese coast demonstrates that C. angulata has a wide distribution in southern China, with the Yangtze River separating it from C. gigas in northern China8,9. Taiwan, the largest continental island situated at the border between the northern and southern Chinese coasts, possesses climates ranging from tropical to subtropical/temperate. The Taiwanese coast provides a variety of marine environments that host a relatively highly diverse fauna17,18, thus C. angulata and C. gigas might prove to coexist along the Taiwan coast if intensively sampled. In addition, geological events such as the last glacial maximum (LGM~20 ka BP) are speculated to have affected the phylogeography of marine biota in the western Pacific19,20,21 and might play an important role in shaping the demographic history of Crassostrea species in Taiwan.
DNA barcoding based on the mitochondrial cytochrome oxidase I (COI) fragment has been a standard way to discriminate between closely related species, identify new, cryptic, or invasive species and assess species assemblages of communities across many animal phyla. The general rule in DNA barcoding is that intraspecific COI variation is <1%, whereas interspecific divergence is normally >2%22. Here, we applied DNA barcoding of a COI fragment to examine the status of cultivated Crassostrea species from eight populations in Taiwan and four along the southern China coast (Supplementary Fig. 1S). In addition, the published COI DNA sequences of C. gigas from the northwestern Pacific (China, Japan and Korea) were retrieved from GenBank20 for phylogenetic and phylogeographic comparisons.
Results
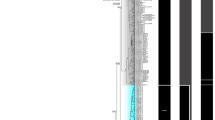
We obtained COI sequences from 313 Crassostrea samples collected from 12 locations along the Taiwanese and Chinese coasts. In a total of 43 haplotypes with 41 variable sites, 14 phylogenetically informative sites were identified for Crassostrea. The COI phylogenies constructed using ML and Bayesian methods produced identical topologies (Fig. 1). Instead of grouping with C. gigas, all 43 COI haplotypes formed a monophyletic group with the Portuguese oyster, C. angulata (DQ659374), with high bootstrap support (99) and Bayesian posterior probability (100). ITS-1 phylogeny showed that all oyster samples collected in this study were of C. angulata, not C. gigas (Supplementary Fig. 2S). Our results confirmed that C. gigas does not occur along the coasts of Taiwan and southern China, regardless of whether the oysters were from marine farms or natural populations. Therefore, C. angulata was used herein as the species epithet in the following analyses to infer phylogenetic, population genetic structure and demographic history.
Phylogenetic tree constructed for Taiwan Crassostrea oysters and reference sequences based on the COI gene.
Branch support values estimated by bootstrap pseudo-replicates in maximum-likelihood and Bayesian inference, respectively, are shown above each branch. A minus sign (−) indicates that bootstrap values and Bayesian posterior probabilities were <75%. 46 sequences of C. gigas from the northwestern Pacific were retrieved from GenBank20 (KP099007- KP099052). The accompanied photos showed the shell morphology of these two species used in this study. C. angulata were collected from (a) Hsinchu, (b) Changhua, (C) Tainan and (d) Pingtung in Taiwan. C. gigas purchased from market were imported from (a) Australia, (b) France, (c) South Africa, (d) and Ireland to Taiwan.
The nucleotide composition of the COI gene fragment in C. angulata was A + T-rich (A, 22.4%; T, 38%) and variations consisted predominantly of transition substitutions (Ti: Tv = 11.78). The number of haplotypes (nh) ranged from 4 in Beihai to 14 in Chiayi. The number of endemic haplotypes (neh) was significantly higher in populations along the Taiwanese coast than from the Chinese coast (X2-test = 72.921, p < 0.001). Haplotypic diversity (h) was high in all populations, with a mean value of 0.882 ± 0.012. In contrast, nucleotide diversity (π) was low, with a mean value of 0.00396 ± 0.00017 (Table 1).
Relationships between haplotypes are represented on a median-joining network (MJN). The MJN analysis showed a complex star-like network of six haplotypes accounted for 79.87% of the COI DNA sequences collected from C. angulata (Fig. 2a). In contrast, the MJN of C. gigas was a typical star-like network dominated by one haplotype (Fig. 2b). The MJN showed no obvious haplotype clustering with respect to Taiwanese and Chinese coast populations. The high mean haplotypic diversity (0.882 ± 0.012) relative to low mean nucleotide diversity (0.00396 ± 0.00017) in C. angulata is indicative of a population bottleneck followed by rapid population growth and accumulation of mutations23,24. Summary statistics for testing drift-mutation equilibrium (Tajima’s D and Fu’ Fs) showed that the two C. angulata populations along the Taiwanese and Chinese coasts experienced different demographic histories (Table 2). The Tajima’s D (−2.03279) and Fu’s Fs (−27.30487) values along the Taiwanese coast were significantly negative (p < 0.05), indicating these populations had a sudden population expansion, although most individuals were not statically significant in these two parameters. In contrast, Tajima’s D (0.01183) was positive and Fu’s Fs (−1.11287) along the Chinese coast was not significantly negative, suggesting that these populations have suffered a recent bottleneck or expansion. The different demographic history across the two sides of the Taiwan Strait is also illustrated by mismatched distributions (Fig. 3). The combined data from Chinese and Taiwanese populations showed a unimodal frequency distribution similar to an analysis of the Taiwan-only population (Fig. 3a,b). In contrast, the Chinese population had a negative binomial frequency distribution (Fig. 3c). Tau values (τ = 2.865) can provide a rough estimate of when the rapid population expansion began. Using a mutation rate of 2.8% per myr for the C. angulata15 COI, the time of demographic expansion for overall populations was estimated at ~91 Kya (95% CI: 35–138 Kya), for Chinese populations at ~118 Kya (95% CI: 33–196 Kya) and for Taiwanese populations at ~64 Kya (95% CI: 37–79 Kya). In contrast, a Bayesian skyline plot (BSP) showed that the initiation of overall C. angulata population growth was approximately 12,000 years ago and reached a stable effective population size approximately 3,000 years ago (Fig. 4a). When estimations were conducted on Taiwanese and Chinese populations, a BSP showed that Taiwanese C. angulata population growth began approximately 16,000 years ago and reached a stable effective population size approximately 4,000 years ago (Fig. 4b). In contrast, Chinese population growth began 6,000 years ago and reached a stable effective population size less than 1,000 years ago (Fig. 4c). In contrast, a BSP showed that C. gigas population grew and remained a stable effective population size before 45,000 years ago (Fig. 4d).
Mismatch distribution for the mtCOI of (a) overall Crassostrea angulata; (b) C. angulata, of Taiwanese coast; (c) C. angulata, of Chinese coast; (d) C. gigas20. The bar represents observed frequency and the line represents modeled frequency.
Bayesian skyline plotting of (a) overall Crassostrea angulata; (b) C. angulata, of Taiwanese coast; (c) C. angulata, of Chinese coast; (d) C. gigas20.The Bayesian skyline plot is derived from mtDNA COI sequences, where the x-axis is time in years and the y-axis is the product of effective population size (Ne) and generation time (T). The black line indicates the median estimate and dashed lines indicate the 95% highest posterior density (HPD) region.
All the pairwise Fst values between C. angulata populations along the Chinese coast and 20 of 28 pairwise Fst between populations along the Taiwanese coast were not significant (Table 3). In contrast, 28 of 32 pairwise comparisons between Taiwanese and Chinese populations showed significant Fst values (p < 0.05). The overall population subdivision between Chinese and Taiwanese coasts was highly significantly different (Fst = 0.10167, p < 0.001), suggesting a limited gene flow across the Taiwan Strait. An analysis of molecular variance (AMOVA) supported this tendency with a significant among-group ΦCT (0.09787) between Taiwanese and Chinese coast populations, although 88.26% of the variance existed within populations (Supplementary Table 1S).
Discussion
By intensively sampling cupped oyster populations island-wide, we confirmed that the Crassostrea species in Taiwan is C. angulata and that no C. gigas occurs in the natural environments of Taiwan and southern China sea waters. Several studies on the origin of C. angulata in Europe have used small numbers of Crassostrea previously identified as C. gigas from a single population in Taiwan as Asian references. Our result shows that C. angulata has an Asian origin and probably was introduced to Portugal from Taiwan in the 16th century3,4,25. Further surveys of cultured cupped oysters with five samples from Chiayi, Taiwan, identified C. angulata as the main species in the southern China, although it was suggested that C. angulata is a subspecies of C. gigas in this region9. Although the species/subspecies status between C. gigas and C. angulata remains unsettled as they can hybridize in hatcheries and in Southern Portugal9,26, 2.67% sequence divergence of COI fragment demonstrates that these two species are genetic distinct based on the criteria of interspecific divergence normally >2% for the standard DNA barcoding22. Indeed, species phylogeny generated by 31 Asian Crassostrea mitochondrial genomes from Chinese coast clearly demonstrates that C. angulata and C. gigas are closely related but independent species that probably diverged about 2.7 mya15,16 during the late Pliocene when sea levels and sea surface temperatures (SST) were higher and warmer but starting to drop during the onset of the “age of ice-ages” in the Pleistocene27. We hypothesized that the common ancestor of C. angulata and C. gigas might represent a species (resembling C. angulata) that was adapted to the warmer SSTs along the West Pacific coast in the late Pliocene. When glacial-interglacial cycles emerged during the Pleistocene, dramatic changes in the area and configuration of the coastline along the West Pacific28 might have isolated ancestor populations. Isolated populations in West Pacific high latitudes evolved into cold-adapted species resembling C. gigas. Haplotypic diversity, nucleotide diversity and demographic analyses of the COI gene also supported C. angulata and C. gigas being distinct species and that C. angulata is an older lineage than C. gigas. C. angulata had twice as much haplotypic diversity (h: 0.882) and four times the nucleotide diversity (π: 0.00396) of C. gigas (h: 0.457 and π: 0.00097)20. While the median-joining network (MJN) of C. angulata was complex and star-like, being complicated by multiple connections and high-frequency internal haplotypes, the MJN of C. gigas from the northwestern Pacific was simple and star-like, which is the most common and widespread haplotype and the one that is typically assumed to be the most ancestral24. Paleoclimatic fluctuations could also account for the present-day distributions of C. angulata in the warm, subtropical southwestern Pacific and C. gigas in the cool, temperate northwestern Pacific, with the Yangtze River as a natural biogeographic barrier to separate not only these two cupped oyster species but also other coastal marine species along the Asian continent8,9,19,21.
Significant population expansion was identified for C. angulata along the coast of Taiwan and southern China based on different lines of evidence including network, mismatch distribution analysis (MDA) and neutral tests. For example, populations in Taiwan and southern China experienced different expansion models, with the former having a sudden population expansion and unimodal frequency distribution and the latter with a recent bottleneck or expansion and a negative binomial frequency distribution. The different expansion history of C. angulata, as indicated by MDA, is that the Chinese population expanded around 118 Kya and the Taiwanese population around 64 Kya. Beginning 140 Kya during the last interglacial period, Earth’s air temperature rose to approximately 2 °C above the present level and sea level rose to 5–6 meters or more above present ocean levels by 130–127 Kya29; therefore, ancestral C. angulata populations might have had the chance to colonize and expand along the coast of southern China as well as western Taiwan. However, this scenario was not supported because, following this interglacial high stand and several more oscillations, sea level dropped 120–130 m below the present level during the last glacial maximal (LGM; 21 Kya). Thus, C. angulata populations must be extinct because southern China and western Taiwan became connected when the sea level dropped during the LGM. In contrast, the Bayesian sky plot (BSP) matched the sequences of demographic histories for Taiwanese and southern Chinese populations to the effects of the LGM. That is, the BSP fits the time needed for their expansion to stable population sizes after another rapid rise in sea level (over a 12 Kya period) stabilized close to the present level 7,000 years ago30. However, caution should be taken in interpreting the effects of interglacial-glacial sea-level fluctuations on the population expansion history of C. angulata in this region because in consistent estimations of demographic expansion between MDA versus the BSP analysis have been noticed in several marine species in the marginal seas of the northwestern Pacific19,21. Methodological difference between MDA and BSP single-locus phylogeny using mtDNA and the uncertainty of the mtDNA mutation rate have been proposed to account for the estimation Bayesian determining expansion times21,31. Future multi-locus analyses (e.g., microsatellites, SNPs) should be considered for confirming the population expansion of C. angulata along the Taiwanese and southern Chinese coasts.
Although caution should be taken to interpret genetic connectivity based on single-locus mtDNA fragments such as the COI gene, the pairwise Fst between Taiwanese and southern Chinese populations of C. angulata is still highly significant, suggesting that cross-strait gene flow is limited. The Taiwan Strait, situated between the north end of the South China Sea (SCS) and south side of the East China Sea (ECS), has been recognized as a corridor for the migration of marine species between the two seas, aided by complex ocean currents around Taiwan (reviewed in ref. 18). Most studies of marine species in this region have focused on the phylogeography and connectivity between the SCS and ECS (reviewed in refs 19 and 21) and little attention has been paid to connectivity between the two sides of the Taiwan Strait. On the contrary, seasonal changes in northward and southward currents, such as the South China Sea Surface Current in the summer and China Coastal Water in the winter, might create a boundary that blocks cross-strait gene flow in C. angulata. A similar pattern was observed in the horseshoe crab, Tachypleus tridentatus, with gene flow being strong along the Chinese coast but limited in its ability to cross the Taiwan Strait32. Further studies using multi-locus population genetic analyses of C. angulata and more common benthic species should be done to confirm this observed pattern of restricted gene flow across the Taiwan Strait.
Confirmation of C. angulata in Taiwan is important not only to science but also to the oyster farming industry that plays a significant role in aquacultural economics2. For example, crosses between the Japanese C. gigas and Taiwanese cultivated Crassostrea and breeding of triploids have both been attempted but eventually failed in the last few decades in Taiwan33. Using standard COI DNA barcoding, we have demonstrated that the cultivated cupped oyster in Taiwan is C. angulata, not C. gigas. Thus, the failure to produce viable hybrid offspring from Japanese C. gigas and Taiwanese C. angulata is not unexpected even though natural hybrids between these two cupped oyster species have been detected by a microsatellite locus in southern Europe7. In addition, significant differences in demographic history, mtDNA genetic diversity and limited gene flow across the Taiwan Strait highlight the importance of conserving the uniqueness of the Taiwanese C. angulata population, particularly under the current impacts of large-scale environmental disturbances that include marine pollution, habitat destruction and climate change around Taiwan.
Materials and Methods
Ethics Statement
No specific permits were required for the described studies and no specific permissions were required for these locations/activities.
Sampling and DNA sequencing
In total, 313 individual oyster specimens were collected from 12 locations that were subdivided into two geographical regions (Taiwanese coast and Chinese coast) (Supplementary Fig. 1S; Supplementary Table 1S). Eight foreign (e.g., Japan, Korea, Australia and America) C. gigas samples were purchased from importers and restaurants. Ten additional species of Crassostrea were selected as outgroups (COI: C. virginica EU007484, C. cuttackensis FJ262983, C. belcheri AY038077, C. gryphoides EU007491, C. madrasensis EU007462, C. iredalei AY038078, C. hongkongensis AY632557, C. nippona AF300616, C. ariakensis AY160752, C. sikamea EU007475; ITS-1: C. virginica EU072460, C. rivularis DQ785895, C. ariakensis FJ356683, C. hongkongensis EU073317, C. nippona FJ356681, C. madrasensis EU073241, C. iredalei EU073329, C. gryphoides EU073247, C. belcheri EU073259 and C. sikamea AB735523). Collected samples were frozen and transported to the laboratory. Adductor muscle tissue was cut into pieces and preserved in 95% alcohol. All reference samples are preserved in the Molecular Systematic Laboratory, Marine Fisheries Division, Fisheries Research Institute.
For initial species diagnosis, COI was selected as the molecular marker. Total genomic DNA was extracted from adductor muscle tissue using a commercial DNA isolation kit (Gentra, Minneapolis, MN, USA). The LCO-1490 (forward primer, 5′-GGT CAA CAA ATC ATA AAG ATA TTG G-3′) and HCO-2198 (reverse primer, 5′-TAA ACT TCA GGG TGA CCA AAA AAT CA-3′) primer set that amplifies COI was used34.The PCR reaction mixture included 5 μL of 10 × PCR buffer (Perkin-Elmer, Foster City, CA, USA), 4 μL of dNTPs (2.5 mM each), 2 μL of each primer (5 μM), 0.5 μL of 1.25-unit TaKaRa Taq (Takara Bio, Shiga, Japan) and 1.0 μL of a template containing approximately 5 ng of DNA; ddH2O was added to make a volume of 50 μL. A model 2400 thermal cycler (Perkin-Elmer) was used for PCR and conditions were set as follows: one cycle at 95 °C for 4 min, 40 cycles at 95 °C for 1 min, at 50 °C for 1 min, at 72 °C for 3 min and a final elongation at 72 °C for 7 min. The reaction was stopped and products were preserved at 4 °C.
PCR products were examined on 1% agarose gel by electrolysis to confirm that the correct fragment length had been obtained and then eluted using the QIAquick Gel Extraction kit (QIAGEN, Hilden, Germany). Purified PCR products were sent to Mission Biotech (Taipei, Taiwan) for DNA sequencing.
To reinforce the accuracy of species diagnosis and to determine whether C. gigas and hybrids (C. gigas × C. angulata) occur in Taiwan, we also sequenced the nuclear ITS-1 region with primers described by Hedgecock et al.35. We amplified the samples, which were haplotypes recognized from COI sequences and used two or more replicates of each. The cycling conditions were one cycle at 95 °C for 4 min, 40 cycles at 95 °C for 1 min, at 55 °C for 1 min, at 72 °C for 3 min and a final elongation at 72 °C for 7 min. PCR products were electrophoresed on 1% agarose gel to check their lengths. Amplified fragments of ITS-1 region were cloned into the yT&A cloning vector (Yeastern Biotech, Taipei, Taiwan) and transformed into Escherichia coli JM109 according the manufacturer’s protocol. Plasmids were screened for inserts and inserts were sent to Mission Biotech (Taipei, Taiwan) for DNA sequencing.
Phylogenetic and haplotype network analyses
COI and ITS-1 sequences were aligned by ClustalW in MEGA 6.036. DnaSP 5.1037 was used to identify haplotypes and polymorphic sites. Nucleotide sequence data for the haplotypes used in this paper were deposited in GenBank under accession numbers KU726888– KU726939. Maximum likelihood (ML) trees were reconstructed with a Kimura two-parameter model using MEGA 6.0 for both COI and ITS-1 sequences. The evaluation of statistical confidence was based on bootstrapping with 1,000 pseudo-replicates for ML38. Bayesian analysis was performed using MrBayes version 3.1.239. The best-fit model of nucleotide substitution was the HKY + I model, which was selected using jModelTest version 2.1.740. This model allows for a different rate of transitions and transversions and unequal frequencies of the four nucleotides, with a proportion of invariable site, I = 0.80. COI analyses were run with the best-fit model for 2.3 × 107 generations and a sampling frequency of 100 generations (standard deviation of split frequencies = 0.008835) and 3 × 106 generations for the ITS-1 analysis (standard deviation of split frequencies = 0.009508. Phylogenetic trees were visualized and edited with FigTree (http://tree.bio.ed.ac.uk/software/figtree/).
Gene diversity was described as haplotype diversity (h), endemic haplotypes (eh) and nucleotide diversity (π) for each population using DnaSP 5.10. Pairwise genetic divergence between populations was estimated by using Fst values and significance was verified through 10,000 permutations and adjusted by sequential Bonferroni correction41. Historical demographic history was obtained with neutrality tests, mismatch distributions and a Bayesian Skyline Plot based on COI data. As for the neutrality test, Tajima’s D test42 and Fu’s Fs test43 were calculated using Arlequin 3.544 with 10,000 permutations. Mismatch distribution was constructed for each geographic population to test a model of exponential population growth45. A goodness of fit test was performed to test the validity of the sudden expansion model using a parametric bootstrap approach based on the sum of square deviations (SSD) between observed and expected mismatch distributions. The demographic expansion parameter (τ) was calculated by Arlequin 3.5.
To detect any differences in genetic structure among populations using analysis of molecular variances (AMOVA), we grouped populations into three types: (a) those along Taiwanese (NTC, HC, CH, CY, TN, PT, KL, TT) and Chinese coasts (WZ, MT, KM, BH), (b) those along the Taiwanese (NTC, HC, CH, CY, TN, PT, KL, TT, MT, KM) and Chinese coasts(WZ, BH) and (c) those along Taiwan’s east (YL, TT) and west coasts (NTC, HC, CH, CY, TN, PT) and the China coast (WZ, MT, KM, BH). This first analysis was carried out to determine whether there were any differences in genetic structure across the Taiwan Strait. Oyster spats are regularly transported between Taiwanese coast and two islands, MT and KM, along the Chinese coast for marine culture46, therefore, the second analysis was carried out to examine whether the oyster spat transportation affected the genetic structure of oyster populations along the Chinese coast. The third analysis was carried out to determine whether there were any differences in genetic structure between Taiwan east coast and west coast populations. Pairwise genetic divergence between populations was estimated using the fixation index Φst47. Demographic history and divergence time for the C. angulata COI sequence were further estimated using Bayesian evolutionary analysis sampling trees (BEAST) ver. 2.1.348. Changes in effective population size (Ne) across time were inferred using Bayesian skyline analyses. To infer demographic history, coalescence methods require an initial demographic model to be specified. Where evidence of population expansion was found, the timing of expansion in generations (t) was estimated from τ = 2 μt, where τ (tau) is a parameter of the time to expansion in units of mutations and μ is the mutation rate per generation for the DNA sequence under study. The mutation rate of C. angulata was calculated as 2.785% per nucleotide per Myr since the divergence time was about 2.7 Myr for C. angulata and C. gigas15. The analysis was run with 50 million steps in a Markov chain Monte Carlo (MCMC) simulation under the HKY + I model, relaxed molecular clock model (uncorrelated log-normal). Other operators were optimized automatically. Results were viewed with Tracer v1.549.
The haplotype network of the COI gene was inferred using the median-joining algorithm50. Before the calculation, the star contraction method with a maximum star radius value of 10 was used to simplify the data matrix. The MJN was drawn by Network, version 4.6.1.0 (Fluxus Technology, U.K.).
Additional Information
How to cite this article: Hsiao, S.-T. et al. DNA barcoding reveals that the common cupped oyster in Taiwan is the Portuguese oyster Crassostrea angulata (Ostreoida; Ostreidae), not C. gigas. Sci. Rep. 6, 34057; doi: 10.1038/srep34057 (2016).
References
FAO. Global Forest Resources Assessment (Food and Agriculture Organization of the United Nations, Rome, 2015); [accessed 2016 Jun 1]. URL: http://www.fao.org/fishery/culturedspecies/Crassostrea_gigas/en.
Fisheries Administration: Taiwan Fisheries Yearbook. Council of Agriculture, Executive Yuan, Taipei, Republic of China (2013).
Boudry, P., Heurtebise, S., Collet, B., Cornette, F. & Gérard, A. Differentiation between populations of the Portuguese oyster, Crassostrea angulata (Lamark) and the Pacific oyster, Crassostrea gigas (Thunberg), revealed by mtDNA RFLP analysis. Journal of Experimental Marine Biology and Ecology 226, 279–291 (1998).
O’Foighil, D., Gaffney, P. M., Wilbur, A. E. & Hilbish, T. J. Mitochondrial cytochrome oxidase I gene sequences support an Asian origin for the Portuguese oyster Crassostrea angulata. Marine Biology 131, 497–503 (1998).
Huvet, A., Balabaud, K., Bierne, N. & Boudry, P. Microsatellite analysis of 6-hour-old embryos reveals no preferential intraspecific fertilization between cupped oysters Crassostrea gigas and Crassostrea angulata. Marine Biotechnology 3, 448–453 (2001).
Huvet, A. et al. Is fertility of hybrids enough to conclude that the two oysters Crassostrea gigas and C. angulata are the same species? Aquatic Living Resources 15, 45–52 (2002).
Huvet, A., Fabioux, C., McCombie, H., Lapègue, S. & Boudry, P. Natural hybridization between genetically differentiated populations of Crassostrea gigas and C. angulata highlighted by sequence variation in flanking regions of a microsatellite locus. Marine Ecology Progress Series 272, 141–152 (2004).
Wang, H. & Guo, X. Identification of Crassostrea ariakensis and related oysters by multiple species-specific PCR. Journal of Shellfish Research 27, 481–487 (2008).
Wang, H., Qian, L., Liu, X., Zhang, G. & Guo, X. Classification of a common cupped oyster from southern China. Journal of Shellfish Research 29, 857–866 (2010).
Mattiucci, S. & Villani, F. Allozyme study in oysters classified as Crassostrea gigas (Thunberg, 1793) and Crassostrea angulata (Lamark, 1819) (Mollusca: Ostreidae). Parasitologia 25, 21–27 (1983).
Leitão, A., Thiriot-Quiévreux, C., Boudry, P. & Malheiro, I. A. “G” chromosome banding study of three cupped oyster species: Crassostrea gigas, Crassostrea angulata and Crassostrea virginica (Mollusca: Bivalvia). Genetics, Selection and Evolution 31, 519–527 (1999a).
Leitão, A., Boudry, P., Labat, J. P. & Thiriot-Quiévreux, C. Comparative kariological study of cupped oyster species. Malacologia 41, 175–186 (1999b).
Leitão, A., Chaves, R., Santos, S., Guedes-Pinto, H. & Boudry, P. Restriction enzyme digestion chromosome banding in Crassostrea and Ostrea species: comparative karyological analysis within Ostreidae. Genome 47, 781–788 (2004).
Leitao, A., Chaves, R., Santos, S., Guedes-Pinto, H. & Boudry, P. Interspecific hybridization in oysters: restriction enzyme digestion chromosome banding confirms Crassostrea angulata x Crassostrea gigas F1 hybrids. Journal of Experimental Marine Biology and Ecology 343, 253–260 (2007).
Ren, J., Liu, X., Jiang, F., Guo, X. & Liu, B. Unusual conservation of mitochondrial gene order in Crassostrea oysters: evidence for recent speciation in Asia. BMC Evolutionary Biology 10, 394 (2010).
Ren, J. et al. Intraspecific variation in mitogenomes of five crassostrea species provides insight into oyster diversification and speciation. Marine Biotechnology 18, 242–254 (2016).
Chen, C. Analysis of scleractinian distribution in Taiwan indicating a pattern congruent with sea surface temperatures and currents: examples from Acropora dn Faviidae. Zoological Studies 38, 119–129 (1999).
Chen, C. A. & Keshavmurthy, S. Taiwan as a connective stepping-stone in the kuroshio triangle and the conservation of coral ecosystems under the impacts of climate change. Kuroshio Science 3, 15–22 (2009).
Ni, G., Li, Q., Kong, L. & Yu, H. Comparative phylogeography in marginal seas of the northwestern Pacific. Molecular Ecology 23, 534–548 (2014).
Li, S., Li, Q., Yu, H., Kong, L. & Liu, S. Genetic variation and population structure of the Pacific oyster Crassostrea gigas in the northwestern Pacific inferred from mitochondrial COI sequences. Fisheries Science 81, 1071–1082 (2015).
Wang, J., Tsang, L. M. & Dong, Y. W. Causations of the phylogeographical barrier of some rocky shore species along the Chinese coastline. BMC Evolutionary Biology 15, 114 (2015).
Ratnasingham, S. & Hebert, P. D. N. bold: The Barcode of Life Data System (http://www.barcodinglife.org). Molecular Ecology Notes 7, 355–364 (2007).
Grant, W. & Bowen, B. W. Shallow population histories in deep evolutionary lineages of marine fishes: insights from sardines and anchovies and lessons for conservation. Journal of Heredity 89, 415–426 (1998).
Avise, J. C. Phylogeography: The history and formation of species. Harvard University Press, Cambridge, MA (2000).
Lapègue, S., Batista, F. M., Heurtebise, S., Yu, Z. & Boudry, P. Evidence for the presence of the Portuguese oyster, Crassostrea angulata, in Northern China. Journal of Shellfish Research 23, 759–763 (2004).
Reece, K. S., Cordes, J. F., Stubbs, J. B., Hudson, K. L. & Francis, E. A. Molecular phylogenies help resolve taxonomic confusion with Asian Crassostrea oyster species. Marine Biology 153, 709–721 (2008).
Miller, K. G. et al. High tide of the warm Pliocene: Implications of global sea level for Antarctic deglaciation. Geology 40, 407–410 (2012).
Hall, R. & Holloway, J. D. Biogeography and Geological Evolution of SE Asia. Backhuys Publishers, Leiden (1998).
Waelbroeck, C. et al. Sea-level and deep water temperature changes derived from benthic foraminifera isotopic records. Quaternary Science Reviews 21, 295–305 (2002).
Kimura, M. Paleogeography of the Ryukyu Islands. Tropics 10, 5–24 (2000).
Marko, P. B. et al. The ‘Expansion-Contraction’ model of Pleistocene biogeography: rocky shores suffer a sea change? Molecular Ecology 19, 146–169 (2010).
Yang, M. C., Chen, C. P., Hsieh, H. L. & Chen, C. A. Is a living fossil about to go locally extinct? No mitochondrial genetic variation in horse shoe crab juveniles Tachypleus tridentatus at haomeiliao nature reserve, Budai, Taiwan. Zoological Studies 48, 737 (2009).
Chao, N. H. et al. Induction and performance of triploid oyster, Crassostrea gigas, in Taiwan. COA (Council of Agriculture, Taiwan). Fisheries Series 65, 99–115 (1999).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology 3, 294–299 (1994).
Hedgecock, D., Li, G., Banks, M. A. & Kain, Z. Occurrence of the Kumamoto oyster Crassostrea sikamea in the Ariake Sea, Japan. Marine Biology 133, 65–68 (1999).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Molecular Biology and Evolution 30, 2725–2729 (2013).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Ronquist, F. et al. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology 61, 539–542 (2012).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nature Methods 9, 772 (2012).
Rice, W. R. Analyzing tables of statistical tests. Evolution 43, 223–225 (1989).
Tajima, F. Statistical methods to test for nucleotide mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Fu, Y. X. Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147, 915–925 (1997).
Excoffier, L. & Lischer, H. E. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources 10, 564–567 (2010).
Rogers, A. R. & Harpending, H. Population growth makes waves in the distribution of pairwise genetic differences. Molecular Biology and Evolution 9, 552–569 (1992).
Huang, S. C., Hsieh, H. L. & Chen, C. P. Effects of winter Monsoon on the growth, mortality and metabolism of adult oysters in Kinmen Island, Taiwan. Aquaculture 251, 256–270 (2006).
Excoffier, L., Smouse, P. E. & Quattro, J. M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131, 479–491 (1992).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Molecular Biology and Evolution 29, 1969–1973 (2012).
Rambaut, A., Suchard, M. & Drummond, A. J. Tracer v1.5. Available from http://tree.bio.ed.ac.uk/software/tracer/) [accessed Nov 30] 2009.
Bandelt, H. J., Forster, P. & Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution 16, 37–48 (1999).
Acknowledgements
The authors thank Mr. Min-Yu Hwang and Mr. Hong-Yen Liang for collecting tissue samples of oysters from markets. We are very grateful to Dr. Wei-Cheng Su and Dr. Don-Chung Liu for their encouragement and support. This study was supported by the Council of Agriculture (97AS-11.1.2-AI-A2, 98AS-8.1.4-AI-A1), Taiwan, ROC.
Author information
Authors and Affiliations
Contributions
S.-T.H. conceived, designed and performed the experiments. S.-T.H., S.-C.C. and K.-S.C. performed the statistics. S.-T.H. and C.A.C. wrote the paper. P.-H.H. and C.A.C. made intellectual contributions. C.-L.W. a division chief who provided general support.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hsiao, ST., Chuang, SC., Chen, KS. et al. DNA barcoding reveals that the common cupped oyster in Taiwan is the Portuguese oyster Crassostrea angulata (Ostreoida; Ostreidae), not C. gigas. Sci Rep 6, 34057 (2016). https://doi.org/10.1038/srep34057
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34057
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.