Abstract
The diverse and complex developmental mechanisms of segmentation have been more thoroughly studied in arthropods, vertebrates and annelids—distantly related animals considered to be segmented. Far less is known about the role of “segmentation genes” in organisms that lack a segmented body. Here we investigate the expression of the arthropod segment polarity genes engrailed, wnt1 and hedgehog in the development of brachiopods—marine invertebrates without a subdivided trunk but closely related to the segmented annelids. We found that a stripe of engrailed expression demarcates the ectodermal boundary that delimits the anterior region of Terebratalia transversa and Novocrania anomala embryos. In T. transversa, this engrailed domain is abutted by a stripe of wnt1 expression in a pattern similar to the parasegment boundaries of insects—except for the expression of hedgehog, which is restricted to endodermal tissues of the brachiopod embryos. We found that pax6 and pax2/5/8, putative regulators of engrailed, also demarcate the anterior boundary in the two species, indicating these genes might be involved in the anterior patterning of brachiopod larvae. In a comparative phylogenetic context, these findings suggest that bilaterians might share an ancestral, non-segmental domain of engrailed expression during early embryogenesis.
Similar content being viewed by others
Introduction
Annelids (e.g. earthworms), arthropods (e.g. insects) and vertebrates (e.g. humans) are bilaterally symmetric animals that have a major part of their adult body organized into segments—sophisticated morphological units repeated along the anteroposterior axis1. The developmental mechanisms of segmentation are diverse and complex, but arthropods and vertebrates do share some molecular similarities in the patterning of their body segments2,3,4,5,6,7. These findings stimulated a debate about the evolution of segmentation that resulted in two conflicting evolutionary hypotheses. Either the similarities are the outcome of common descent, and thus support the homology of bilaterian body segments2,8,9,10, or they represent evolutionary convergences11,12,13.
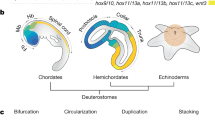
These hypotheses, however, are based on the examination of distantly related groups. Annelids, arthropods and vertebrates belong to distinct branches of the Bilateria, Spiralia, Ecdysozoa and Deuterostomia, respectively (Fig. 1a). This sole comparison over long evolutionary distance can be misleading to reconstruct the evolution of segmentation, because the ancestral conditions within more closely related taxa remain unknown. Thus, distinguishing homology from convergence requires a robust phylogenetic framework and dense taxonomic sampling11,14. To better understand the evolution of segmentation, it is necessary to investigate the related groups of annelids, arthropods and vertebrates. Most of these lineages do not have body segments, but can display a variety of serially repeated structures, such as neural ganglia, excretory organs, coeloms, and others1,12,15. Since segmentation is not an “all-or-nothing” trait15,16, further data on the morphology and gene networks in these lineages can help to elucidate the evolution of the developmental mechanisms of segmentation.
(a) Phylogenetic tree based on recent data29,60,61,74 highlighting the three major bilaterian clades, Deuterostomia, Ecdysozoa and Spiralia. Groups traditionally regarded as truly segmented are marked in bold. Brachiopoda is highlighted in orange. (b) Ectodermal and mesodermal larval traits of the brachiopods T. transversa and N. anomala, as representatives of the Rhynchonelliformea and Craniiformea47.
A crucial mechanism of segmentation is the ability to establish tissue boundaries during animal development17. One of the best studied developmental boundaries are the parasegmental boundaries of the fruit fly Drosophila melanogaster. Fly segmentation occurs by the simultaneous division of the trunk into fourteen parasegments, developmental regions that lie out of register with the larval segments18. At the molecular level, the parasegment boundary is established by the abutting domains of engrailed (en) and wingless (wnt1)19, and is maintained by a positive feedback loop mediated by hedgehog (hh)20. Therefore, en, wnt1 and hh are commonly referred to as segment polarity genes21,22,23, and their expression and function is conserved among the body segments of other arthropods7,24,25,26. The expression of segment polarity genes in annelids is more variable and, in general, do not suggest a role in the patterning of body segments5,27. So far, the only exception is the annelid Platynereis dumerilii, where the expression of en, wnt1 and hh correlates with the segmental boundaries10,28. Thus, to better understand the ancestral developmental functions of the segment polarity genes in the spiralians, it is important to investigate gene expression in groups more closely related to annelids.
Brachiopods are bivalved marine invertebrates with close relations to other lophophorates, nemerteans, annelids and molluscs29 (Fig. 1a). Even though adult brachiopods do not have body segments, the larval stages show putative repeated structures in ectodermal and mesodermal tissues. Externally, brachiopod larvae exhibit two to four lobes along the anteroposterior axis that are divided by transverse ectodermal boundaries. These lobes were once homologized to annelid segments30, but the observation that the ectodermal boundaries do not involve the underlying mesoderm weakened this hypothesis31. The mesoderm morphology is variable between brachiopod species, it can be partitioned into two or four pairs of coelomic sacs32,33 or not subdivided. The discovery of a brachiopod larvae with serially arranged coelomic sacs33 has revived the idea that brachiopods had a segmented ancestor34,35. Therefore, these putative segmented structures, and the closer phylogenetic position to annelids, place brachiopods as an interesting group to test the involvement of the segment polarity genes in other developmental boundaries, and to better comprehend the evolution of segmentation mechanisms in protostomes.
To investigate the developmental and molecular features of the boundaries in brachiopod embryos, we studied the trilobed larva of Terebratalia transversa36, and the bilobed larva of Novocrania anomala33,37, species that belong to distinct brachiopod lineages (Fig. 1b). In these two species, we analyzed the expression of the segment polarity genes en, wnt1, and the core components of the Hedgehog signaling pathway to test whether their expression correlate with the development of the ectodermal and mesodermal boundaries of T. transversa and N. anomala larvae. Furthermore, we examined upstream control genes of en and discovered similarities between the molecular profile of a brachiopod larval boundary and the embryonic patterning of anterior boundaries in deuterostomes, such as the hemichordate collar/trunk boundary and the vertebrate fore/midbrain boundary, further suggesting a non-segmental ancestral role of en for bilaterians.
Results
Ectodermal and mesodermal boundaries of larval brachiopods
In T. transversa, gastrulation occurs by invagination and results in a radially symmetric gastrula that elongates in the anteroposterior axis (Fig. 2a–c). A transverse ectodermal furrow (indentation of the epidermis) appears above the midline of the bilateral gastrula stage, extending from the blastopore lips to the dorsal side (Fig. 2c), with epidermal cells posterior to the indentation being more elongated (Supplementary Fig. S1). The ectodermal furrow and cell shape differences at the bilateral gastrula are the first morphological manifestation of the boundary between the future apical and mantle lobes in T. transversa (apical/mantle boundary). At subsequent developmental stages, this furrow deepens and clearly divides the apical lobe from the remainder of the embryo (Fig. 2d).
Developmental stages of the trilobed larva of T. transversa (a–e) and the bilobed larva of N. anomala (f–j). Each panel shows a ventral view (top) and lateral view (bottom) of a maximum intensity projection of embryos stained with DAPI. Anterior is top in all panels and ventral is to the right in all lateral views. White arrowheads mark the apical/mantle boundary and black arrowheads mark the mantle/pedicle boundary. al: apical lobe, ml: mantle lobe, pl: pedicle lobe. Scale bars = 20 μm.
T. transversa exhibits a third lobe at the posterior end, the pedicle lobe (Fig. 2e). The boundary between the mantle and pedicle lobes (mantle/pedicle boundary) is identified by the narrowing of the posterior portion of the embryo (Fig. 2d) and by the subsequent outgrowth of the mantle lobe (Fig. 2e). The morphology of the mantle/pedicle boundary differs from the apical/mantle boundary because it is demarcated by an ectodermal fold, rather than by an indentation furrow. For this reason, the ectodermal boundaries of the trilobed brachiopod larva are not repeated, but unique structures of the larval body. In N. anomala, the apical and mantle lobes are also demarcated by an ectodermal furrow, as seen in T. transversa (Fig. 2f–i), but its larva does not form a pedicle lobe (Fig. 2i). Thus, the morphology of the apical/mantle boundary is conserved between both species of brachiopods.
The mesoderm of T. transversa has prominent projections associated to the chaetae sacs in the mantle lobe, but there are no partitions (Supplementary Fig. S2). In contrast, the mesoderm of N. anomala has lateral constrictions that individualize four pairs of coelomic sacs, but the posterior pouches are also tightly associated to the three pairs of dorsal chaetae sacs, while remaining interconnected more medially in the ventral side (Supplementary Fig. S2).
Expression of the segment polarity genes hh, en and wnt1
To compare the ectodermal and mesodermal boundaries of brachiopods with the segment boundaries of arthropods and annelids, we analyze the expression of the segment polarity genes en, wnt1 and hh during the embryonic development of T. transversa and N. anomala. Expression of hh localizes to the blastoporal lip and invaginating endomesoderm during early gastrulation, and is restricted to the endoderm in later stages (Figs 3a and 4a). In N. anomala, we detected an additional transverse ventral domain of hh near the animal pole that disappears in the bilobed stage (Fig. 4a). Since the Hedgehog ligand can signal across embryonic layers, we further analyzed the expression of the Hedgehog receptors ptc and smo and the transcription factor gli. Transcripts are expressed in the mesoderm of T. transversa and N. anomala, as well as in the ectodermal apical and mantle territories (Supplementary Figs S3 and S4). Altogether, the expression of hh and downstream pathway components does not correlate spatially or temporally with the development of the larval ectodermal boundaries of T. transversa and N. anomala.
Anterior is top in all panels and ventral is to the right in all lateral views. (a) Expression of hh. (b) Expression of en. (c) Expression of wnt1. Line drawings represent expression at the bilateral gastrula stage. White arrowheads mark the apical/mantle boundary and black arrowheads mark the mantle/pedicle boundary. Double arrowheads mark the ectodermal expression of en. Asterisk marks area of unspecific staining due to the rudiment of the larval shell (b, trilobed). cs: chaetae sacs. Scale bars = 20 μm.
We first detect transcripts of en in T. transversa in the radial gastrula stage. The expression forms an almost-complete ring in the midregion of the embryo and, progressively fades from the anterior and posterior end (Supplementary Fig. S5), leaving a pair of lateral domains of en in the asymmetric gastrula stage (Fig. 3b). These domains extend ventrally and dorsally without reaching the blastopore on the ventral side of the bilateral gastrula stage (Fig. 3b). Transcripts of en border the apical/mantle furrow in the subsequent stage and fade in the trilobed larvae (Fig. 3b). In N. anomala a single dorsal ectodermal stripe of en expression occurs in the radial gastrula (Supplementary Fig. S5) and a second stripe emerges at the posterior end of the asymmetric gastrula (Fig. 4e). Similar to T. transversa, en stripes extend ventrally without reaching the embryo midline and fade in the developed larva (Fig. 4e). Therefore, expression of en is consistent between the two species, with the apical/mantle furrow forming immediately anterior to a pair of en stripes.
In addition to the striped domains demarcating the apical/mantle boundary, we identified two more posterior en territories on the ventral and dorsal regions of larval brachiopods, localized in the anterior-most region of the pedicle lobe. One is located on the ventral side of T. transversa and consists of a pair of en ectodermal bands associated with the blastopore in the bilateral gastrula stage (Fig. 3b). The other is located on the opposite side, at the center of the dorsal surface, as a triangular-shaped domain of en expression (Supplementary Fig. S5). A correspondent domain also occurs on the dorsal side of N. anomala and localizes to the shell rudiment in the bilobed larva stage (Supplementary Fig. S5).
Mesodermal expression of en initiates later than the ectodermal domains. In T. transversa, transcripts of en form a pair of bands in the pedicle mesoderm that are connected to the ectodermal domains, while the mesodermal expression in N. anomala localizes directly inner to the lateral ectodermal domains of en (Supplementary Fig. S6). Expression of en is located to the posterior portion of the second and third pair of coelomic sacs, but not in the first or fourth (Supplementary Fig. S6). Overall, the mesodermal expression of en occurs in contiguity with the preceding ectodermal domains in both species.
Expression of wnt1 in the asymmetric gastrula of T. transversa is associated with the posterior portion of the blastopore (Fig. 3c) and shows no spatial correlation to the lateral en domains (Fig. 5a). At the bilateral gastrula, a transverse pair of stripes of wnt1 expression appears in the apical lobe of T. transversa, bordering the apical/mantle furrow anteriorly (Fig. 3c). The expression abuts en at the apical/mantle boundary without overlap (Fig. 5b). As the apical/mantle furrow deepens in the bilobed larva, wnt1 and en transcripts resolve into well-defined non-overlapping stripes in T. transversa (Fig. 5c). The stripes demarcate precisely the morphological furrow at the apical/mantle boundary of T. transversa with wnt1 positioned anterior and en positioned posterior to the furrow (Fig. 5c). In contrast, the posterior domain of wnt1 develops a tight expression overlap with the pedicle lobe domain of en, on the ventral side of T. transversa (Fig. 5b,c). Such precise coexpression pattern also occurs on the dorsal side, when wnt1 transcripts are first detected within the triangle-shaped territory of en (Fig. 5c).
(a) Asymmetric gastrula. Same specimens of Fig. 2b. (b) Bilateral gastrula. Same specimens of Fig. 2c. (c) Bilobed larva. Ventral view is the same specimen of Fig. 2d. Ventral and lateral views are maximum intensity projections. Optical sections show additional counter staining with DAPI (gray). Merge (yellow) highlights the areas of en and wnt1 co-expression. Scale bars = 20 μm.
N. anomala does not exhibit wnt1 domains in the apical lobe. The transcripts are exclusively at the posterior end, associated with the blastopore (Fig. 4c). We did not detect any coexpression sites for en and wnt1 in N. anomala. Thus, the expression of wnt1 at the apical/mantle boundary is not consistent between brachiopods, and only T. transversa has wnt1 stripes demarcating the morphology of the furrow.
Expression of the putative engrailed regulators pax6, pax2/5/8 and fgf8/17/18
The only gene consistently related to the apical/mantle boundary in both brachiopod larvae is en. Its early expression, however, does not encircle the circumference of the embryo (Fig. 3b), suggesting that there must be upstream factors positioning the apical/mantle boundary. Regulators of en known from D. melanogaster segmentation, such as even-skipped or sloppy paired38,39, do not correlate to the en domains at the apical/mantle boundary of T. transversa40. Therefore, we selected further candidate genes related to the patterning of the vertebrate neuroectoderm, where en plays a crucial role in the establishment of boundaries41. In the mid/hindbrain boundary, en is upregulated by pax2 in a positive feedback loop, represses and is repressed by pax6, and, together with fgf8, is essential to maintain the boundary between the diencephalon and mesencephalon42,43,44,45. Thus, we analyzed the expression of the correspondent brachiopod orthologs pax6, pax2/5/8 and fgf8/17/18 in relation to the expression of en and the development of the apical/mantle boundary.
Transcripts of pax6 form a pair of broad anterior domains that occupy the entire anterior half of T. transversa radial gastrula, and localize to the apical lobe in subsequent stages (Fig. 6a). In N. anomala, pax6 expression initiates in the radial gastrula as an apical ring and also localizes to the apical lobe in later stages (Fig. 7a). In both species the posterior limit of pax6 expression borders the apical/mantle furrow.
(a) Expression of pax6. (b) Expression of pax2/5/8. (c) Expression of fgf8/17/18. Anterior is top in all panels and ventral is to the right in all lateral views. Line drawings represent expression at the bilateral gastrula stage. White arrowheads mark the apical/mantle boundary and black arrowheads mark the mantle/pedicle boundary. Scale bars = 20 μm.
T. transversa pax2/5/8 expression begins as a posterior half-ring in the radial gastrula (Fig. 6b) that overlaps with the pax6 territory at the future apical/mantle boundary (Fig. 8a). The transcripts cover the whole extension of the mantle ectoderm in the asymmetric gastrula, except for the chaetae sac primordia and blastoporal lips (Fig. 6b), and contain the lateral domains of en (Fig. 8d). The anterior limit of pax2/5/8 expression crosses the apical/mantle furrow (Fig. 6b) and overlaps with pax6 expression at the posteriormost region of the apical lobe (Fig. 8b,c). We corroborated this anterior limit of pax2/5/8 transcripts by the combined analysis with en expression (Fig. 8e,f); see Supplementary Fig. S7 for additional data. In N. anomala, pax2/5/8 is expressed in a posterodorsal territory in the mantle lobe, limited by the apical lobe (Fig. 7b). Different than T. transversa, we observe mesodermal expression of pax2/5/8 in two pairs of domains between the chaetae sacs of N. anomala (Fig. 7b). In summary, pax6 and pax2/5/8 are expressed in complementary domains, with a narrow overlap, that encircle the whole embryo before the lateral patches of en form transverse stripes. Thus, the apical/mantle furrow in both species is demarcated by the posterior limit of pax6 expression with abutting domains of en in the mantle lobe.
Whole mount double fluorescent in situ hybridization of pax6 + pax2/5/8 (a–c) and en + pax2/5/8 (d–f) in the brachiopod T. transversa. (a,d) Asymmetric gastrula in lateral view. (b,e) Bilateral gastrula in ventral view. (c,f) Bilobed larva in ventral view. Anterior is top in all panels and ventral is to the right in all lateral views. Side panels show the relation between gene expression and the apical/mantle boundary (striped yellow line). White arrowheads mark the apical/mantle boundary and black arrowheads mark the mantle/pedicle boundary. Scale bars = 20 μm.
Transcripts of fgf8/17/18 form a pair of transverse ventral bands in the apical lobe of T. transversa asymmetric gastrula (Fig. 6c). These bands do not border the stripes of en posterior to the furrow (Supplementary Fig. S7). In addition, developing chaetae sacs express fgf8/17/18 (Fig. 6c) and are cleared from en and pax2/5/8 expression (Supplementary Fig. S7). In N. anomala, fgf8/17/18 expression initiates in the asymmetric gastrula with a similar anterior ventral band and a posterior domain encircling blastopore (Fig. 7c). This posterior demarcates the anterior-most region of the mantle lobe at the bilateral gastrula but fades in the bilobed larva (Fig. 7c). Similarly to T. transversa, chaetae primordia express fgf8/17/18 (Fig. 7c). Thus, the expression of fgf8/17/18 is mainly associated with the chaetae sacs in both species and does not show a clear correlation with ectodermal boundaries.
Over-activation of the Wnt pathway in T. transversa
The pervading role of Wnt signaling in the axial patterning of metazoans46, led us to investigate the role of the canonical Wnt pathway in the development of T. transversa. We found that the over-activation of Wnt signaling prevents the formation of the apical/mantle furrow (Supplementary Fig. S8). The posterior larval structures are expanded, the mantle lobe is not formed, and the expression of en, wnt1 and pax6 shifts more anteriorly (Supplementary Fig. S9). Early treatments entirely abolish the anterior domains of en and wnt1, but do not suppress the expression of pax6 (Supplementary Fig. S9).
Discussion
The evolutionary lineages of T. transversa (Rhynchonelliformea) and N. anomala (Craniiformea) diverged at least 500 million years ago47. It has been hypothesized that planktotrophy is ancestral for the Brachiopoda48 and that larvae of rhynchonelliforms and craniiforms might have evolved a lecithotrophic mode of development independently49,50. However, there are no extant planktotrophic larvae in the rhynchonelliforms, and for this reason, it is not possible to infer if the planktotrophy found in the larvae of extant linguliforms—the sister group of the craniiforms—is ancestral or derived. In our study, we compared species from the rhynchonelliforms and craniiforms to determine which traits are conserved or derived. We found that the trilobed larva of T. transversa and the bilobed larva of N. anomala share an apical/mantle boundary with a conserved ectodermal furrow morphology. A similar ectodermal furrow also delimits the apical lobe of the planktotrophic larva of Lingula anatina51, suggesting the apical/mantle boundary is an ancestral feature of brachiopod embryogenesis.
We found that T. transversa and N. anomala larvae show surprisingly consistent patterns of gene expression. Most expression domains in a species have a reciprocal, similarly positioned territory in the other species. In the trilobed larva of T. transversa, for example, en is expressed in a pair of anterior domains that correlate closely with the apical/mantle boundary, and in a pair of posterior domains bordering the mantle/pedicle boundary. The bilobed larva of N. anomala not only exhibits the correspondent en domains lining its apical/mantle boundary, but also the reciprocal posterior pair. We do not interpret these domains in N. anomala as a vestigial expression of en. First, in known cases of vestigial expression, an incipient aspect of the morphology is still present52,53. But there is no morphological evidence of a differentiated pedicle lobe in N. anomala. Second, the evolutionary origin of the pedicle lobe remains unresolved and it is unclear if it is an ancestral feature of brachiopod larvae. Nonetheless, the consistent expression patterns between T. transversa and N. anomala allow us to suggest that the presence of two pairs of lateral en domains is the ancestral condition for brachiopods, independent of the trilobed or bilobed larval morphology.
In order to better understand the evolution of en expression across bilaterians, we compared the brachiopod data with the available data from other animals (Fig. 9). We found that most domains correlate to some kind of epithelial boundary such as segment boundaries of annelids, shell borders in molluscs, arthropod segment boundaries, seam cells in nematodes, collar/trunk boundary in hemichordates, brain boundaries in vertebrates and somite boundaries in cephalochordates (Supplementary Table S1). A closer look reveals that at its earliest instance of embryonic expression, most groups have an ectodermal expression of en in laterodorsal domains (Fig. 9 and Supplementary Table S1). In annelids and brachiopods, a pair of lateral en domains are located at the head/trunk and apical/mantle boundary, respectively. In D. melanogaster the first stripe of en occurs at the cephalic furrow54 and in hemichordates en is expressed at the collar/trunk boundary55. This comparative data suggests that en expression is mainly ectodermal and might have been associated to the interface between an anterior (e.g. head) and a posterior region of the embryo (e.g. trunk), a division shared by most protostomes and deuterostomes.
Phylogenetic tree based on recent data29,60,61,74 and gene expression compiled on the Supplementary Table S1. Boxes filled with color (vermillion) indicate the presence of en expression, either in the ectoderm (left column labeled with “e”) or mesoderm (right column labeled with “m”). If en is expressed in both ectoderm and mesoderm, asterisks indicate the tissue where en is first expressed during embryogenesis. Empty boxes indicate the absence of en expression and boxes with a question mark (“?”) indicate groups where the expression of en is unknown. Drawings show the expression of en (vermillion) in embryonic stages of representative groups. Early and late expression of en was depicted for Onychophora, Arthropoda, Annelida and Brachiopoda. Structures associated with en expression in each group are listed on the right.
In addition, our data indicates that the early patterning of the apical/mantle boundary of brachiopods is characterized by the overlapping expression of pax6 and pax2/5/8 (Fig. 10a). Interestingly, the complementary patterns of pax6 and pax2/5/8 expression during early embryogenesis occur in the ectoderm of hemichordates55, cephalochordates56,57 and in the neuroectoderm of vertebrates43,44 (Fig. 10b), suggesting that deuterostomes share a similar arrangement of gene products along the anteroposterior axis58. No nervous structures have been described at the apical/mantle boundary of larval brachiopods, so far40,59. Thus, the similarity to the brachiopod pattern could indicate that such genes were also involved in the axial patterning of the last common protostome ancestor. Or simply, that they have been independently coopted to these structures. Unfortunately, data on the expression of pax6 and pax2/5/8, specially during early embryogenesis, are still scarce outside arthropods (Supplementary Table S1). Thus, it remains to be determined if the expression patterns of these genes in brachiopods are ancestral to the protostomes.
(a) Expression of genes related to the apical/mantle boundary with consistent expression between the brachiopods T. transversa and N. anomala (pax6, pax2/5/8 and en). (b) Spatial relationship between the expression of pax6, pax2/5/8 and en in the fore/midbrain boundary of the vertebrate brain (chicken)43,44, the collar/trunk boundary in the hemichordate (Saccoglossus kowalevskii)55 and the apical/mantle boundary in the brachiopod larval body (T. transversa).
Overall, the spatial arrangement and temporal expression patterns of our candidate genes suggest the apical/mantle boundary of brachiopods is not patterned by a segment polarity mechanism (Supplementary Fig. S10). Nevertheless, we show that abutting stripes of en and wnt1 are not exclusive for the segment and parasegment boundaries of P. dumerilii28 and arthropods7,19,24,25,26, respectively—it also occurs in a non-segmental boundary of a larval brachiopod. Since the expression of segment polarity genes in annelids is variable and does not support a role in segment formation5, and recent phylogenies suggest the protostome ancestor did not have a segmented trunk29,60, the segment polarity genes might be an arthropod-specific feature. Therefore, we interpret the adjacent expression of en and wnt1 in T. transversa, P. dumerilii and arthropods as the independent recruitment of a common boundary patterning mechanism—not necessarily linked to segmentation—to these developmental boundaries. It remains yet to be found if annelids have a common set of genes patterning their homologous segments. Identifying such annelid-specific segmentation genes, and investigating their expression in other spiralians, will be essential to elucidate the evolution of segmentation mechanisms in the Spiralia, and how it compares to arthropod segmentation.
In conclusion, the conserved expression of pax6, pax2/5/8 and en between T. transversa and N. anomala, and their correlation with the furrow morphology, suggests these genes might be involved in the patterning of the apical/mantle boundary of brachiopods. A broader comparison among bilaterians indicates the ancestral expression of en during early development was non-segmental and putatively related to the embryonic head/trunk boundary. We thus propose that en might have had a role in the early axial patterning of the protostome/deuterostome ancestor. Comparative expression data in the Xenacoelomorpha—the sister group to all remaining bilaterians61—as well as in less-studied protostome groups (e.g. gastrotrichs, rotifers, chaetognaths, nemerteans and priapulids) will be crucial to test this hypothesis.
Materials and Methods
Sample collection
We collected adult brachiopods by dredging rocky ocean floor in Friday Harbor, USA (T. transversa) and Raunefjorden near Bergen, Norway (N. anomala) during reproductive season, December/January and September/October respectively. Fertilization of ripe gametes was conducted in the laboratory as described previously37,62 and the representative developmental stages were fixed in 4% formaldehyde for 1 h, washed in PTw (1× PBS + 0.1% Tween-20) and kept in PTw at 4 °C for antibody staining and in 100% methanol at −20 °C for in situ hybridization.
Immunohistochemistry
We permeabilized the embryos with several washes in PTx (1x PBS + 0.2% Triton X-100) for 2 h and blocked with two washes of 1 h in PTx + 0.1% BSA (Bovine Serum Albumin) succeeded by 1 h incubation in PTx + 5% NGS (Normal Goat Serum). Samples were incubated with the primary antibodies (mouse anti-Tyrosinated Tubulin 1:500 and rabbit anti-Synapsin II 1:500) stored overnight at 4 °C on a nutator. We removed the primary antibodies with three 5 min and four 30 min washes in PTx + 0.1% BSA, blocked in PTx + 5% NGS for 1 h and incubated nutating overnight at 4 °C with the secondary antibodies (Alexa Fluor 594 anti-mouse and Alexa Fluor 647 anti-rabbit 1:200). Secondary antibodies were removed with three 5 min followed by several washes in PTx + 0.1% BSA for 2 h. We stained nuclei by incubating permeabilized embryos in DAPI 1:500 or Sytox Green 1:1000 for 2 h. Nuclei staining was combined with f-actin staining by the addition of BODIPY FL Phallacidin 5 U/mL previously evaporated to remove methanol.
Gene cloning and orthology
We identified orthologous genes by reciprocal BLAST searches using known sequences against the transcriptomes of T. transversa and N. anomala. We performed PCR gene specific primer pairs on cDNA of each brachiopod species synthesized with the SMARTer RACE cDNA Amplification kit (Clontech). We used RACE primers to clone T. transversa en and pax6. All other T. transversa and N. anomala genes were cloned with regular primer pairs (Supplementary Table S2).
Primers were designed with Primer363. Orthology was assigned by aligning amino acid sequences of brachiopods against annotated genes using MAFFT 7.21564, retaining only informative portions of the alignment with GBlocks 0.91b with relaxed parameters65 and running a Maximum Likelihood phylogenetic analysis with RAxML 8.1.1766 using automatic model recognition and rapid bootstrap (Supplementary Fig. S11).
Alignments were manually verified using UGENE67. Resulting trees from the maximum likelihood analysis were rendered into cladograms using the ETE Toolkit68. Source files, alignments and scripts are available online at http://dx.doi.org/10.6084/m9.figshare.1473087.
In situ hybridization
We synthesized antisense DIG-labeled riboprobes with MEGAscript kit (Ambion) and performed colorimetric in situ hybridization according to an established protocol69. Whole mount double fluorescent in situ hybridization was performed with the above protocol, but hybridizing the samples with a DIG-labeled and a DNP-labeled riboprobes. Samples were first incubated overnight at 4 °C with Anti-DIG-POD conjugate diluted 1:250 in blocking buffer. After PTw washes, we developed the reaction with the TSA reagent kit Cy3 (Perkin Elmer). POD activity was inactivated by incubating 45 min in 0.1% H2O2 in PTw at room temperature followed by a 15 min incubation at 67 °C in a detergent solution (50% formamide, 2x SSC, 1% SDS) and incubated overnight at 4 °C with Anti-DNP-POD conjugate diluted 1:100 in blocking buffer. Second probe was developed in the same manner with the TSA reagent kit Cy5 (PerkinElmer).
Imaging and image processing
Specimens were mounted in 80% Glycerol in PBS, 97% 2,2′-Thiodiethanol70,71 or Murray’s Clear (2:1 benzyl benzoate and benzyl alcohol solution) after a quick dehydration series in isopropanol (70%, 85%, 95%, 100%). After colorimetric in situ hybridization we imaged samples with a Zeiss AxioCam HRc mounted on a Zeiss Axioscope A1 using differential interference contrast technique (Nomarski). Fluorescent in situ hybridization and immunostainings were imaged in a Confocal Leica TCS SP5 and the resulting confocal stacks were processed in Fiji72. LUTs are available at http://github.com/nelas/color-blind-luts. We adjusted the levels of the final panels to improve the contrast using Fiji for confocal stacks and GIMP for photomicrographs. We created vector graphics and assembled the figure plates using Inkscape.
Inhibitor experiments with 1-azakenpaullone
We sampled developing embryos of T. transversa from wild type cultures and incubated with a final concentration of 1 and 10 μM 1-azakenpaullone73 diluted in seawater. Embryos were picked at the mid-blastula and radial gastrula stage and fixed for immunohistochemistry and in situ hybridization at the bilateral gastrula and trilobed larval stage. Controls were treated with the highest concentration of dimethyl sulfoxide (DMSO) contained in the experimental samples (1% in seawater).
Additional Information
How to cite this article: Vellutini, B. C. and Hejnol, A. Expression of segment polarity genes in brachiopods supports a non-segmental ancestral role of engrailed for bilaterians. Sci. Rep. 6, 32387; doi: 10.1038/srep32387 (2016).
References
Scholtz, G. The Articulata hypothesis – or what is a segment? Org. Divers. Evol. 2, 197–215 (2002).
Kimmel, C. B. Was Urbilateria segmented? Trends Genet. 12, 329–331 (1996).
Davis, G. K. & Patel, N. H. The origin and evolution of segmentation. Trends Cell Biol. 9, M68–72 (1999).
Patel, N. H. The ancestry of segmentation. Dev. Cell 5, 2–4 (2003).
Seaver, E. C. Segmentation: Mono- or polyphyletic? Int. J. Dev. Biol. 47, 583–595 (2003).
Tautz, D. Segmentation. Dev. Cell 7, 301–312 (2004).
Damen, W. G. M. Evolutionary conservation and divergence of the segmentation process in arthropods. Dev. Dyn. 236, 1379–1391 (2007).
De Robertis, E. M. Evolutionary biology. The ancestry of segmentation. Nature 387, 25–26 (1997).
De Robertis, E. M. The molecular ancestry of segmentation mechanisms. Proc. Natl. Acad. Sci. USA 105, 16411–16412 (2008).
Dray, N. et al. Hedgehog signaling regulates segment formation in the annelid Platynereis . Science 329, 339–342 (2010).
Abouheif, E. et al. Homology and developmental genes. Trends Genet. 13, 432–433 (1997).
Minelli, A. & Fusco, G. Evo-devo perspectives on segmentation: Model organisms, and beyond. Trends Ecol. Evol. 19, 423–429 (2004).
Chipman, A. D. Parallel evolution of segmentation by co-option of ancestral gene regulatory networks. Bioessays 32, 60–70 (2010).
Hejnol, A. & Lowe, C. J. Embracing the comparative approach: How robust phylogenies and broader developmental sampling impacts the understanding of nervous system evolution. Philos. Trans. R. Soc. Lond. B Biol. Sci. 370, (2015).
Budd, G. E. Why are arthropods segmented? Evol. Dev. 3, 332–342 (2001).
Scholtz, G. Deconstructing morphology. Acta Zool. 91, 44–63 (2010).
Dahmann, C., Oates, A. C. & Brand, M. Boundary formation and maintenance in tissue development. Nat. Rev. Genet. 12, 43–55 (2011).
Martinez-Arias, A. & Lawrence, P. A. Parasegments and compartments in the Drosophila embryo. Nature 313, 639–642 (1985).
Ingham, P. W., Baker, N. E. & Martinez-Arias, A. Regulation of segment polarity genes in the Drosophila blastoderm by fushi tarazu and even skipped . Nature 331, 73–75 (1988).
Ingham, P. W. & Martinez Arias, A. Boundaries and fields in early embryos. Cell 68, 221–235 (1992).
Martinez Arias, A., Baker, N. E. & Ingham, P. W. Role of segment polarity genes in the definition and maintenance of cell states in the Drosophila embryo. Development 103, 157–170 (1988).
Ingham, P. W. Segment polarity genes and cell patterning within the Drosophila body segment. Curr. Opin. Genet. Dev. 1, 261–267 (1991).
Tabata, T. & Kornberg, T. B. Hedgehog is a signaling protein with a key role in patterning Drosophila imaginal discs. Cell 76, 89–102 (1994).
Nagy, L. M. Insect segmentation. A glance posterior. Curr. Biol. 4, 811–814 (1994).
Hughes, C. L. & Kaufman, T. C. Exploring myriapod segmentation: The expression patterns of even-skipped, engrailed, and wingless in a centipede. Dev. Biol. 247, 47–61 (2002).
Mellenthin, K. et al. Wingless signaling in a large insect, the blowfly Lucilia sericata: A beautiful example of evolutionary developmental biology. Dev. Dyn. 235, 347–360 (2006).
Seaver, E. C. & Kaneshige, L. M. Expression of ’segmentation’ genes during larval and juvenile development in the polychaetes Capitella sp. i and H. elegans . Dev. Biol. 289, 179–194 (2006).
Prud’homme, B. et al. Arthropod-like expression patterns of engrailed and wingless in the annelid Platynereis dumerilii suggest a role in segment formation. Curr. Biol. 13, 1876–1881 (2003).
Laumer, C. E. et al. Spiralian phylogeny informs the evolution of microscopic lineages. Curr. Biol. 25, 2000–2006 (2015).
Balfour, F. M. A treatise on comparative embryology. (Macmillan; Company, 1880).
Conklin, E. G. The embryology of a brachiopod, Terebratulina septentrionalis Couthouy. Proc. Am. Philos. Soc. 41, 41–76 (1902).
Hyman, L. H. In The invertebrates: Smaller coelomate groups V, 568 (McGraw-Hill Book Company, Inc, 1959).
Nielsen, C. The development of the brachiopod Crania (Neocrania) anomala (O. F. Müller) and its phylogenetic significance. Acta Zool. 72, 7–28 (1991).
Balavoine, G. & Adoutte, A. The segmented Urbilateria: A testable scenario. Integr. Comp. Biol. 43, 137–147 (2003).
Temereva, E. N. & Malakhov, V. V. The evidence of metamery in adult brachiopods and phoronids. Invert Zool 8, 91–112 (2011).
Long, J. A. & Stricker, S. A. In Reproduction of marine invertebrates (eds. Giese, A. C., Pearse, J. S. & Pearse, V. B. ) IV, 47–84 (The Boxwood Press, 1991).
Freeman, G. Regional specification during embryogenesis in the craniiform brachiopod Crania anomala . Dev. Biol. 227, 219–238 (2000).
Cadigan, K. M., Grossniklaus, U. & Gehring, W. J. Localized expression of sloppy paired protein maintains the polarity of Drosophila parasegments. Genes Dev. 8, 899–913 (1994).
Fujioka, M., Yusibova, G. L., Patel, N. H., Brown, S. J. & Jaynes, J. B. The repressor activity of Even-skipped is highly conserved, and is sufficient to activate engrailed and to regulate both the spacing and stability of parasegment boundaries. Development 129, 4411–4421 (2002).
Santagata, S., Resh, C., Hejnol, A., Martindale, M. Q. & Passamaneck, Y. J. Development of the larval anterior neurogenic domains of Terebratalia transversa (Brachiopoda) provides insights into the diversification of larval apical organs and the spiralian nervous system. Evodevo 3, 3 (2012).
Rhinn, M. & Brand, M. The midbrain–hindbrain boundary organizer. Curr. Opin. Neurobiol. 11, 34–42 (2001).
Okafuji, T., Funahashi, J. & Nakamura, H. Roles of Pax-2 in initiation of the chick tectal development. Brain Res. Dev. Brain Res. 116, 41–49 (1999).
Araki, I. & Nakamura, H. Engrailed defines the position of dorsal di-mesencephalic boundary by repressing diencephalic fate. Development 126, 5127–5135 (1999).
Matsunaga, E., Araki, I. & Nakamura, H. Pax6 defines the di-mesencephalic boundary by repressing En1 and Pax2 . Development 127, 2357–2365 (2000).
Scholpp, S., Lohs, C. & Brand, M. Engrailed and Fgf8 act synergistically to maintain the boundary between diencephalon and mesencephalon. Development 130, 4881–4893 (2003).
Petersen, C. P. & Reddien, P. W. Wnt signaling and the polarity of the primary body axis. Cell 139, 1056–1068 (2009).
Bitner, M. A. & Cohen, B. L. In eLS (John Wiley & Sons, Ltd, 2013).
Zimmer, R. L. In Embryology: Constructing the organism (eds. Gilbert, S. F. & Raunio, A. M. ) 279–305 (Sinauer Associates, Inc., 1997).
Freeman, G. & Lundelius, J. W. Changes in the timing of mantle formation and larval life history traits in linguliform and craniiform brachiopods. Lethaia 32, 197–216 (1999).
Freeman, G. & Lundelius, J. The transition from planktotrophy to lecithotrophy in larvae of lower palaeozoic rhynchonelliform brachiopods. Lethaia 38, 219–254 (2005).
Yatsu, N. On the development of Lingula anatina . Journal of the College of Science 17, 1–136 (1902).
Scholtz, G. Expression of the engrailed gene reveals nine putative Segment-Anlagen in the embryonic pleon of the freshwater crayfish Cherax destructor (Crustacea, Malacostraca, Decapoda). Biol. Bull. 188, 157–165 (1995).
Hejnol, A. & Scholtz, G. Clonal analysis of Distal-less and engrailed expression patterns during early morphogenesis of uniramous and biramous crustacean limbs. Dev. Genes Evol. 214, 473–485 (2004).
Fjose, A., McGinnis, W. J. & Gehring, W. J. Isolation of a homoeo box-containing gene from the engrailed region of Drosophila and the spatial distribution of its transcripts. Nature 313, 284–289 (1985).
Pani, A. M. et al. Ancient deuterostome origins of vertebrate brain signalling centres. Nature 483, 289–294 (2012).
Glardon, S., Holland, L. Z., Gehring, W. J. & Holland, N. D. Isolation and developmental e7xpression of the amphioxus Pax-6 gene (AmphiPax-6): Insights into eye and photoreceptor evolution. Development 125, 2701–2710 (1998).
Kozmik, Z. et al. Characterization of an amphioxus paired box gene, AmphiPax2/5/8: Developmental expression patterns in optic support cells, nephridium, thyroid-like structures and pharyngeal gill slits, but not in the midbrain-hindbrain boundary region. Development 126, 1295–1304 (1999).
Lowe, C. J., Clarke, D. N., Medeiros, D. M., Rokhsar, D. S. & Gerhart, J. The deuterostome context of chordate origins. Nature 520, 456–465 (2015).
Altenburger, A. & Wanninger, A. Neuromuscular development in Novocrania anomala: Evidence for the presence of serotonin and a spiralian-like apical organ in lecithotrophic brachiopod larvae. Evol. Dev. 12, 16–24 (2010).
Struck, T. H. et al. Platyzoan paraphyly based on phylogenomic data supports a noncoelomate ancestry of Spiralia. Mol. Biol. Evol. 31, 1833–1849 (2014).
Cannon, J. T. et al. Xenacoelomorpha is the sister group to Nephrozoa. Nature 530, 89–93 (2016).
Freeman, G. Regional specification during embryogenesis in the articulate brachiopod Terebratalia . Dev. Biol. 160, 196–213 (1993).
Untergasser, A. et al. Primer3–new capabilities and interfaces. Nucleic Acids Res. 40, e115–e115 (2012).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577 (2007).
Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Okonechnikov, K., Golosova, O., Fursov, M. & UGENE team. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics 28, 1166–1167 (2012).
Huerta-Cepas, J., Dopazo, J. & Gabaldón, T. ETE: A python Environment for Tree Exploration. BMC Bioinformatics 11, 24 (2010).
Martín-Durán, J. M., Janssen, R., Wennberg, S., Budd, G. E. & Hejnol, A. Deuterostomic development in the protostome Priapulus caudatus . Curr. Biol. 22, 2161–2166 (2012).
Staudt, T., Lang, M. C., Medda, R., Engelhardt, J. & Hell, S. W. 2,2′-thiodiethanol: A new water soluble mounting medium for high resolution optical microscopy. Microsc. Res. Tech. 70, 1–9 (2007).
Asadulina, A., Panzera, A., Verasztó, C., Liebig, C. & Jékely, G. Whole-body gene expression pattern registration in Platynereis larvae. Evodevo 3, 27 (2012).
Schindelin, J. et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Kunick, C., Lauenroth, K., Leost, M., Meijer, L. & Lemcke, T. 1-azakenpaullone is a selective inhibitor of glycogen synthase kinase-3 beta. Bioorg. Med. Chem. Lett. 14, 413–416 (2004).
Dunn, C. W., Giribet, G., Edgecombe, G. D. & Hejnol, A. Animal phylogeny and its evolutionary implications. Annu. Rev. Ecol. Evol. Syst. 45, 371–395 (2014).
Acknowledgements
We thank Chema Martín-Durán for crucial discussions and help with the in situ hybridization experiments, Carmen Andrikou and Kevin Pang for improving the manuscript, Yale Passamaneck, Yvonne Müller, Jonas Bengtsen, Daniel Thiel and Anlaug Boddington for the help with the collections, and Aina Børve for the laboratory guidance. We are thankful to the staff of Friday Harbor Labs and Espeland Marine Biological Station for their logistic support with the brachiopod collections. We also thank Alessandro Minelli for the comments on an earlier draft and two anonymous reviewers for the constructive feedback. The study was funded by the core budget of the Sars Centre and received support from the Meltzer Research Fund.
Author information
Authors and Affiliations
Contributions
B.C.V. and A.H. designed the study and collected the material. B.C.V. performed the experiments and analyzed the data. B.C.V. and A.H. and wrote the manuscript. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Vellutini, B., Hejnol, A. Expression of segment polarity genes in brachiopods supports a non-segmental ancestral role of engrailed for bilaterians. Sci Rep 6, 32387 (2016). https://doi.org/10.1038/srep32387
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep32387
This article is cited by
-
Brachiopod and mollusc biomineralisation is a conserved process that was lost in the phoronid–bryozoan stem lineage
EvoDevo (2022)
-
The evolution of the metazoan Toll receptor family and its expression during protostome development
BMC Ecology and Evolution (2021)
-
Molecular and morphological analysis of the developing nemertean brain indicates convergent evolution of complex brains in Spiralia
BMC Biology (2021)
-
Hox gene expression in postmetamorphic juveniles of the brachiopod Terebratalia transversa
EvoDevo (2019)
-
Embryonic expression of priapulid Wnt genes
Development Genes and Evolution (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.