Abstract
Florfenicol is extensively used in livestock to prevent or cure bacterial infections. However, it is not known whether the administration of florfenicol has resulted in the emergence and dissemination of florfenicol resistance genes (FRGs, including fexA, fexB, cfr, optrA, floR and pexA) in microbial populations in surrounding farm environments. Here we collected soil samples for the detection of FRGs and the residue of florfenicol from six swine farms with the record of florfenicol usage. Quantitative polymerase chain reaction and metagenomic sequencing revealed a significantly higher relative abundance of FRGs in the soils adjacent to the three swine farms where florfenicol was heavily used compared with the other sites. Meanwhile, the detectable levels of florfenicol were also identified in soils from two of these three farms using ultra-performance liquid chromatography tandem mass spectrometry. It appears that amount of florfenicol used on swine farms and the spreading of soils with swine waste could promote the prevalence and abundance of FRGs, including the linezolid resistance genes cfr and optrA, in adjacent soils and agricultural application of swine manure with florfenicol may have caused a residual level of florfenicol in the soils.
Similar content being viewed by others
Introduction
The growing number of bacterial strains carrying antibiotic-resistance genes (ARGs) poses a significant risk to both animal and human health. It is widely believed that the increased abundance of ARGs in the environment is contributing to the emergence of multidrug-resistant pathogens, which might lead to the failure of antibiotic treatment of bacterial infections1. In China, approximately 210,000 metric tons of antibiotics are produced per year, of which, 97,000 metric tons are used for therapy and growth promotion in animal husbandry2. It is estimated that approximately more than half of antibiotics are not absorbed in the animal gut and subsequent selection has given rise to increasing numbers of resistant bacteria in the gastrointestinal tract, providing a potential reservoir for antibiotic resistance genes3. Furthermore, it is likely that many of the unabsorbed antibiotics, along with bacterial antibiotic resistance genes, are excreted into the environment via feces from livestock animals3. These antibiotics and their associated antibiotic resistance genes may accumulate in soils after repeated application of manure3,4. The residue of antibiotics even at low concentrations in the environment is likely to impose selective pressures on environmental microorganisms, which might induce the emergence of diverse ARGs and promote the evolution of novel genes conferring certain antibiotic resistance mechanisms4,5,6,7,8.
Soil is the predominant reservoir for bacteria harboring genes associated with the antibiotic resistance, with a number of antibiotic resistance determinants identified from soil bacteria9 and various resistance bacteria were cultured from soil samples10. Growing evidence shows that antibiotics, along with considerable numbers of antibiotic-resistant bacteria, ARGs and associated mobile genetic elements, are being disseminated into agricultural soils through frequent manure waste application and contamination11,12,13. Fang et al. reported that the contamination of antibiotics and ARGs were detected in soils and the samples were collected from the actual field which had been treated with chicken manure for long term6. Moreover, the abundance of antibiotics and ARGs increased with the extension of greenhouse planting years6. This is likely to result in the ubiquitous pollution of antibiotic resistance in agricultural soils, while posing significant potential risks to the environment and public health. In addition, an ever-increasing number of novel functional resistance genes are being identified from environmental samples using widely available functional metagenomic methodologies14,15,16,17.
Florfenicol, a fluorinated thiamphenicol derivative, is a broad-spectrum antimicrobial agent exclusively approved for use in veterinary medicine18. It has been licensed in China since 1999 for the control of respiratory tract diseases and enteric infections in food-producing animals. However, the excessive use of florfenicol as an antimicrobial chemotherapeutic agent has resulted in bacterial species acquiring resistance to this agent. Since the identification of florfenicol resistance gene floR in the fish pathogen Pasteurella piscicida in 199619, several specific phenicol resistance genes have been reported in florfenicol-resistant bacteria of animal origin. These genes include the phenicol-specific exporter genes fexA, fexB and floR and the multidrug resistance gene cfr, which encodes a 23S rRNA methyltransferase that confers resistance to phenicols as well as four other structurally unrelated antimicrobial groups (lincosamides, oxazolidinones, pleuromutilins and streptogramin A)20. More recently, a novel ATP-binding cassette (ABC) transporter gene, optrA, which confers resistance to phenicols and oxazolidinones, was identified in Enterococcus and Staphylococcus species of both animal and human origin21,22. It is noteworthy that both cfr and optrA confer transferable resistance to linezolid. Linezolid was the first oxazolidinone to be introduced into clinical medicine to treat infections caused by vancomycin-resistant enterococci and methicillin-resistant Staphylococcus aureus23. optrA also confers resistance to tedizolid, which is a newly approved oxazolidinone for the management of human infections associated with Gram-positive pathogens, including linezolid-resistant strains (especially those carrying cfr)24. Although oxazolidinones have not been approved for use in the livestock or aquaculture industries, cfr and optrA are commonly detected in florfenicol-resistant bacteria of animal origin21,25. In addition to fexA, fexB, cfr, optrA and floR, phenicol exporter gene pexA was identified in metagenomic libraries of cloned DNA isolated from Alaskan soils14. Interestingly, all of these genes, apart from pexA14, coexist with bacterial mobile genetic elements such as plasmids, transposons, or integrons21,25,26,27,28, which aid the horizontal transfer of florfenicol resistance genes (FRGs) to numerous bacterial species and genera.
Previous publications have reported the occurrence of chloramphenicol resistance genes, including fexA, fexB, cfr and floR, in association with chloramphenicol residue in wastewater effluent from swine farm operations and corresponding wastewater-irrigated agricultural fields29. However, more than one decade ago, the usage of chloramphenicol in the livestock and aquaculture industries has been completely banned and florfenicol became the only available antimicrobial agent from the phenicol class in China. It is not clear whether the use of florfenicol in agriculture has contributed to the environmental accumulation of florfenicol and FRGs, especially the cfr and optrA genes also conferring resistance to other antimicrobial agents, which are critically important in the human medicine. It is very likely that the frequency of florfenicol usage in swine farms could affect the abundance and prevalence of FRGs and the accumulation of florfenicol in adjacent soils. Thus, we quantified six FRGs (fexA, fexB, cfr, optrA, floR and pexA) using quantitative polymerase chain reaction (qPCR) via a culture-independent method and detected the florfenicol residue concentrations using ultra performance liquid chromatography-tandem mass spectrometry (UPLC–MS/MS) in soils from different farms.
Results
Abundance of FRGs
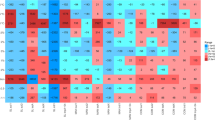
The florfenicol had been used to treat respiratory diseases only in farms HN-S-4, HN-S-5 and HN-S-6, while the florfenicol had been extensively used for prevention purposes in long-standing farms HN-S-1, HN-S-2 and HN-S-3. The approximate annual dosage of florfenicol used in farms HN-S-1, HN-S-2 and HN-S-3 in past three years is much more than that in farms HN-S-4, HN-S-5 and HN-S-6 (Table 1). Correspondently, our quantitative PCR result showed the relative abundance of FRGs from the farms HN-S-4, HN-S-5 and HN-S-6 was much lower than that from the farms HN-S-1, HN-S-2 and HN-S-3 (Fig. S1) and no FRGs was detected in control samples. Of the six farms, the abundance of cfr, optrA and fexA was significantly higher in soils from HN-S-2 and HN-S-3 compared with the other four farms (p < 0.001). Interestingly, fexB was only detected in soils from HN-S-2 and HN-S-3 (Fig. 1), the only two farms with detectable florfenicol residue. Significantly higher levels of floR were detected in samples from the three long-standing farms compared with samples from the newly established farms (p < 0.001), with the highest abundance of floR detected in soils from HN-S-1 (Fig. 1). Additionally, the newly identified gene optrA, which confers transferable resistance to florfenicol and linezolid, was present across all soils adjacent to swine feedlots. Conversely, phenicol exporter gene pexA was not detected in any of the soil samples. Importantly, the relative abundance of the multidrug-resistance genes cfr and optrA was significantly higher in soils from HN-S-2 and HN-S-3 than in the other farms analyzed (p < 0.001).
Relative abundance of five florfenicol resistance genes (a: fexA, b: fexB, c: cfr, d: optrA and e: floR) in the six soil samples (target gene copies/16S rRNA gene copies).
Bars represent the relative abundance of a single florfenicol resistance gene and values shown are mean ± SE of three analytical replicates.
Metagenomic Analysis
The absolute abundance of the six FRGs was further validated using high-throughput sequencing-based metagenomic analysis. In these six soil samples adjacent to swine farms, the number of reads identified as florfenicol resistance gene sequences was used to determine the absolute abundance. There was a significant difference in the log2 values for transformed FRG reads between the two different groups of farms (p < 0.05), which was consistent with the qPCR assays (Fig. 2a,b). Pearson correlation analyses were conducted to investigate potential relationships between the abundance of FRGs and transposases in these soil samples. Transposases, including IS256, IS6100, IS26, IS1216, ISEnfa4 and ISEnfa5, which play an important role in the horizontal transfer of FRGs between animal-associated bacteria and human clinical isolates, were identified in all soil samples. The total number of FRGs in each sample and the abundance of these genes were highly correlated with the levels of transposases in the soil samples (Fig. 2c, r2 = 0.9833, p < 0.0001). Furthermore, the phenicol exporter gene floR was highly correlated with the insertion sequence IS6100 (Fig. 2d, r2 = 0.9912, p < 0.0001).
Abundance of florfenicol resistance genes (fexA, fexB, cfr, optrA, floR) and transposases.
(a) Number of total reads of each of the genes in different soil samples. Red bars represent the sum of florfenicol resistance genes and purple represents transposases. (b) Reads associated with floR and IS6100 gene sequences in each soil sample. Red bars represent floR genes and purple represents IS6100. (c) Correlation between total abundance of florfenicol resistance genes and associated transposases. (d) Correlation of the abundance of floR and IS6100. Sequences were analyzed using BLAT software. All figures and correlation analyses were generated using GraphPad Prism.
Florfenicol Concentrations in Soil Samples
Methods were validated prior to sample analysis. To conform with the relevant validation criteria, the following parameters were used: (a) the matrix-matched calibration curves were linear in the concentration range of 0.25–20 ng g−1, with correlation coefficients (r2) greater than 0.999; (b) the limits of detection (LOD) and quantification (LOQ) were 0.08 ng g−1 and 0.25 ng g−1, respectively; (c) the average recoveries for florfenicol were in the range of 94.20–96.28% with relative standard deviation less than 10.5% (n = 6). Following method validation, florfenicol was detected in the soil samples from farms HN-S-2 (3.730 ± 0.327 ng g−1) and HN-S-3 (0.359 ± 0.028 ng g−1), while concentrations were below the LOQ for all other samples.
Discussion
Our results indicated that the usage of florfenicol in the farms affect the prevalence of FRGs in the soil irrigated with the farm waste. This conclusion could be supported by the following observations: first, both the metagenomic and qPCR via culture-independent methods revealed that FRGs can be detected in the soils adjacent to the farms with florfenicol usage, but not in the control samples. Second, the prevalence and abundance of FRGs maybe affected by the amount of florfenicol used in the farms, as the farms consumed more florfenicol resulted in more detected FRGs in soils. The detectable level of florfenicol in the soils adjacent to farms HN-S-2 and HN-S-3 imply that the unabsorbed florfenicol could be excreted into the environment via manure from livestock animals.
In addition to the usage of florfenicol, many other factors could contribute to prevalence of ARGs in soils. It has been reported that the variety of ARGs and antibiotic residues in agricultural soils are correlated with soil type, manure application rate and environment conditions30,31. The long-term fertilization of antibiotic-contaminated chicken manure in greenhouse soils led to higher levels of ARGs and antibiotic residues in greenhouse soils compared with field soils6. The occurrence of chloramphenicol resistance genes as environmental pollution were found in manured soils29. Several studies also have reported that the abundance and diversity of ARGs have a marked increase in soils after repeated manure application6,32,33, such as multidrug resistance (MDR) genes, macrolide-lincosamide-streptogramin (MLS) genes and sulfonamide resistance genes. Interestingly, following the use of florfenicol in these swine farms, the linezolid resistance gene optrA was detected in all soil samples; and the abundance of optrA was greater in soil samples obtained from farms HN-S-2 and HN-S-3. These were also the only two farms with florfenicol soil concentrations above the limit of quantification. However, oxazolidinones have not been approved for veterinary applications worldwide and florfenicol is the only phenicol-derived antimicrobial agent exclusively used in selected swine farms21. In addition, the results imply that prior to being introduced to the farm environment, in vivo florfenicol can directly select for the propagation of optrA in bacterial isolates of animal origin. This is supported by the finding that this gene is more frequently detected in enterococci isolated from animals than from humans21.
Meanwhile, a rapid UPLC-MS/MS method with a lower LOD (0.08 ng g−1) and LOQ (0.25 ng g−1) values compared with previous methods34,35 was established to determine the levels of florfenicol in soil samples. To date, a number of reports have described the detection of various antibiotic residues, including tetracyclines, (fluoro)quinolones and sulfonamides, in agricultural soils30,31. However, none of these studies described a method that is suitable for the determination of florfenicol in soil using UPLC-MS/MS. To the best of our knowledge, this is the first study to quantify florfenicol residue in soil samples collected from swine production facilities using UPLC-MS/MS. The results showed that florfenicol levels were above the LOQ on two of the long-standing farms (HN-S-2 and HN-S-3). However, the observed lower concentrations of florfenicol in soils was consistent with that published previously35, may be due to the adsorption, biodegradation, photolysis, infiltration of florfenicol6 and the dilution of florfenicol by swine manure in soils. Notably, Subbiah et al. confirmed that the biologically active of florfenicol could remain long-time in soils36 and exert a selective pressure for resistance genes in the environment. In addition, a previous study34 looking at the relative robustness of florfenicol showed that the half-life of florfenicol in native soil is approximately 8 days and the degradation behavior of florfenicol in the soils was connected with the activity of microbe community. Considering the rapid degradation rate of florfenicol in native soils, it is possible that the detection of florfenicol in these soil samples depend on whether the sampling sites were irrigated with florfenicol contaminated manure or not in recent days.
In general, the emergence and spread of antibiotic resistance genes are associated with mobile genetic elements, such as plasmids, integrases and transposases37,38,39. To date, several insertion sequences responsible for the mobility of FRGs, including IS6100, IS26, IS1216, IS256, ISEnfa4 and ISEnfa5, have been widely detected in different Gram-positive and Gram-negative bacteria21,25,26,27,28. This has likely allowed the translocation of resistance genes between different plasmids, as well as mediating their integration into chromosomal DNA25. Importantly, in this current study, according to metagenomic analysis, the abundance of total FRGs (fexA, fexB, cfr, optrA and floR) was significantly correlated with the abundance of insertion sequences (Fig. 2a,c). Of the transposases investigated as part of this study, IS1216 is commonly located adjacent to the linezolid resistance gene optrA in enterococci40, while IS26, IS1216, ISEnfa4 and IS256 have been reported to play an important role in the mobility of the multiresistance gene cfr25. In addition, the most frequently detected transposase, the IS6100 family element, commonly coexists with complex integrons in Gram-negative bacteria and is typically located in the flanking regions of several resistance genes, including floR and tetR27,38,41. We also observed a strong positive correlation between the abundance of floR and the presence of IS6100 in soil samples from each of the tested farms (Fig. 2b,d). All of these results suggest that the amount of florfenicol used in the farm not only play a considerable role in the abundance and diversity of FRGs, but also simultaneously leads to the enrichment of horizontally mobile genetic elements in adjacent soils. Therefore, the effects of the application of biogas slurry and the build-up of manure generated through livestock maintenance on the microbiota of agricultural soils should not be ignored when considering the spread of antibiotic resistance genes and insertion sequences (e.g. IS6100). Indeed, these factors are likely to assist in the migration of FRGs through soil-dwelling bacteria and horizontally transferred mobile genetic elements, facilitating the spread to human-associated bacteria. In this study, the strong correlation between transposase enrichment and the abundance of FRGs suggests that horizontal gene transfer may have aided in the enrichment of resistance genes through the transfer of mobile genetic elements among soil bacteria.
To the best of our knowledge, this is the first study revealing the impact of florfenicol administration on the occurrence of FRGs in soil samples collected from swine farms. Our findings indicate that the amount of florfenicol used in swine husbandry could have played a considerable role in the abundance and diversity of FRGs in adjacent soils via application of swine waste. It is also the first report detailing the occurrence of the florfenicol and linezolid resistance gene optrA in soil samples. The results of this study suggest that soils containing optrA, cfr and other phenicol resistance genes may act as a reservoir for florfenicol resistance. Therefore, the transfer of FRGs is also likely to affect humans that come into contact with the affected livestock, either through the food chain or through further environmental dissemination. It should be note that the selection of florfenicol and linezolid resistance genes optrA and cfr in soils, following the application of manure in swine farms with a history of administration of florfenicol, pose a significant risk to public health.
Methods
Sample Collection
Soil samples were collected from agricultural fields surrounding six swine feedlots in November 2014. The antibiotic use records of the six farms indicated that florfenicol had been used at each of the farms. All of the field soils were irrigated with manure every few days and had been planted with vegetable crops for human consumption. The swine farms were located in six different cities in Henan Province, China (Fig. S2) and general information regarding these pig farms is listed in Table 1. Of the six farms analyzed, HN-S-4, HN-S-5 and HN-S-6 were relatively new farms (less than 3–4 years old), while HN-S-1, HN-S-2 and HN-S-3 had been producing sows for almost two decades. Overall, the long-standing farms HN-S-1/2/3 used more florfenicol (ranging from 0.63–2.88 metric tons per year) for preventing usually once a month in the past three year than other farms HN-S-4/5/6 (ranging from 0.11–0.30 metric tons per year) for treating respiratory diseases but not prevention (Table 1). Five soil samples were collected from different locations (near the pigsty) at each farm at depths of 5–10 cm4 and the soils were irrigated with manure repeatedly in the past few years. Additionally, control soil sample was collected at sites >5km away from the swine farm which was manure/antibiotic free. All samples were stored in an icebox during transfer to the laboratory and were then stored at −20 °C long term. Soil samples collected from the same farm were mixed to form a composite sample and subsequently sieved through a 2.0-mm mesh and frozen at −80 °C for further processing.
Quantification of FRGs
Metagenomic DNA was extracted from 0.25 g of homogenized soil using a PowerSoil DNA Isolation Kit (MO BIO Laboratories Inc., CA, USA) according to the manufacturer’s instructions and stored at −20 °C until use. The DNA extracted from three independent replications of soil samples from each site to minimize any potential DNA extraction bias. qPCR assays were used to detect the presence of six FRGs (fexA, fexB, cfr, optrA, floR and pexA), along with 16S rRNA, as described previously42. The specific primers used to amplify these gene fragments were designed using Primer 3 Plus (Table S1). Positive and negative controls were performed for each PCR reaction. The positive products were purified and ligated into vector pMD19-T (Takara, Dalian, China) and then transformed into Escherichia coli DH5α (Takara). DNA from recombinant plasmids containing target gene inserts was extracted using a Qiagen DNA Mini Kit (Qiagen, Hilden, Germany). The presence of the desired inserts was verified by PCR and sequencing and then the positive plasmids were used to generate a standard curve for qPCR analysis. The qPCR assay was conducted using a QuantStudio 7 Flex Real-Time PCR System (Life Technologies, Carlsbad, CA, USA) with SYBR Premix Ex Taq II (Takara). The 20-μL reaction volume contained the following: 10 μL of SYBR Premix Ex Taq II (Til RNaseH Plus) 2*, 0.8 μL of each primer (10 nmol L−1), 1 μL of template DNA and 7.4 μL of RNase-Free H2O (Takara). The qPCR conditions were: 95 °C for 3 min, followed by 40 cycles of 30 s at 95 °C, 30 s at 60 °C and 30 s at 72 °C. The qPCR amplification efficiency was examined using R2 values (>0.999) for each calibration curve (Table S2). The amplification specificity was verified by performing a melting curve analysis (95 °C for 15 s, 60 °C for 1 min, 95 °C for 15 s) for each qPCR reaction, along with gel electrophoresis. The copy numbers of the target FRGs were quantified using a standard curve. Presence of the 16S rRNA gene was also quantified on the same plate and the results are shown as relative abundance. To ensure reproducibility, three technique replicates for each sample were performed in parallel in each qPCR assays.
Metagenomic Analyses
For the metagenomic analysis, six prepared soil DNA samples were sent to Berry Genomics Company (Beijing, China) for high-throughput sequencing using the HiSeq 2500 platform. The raw data were filtered following removal of low-quality reads. The filtered data were then searched against the nucleotide sequences of six florfenicol resistance genes (fexA, fexB, cfr, optrA, floR and pexA) with a minimum 50-bp overlap length and 95% identity using BLAT software43. The reads that matched the florfenicol resistance gene sequences were extracted, counted and normalized from the total reads for each sample. The reads matching major transposase genes such as IS6100, IS26, IS256, ISEnfa5, ISEnfa4 and IS1216 were also extracted25,27,28,38. The flanking insert sequences were measured following metagenomic sequencing and analysis, as was florfenicol resistance gene abundance.
Extraction Procedures and Sample Preparation
In this study, a method was developed to quantify florfenicol residues in the soil using UPLC-MS/MS. First, soil samples were thawed at room temperature. A 5-g aliquot of soil was then weighed and 4 mL of ammonia-ethyl acetate (2 + 98, v/v) were added to extract the drug. The mixture was mixed vigorously for 2 min using a vortex and then centrifuged at 10,000 rpm for 10 min at 4 °C. The supernatant was then transferred to a 10-mL centrifuge tube containing 500 μL of an acetic acid-water solution (5 + 95, v/v) and the extraction was repeated. The supernatants were subsequently combined and evaporated at 45 °C using a gentle stream of nitrogen until the combined volume was less 500 μL. The residue was reconstituted in 2 mL of an acetic acid-water solution (5 + 95, v/v). After vortexing for 1 min, the solution was transferred to an MCX cartridge (60 mg, 3 cc, Waters, Milford, MA, USA), which was sequentially preconditioned with 3 mL of methanol and 3 mL of water. The cartridge was washed with 1 mL of the acetic acid-water solution and eluted in 3 mL of ammonia-methanol (1 + 9, v/v). The eluate was collected and evaporated completely using a gentle stream of nitrogen. The residue was subsequently reconstituted in 500 μL of acetonitrile-water (1 + 1, v/v) and centrifuged at 12,000 rpm for 15 min at 4 °C. The supernatant was filtered through a 0.22-μm nylon membrane filter and transferred into a 2-mL vial. Finally, 10 μL of the solution were injected into the UPLC-MS/MS system.
UPLC-MS/MS Analysis
An UPLC system coupled with a Quattro LC triple quadrupole tandem mass spectrometer (Waters) equipped with electrospray ionization (ESI) was used to determine the presence and subsequently quantify, florfenicol in the soil samples. Chromatographic separation was achieved using an Acquity UPLC BEH Shield RP18 column (50 mm × 2.1 mm, 1.7 μm) at 35 °C. Samples were separated using a mobile phase consisting of 0.1% formic acid in water (eluent A) and acetonitrile (eluent B) at a flow rate of 0.3 mL min−1. A linear gradient of eluent B (5–100%) was used in the total run time of 4 min (Table S3). The mass spectrometer was operated in the negative ESI mode with the following parameters: capillary voltage, 3.2 kV; cone voltage, 25 V; source temperature, 100 °C; desolvation temperature, 300 °C. Direct infusion was performed to optimize multiple reaction monitor transitions and associated acquisition parameters. The optimized conditions were as follows: m/z 356 > 336.1 (quantitative transition, collision energy, 13 eV), m/z 356 > 185.1 (qualitative transition, collision energy, 11 eV). Each sample was replicated three times for the determination of florfenicol residues.
Statistical Analysis
The difference in the relative abundance of FRGs (target gene copies/16S rRNA gene copies) was tested in all of the soil samples using t-test (non-parametric test), with further comparison of each soil sample carried out using Unpaired t-test.
Additional Information
How to cite this article: Zhao, Q. et al. Prevalence and Abundance of Florfenicol and Linezolid Resistance Genes in Soils Adjacent to Swine Feedlots. Sci. Rep. 6, 32192; doi: 10.1038/srep32192 (2016).
References
Jechalke, S., Heuer, H., Siemens, J., Amelung, W. & Smalla, K. Fate and effects of veterinary antibiotics in soil. Trends Microbiol 22, 536–545 (2014).
Li, C. et al. Occurrence of antibiotics in soils and manures from greenhouse vegetable production bases of Beijing, China and an associated risk assessment. Sci Total Environ 521–522, 101–107 (2015).
Chee-Sanford, J. C. et al. Fate and transport of antibiotic residues and antibiotic resistance genes following land application of manure waste. J Environ Qual 38, 1086–1108 (2009).
Wu, N., Qiao, M., Zhang, B., Cheng, W. D. & Zhu, Y. G. Abundance and diversity of tetracycline resistance genes in soils adjacent to representative swine feedlots in China. Environ Sci Technol 44, 6933–6939 (2010).
Wright, G. D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat Rev Microbiol 5, 175–186 (2007).
Fang, H., Wang, H., Cai, L. & Yu, Y. Prevalence of antibiotic resistance genes and bacterial pathogens in long-term manured greenhouse soils as revealed by metagenomic survey. Environ Sci Technol 49, 1095–1104 (2015).
Martinez, J. L. Environmental pollution by antibiotics and by antibiotic resistance determinants. Environ Pollut 157, 2893–2902 (2009).
Joy, S. R. et al. Fate and transport of antimicrobials and antimicrobial resistance genes in soil and runoff following land application of swine manure slurry. Environ Sci Technol 47, 12081–12088 (2013).
Nesme, J. & Simonet, P. The soil resistome: a critical review on antibiotic resistance origins, ecology and dissemination potential in telluric bacteria. Environ Microbiol 17, 913–930 (2015).
Edrington, T. S. et al. Pathogen prevalence and influence of composted dairy manure application on antimicrobial resistance profiles of commensal soil bacteria. Foodborne Pathog Dis 6, 217–224 (2009).
Heuer, H., Schmitt, H. & Smalla, K. Antibiotic resistance gene spread due to manure application on agricultural fields. Curr Opin Microbiol 14, 236–243 (2011).
Looft, T. et al. In-feed antibiotic effects on the swine intestinal microbiome. Proc Natl Acad Sci USA 109, 1691–1696 (2012).
Chantziaras, I., Boyen, F., Callens, B. & Dewulf, J. Correlation between veterinary antimicrobial use and antimicrobial resistance in food-producing animals: a report on seven countries. J Antimicrob Chemother 69, 827–834 (2014).
Lang, K. S. et al. Novel florfenicol and chloramphenicol resistance gene discovered in Alaskan soil by using functional metagenomics. Appl Environ Microbiol 76, 5321–5326 (2010).
Forsberg, K. J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012).
Su, J. Q., Wei, B., Xu, C. Y., Qiao, M. & Zhu, Y. G. Functional metagenomic characterization of antibiotic resistance genes in agricultural soils from China. Environ Int 65, 9–15 (2014).
Lopez-Perez, M., Mirete, S., Jardon-Valadez, E. & Gonzalez-Pastor, J. E. Identification and modeling of a novel chloramphenicol resistance protein detected by functional metagenomics in a wetland of Lerma, Mexico. Int Microbiol 16, 103–111 (2013).
Schwarz, S., Kehrenberg, C., Doublet, B. & Cloeckaert, A. Molecular basis of bacterial resistance to chloramphenicol and florfenicol. FEMS Microbiol Rev 28, 519–542 (2004).
Kim, E. & Aoki, T. Sequence analysis of the florfenicol resistance gene encoded in the transferable R-plasmid of a fish pathogen, Pasteurella piscicida. Microbiol Immunol 40, 665–669 (1996).
Long, K. S., Poehlsgaard, J., Kehrenberg, C., Schwarz, S. & Vester, B. The Cfr rRNA methyltransferase confers resistance to Phenicols, Lincosamides, Oxazolidinones, Pleuromutilins and Streptogramin A antibiotics. Antimicrob Agents Chemother 50, 2500–2505 (2006).
Wang, Y. et al. A novel gene, optrA, that confers transferable resistance to oxazolidinones and phenicols and its presence in Enterococcus faecalis and Enterococcus faecium of human and animal origin. J Antimicrob Chemother 70, 2182–2190 (2015).
Li, D. et al. Co-location of the oxazolidinone resistance genes optrA and cfr on a multiresistance plasmid from Staphylococcus sciuri. J Antimicrob Chemother, Epub ahead of print. 10.1093/jac/dkw040 (2016).
Bozdogan, B. & Appelbaum, P. C. Oxazolidinones: activity, mode of action and mechanism of resistance. Int J Antimicrob Agents 23, 113–119 (2004).
Locke, J. B., Zurenko, G. E., Shaw, K. J. & Bartizal, K. Tedizolid for the management of human infections: in vitro characteristics. Clin Infect Dis 58 Suppl 1, S35–S42 (2014).
Shen, J., Wang, Y. & Schwarz, S. Presence and dissemination of the multiresistance gene cfr in Gram-positive and Gram-negative bacteria. J Antimicrob Chemother 68, 1697–1706 (2013).
Kehrenberg, C. & Schwarz, S. Florfenicol-chloramphenicol exporter gene fexA is part of the novel transposon Tn558. Antimicrob Agents Chemother 49, 813–815 (2005).
Coyne, S., Courvalin, P. & Galimand, M. Acquisition of multidrug resistance transposon Tn6061 and IS6100-mediated large chromosomal inversions in Pseudomonas aeruginosa clinical isolates. Microbiology 156, 1448–1458 (2010).
Liu, H. et al. A novel phenicol exporter gene, fexB, found in enterococci of animal origin. J Antimicrob Chemother 67, 322–325 (2012).
Li, J., Shao, B., Shen, J., Wang, S. & Wu, Y. Occurrence of chloramphenicol-resistance genes as environmental pollutants from swine feedlots. Environ Sci Technol 47, 2892–2897 (2013).
Li, J. et al. Plasmid-mediated quinolone resistance genes and antibiotic residues in wastewater and soil adjacent to swine feedlots: potential transfer to agricultural lands. Environ Health Perspect 120, 1144–1149 (2012).
Ho, Y. B., Zakaria, M. P., Latif, P. A. & Saari, N. Occurrence of veterinary antibiotics and progesterone in broiler manure and agricultural soil in Malaysia. Sci Total Environ 488–489, 261–267 (2014).
Heuer, H. et al. Accumulation of sulfonamide resistance genes in arable soils due to repeated application of manure containing sulfadiazine. Appl Environ Microbiol 77, 2527–2530 (2011).
Fahrenfeld, N. et al. Effect of manure application on abundance of antibiotic resistance genes and their attenuation rates in soil: field-scale mass balance approach. Environ Sci Technol 48, 2643–2650 (2014).
Xu, M. et al. Simultaneous determination of florfenicol with its metabolite based on modified quick, easy, cheap, effective, rugged and safe sample pretreatment and evaluation of their degradation behavior in agricultural soils. J Sep Sci 38, 211–217 (2015).
Zhou, L. J. et al. Excretion masses and environmental occurrence of antibiotics in typical swine and dairy cattle farms in China. Sci Total Environ 444, 183–195 (2013).
Subbiah, M., Mitchell, S. M., Ullman, J. L. & Call, D. R. beta-lactams and florfenicol antibiotics remain bioactive in soils while ciprofloxacin, neomycin and tetracycline are neutralized. Appl Environ Microbiol 77, 7255–7260 (2011).
Zhu, Y. G. et al. Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci USA 110, 3435–3440 (2013).
Targant, H., Doublet, B., Aarestrup, F. M., Cloeckaert, A. & Madec, J. Y. IS6100-mediated genetic rearrangement within the complex class 1 integron In104 of the Salmonella genomic island 1. J Antimicrob Chemother 65, 1543–1545 (2010).
Zhang, W. J. et al. The new genetic environment of cfr on plasmid pBS-02 in a Bacillus strain. J Antimicrob Chemother 66, 1174–1175 (2011).
He, T. et al. Genetic environment of the transferable oxazolidinone/phenicol resistance gene optrA in Enterococcus faecalis isolates of human and animal origin. J Antimicrob Chemother, Epub ahead of print 10.1093/jac/dkw016 (2016).
He, T., Shen, J., Schwarz, S., Wu, C. & Wang, Y. Characterization of a genomic island in Stenotrophomonas maltophilia that carries a novel floR gene variant. J Antimicrob Chemother 70, 1031–1036 (2015).
Gaze, W. H. et al. Impacts of anthropogenic activity on the ecology of class 1 integrons and integron-associated genes in the environment. ISME J 5, 1253–1261 (2011).
Kent, W. J. BLAT–the BLAST-like alignment tool. Genome Res 12, 656–664 (2002).
Acknowledgements
This research was financially supported by grants from the National Natural Science Foundation of China (31370046) and the National Basic Research Program of China (2013CB127200).
Author information
Authors and Affiliations
Contributions
Q.Z., Y. Wu, Y. Wang and J.S. designed the experiment. Q.Z., Z.W., B.Z. and X.-d.D. performed experiments. Q.Z., S.W and Z.S. contributed to analysis the experimental data. Q.Z., Y. Wu and S.W. wrote the manuscript. Y. Wu, Y. Wang and J.S. supported and designed the project; Z.S., H.J., C.W., X.X., S.D., Y. Wang and J.S. critically revised the manuscript. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhao, Q., Wang, Y., Wang, S. et al. Prevalence and Abundance of Florfenicol and Linezolid Resistance Genes in Soils Adjacent to Swine Feedlots. Sci Rep 6, 32192 (2016). https://doi.org/10.1038/srep32192
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep32192
This article is cited by
-
IS6 family insertion sequences promote optrA dissemination between plasmids varying in transfer abilities
Applied Microbiology and Biotechnology (2024)
-
Occurrence and prevalence of antibiotic resistance genes in apple orchard after continual application of anaerobic fermentation residues of pig manure
Environmental Science and Pollution Research (2022)
-
Characterization of florfenicol resistance genes in the coagulase-negative Staphylococcus (CoNS) isolates and genomic features of a multidrug-resistant Staphylococcus lentus strain H29
Antimicrobial Resistance & Infection Control (2021)
-
A TaqMan-based multiplex real-time PCR assay for the rapid detection of tigecycline resistance genes from bacteria, faeces and environmental samples
BMC Microbiology (2020)
-
Impacts of florfenicol on the microbiota landscape and resistome as revealed by metagenomic analysis
Microbiome (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.