Abstract
Dairy Propionibacterium freudenreichii is a candidate non-lactic acid probiotic. However, little information is available on the effect of P. freudenreichii on lifespan extension in humans. The aim of this study was to evaluate the effects of P. freudenreichii on lifespan extension and to elucidate the mechanism of P. freudenreichii-dependent lifespan extension in Caenorhabditis elegans. The results showed that P. freudenreichii significantly (p < 0.05) extended the lifespan of C. elegans compared with Escherichia coli OP50, a standard food for the worm. Analysis of age-related biomarkers showed that P. freudenreichii retards ageing. Moreover, P. freudenreichii increased resistance against a human pathogen, Salmonella typhimurium, through the activation of skn-1, which is involved in pathogen resistance in C. elegans. Furthermore, P. freudenreichii-fed daf-16, jnk-1, skn-1 or daf-7 loss-of-function mutants showed an extended mean lifespan compared with E. coli OP50-fed worms. However, the increase in lifespan was not observed in pmk-1, sek-1, mek-1, dbl-1, daf-12 or daf-2 mutants, which suggests potential roles for these genes in P. freudenreichii-induced longevity in C. elegans. In conclusion, P. freudenreichii extends the lifespan of C. elegans via the p38 MAPK pathway involved in stress response and the TGF-β pathways associated with anti-inflammation processes in the immune system.
Similar content being viewed by others
Introduction
Probiotics play an important role in host intestinal health and directly and/or indirectly regulate intestinal microflora. Probiotics are effective in decreasing the colonization of pathogenic bacteria in the intestines, increasing the immune response and regulating mucosal immune signalling1,2. Propionibacterium freudenreichii is a candidate non-lactic acid probiotic3. P. freudenreichii has been used in the fermentation of Swiss Emmental cheese and is found in dairy products, such as milk and cheese. The short-chain fatty acid (SCFA)-producing bacterium, P. freudenreichii, predominantly ferments lactate to acetate, propionate and carbon dioxide. Acetate and propionate have been shown to enhance human gut immunity4. Recently, the benefits of P. freudenreichii intake, such as modulating immunity, reducing inflammation and protecting against pathogens, have been reported5. P. freudenreichii also functions as an immunomodulator of the human immune system6.
Caenorhabditis elegans is a small, free-living soil nematode commonly used as a model experimental animal because it is easy to treat, has a short lifespan, can be safely used in the laboratory and propagates through self-fertilization7. In particular, C. elegans is frequently used in studies on longevity, immunity, neurodegenerative diseases, fat storage, DNA damage responses and apoptosis8. C. elegans is a good research model because two-thirds of the genes related to human diseases are conserved in the nematode8,9,10. Several human pathogens also infect the intestine of C. elegans; thus, immune response against pathogens can be studied in C. elegans11. The immune system of C. elegans has evolved into the highly complex immunity of vertebrates. The relatively simple immunity of C. elegans might be experimentally advantageous in studying the immune signalling pathways. Notably, C. elegans research has elucidated entire genomes and structures of immune genes; therefore, many studies have used nematodes for studying the immune response12. Ageing and immunity are closely connected in C. elegans13. The C. elegans immune system is regulated by three major pathways of the defence system against pathogens: the transforming growth factor-beta (TGF-β) pathway, the p38 mitogen-activated protein kinase (MAPK) pathway and the Daf-2/DAF-16 pathway14. In the DBL/TGF-β pathway, following the binding of the TGF-β-like ligand encoded by the dbl-1 gene to TGF-β receptors in cell membranes, SMAD proteins (SMA-2, SMA-3 and SMA-4) are phosphorylated and activate TGF-β target genes related to antibacterial defence. In the MAPK pathway, SEK-1 and MEK-1 activate PMK-1, which is involved in innate immunity in C. elegans, but the downstream signalling of PMK-1 has not been elucidated. The DAF-2/DAF-16 pathway is involved in antibacterial defence and regulates lifespan in C. elegans. DAF-16 increases stress resistance in C. elegans in response to insulin/insulin-like growth factor-1 (IGF-1) signalling (IIS), which plays an important role in ageing. In C. elegans, the IIS receptor DAF-2 inhibits the transcription factor DAF-16; thus, decreased IIS increases DAF-16 activity, resulting in extended mean lifespan in C. elegans. DAF-2 regulates LYS-7, which is encoded by an antibacterial gene, lys-7 15. A longevity-promoting factor, SKN-1, increases stress tolerance and extends lifespan in an IIS-dependent manner. SKN-1 is inhibited by DAF-2 in a DAF-16-independent manner16. Longevity is also affected by immune dysfunction (immunosenescence) in nematodes17. According to a previous report, probiotics (such as lactic acid bacteria), compared with a standard food (such as E. coli OP50), may enhance immunity and longevity in C. elegans18,19. Lactic acid bacteria, such as Weissella koreensis and W. cibaria, extend the lifespan of C. elegans and improve biomarkers of ageing, such as a reduction in lipofuscin accumulation, decreases in body size and enhancement of body movement20. Small body size is often associated with increased longevity and a well-known factor regulating body size is diet-restriction that induces longevity in C. elegans21. Diet-restricted worms exhibit small body size as well as increased lifespan. Although it has been reported that surface proteins and 1,4-dihydroxy-2-naphthoic acid from P. freudenreichii are involved in their anti-inflammatory effects22,23, little information, including the mechanisms of immunity and longevity in C. elegans, is available on SCFA-producing dairy bacteria. In this study, the effect of P. freudenreichii on the lifespan extension of C. elegans and enhancement of immune responses were investigated, and the mechanism underlying P. freudenreichii-mediated lifespan extension in C. elegans was elucidated using loss-of-function mutants.
Results
P. freudenreichii extends the mean lifespan of C. elegans
C. elegans fed a lawn of P. freudenreichii, compared with the worms fed the standard E. coli OP50 lawn, showed a significantly (p < 0.001) increased mean lifespan (MLS) (Table 1). The MLS of P. freudenreichii-fed C. elegans, compared with that of E. coli OP50-fed worms, increased by approximately 13%. The survival rates were similar in both the P. freudenreichii- and E. coli OP50-fed C. elegans until day 13. After day 13, the two groups showed a significant difference in the survival rate (Fig. 1).
The effects of P. freudenreichii on age-related biomarkers in C. elegans
Three-day-old worms with similar sizes were transferred to nematode growth medium (mNGM) plates seeded with E. coli OP50 or P. freudenreichii, and body size was measured for 4 days. Body size increased with age in all groups, and the P. freudenreichii-fed worms were significantly (p < 0.001) smaller than the E. coli-fed worms (Fig. 2).
Young adult nematodes were transferred to mNGM plates seeded with P. freudenreichii or E. coli OP50. Body size was determined with 12 worms for each bacterial strain. Significant differences shown are relative to E. coli OP50. The results are expressed as the mean ± standard deviation; ***p < 0.001.
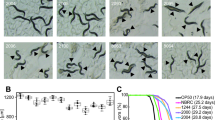
Body movement is related to age and the gut muscle. As the nematodes age, body movement becomes blunter and weaker. The locomotory rate of C. elegans on days 4, 7, 10, 13 and 16 was measured. During the experimental period, the proportion of worms displaying sinusoidal locomotion (class A) was always higher in P. freudenreichii-fed C. elegans than in E. coli OP50-fed worms (Fig. 3).
Young adult worms fed E. coli OP50 for 3 days after hatching were transferred to fresh mNGM plates with 50 mg of E. coli OP50 or P. freudenreichii lawn. Individuals were assigned to four classes based on locomotion: class (A) normal coordinated sinusoidal locomotion; class (B) uncoordinated and/or sluggish movement; class (C) no movement except head or tail in response to prodding; and class (D) dead worms. The bars indicate the proportion of each class at the designated time.
Lipofuscin accumulation is a biomarker of ageing in C. elegans. The accumulation of lipid granules was determined by autofluorescence. The intensity of 4′,6-diamidino-2-phenylindole (DAPI) decreases when nematode ageing is delayed18,24. The autofluorescence of worms fed P. freudenreichii was significantly (p < 0.001) decreased compared with that of worms fed E. coli OP50 (Fig. 4A). The lipofuscin level in P. freudenreichii-fed worms was approximately 50% of the level observed in E. coli OP50-fed worms (Fig. 4B).
L4 stage worms were transferred to the mNGM plates with E. coli OP50 or P. freudenreichii and fed bacteria for 14 days. After 14 days, lipofuscin was measured by assessing autofluorescence using a confocal microscope. (A) Fluorescence of lipofuscin in worms fed E. coli OP50 and P. freudenreichii on day 14. (B) Fluorescence was quantified using ImageJ software. The graph depicts the mean percentage in arbitrary units relative to that of control worms fed E. coli OP50 on day 14; ***p < 0.001. Scale bar = 100 μm.
Expression of lifespan extension-related genes
Quantitative real-time polymerase chain reaction (qPCR) analysis was conducted to investigate the expression of genes related to lifespan extension and immune response in C. elegans fed E. coli OP50 or P. freudenreichii. The expression levels of the daf-2, daf-16, pmk-1, sek-1, mek-1, dbl-1, daf-7, sma-3, skn-1 and daf-12 genes were measured. All genes except daf-16 and skn-1 were significantly up-regulated (p < 0.05) (Fig. 5). All genes involved in the p38 MAPK pathway and the TGF-β pathway assessed in this study were significantly up-regulated (p < 0.05). Notably, daf-12 expression increased 3.1-fold compared with that in the control. In addition, lys-7 and lys-8, antimicrobial peptide genes, were significantly overexpressed (p < 0.05).
Lifespan extension in C. elegans mutants
To determine whether MLS extension is related to enhanced innate immunity, C. elegans loss-of-function mutants for genes related to innate immunity and lifespan extension were fed P. freudenreichii or E. coli OP50, and their longevity was measured. The mutants defective in innate immunity were selected based on previous studies13,14,25 and our qPCR results (Fig. 5). P. freudenreichii failed to extend the lifespan of the pmk-1, sek-1 and mek-1 mutants, which are defective in genes involved in the p38 MAPK pathway; the dbl-1 mutant, which is defective in a gene involved in the TGF-β pathway; and the daf-2 mutant, which contains a mutation in the insulin/IGF receptor orthologue (Table 1, Fig. S1). However, the jnk-1, daf-16, skn-1 and daf-7 mutants had a significantly extended MLS after feeding on P. freudenreichii (p < 0.05). Interestingly, the daf-12 mutant, which is defective for a gene encoding a nuclear hormone receptor that acts as a lipid transcription factor activated by steroids and fatty acids26, had a significantly decreased MLS (p < 0.001). The results suggested that P. freudenreichii may activate the p38 MAPK and TGF-β pathways, which are related to innate immunity and lifespan extension.
Food preference of C. elegans
To confirm that the lifespan extension effect of P. freudenreichii was not derived from a preference between P. freudenreichii and E. coli OP50, binary choice assay was carried out. C. elegans showed positive food preference (CI = +0.13) for P. freudenreichii, which means that C. elegans did not show any preference between E. coli OP50 and P. freudenreichii (Fig. S2). The results suggest that the lifespan extension effect of P. freudenreichii is not due to dietary restriction.
C. elegans fed P. freudenreichii exhibits resistance to Salmonella typhimurium
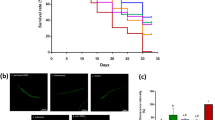
When worms are exposed to pathogens, the skn-1 gene is expressed via the activation of the PMK-1 pathway27. To determine whether the resistance of worms to pathogenic bacteria is enhanced, C. elegans wild type (N2) and the skn-1 mutant were exposed to S. typhimurium, and the survival rate was measured. Compared to the skn-1 mutant, C. elegans wild type (N2) worms exhibited increased resistance to S. typhimurium (p < 0.001) (Fig. 6A). The survival rates of C. elegans were similar until day 7, but after day 7, P. freudenreichii-fed worms had a higher survival rate than E. coli OP50-fed worms. However, the skn-1 mutant failed to show an extended MLS when fed P. freudenreichii (Fig. 6B). The MLS of P. freudenreichii-fed C. elegans (N2), compared with that of the E. coli OP50-fed worms, increased by approximately 11% (Fig. 6C). The MLS decreased approximately 30% in both E. coli OP50- and P. freudenreichii-fed skn-1 mutants compared with C. elegans (N2). The results suggested that P. freudenreichii activates the defence system of C. elegans during infection via the p38 MAPK pathway.
(A) L4 stage C. elegans (N2) and (B) skn-1 mutants were fed E. coli OP50 or P. freudenreichii for 4 days and transferred to fresh NGM/FUdR seeded with S. typhimurium. The survival rate was measured every 24 h after transferring the worms to the plate seeded with the pathogen. (C) Mean lifespan of C. elegans (N2) and skn-1 mutants infected with S. typhimurium. All experiments were performed at least three times; ***p < 0.001.
Discussion
Several factors, such as body size, lipofuscin and locomotory activity, change during ageing13,18,24, and thus, they are used as biomarkers of ageing. Feeding P. freudenreichii, compared E. coli OP50, to C. elegans reduced lipofuscin accumulation, decreased body size and increased locomotory activity, which suggests that P. freudenreichii extends the MLS of C. elegans. Additionally, compared with E. coli OP50-fed worms, P. freudenreichii-fed C. elegans showed a significantly increased MLS.
Genes related to immunity, neuroendocrine signalling, dietary restriction and mitochondrial function modulate the lifespan of C. elegans28. The results of food preference test suggest that dietary restriction is not linked to the lifespan extension of P. freudenreichii-fed C. elegans. C. elegans has a deficient adaptive immunity system and an NF-κB-based defence system; nematodes are dependent on the innate immune system for immune responses29. TGF-β signalling plays an important role in the regulation of cell growth, differentiation and development in many biological systems. The TGF-β pathway is composed of the DBL-1/SMA pathway and the DAF-7 pathway. The DBL-1 ligand is related to innate immunity and body morphology in C. elegans, while DAF-7 is the TGF-β-related ligand for the dauer pathway. P. freudenreichii did not extend the MLS of dbl-1 mutants, but it increased the MLS of daf-7 mutants. SMA-3 plays in a role in innate immunity30. The expression level of the sma-3 gene, which is located downstream of dbl-1, significantly increased. The results suggest that P. freudenreichii affects the MLS of C. elegans via the TGF-β pathway by activating DBL-1 but not by activating DAF-7. lys-8, a broad-spectrum antimicrobial peptide, encodes a lysozyme and is regulated by the DBL-1 pathway25. Lysozyme plays an important role in innate immunity and acts directly in the nematode defence system against infection25. In this study, P. freudenreichii influenced the expression of the lys-8 gene. Feeding P. freudenreichii to C. elegans up-regulated infection-inducible genes, such as sma-3 and lys-8, under the control of the DBL-1/TGF-β pathway, which may increase the resistance to infection in C. elegans.
p38 mitogen-activated protein kinase (MAPK) is involved in innate immunity in C. elegans and influences lifespan extension in C. elegans31. pmk-1, sek-1, mek-1 and skn-1 are components of the MAPK pathway in C. elegans. The pmk-1, sek-1 and mek-1 mutants failed to show increases in MLS of nematodes fed P. freudenreichii compared with worms fed E. coli OP50. The p38 MAPK pathway in C. elegans is similar to the mammalian p38 MAPK pathway that regulates the production of antimicrobials and inflammatory cytokines in response to lipopolysaccharide31. SKN-1, a transcription factor in the oxidative stress response, is phosphorylated by the p38 MAPK orthologue PMK-1 and is expressed during pathogen exposure27,32. skn-1 mutants fed P. freudenreichii had an increased MLS. However, interestingly, when exposed to a pathogen, P. freudenreichii feeding, compared with E. coli OP50 feeding, increased the survival of C. elegans (N2); however, the lifespans of P. freudenreichii- and E. coli OP50-fed skn-1 mutants decreased significantly compared with that of C. elegans (N2), without a significant difference between them. These results suggest that P. freudenreichii activates the p38 MAPK pathway transcription factor skn-1 during pathogenic infection in C. elegans. Although SKN-1 did not affect the longevity of C. elegans mediated by P. freudenreichii in normal conditions, SKN-1 might play an important role in pathogen resistance and survival during infection.
DAF-2 is the upstream receptor of the IIS signal and controls many downstream genes that contribute to nematode survival13. DAF-16, an orthologue of the FOXO transcription factor and the IIS transcription factor, is also a key player in innate immunity in C. elegans33. The daf-16 mutant shows no resistance to pathogens13. daf-16 acts in the insulin/IGF-1-mediated signalling pathway that regulates longevity in dietary restricted conditions, detoxification and oxidative stress. The daf-16 mutant showed significantly increased MLS of worms fed P. freudenreichii compared with worms fed E. coli OP50, and the expression of the daf-16 gene in P. freudenreichii-fed C. elegans, compared with that in E. coli OP50-fed worms, did not significantly increase. Mutation in daf-2 activated DAF-16, resulting in the extension of lifespan in C. elegans. In this study, the daf-2 mutant did not prolong lifespan, but the daf-16 mutant did extend the MLS of C. elegans, indicating that the longevity effect of P. freudenreichii is not related to the daf-2/daf-16 pathway. The results suggest that P. freudenreichii influences C. elegans innate immunity through the TGF-β and p38 MAPK pathways but not through the IIS pathway.
lys-7 encodes an enzyme homologous to an antimicrobial lysozyme that functions in the innate immune response, indicating that lys-7 is involved in pathogen resistance31. lys-7 is a DAF-16-dependent gene and is down-regulated by DAF-2. Curiously, in this study, although P. freudenreichii did not stimulate the expression of daf-16 and induced daf-2 expression, lys-7 was significantly (p < 0.05) expressed in worms fed P. freudenreichii compared with worms fed E. coli OP50. DAF16-independent alternate pathway(s) may contribute to lys-7 expression in worms fed P. freudenreichii.
DAF-12 is a well-known nuclear hormone receptor and affects the innate immune system by inducing the production of antimicrobial peptides. DAF-12 is correlated with the p38 MAPK pathway34,35. daf-12 directly regulates skn-1, a component of the PMK-1 immune signalling pathway, during pathogen infection. It was reported that DAF-12 is involved in the innate immune system via crosstalk with the p38/PMK-1 MAPK pathway34. However, a clear relationship between pmk-1 and daf-12 has not been elucidated. In this study, we found that pmk-1 and daf-12 mutants failed to extend the MLS of P. freudenreichii-fed C. elegans, while the skn-1 mutant extended the MLS of P. freudenreichii-fed worms. These results indicate that there may be an alternative pathway between PMK-1 and DAF-12 that does not involve SKN-1 implicated in the MLS extension of C. elegans by P. freudenreichii. daf-12 is a putative target gene of DAF-7/TGF-β (dauer pathway); however, DAF-12 might act in a DAF-7-independent manner to alter longevity in P. freudenreichii-fed C. elegans. In addition, DAF-12 plays an important role in longevity through the integration of both germline and DAF-2 signalling36. DAF-12, similar to DAF-16, is negatively regulated by DAF-2. However, in this study, DAF-12, in contrast to DAF-16, affected the longevity of P. freudenreichii-fed C. elegans. The results suggest that DAF-12 is involved in the longevity of P. freudenreichii-fed C. elegans through an alternative pathway in a DAF-2- and DAF-7-independent manner.
Based on the gene expression and mutant survival data obtained in this study, we developed a model of the pathways involved in the MLS extension of P. freudenreichii-fed C. elegans (Fig. 7). The molecular mechanisms that modulate ageing and immune response are linked in C. elegans. DAF-2–mediated insulin signalling pathway is universal in regulating ageing and p38 MAPK regulates the innate immune function in C. elegans, which are conserved among living organisms including human13. Therefore, it is possible that the mechanisms identified in this study may apply to other species including humans. In conclusion, dairy P. freudenreichii extends the MLS of C. elegans by activating the innate immune system via the p38 MAPK pathway- and the TGF-β pathway-dependent but IIS pathway-independent manner.
DAF-16, the primary transcription factor required for lifespan extension, is not related to lifespan extension of P. freudenreichii-fed C. elegans. DBL-1 activates SMA-3 through the TGF-β pathway, which produces antimicrobial peptides. MEK-1, SEK-1 and PMK-1, the components of the MAPK pathway, are regulated to extend lifespan in P. freudenreichii-fed C. elegans. A nuclear hormone receptor, DAF-12, is also activated by P. freudenreichii and is involved in the extension of lifespan in C. elegans. The solid lines represent potential pathways and putative proteins activated by P. freudenreichii are shown in a dark circle, and dashed lines indicate speculated pathways that are not involved in the lifespan extension in P. freudenreichii-fed C. elegans.
Methods
Bacteria culture conditions and strains
P. freudenreichii KCTC 1063 was obtained from the Korean Collection for Type Cultures (KCTC), and Escherichia coli OP50 was provided by the Caenorhabditis Genetics Centre (CGC) at the University of Minnesota and used as a standard food for C. elegans. E. coli OP50 was grown in Luria-Bertani (LB) broth (Difco, Detroit, MI, USA) at 37 °C with shaking overnight. P. freudenreichii was cultured using reinforced clostridial medium (RCM) broth (Oxoid, Hampshire, United Kingdom) for 48–72 h in anaerobic conditions using a BD GasPak (Becton Dickinson, Franklin Lakes, NJ, USA) at 30 °C. E. coli OP50 and P. freudenreichii were collected by centrifugation and washed twice with M9 buffer. Then, the bacteria were diluted to a final concentration of 0.1 mg (wet weight) per mL in M9 buffer37.
Nematode preparation
C. elegans Bristol strain N2 was used as the wild-type strain, and the mutant strains were provided by CGC. The mutants used in this study were CF1038 daf-16 (mu86), CB1370 daf-2 (e1370), EU1 skn-1 (zu67), VC8 jnk-1 (gk7), FK171 mek-1 (ks54), KU25 pmk-1 (km25), KU4 sek-1 (km4), RB2302 daf-7 (ok3125), NU3 dbl-1 (nk3) and DR20 daf-12 (m20). Worms were maintained and propagated in peptone-free mNGM (modified nematode growth medium) at 25 °C using standard techniques38. Some bacteria consume media components, which may affect nematode survival18. E. coli OP50 and P. freudenreichii suspended in M9 buffer were seeded on mNGM in 90 mm diameter petri dishes to feed the worms. C. elegans was exposed to a sodium hypochlorite-sodium hydroxide solution as previously described to obtain viable eggs39. The eggs were incubated overnight in M9 buffer at 25 °C for hatching, and the suspension of L1-stage worms was centrifuged at 1,200 × g for 2 min. After eliminating the supernatant, the remaining larvae were transferred to fresh mNGM plates seeded with E. coli OP50 and incubated at 25 °C. The worms were allowed to grow on E. coli OP50 until the L4 stage18. All experiments used 3-day-old worms (1 day adult) to regulate the reproductive system in C. elegans, with the exception of the mutants40.
Determination of the mean lifespan of C. elegans
The C. elegans lifespan assay was performed by transferring 15 young adult (L4 stage) worms per bacterial species to three mNGM/FUdR plates. 5-Fluoro-2′-deoxyuridine (FUdR, Sigma Aldrich, St. Louis, MO, USA) (50 μM) was added to the plates41, and E. coli OP50 and P. freudenreichii were seeded on them. The plates were incubated at 25 °C, and the numbers of live or dead worms were scored every 24 h. Worms were considered “dead” when they failed to respond to a gentle touch with a worm picker. Nematodes that crawled off the plates and showed non-natural deaths, such as bagging or adhering to the wall of the plate, were censored42. For the first 3 days, worms were transferred to fresh mNGM/FUdR every day, and then they were transferred once per week for the rest of the experiment to maintain a sufficient food source. Experiments were performed at least in triplicate, and more than 100 worms for each bacterial species were used in the longevity assay.
The mean lifespan was estimated using the following formula43:

where j is the age category (day), dj is the number of worms that died in the age interval (xj, xj1), and n is the total number of worms. The standard error (SE) of the MLS estimate was calculated using the following equation:

Measurement of body size
Young adult worms (L4 stage) were transferred to mNGM plates (60 mm plate) covered with 5 mg (wet weight) of E. coli OP50 or P. freudenreichii cells in M9 buffer. The plates were incubated at 25 °C, and the body size of the live worms was measured every 24 h until 7 days of age. In total, 12 worms per bacterial species were measured. Images of C. elegans were taken with a stereomicroscope (Olympus SZ61) and a ToupCam (UCMOS05100KPA), and the images were analysed using ToupCam software. For this experiment, the area of the worm’s projection was estimated automatically and used as an index of body size19. Three independent experiments were performed for each bacterial species.
Lipofuscin accumulation
The autofluorescence of intestinal lipofuscin was measured as an index of senescence of day 14 adults. Randomly selected worms from each bacterial lawn were washed three times with M9 buffer. Then, the worms were placed onto a 5% agar pad coated with 10 mM sodium azide in M9 buffer to be paralyzed. Lipofuscin autofluorescence images were taken using blue excitation light (405−488 nm) with DAPI (4′,6-diamidino-2-phenylindole) and laser confocal scanning microscopy (Olympus Ix81-FV1000, Tokyo, Japan)18. Fluorescence was quantified using ImageJ software (National Institutes of Health, Bethesda, MD, USA) to determine the lipofuscin levels. Three independent experiments were performed with more than 30 worms for each bacterial species on each day.
Locomotory scoring
The locomotory assay of worms at different ages was performed using a scoring method described in previous reports18. Nematodes were classified as class “A” when they showed spontaneous movement or vigorous locomotion in response to prodding, class “B” worms did not move unless touched or appeared to have uncoordinated movement, class “C” worms moved only their head and/or tail in response to prodding, and class “D” worms were dead. Experiments were conducted at least three times (n > 100) independently for each bacterial species.
Binary choice assay
To examine the preference of worms towards P. freudenreichii, we conducted binary choice assay compared with E. coli OP50 44. Assays were done on NGM medium in 90 mm plates. Bacterial food was seeded at the same distance from the sides of the plates with a diameter of 0.5 cm (Fig. S3), and L4 stage nematodes (30 worms) were put on the centre of the plates. The number of worms in each bacterial lawn was counted after 3 h of incubation at 25 °C. Three independent experiments were performed. The choice index (CI) was calculated as follows:

If, CI = −1.0 represents complete preference for E. coli OP50. +1.0 represents complete preference for the test bacterium.
0.0 represents an equal distribution.
Resistance against Salmonella infection
After hatching, the nematodes were grown on E. coli OP50 until they were 3-day-old adults. Then, the worms were allocated to a control group that continued to be fed E. coli OP50 or the test group that received P. freudenreichii for 4 days. The 7-day-old worms were transferred to NGM agar plates with Salmonella typhimurium ATCC 13311 cell suspensions (concentration of 0.1 mg/mL). The bacterial suspensions were incubated at room temperature for 12 h before the nematodes were transferred to the NGM plates45. Then, 45 worms from each bacterial group were divided into three plates that were seeded with Salmonella and incubated at 25 °C. The numbers of live and dead nematodes were scored every 24 h. The survival rate of C. elegans exposed to Salmonella was compared with that of worms without Salmonella19. Each test was performed with at least 100 worms.
RNA isolation and qPCR
Worms fed E. coli OP50 or P. freudenreichii for 24 h were collected and washed twice with sterilized M9 buffer. Then, total mRNA from whole worms was isolated using TRIzol as previously described46. Total RNA was converted to cDNA using a RevertAid First Strand cDNA Synthesis Kit according to the manufacturer’s instructions (Thermo Scientific, Wilmington, DE, USA), followed by qPCR using SYBR Green (KAPA Biosystems, Wilmington, MA, USA) and a QuantStudio 6 Flex Real Time PCR machine (Applied Biosystems, Foster City, CA, USA). The primer sequences are listed in Table S1. The reactions had an initial step at 95 °C, followed by 40 cycles of 95 °C for 20 s, 60 °C for 20 s and 72 °C for 30 s, followed by melt curve analysis. The experiments were independently performed three times, and relative expression levels were calculated using the 2−ΔΔCT method46. The control gene act-1 was used to normalize gene expression data.
Statistical analysis
The MLS of C. elegans was calculated using the Kaplan-Meier method, and the p-value of survival differences was tested using the log-rank test37. In other experiments, the significance of comparisons between E. coli OP50 and P. freudenreichii was determined using Student’s t-test. Significance was defined as a p-value under 0.05 in all experiments. If the data were not normally distributed, the Mann-Whitney U test was used18.
Additional Information
How to cite this article: Kwon, G. et al. Dairy Propionibacterium extends the mean lifespan of Caenorhabditis elegans via activation of the innate immune system. Sci. Rep. 6, 31713; doi: 10.1038/srep31713 (2016).
References
Zhou, M. et al. Investigation into in vitro and in vivo models using intestinal epithelial IPEC‐J2 cells and Caenorhabditis elegans for selecting probiotic candidates to control porcine enterotoxigenic Escherichia coli. Journal of Applied Microbiology 117, 217−226 (2014).
Gaggìa, F., Mattarelli, P. & Biavati, B. Probiotics and prebiotics in animal feeding for safe food production. International Journal of Food Microbiology 141, S15−S28 (2010).
Holzapfel, W. H., Haberer, P., Snel, J., Schillinger, U. & in’t Veld, J. H. J. H. Overview of gut flora and probiotics. International Journal of Food Microbiology 41, 85−101 (1998).
Cousin, F. J., Mater, D. D., Foligne, B. & Jan, G. Dairy propionibacteria as human probiotics: a review of recent evidence. Dairy Science & Technology 91, 1−26 (2011).
Foligne, B. et al. Promising immunomodulatory effects of selected strains of dairy propionibacteria as evidenced in vitro and in vivo . Applied and Environmental Microbiology 76, 8259−8264 (2010).
Thierry, A. et al. New insights into physiology and metabolism of Propionibacterium freudenreichii. International Journal of Food Microbiology 149, 19−27 (2011).
Riddle, D. L., Blumenthal, T., Meyer, B. J., Preiss, J. & Pettitt, J. C. Elegans II. Trends in Cell Biology 8, 92 (1998).
Zhang, R. & Hou, A. Host-microbe interactions in Caenorhabditis elegans. ISRN Microbiology 2013, 356451 (2013).
Chavez, V., Mohri-Shiomi, A. & Garsin, D. A. Ce-Duox1/BLI-3 generates reactive oxygen species as a protective innate immune mechanism in Caenorhabditis elegans. Infection and Immunity 77, 4983−4989 (2009).
Finley, J. W. et al. Legumes reduced intestinal fat deposition in the Caenorhabditis elegans model system. Journal of Functional Foods 5, 1487−1493 (2013).
Ermolaeva, M. A. & Schumacher, B. Insights from the worm: The C. elegans model for innate immunity. Seminars in Immunology 26, 303−309 (2014).
Flajnik, M. F. & Du Pasquier, L. Evolution of innate and adaptive immunity: can we draw a line? Trends in Immunology 25, 640−644 (2004).
Kurz, C. L. & Tan, M. W. Regulation of aging and innate immunity in C. elegans. Aging Cell 3, 185−193 (2004).
Engelmann, I. & Pujol, N. In Invertebrate Immunity, Innate immunity in C. elegans 105–121, Springer (2010).
Murphy, C. T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424, 277−283 (2003).
Tullet, J. M. et al. Direct inhibition of the longevity-promoting factor SKN-1 by insulin-like signaling in C. elegans. Cell 132, 1025−1038 (2008).
Gill, H. S., Cross, M. L., Rutherford, K. J. & Gopal, P. K. Dietary probiotic supplmentation to enhance cellular immunity in the elderly. British Journal of Biomedical Science 58, 94 (2001).
Komura, T., Ikeda, T., Yasui, C., Saeki, S. & Nishikawa, Y. Mechanism underlying prolongevity induced by bifidobacteria in Caenorhabditis elegans. Biogerontology 14, 73−87 (2013).
Ikeda, T., Yasui, C., Hoshino, K., Arikawa, K. & Nishikawa, Y. Influence of lactic acid bacteria on longevity of Caenorhabditis elegans and host defense against salmonella enterica serovar enteritidis. Applied and Environmental Microbiology 73, 6404−6409 (2007).
Lee, J., Kwon, G. & Lim, Y.-H. Elucidating the Mechanism of Weissella-dependent Lifespan Extension in Caenorhabditis elegans. Scientific Reports 5 (2015).
Bishop, N. A. & Guarente, L. Two neurons mediate diet-restriction-induced longevity in C. elegans. Nature 447, 545−549 (2007).
Okada, Y. et al. 1,4-Dihydroxy-2-naphthoic acid from Propionibacterium freudenreichii reduces inflammation in interleukin-10-deficient mice with colitis by suppressing macrophage-derived proinflammatory cytokines. Journal of Leukocyte Biology 94, 473−480 (2013).
Le Marechal, C. et al. Surface proteins of Propionibacterium freudenreichii are involved in its anti-inflammatory properties. Journal of Proteomics 113, 447−461 (2015).
Pincus, Z. & Slack, F. J. Developmental biomarkers of aging in Caenorhabditis elegans. Developmental Dynamics 239, 1306−1314 (2010).
Millet, A. C. M. & Ewbank, J. J. Immunity in Caenorhabditis elegans. Current Opinion in Immunology 16, 4−9 (2004).
Gerisch, B. et al. A bile acid-like steroid modulates Caenorhabditis elegans lifespan through nuclear receptor signaling. Proceedings of the National Academy of Sciences of the United States of America 104, 5014−5019 (2007).
Papp, D., Csermely, P. & Soti, C. A role for SKN-1/Nrf in pathogen resistance and immunosenescence in Caenorhabditis elegans. PLoS Pathogen 8, e1002673 (2012).
Evans, E. A., Chen, W. C. & Tan, M. W. The DAF‐2 insulin‐like signaling pathway independently regulates aging and immunity in C. elegans. Aging Cell 7, 879−893 (2008).
Kurz, C. L. & Ewbank, J. J. Caenorhabditis elegans: an emerging genetic model for the study of innate immunity. Nature Reviews Genetics 4, 380−390 (2003).
Mallo, G. V. et al. Inducible antibacterial defense system in C. elegans. Current Biology 12, 1209−1214 (2002).
Troemel, E. R. et al. p38 MAPK regulates expression of immune response genes and contributes to longevity in C. elegans. PLoS Genetics 2, e183 (2006).
Hoeven, R., McCallum, K. C., Cruz, M. R. & Garsin, D. A. Ce-Duox1/BLI-3 generated reactive oxygen species trigger protective SKN-1 activity via p38 MAPK signaling during infection in C. elegans. PLoS Pathogens 7, e1002453 (2011).
Singh, V. & Aballay, A. Regulation of DAF-16-mediated Innate Immunity in Caenorhabditis elegans. The Journal of Biological Chemistry 284, 35580−35587 (2009).
Liu, F. et al. Nuclear hormone receptor regulation of microRNAs controls innate immune responses in C. elegans. PLoS Pathogens 9, e1003545 (2013).
Mooijaart, S. P. et al. C. elegans DAF-12, nuclear hormone receptors and human longevity and disease at old age. Ageing Research Reviews 4, 351−371 (2005).
Fisher, A. L. & Lithgow, G. J. The nuclear hormone receptor DAF-12 has opposing effects on Caenorhabditis elegans lifespan and regulates genes repressed in multiple long-lived worms. Aging Cell 5, 127−138 (2006).
Zhao, Y. et al. Lactobacillus salivarius strain FDB89 induced longevity in Caenorhabditis elegans by dietary restriction. Journal of Microbiology 51, 183−188 (2013).
Stiernagle, T. Maintenance of C. elegans. Wormbook 1−11 (2006).
John, S. & Jonathan, H. In The nematode Caenorhabditis elegans. Vol. 17 (ed Wood, William B. ) 587−606, Cold Spring Harbor Monograph Series (1988).
Hsin, H. & Kenyon, C. Signals from the reproductive system regulate the lifespan of C. elegans. Nature 399, 362−366 (1999).
Gruber, J., Ng, L. F., Poovathingal, S. K. & Halliwell, B. Deceptively simple but simply deceptive–Caenorhabditis elegans lifespan studies: considerations for aging and antioxidant effects. FEBS Letters 583, 3377−3387 (2009).
Schmeisser, S. et al. Neuronal ROS signaling rather than AMPK/sirtuin-mediated energy sensing links dietary restriction to lifespan extension. Molecular Metabolism 2, 92−102 (2013).
Wu, D., Rea, S. L., Yashin, A. I. & Johnson, T. E. Visualizing hidden heterogeneity in isogenic populations of C. elegans. Experimental Gerontology 41, 261−270 (2006).
Abada, E. A. et al. C. elegans behavior of preference choice on bacterial food. Molecules and cells 28, 209−213 (2009).
Marsh, E. K., van den Berg, M. C. W. & May, R. C. A two-gene balance regulates Salmonella typhimurium tolerance in the nematode Caenorhabditis elegans. PLoS ONE 6, e16839 (2011).
Greer, E. L. et al. An AMPK-FOXO pathway mediates longevity induced by a novel method of dietary restriction in C. elegans. Current Biology 17, 1646−1656 (2007).
Author information
Authors and Affiliations
Contributions
Y.-H.L. and G.K. designed the research, and G.K., J.L. and Y.-H.L. performed the experiments. Y.-H.L. and G.K. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kwon, G., Lee, J. & Lim, YH. Dairy Propionibacterium extends the mean lifespan of Caenorhabditis elegans via activation of the innate immune system. Sci Rep 6, 31713 (2016). https://doi.org/10.1038/srep31713
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep31713
This article is cited by
-
Lactobacillus paracasei HII01 enhances lifespan and promotes neuroprotection in Caenorhabditis elegans
Scientific Reports (2023)
-
Geroprotective potential of microbiome modulators in the Caenorhabditis elegans model
GeroScience (2023)
-
Weissella confusa CGMCC 19,308 Strain Protects Against Oxidative Stress, Increases Lifespan, and Bacterial Disease Resistance in Caenorhabditis elegans
Probiotics and Antimicrobial Proteins (2022)
-
Oligosaccharides from agar extends lifespan through activation of unfolded protein response via SIR-2.1 in Caenorhabditis elegans
European Journal of Nutrition (2022)
-
Dairy associations for the targeted control of opportunistic Candida
World Journal of Microbiology and Biotechnology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.