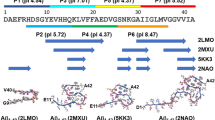
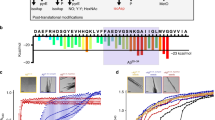
Abstract
Fibrillation of differently modified amyloid β peptides and deposition as senile plaques are hallmarks of Alzheimer’s disease. N-terminally truncated variants, where the glutamate residue 3 is converted into cyclic pyroglutamate (pGlu), form particularly toxic aggregates. We compare the molecular structure and dynamics of fibrils grown from wildtype Aβ(1–40) and pGlu3-Aβ(3–40) on the single amino acid level. Thioflavin T fluorescence, electron microscopy, and X-ray diffraction reveal the general morphology of the amyloid fibrils. We found good agreement between the 13C and 15N NMR chemical shifts indicative for a similar secondary structure of both fibrils. A well-known interresidual contact between the two β-strands of the Aβ fibrils could be confirmed by the detection of interresidual cross peaks in a 13C-13C NMR correlation spectrum between the side chains of Phe 19 and Leu 34. Small differences in the molecular dynamics of residues in the proximity to the pyroglutamyl-modified N-terminus were observed as measured by DIPSHIFT order parameter experiments.
Similar content being viewed by others
Introduction
Fibrillation of amyloid β (Aβ) peptides of different length and degrees of modification and brain deposition as senile plaques represent a hallmark of Alzheimer’s disease. The N-terminally truncated variant, which forms a cyclic pyroglutamate residue at position 3, plays a major role in the development of the disease by forming very toxic aggregates1,2. For pyroglutamated Aβ variants (pGlu-Aβ) increased oligomerization3,4,5, enhanced fibrillation6,7,8, and increased lipid peroxidation accompanied with a loss of plasma membrane integrity6 were reported. These properties can probably explain the enhanced cytotoxic effects of pGlu-Aβ1,4,9. Indeed, pharmacological inhibition of the enzyme that catalyzes the production of the pGlu lactam ring (see Fig. 1) has been reported to reduce the deposition of amyloid plaques and retards the memory decline in mice10.
(A) ThT fluorescence intensity of pGlu3-Aβ(3–40) (red) and Aβ(1–40) (black) as a function of time. (B) Scanning transmission electron micrograph of the pGlu3-Aβ(3–40) fibrils after 3 weeks of incubation. The scale bar represents 1.5 μm. (C) X-ray diffraction pattern of pGlu3-Aβ(3–40) fibrils. Above, a sketch of the structural modification from glutamic acid to pyroglutamate is shown.
The molecular structure of pGlu-Aβ(3–40) has only been studied for peptides dissolved in TFE-containing buffers by solution NMR8,11,12, or in aqueous media by CD- and FTIR spectroscopy6,7,8,13. Furthermore, H/D exchange was monitored by solution NMR6. Compared to unmodified Aβ(1–40) or Aβ(1–42), some studies claimed a higher propensity of pGlu-Aβ to from β-sheet structure7,12, while another paper reported a higher α-helical content and a decreased β-sheet propensity13.
A direct comparison of the molecular structure on the amino acid level in the fibrillar state of pGlu-Aβ(3–40) and wildtype (WT) Aβ fibrils is missing. For the study of the molecular structure of fibrils, solid-state NMR spectroscopy is the method of choice14 to provide details of the molecular structure of amyloid fibrils in different stages of fibrillation. Overall and in relatively good agreement, the models have identified two β-strand regions in the Aβ(1–40) fibrils (mainly around residues 10–22 and 30–38), which are connected by a short hairpin so that U-shaped monomers form the fibrils, while the N-terminus of the peptide is unstructured and more dynamic15,16,17. Here, we report data on the secondary structure and tertiary interresidual contacts of mature Aβ fibrils grown from pGlu3-Aβ(3–40) peptides on the level of selected amino acids.
Results and Discussions
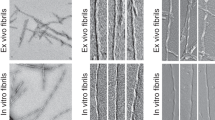
The fibrillation kinetics of pGlu3-Aβ(3–40) in comparison to WT Aβ(1–40) was monitored by standard thioflavin T (ThT) fluorescence spectroscopy. Figure 1A shows the maximum of the ThT fluorescence intensity as a function of time. In agreement with previous results6, it is observed that also under our conditions the fibrillation of pGlu3-Aβ(3–40) has a significantly shorter lag time and is overall faster than for WT Aβ(1–40). The lag time for pGlu3-Aβ(3–40) is 7 ± 1 h, while it is 43 ± 2 h for Aβ(1–40). The morphology of the pGlu3-Aβ(3–40) fibrils was studied by electron microscopy (EM). Figure 1B shows a typical EM micrograph of pGlu3-Aβ(3–40) fibrils after 3 weeks of incubation, displaying fibrils of homogeneous morphology. These fibrils have a width of 12.8 ± 2.1 nm (n = 20), which is slightly larger than the mean diameter of WT Aβ(1–40) fibrils (10.0 ± 1.6 nm)18. A preferential shorter fibril length for pGlu3-Aβ(3–40) as reported in6 could not be observed under our fibrillation conditions. Fibrils of pGlu3-Aβ(3–40) exhibit a mean length of 850 ± 300 nm WT Aβ(1–40) fibrils of 500 ± 200 nm (n = 30). Also, the X-ray diffraction pattern of pGlu3-Aβ(3–40) fibrils (Fig. 1C) exhibit the typical cross-β structure as observed for all other amyloid fibrils. The measured main X-ray reflections correspond to repeat spacings of 4.7 Å and 10.3 Å, which represent the typical values for the interstrand spacing and the intersheet distance, respectively19.
To obtain insights into the secondary structure on the level of individual amino acids, solid-state NMR spectra of pGlu3-Aβ(3–40) fibrils with uniformly 13C/15N-labeled amino acids (for the labeling scheme see experimental section) were measured. In choosing the sites for isotopic labeling, we paid special attention to the N-terminus of pGlu3-Aβ(3–40) to study possible differences in the structures due to the pGlu modification in position 3 and to probe the extent of the known secondary structure elements. To assign the 13C and 15N chemical shifts, 13C-13C DARR and 15N-13Cα NMR correlation spectra were conducted under magic-angle spinning (MAS) conditions in dual acquisition mode20. Supplementary Figure S1 shows a 13C-13C DARR and a 15N-13Cα NMR spectrum as examples. The chemical shifts values for all labeled amino acids are listed in Table S1.
Figure 2 reports the 13Cα and 13Cβ chemical shifts of pGlu3-Aβ(3–40) and mature WT Aβ(1–40) fibrils (data taken from ref. 21) as differences from random coil values reported in the literature22. Since NMR chemical shifts are sensitive to the secondary structure, values close to zero correspond to random coil regions, while negative values for Cα and positive values for Cβ report β-strand conformations22. One can clearly see that most of the chemical shift values of both fibrillar species are very similar. Some alterations are observed for the Cβ signal of Phe4, which may result from the direct vicinity to the chemically modified pGlu3. For Phe19 Cβ and Gly29 Cα, two chemical shift values were observed. Such structural polymorphism of Aβ fibrils has been observed before in several preparations of WT Aβ fibrils15,23,24,25. Also the larger line width up to 3 ppm (FWHM) for some signals (e.q. Val12 or Val36) could be a result of structural polymorphism.
Comparison of the secondary 13C MAS NMR chemical shifts of pGlu3-Aβ(3–40) (red) and mature WT Aβ(1–40) (black) fibrils for 13Cα (squares) and 13Cβ (circles).
Data are given as the difference of a measured chemical shift to random coil chemical shifts taken from the literature22.
Overall, a striking structural similarity between pGlu3-Aβ(3–40) and WT Aβ(1–40) fibrils is observed and one has to conclude that the typical secondary structure elements of WT Aβ(1–40) with an unstructured N-terminus and two β-strand regions comprising amino acids 10–22 and 30–38, which are connect by a short unstructured region, holds also true for pGlu3-Aβ(3–40).
For a systematic comparison of the secondary structure of pGlu3-Aβ(3–40) fibrils to different preparations of WT Aβ fibrils in different stages of the fibrillation process, Figure 3 shows correlation plots of the differences in the chemical shift values (13Cα-13Cβ), which are very sensitive to secondary structure and have the advantage of being independent of chemical shift referencing, which may vary between the different laboratories. On the secondary structure level, a very good correlation of the pGlu3-Aβ(3–40) fibrils with the results for WT Aβ fibrils is obtained (A–C). Only slightly smaller correlation coefficients are obtained when pGlu3-Aβ(3–40) fibrils are compared to protofibrils (D) and oligomers (E, F). This indicates that the secondary structure of the pGlu3-Aβ(3–40) fibrils is very similar to mature fibrils, but also to oligomers and protofibrils.
Comparison of fibrils of pGlu3-Aβ(3–40) with published chemical shift data of WT Aβ preparations.
Correlation plots of the chemical shift differences (13Cα-13Cβ) for all labeled amino acids of pGlu3-Aβ(3–40) (x axis) and Aβ fibrils in different stages of the fibrillation process (y axis) are given. (A) mature fibrils from24, (B) mature fibrils16, (C) mature fibrils21, (D) protofibrils35, (E) oligomers43, and (F) oligomers44. The Pearson correlation coefficient is given for each plot. Note that Gly9 and Gly29 are not part of this comparison as 13Cα-13Cβ chemical shift differences are analyzed.
Tertiary structure information for Aβ(1–40) fibrils represents a field of some controversy and is highly dependent on the number and precision of the structural constraints available, which has led to different structural models16,17,26,27. To obtain some insight into the spatial relationship of the two β-strands and the tertiary structure of the pGlu3-Aβ(3–40) peptides in the fibrils, 13C-13C DARR NMR experiments were conducted with a long mixing time of 500 ms. This allows to observe interresidual contacts between carbons in spatial proximity of up to ~6 Å, via the detection of cross peaks between carbons of different amino acids. For mature Aβ(1–40) fibrils as well as oligomers and protofibrils, a contact between the side chains of Phe19 and Leu34 has been well-described16,17,18,28,29. This contact indicates the close proximity between the two β-strands of the monomer and the U-shaped structure of the monomers in the fibrils. In the DARR NMR spectrum for pGlu3-Aβ(3–40) fibrils (Figure S1), cross peaks between Phe19 and Leu34 are clearly visible, especially between the aromatic ring carbons of Phe19 and the Cβ signal of Leu34. This suggests a close structural relationship between pGlu3-Aβ(3–40) and WT Aβ(1–40) fibrils also on the tertiary structure level.
It was shown that a molecular contact between Glu22 and Ile31 indicates earlier stages of the fibrillation process in oligomers and protofibrils30,31,32, but this contact is absent in mature Aβ fibrils32. Based on these observations and other data, a model for the reorganization of the hydrogen bonds from intramolecular for oligomers and protofibrils to intermolecular hydrogen bonds for the mature fibrils has been proposed30,31. In our 13C-13C DARR NMR spectrum of the peptide that contains these two amino acids 13C/15N-labeled, such a cross peak indicative of the molecular contact was not observed (Figure S2). This result also confirms that fibrils of pGlu3-Aβ(3–40) exhibit a strong structural similarity to mature Aβ(1–40) fibrils.
Solid-state NMR also offers the possibility to investigate the molecular dynamics of the individual segments of the pGlu3-Aβ(3–40) fibrils. Such measurements can provide information about the different domains in fibrillar peptides or proteins and substantially support the structural data21,33,34. We measured the motionally averaged 1H-13C dipolar couplings for each resolved carbon signal in DIPSHIFT experiments and convert these into molecular order parameters. Figure 4 shows the comparison of the order parameters of the backbone Cα of pGlu3-Aβ(3–40) and Aβ(1–40) (data from ref. 21). For most of the investigated amino acids, the values are similar within the error of the measurement. The deviations for Phe4 and Ser8 may be explained by the close proximity to the pyroglutamyl-modified N-terminus, which likely alters the dynamics of this unstructured part of the fibrils. This results in a somewhat higher order parameter, which corresponds to smaller motional amplitudes of the fluctuations of these residues. One further notable exception is the order parameter of Ile31, which is significantly higher in pGlu3-Aβ(3–40) compared to WT Aβ(1–40). This may suggest some importance of this residue for the formation of oligomers and protofibrils as Ile31 appears to be involved in intramolecular contacts of intermediates, but not in mature fibrils. Interestingly, the correlations of the measured order parameters to the only available datasets for mature Aβ(1–40) fibrils21 and Aβ(1–40) protofibrils35 are not as good as observed for the chemical shifts/secondary structure. This may in part reflect the fact that chemical shifts are measured more precisely than motionally averaged dipolar couplings, but could also indicate a dynamic polymorphism that relates to small packing differences of the individual residues in the fibrils.
(A) Comparison of the 13Cα-1H order parameters of pGlu3-Aβ(3–40) (red) and mature Aβ(1–40) (black) fibrils21. Correlation plots of the 13Cα-1H order parameters of pGlu3-Aβ(3–40) with (B) mature Aβ(1–40)21 and (C) with Aβ(1–40) protofibrils35 are shown as insets. The Pearson correlation coefficient is given for both correlation plots.
Conclusion
Overall, we conclude that on the level of the single amino acid, fibrils formed of pGlu3-Aβ(3–40) exhibit a strong similarity in the molecular structure compared to WT mature Aβ fibrils. Our data agree with a recent study that reported, on the basis of H/D exchange NMR, FTIR, and CD measurements that modified pGlu3-Aβ(3–40) and unmodified Aβ(1–40) comprised similar peptide conformations6. In this study, fibrils of pGlu3-Aβ(3–40) of shorter length have been reported, which was not confirmed by our EM data. Although the pGlu modification on the N-terminus of truncated Aβ peptides significantly accelerates the fibrillation, the end product of this structure forming process shows an astonishing similarity to the well described structural features of all mature Aβ fibrils15,16,23. This suggests once more that the physiological effects of the pGlu peptides must be mediated by transient oligomers, which are very difficult to characterize. However, the N-terminus of the pGlu3-Aβ(3–40) fibrils showed dynamical alterations, that may have an effect of the stability of the intermediates as well as the fibrils as speculated before6.
Methods
Sample preparation
Three pGlu3-Aβ(3–40) peptides with uniformly 13C/15N-labeled amino acids in different positions were synthesized using standard Fmoc protocols. The labeling schemes are as follows. Peptide I: Ser8, Val12 Phe19, and Leu34 labeled; Pepitde II: Phe4, Glu11, Gly29, and Val36 labeled; and Peptide III: Asp7, Gly9, Glu22, and Ile31 labeled. The peptides were solubilized in 50 mM Tris buffer (100 mM NaCl, 0.01% NaN3, pH 8) at a concentration of 0.1 mg/ml. For fibrillation, the peptide solutions were incubated at 37 °C and shaken at 230 rpm for 3 weeks.
ThT fluorescence measurements
The fibrillation kinetics of pGlu3-Aβ(3–40) was followed by ThT fluorescence intensity measurements. Buffer conditions for fibrillation were the same as above with additional 20 μM ThT in the incubation solution. Volumes of 150 μl were pipetted into the wells of a 96-well plate, which was placed in a Tecan infinite M200 microplate reader (Tecan Group AG, Männedorf, Switzerland). The temperature was kept at 37 °C and a kinetic cycle was applied, such that a 2 s shaking time (2 mm shaking amplitude) followed by a 5 min waiting time was repeated four times with one additional 2 s shaking at the end and the subsequent fluorescence measurement. Fluorescence excitation was set to 440 nm and emission was measured at 482 nm. The fluorescence intensity was measured in increments of 30 min for an overall time period of 65 h. For comparison, also the kinetics of WT Aβ(1–40) fibrillation was recorded under the same conditions. Data was analyzed using procedures reported in the literature36.
Electron microscopy
The fibril morphology was checked by electron microscopy (EM). Fibril solutions were diluted 1:1 with pure water and 1 μl droplets of this solution were applied on formvar coated copper grids, allowed to dry for about 1 h and negatively stained with 1% uranyl acetate in pure water. Scanning transmission electron micrographs were recorded using Zeiss SIGMA, (Zeiss NTS, Oberkochen, Germany) equipped with a STEM detector and Atlas Software.
Solid-state MAS NMR spectroscopy
For NMR measurements, fibril solutions were ultracentrifuged at ~200,000× g for 4 h at 4 °C. The pellets were lyophilized, rehydrated to 50 wt% H2O, homogenized by several freeze-thaw cycles and finally transferred into 3.2 mm MAS rotors. All MAS NMR experiments were conducted on a Bruker 600 Avance III NMR spectrometer (Bruker BioSpin GmbH, Rheinstetten, Germany) at a resonance frequency of 600.1 MHz for 1H, 150.9 MHz for 13C, and 60.8 MHz for 15N using a triple channel 3.2 mm MAS probe. Typical pulse lengths were 4 μs for 1H and 13C and 5 μs for 15N. 1H-13C and 1H-15N CP contact time were 1 ms at a spin lock field of ~50 kHz. The relaxation delay was 2.5 s. 1H dipolar decoupling during acquisition with a radio frequency amplitude of 65 kHz was applied using Spinal64. The MAS frequency was 11,777 Hz. 13C chemical shifts were referenced externally relative to TMS.
13C-13C DARR NMR spectra and 13C-15N correlation spectra were acquired simultaneously using dual-acquisition20. In the same experiment, a two dimensional 13C-13C DARR NMR spectrum with a mixing time of 500 ms with 128 data points and four identical 15N-13Cα correlation spectra with 32 data points in the indirect dimensions were measured. The 15N-13Cα spectra were processed using NMRPIPE software37.
To determine 1H-13C dipolar couplings, constant time DIPSHIFT experiments38 were performed. For homonuclear decoupling during dipolar evolution a frequency switched Lee-Goldburg (FSLG)39 with an effective radio frequency field of 80 kHz was used. The MAS frequency for DIPHSIFT experiments was 5 kHz. After Fourier transformation in the direct dimension the signal intensities of the dephasing curve for each resolved carbon was simulated and the determined coupling was divided by the known rigid limit values to obtain the order parameters40,41. The temperature for all NMR experiments was 30 °C.
X-ray diffraction measurements
For X-ray diffraction measurements, fibril samples from the MAS rotors were placed on nylon loops (Hampton Research, Aliso Viejo, CA, USA) and mounted onto the goniometer head of a X-ray source (Rigaku copper rotating anode MM007 with 0.8 kW, Tokyo, Japan). The signals were recorded using an image plate detector (Rigaku, Tokyo, Japan) with an exposure time of 180 s at room temperature. Diffraction images were analyzed using ImageJ42.
Additional Information
How to cite this article: Scheidt, H. A. et al. Fibrils of Truncated Pyroglutamyl-Modified Aβ Peptide Exhibit a Similar Structure as Wildtype Mature Aβ Fibrils. Sci. Rep. 6, 33531; doi: 10.1038/srep33531 (2016).
References
Morawski, M. et al. Glutaminyl cyclase in human cortex: correlation with (pGlu)-amyloid-beta load and cognitive decline in Alzheimer’s disease. J. Alzheimers. Dis. 39(2), 385–400 (2014).
Jawhar, S., Wirths, O. & Bayer, T. A. Pyroglutamate amyloid-beta (Abeta): a hatchet man in Alzheimer disease. J. Biol. Chem. 286(45), 38825–38832 (2011).
Gunn, A. P. et al. Amyloid-beta Peptide Abeta3pE-42 Induces Lipid Peroxidation, Membrane Permeabilization, and Calcium Influx in Neurons. J. Biol. Chem. 291(12), 6134–6145 (2016).
Schlenzig, D. et al. N-Terminal pyroglutamate formation of Abeta38 and Abeta40 enforces oligomer formation and potency to disrupt hippocampal long-term potentiation. J. Neurochem. 121(5), 774–784 (2012).
Schilling, S. et al. On the seeding and oligomerization of pGlu-amyloid peptides (in vitro). Biochemistry 45(41), 12393–12399 (2006).
Wulff, M. et al. Enhanced Fibril Fragmentation of N-Terminally Truncated and Pyroglutamyl-Modified Abeta Peptides. Angew. Chem. Int. Ed Engl. 55(16), 5081–5084 (2016).
He, W. & Barrow, C. J. The A beta 3-pyroglutamyl and 11-pyroglutamyl peptides found in senile plaque have greater beta-sheet forming and aggregation propensities in vitro than full-length A beta. Biochemistry 38(33), 10871–10877 (1999).
Dammers, C. et al. Structural Analysis and Aggregation Propensity of Pyroglutamate Abeta(3–40) in Aqueous Trifluoroethanol. PLoS. One. 10(11), e0143647 (2015).
Nussbaum, J. M. et al. Prion-like behaviour and tau-dependent cytotoxicity of pyroglutamylated amyloid-beta. Nature 485(7400), 651–655 (2012).
Schilling, S. et al. Glutaminyl cyclase inhibition attenuates pyroglutamate Abeta and Alzheimer’s disease-like pathology. Nat. Med. 14(10), 1106–1111 (2008).
Dammers, C. et al. Purification and Characterization of Recombinant N-Terminally Pyroglutamate-Modified Amyloid-beta Variants and Structural Analysis by Solution NMR Spectroscopy. PLoS. One. 10(10), e0139710 (2015).
Sun, N. et al. Structural analysis of the pyroglutamate-modified isoform of the Alzheimer’s disease-related amyloid-beta using NMR spectroscopy. J. Pept. Sci. 18(11), 691–695 (2012).
Matos, J. O., Goldblatt, G., Jeon, J., Chen, B. & Tatulian, S. A. Pyroglutamylated amyloid-beta peptide reverses cross beta-sheets by a prion-like mechanism. J. Phys. Chem. B 118(21), 5637–5643 (2014).
Tycko, R. Solid-state NMR studies of amyloid fibril structure. Annu. Rev. Phys. Chem. 62, 279–299 (2011).
Petkova, A. T. et al. A structural model for Alzheimer’s beta -amyloid fibrils based on experimental constraints from solid state NMR. Proc. Natl. Acad. Sci USA 99(26), 16742–16747 (2002).
Bertini, I., Gonnelli, L., Luchinat, C., Mao, J. & Nesi, A. A New Structural Model of Abeta40 Fibrils. J. Am. Chem. Soc. 133, 16013–16022 (2011).
Paravastu, A. K., Leapman, R. D., Yau, W. M. & Tycko, R. Molecular structural basis for polymorphism in Alzheimer’s beta-amyloid fibrils. Proc. Natl. Acad. Sci. USA 105(47), 18349–18354 (2008).
Adler, J., Scheidt, H. A., Kruger, M., Thomas, L. & Huster, D. Local interactions influence the fibrillation kinetics, structure and dynamics of Abeta(1–40) but leave the general fibril structure unchanged. Phys. Chem. Chem. Phys. 16(16), 7461–7471 (2014).
Sunde, M. et al. Common core structure of amyloid fibrils by synchrotron X-ray diffraction. J. Mol. Biol. 273(3), 729–739 (1997).
Gopinath, T. & Veglia, G. Dual acquisition magic-angle spinning solid-state NMR-spectroscopy: simultaneous acquisition of multidimensional spectra of biomacromolecules. Angew. Chem. Int. Ed Engl. 51(11), 2731–2735 (2012).
Scheidt, H. A., Morgado, I., Rothemund, S. & Huster, D. Dynamics of Amyloid beta Fibrils Revealed by Solid-State NMR. J. Biol. Chem. 287(3), 2017–2021 (2012).
Wishart, D. S. & Sykes, B. D. Chemical shifts as a tool for structure determination. Methods Enzymol. 239, 363–392 (1994).
Paravastu, A. K., Petkova, A. T. & Tycko, R. Polymorphic fibril formation by residues 10-40 of the Alzheimer’s beta-amyloid peptide. Biophys. J. 90(12), 4618–4629 (2006).
Petkova, A. T. et al. Self-propagating, molecular-level polymorphism in Alzheimer’s beta-amyloid fibrils. Science 307(5707), 262–265 (2005).
Sawaya, M. R. et al. Atomic structures of amyloid cross-beta spines reveal varied steric zippers, Nature 447(7143), 453–457 (2007).
Lu, J. X. et al. Molecular Structure of beta-Amyloid Fibrils in Alzheimer’s Disease Brain Tissue. Cell 154(6), 1257–1268 (2013).
Schutz, A. K. et al. Atomic-resolution three-dimensional structure of amyloid beta fibrils bearing the Osaka mutation. Angew. Chem. Int. Ed Engl. 54(1), 331–335 (2015).
Lührs, T. et al. 3D structure of Alzheimer’s amyloid-beta(1–42) fibrils. Proc. Natl. Acad. Sci. USA 102(48), 17342–17347 (2005).
Ahmed, M. et al. Structural conversion of neurotoxic amyloid-beta(1–42) oligomers to fibrils, Nat. Struct. Mol. Biol. 17(5), 561–567 (2010).
Hoyer, W., Gronwall, C., Jonsson, A., Stahl, S. & Härd, T. Stabilization of a beta-hairpin in monomeric Alzheimer’s amyloid-beta peptide inhibits amyloid formation. Proc. Natl. Acad. Sci. USA 105(13), 5099–5104 (2008).
Sandberg, A. et al. Stabilization of neurotoxic Alzheimer amyloid-beta oligomers by protein engineering. Proc. Natl. Acad. Sci. USA 107(35), 15595–15600 (2010).
Scheidt, H. A., Morgado, I. & Huster, D. Solid-State NMR Reveals a Close Structural Relationship between Amyloid β Protofibrils and Oligomers. J. Biol. Chem. 287(26), 22822–22826 (2012).
Sackewitz, M. et al. Structural and dynamical characterization of fibrils from a disease-associated alanine expansion domain using proteolysis and solid-state NMR spectroscopy. J. Am. Chem. Soc. 130(23), 7172–7173 (2008).
Helmus, J. J., Surewicz, K., Surewicz, W. K. & Jaroniec, C. P. Conformational flexibility of Y145Stop human prion protein amyloid fibrils probed by solid-state nuclear magnetic resonance spectroscopy. J. Am. Chem. Soc. 132(7), 2393–2403 (2010).
Scheidt, H. A., Morgado, I., Rothemund, S., Huster, D. & Fändrich, M. Solid-State NMR Spectroscopic Investigation of Aβ Protofibrils: Implication of a β-Sheet Remodeling upon Maturation into Terminal Amyloid Fibrils. Angew. Chem. Int. Ed Engl. 50(12), 2837–2840 (2011).
Nielsen, L. et al. Effect of environmental factors on the kinetics of insulin fibril formation: elucidation of the molecular mechanism. Biochemistry 40(20), 6036–6046 (2001).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes, J. Biomol. NMR 6(3), 277–293 (1995).
Munowitz, M. G., Griffin, R. G., Bodenhausen, G. & Huang, T. H. Two Dimensional Rotational Spin-Echo Nuclear Magentic Resonance in Solids: Correlation of Chemical Shift and Dipolar Interactions. J. Am. Chem. Soc. 103(10), 2529–2533 (1981).
Bielecki, A., Kolbert, A. C. & Levitt, M. H. Frequency-Switched Pulse Sequences: Homonuclear Decoupling and Dilute Spin NMR in Solids. Chemical Physics Letter 155(4-5), 341–346 (1989).
Barre, P., Zschornig, O., Arnold, K. & Huster, D. Structural and dynamical changes of the bindin B18 peptide upon binding to lipid membranes. A solid-state NMR study. Biochemistry 42(27), 8377–8386 (2003).
Huster, D. Investigations of the structure and dynamics of membrane-associated peptides by magic angle spinning NMR. Prog. Nucl. Magn. Reson. Spectrosc. 46, 79–107 (2005).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9(7), 671–675 (2012).
Chimon, S. et al. Evidence of fibril-like beta-sheet structures in a neurotoxic amyloid intermediate of Alzheimer’s beta-amyloid. Nat. Struct. Mol. Biol. 14(12), 1157–1164 (2007).
Sarkar, B. et al. Significant Structural Differences between Transient Amyloid-beta Oligomers and Less-Toxic Fibrils in Regions Known To Harbor Familial Alzheimer’s Mutations. Angew. Chem. Int. Ed Engl. 53(27), 6888–6892 (2014).
Acknowledgements
The authors acknowledge inspiring discussions with Prof. S. Rossner. The study was supported by the DFG (TRR-SFB 102, A06).
Author information
Authors and Affiliations
Contributions
H.A.S. and D.H. designed the study, H.A.S. performed and analyzed the NMR experiments. J.A. performed and analyzed the fluorescence and X-ray experiments. M.K. performed the EM experiments. H.A.S. and D.H. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Scheidt, H., Adler, J., Krueger, M. et al. Fibrils of Truncated Pyroglutamyl-Modified Aβ Peptide Exhibit a Similar Structure as Wildtype Mature Aβ Fibrils. Sci Rep 6, 33531 (2016). https://doi.org/10.1038/srep33531
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33531
This article is cited by
-
Thermophoretic trap for single amyloid fibril and protein aggregation studies
Nature Methods (2019)
-
Effects of Prion Protein on Aβ42 and Pyroglutamate-Modified AβpΕ3-42 Oligomerization and Toxicity
Molecular Neurobiology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.