Abstract
Recent research has highlighted the importance of the gut microbiome in regulating aging, and probiotics are interventions that can promote gut health. In this study, we surveyed several novel lactic acid bacteria to examine their beneficial effect on organismal health and lifespan in C. elegans. We found that animals fed some lactic acid bacteria, including L. acidophilus 1244 and L. paracasei subsp. paracasei 2004, grew healthy. Supplementation with the lactic acid bacterial strains L. acidophilus 1244 or L. paracasei subsp. paracasei 2004 significantly improved health, including food consumption, motility, and resistance to oxidative stressor, hydrogen peroxide. Our RNA-seq analysis showed that supplementation with L. paracasei subsp. paracasei 2004 significantly increased the expression of daf-16, a C. elegans FoxO homolog, as well as genes related to the stress response. Furthermore, daf-16 deletion inhibited the longevity effect of L. paracasei subsp. paracasei 2004 supplementation. Our results suggest that L. paracasei subsp. paracasei 2004 improves health and lifespan in a DAF-16-dependent manner.
Similar content being viewed by others
Introduction
Recent research has revealed the crucial role that the human microbiome plays in regulating the aging process1. Studies have shown that changes in the composition of the microbiome can influence age-related decline in organ function and the onset of age-related diseases, such as Alzheimer’s disease and cardiovascular disease2,3,4,5,6,7,8,9,10. The microbiome supports the immune system, aids in digestion and metabolism, and helps prevent colonization by pathogenic microorganisms11. These functions are essential in older adults because they could help delay age-related changes in the body and improve overall health12,13,14,15. Therefore, understanding the role of the microbiome in aging regulation is critical for developing interventions that can promote healthy aging and prevent age-related diseases.
The diversity of the gut microbiome is established early in life and is shaped by a variety of factors: birth and early life, diet, antibiotic use, age, geography, and lifestyle12,13,14,15. As individuals age, the diversity of the gut microbiota tends to decrease, with certain bacteria becoming more abundant while others decline12,14. This shift in microbial composition has been associated with alterations in gut permeability, inflammation, and immune function, which may contribute to the development of age-related diseases2,6,7,8,10. For example, some studies have found that the gut microbiota of elderly individuals is more pro-inflammatory and less diverse than that of younger individuals12,13,14,15 and that these changes are associated with an increased risk of conditions such as cardiovascular disease and dementia2,8. In Drosophila, transplantation of the microbiota from aged donor flies to young recipient flies changes the microbiota and decreases lifespan16. Conversely, transplantation of the microbiome from young fish to middle-aged fish alters the microbiome composition of the older recipient and significantly increases the lifespan in killifish17. Fecal microbiome transplantation from wild-type to progeria mice recipients enhances health and lifespan18. These lines of evidence suggest a crucial role of the microbiota in lifespan regulation.
Keeping the gut microbiome in good condition could contribute to health and longevity. Interventions to maintain the gut microbiome in good condition are a matter of interest. Probiotics are live microorganisms that are administered to promote gut health adequately: probiotics represent an essential group of beneficial consumed microorganisms found in fermented foods such as yogurt, kefir, and sauerkraut, as well as in supplemental forms19,20. Administration of the probiotic Lactobacillus GKM3 promotes longevity and memory retention in SAMP8 mice, a model of accelerated aging21. Supplementation with Akkermansia muciniphila improves insulin sensitivity and reduces insulinemia and plasma total cholesterol levels in volunteer humans22. Some probiotics have been shown to increase lifespan in Caenorhabditis elegans: lactic acid bacteria (Lactobacillus gasseri SBT2055 and Lactobacillus rhamnosus CNCM I-3690) and non-lactic acid bacteria (Propionibacterium freudenreichii) increase the lifespan of C. elegans23,24,25. Other probiotics might be beneficial for improving health and longevity.
In this study, we surveyed several lactic acid bacteria to find their beneficial effects on health and lifespan. To this end, C. elegans, one of the most used model organisms in aging research, was fed with several lactic acid bacteria after their development, and we examine the effect of lactic acid bacteria administration on growth, health, and longevity.
Results
To identify the lactic acid bacterial strain that positively regulates the lifespan of C. elegans, we first examined the effect of lactic acid bacterial strains on the growth of the animals. Age-synchronized germ-free populations can be obtained by treating animals with bleach. We focused on analyzing the beneficial effects of lactic acid bacteria on health and longevity rather than on their impact on development. Therefore, Day1 adult animals (after completion of development: 3 days after synchronization) were fed lactic acid bacterial strains (L. acidophilus 1244, L. gasseri 2000, 2100, 8063, and 8064, L. paracasei subsp. Paracasei 2004 and 2005, L. gasseri 2093, and L. johnsonii 2095 and 2096), including the lactic acid bacterial strain L. rhamnosus CNCM I-3690, which has been shown to extend the lifespan of wild-type animals25, for 3 days, and the length of the animals was measured. Animals fed lactic acid bacterial strains except for L. rhamnosus CNCM I-3690 and L. paracasei subsp. Paracasei 2004 were significantly shorter than animals fed E. coli OP50; animals fed L. rhamnosus CNCM I-3690 or L. paracasei subsp. Paracasei 2004 was not significantly shorter (Fig. 1A, B and Table S1). Furthermore, animals grew almost normally when fed some lactic acid bacteria (L. rhamnosus CNCM I-3690, L. acidophilus 1244, L. gasseri 2000, and L. paracasei subsp. Paracasei 2004 and 2005), while most animals displayed bags of worms (state of animals where many fertilized eggs, normally laid outside, overly remained in the germline, one of the egg-laying defects26; Fig. 1A, arrowhead) when fed other lactic acid bacteria (L. gasseri 2100, 8063, and 8064, L. gasseri 2093, and L. johnsonii 2095 and 2096). Bags of worms are induced when animals are subjected to harsh environments such as fasting27. We thus focused on the lactic acid bacterial strains that did not induce egg-laying defects and examined their effect on lifespan. Our measurements showed that animals fed lactic acid bacterial strains (L. acidophilus 1244, L. gasseri 2000, and L. paracasei subsp. paracasei 2004 and 2005) as well as the L. rhamnosus CNCM I-3690 strain lived significantly longer than animals fed control food (E. coli OP50) (Fig. 1C and Table S2). These results suggest that some lactic acid bacterial strains have the potential to positively regulate lifespan.
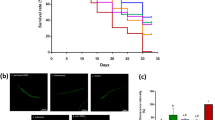
L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 supplementation extends lifespan. (A) Representative images of wild-type animals fed the indicated bacterium for 3 days from Day 1 of adulthood. Feeding some lactic acid bacterial strains (L. acidophilus 1244, L. gasseri 2000, 2100, 8063, and 8064, L. paracasei subsp. paracasei 2004 and 2005, L. gasseri 2093, and L. johnsonii 2095 and 2096) lead to bags of worms, one of the egg-laying defects. Arrowheads indicate animals showing bags of worms. (B) The body length of animals fed the indicated bacterium on Day 4 of adulthood. Feeding lactic acid bacterial strains except for L. rhamnosus CNCM I-3690 and L. paracasei subsp. paracasei 2004 shortened the body compared with E. coli OP50 feeding. See Table S1 for detailed statistics. (C) Survival curves of animals fed the indicated bacterial strains. Feeding lactic acid bacterial strains extended the lifespan compared with E. coli OP50 feeding. See Table S2 for detailed statistics.
We focused on L. acidophilus 1244 and L. paracasei subsp. paracasei 2004, both of which extended the lifespan (Fig. 1C and Table S2), because L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 had the strongest and the weakest effect on body size, respectively, among the strains that extended lifespan (Fig. 1B and Table S1). We then examined the effect of these lactic acid bacterial strains on health as well as lifespan. To this end, animals were fed lactic acid bacteria for 3 days from Day 1 of adulthood and then we examined the pumping rate, the bending rate, and oxidative stress resistance, all of which are indicators of health28, using middle-aged animals. Our measurements showed that L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 feeding significantly increased the pumping rate (Fig. 2A and Table S3), the bending rate (Fig. 2B and Table S4), and oxidative stress resistance (Fig. 2C and Table S5) compared with E. coli OP50 feeding. These results indicate that L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 supplementation improves the animals’ health as well as lifespan.
L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 feeding enhances health. (A) The pumping rate of the animals fed the indicated bacterium on Day 4 of adulthood. L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 feeding improved the pumping rate at middle age. Means not sharing the same letter are significantly different (Tukey test, p < 0.05). See Table S3 for detailed statistics. (B) The bending rate of the animals fed the indicated bacterium on Day 10. L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 feeding improved the bending rate at middle age. Means not sharing the same letter are significantly different (Tukey test, p < 0.05). See Table S4 for detailed statistics. (C) The survival curves of the animals fed the indicated bacterium on Day 10 in 2 mM hydrogen peroxide. L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 feeding improved the oxidative stress resistance at middle age. P-values were calculated by a log-rank test with Bonferroni correction (**, p < 0.01). See Table S5 for detailed statistics. (D) Body size of the animals fed the indicated bacterium on Day 2, Day 3, and Day 4 of adulthood. L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 feeding increased body size on Day 4. See Table S6 for detailed statistics.
Because both L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 feeding affects growth (Fig. 1B and Table S1), it is possible that supplementation with these two lactic acid bacteria causes dietary restriction, which promotes health and lifespan. We thus scrutinized whether L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 feeding could mimic food restriction because of the avoidance of these bacteria or malnutrition under the use of these bacteria as a food resource. We examined the body size every day after lactic acid bacterium feeding. Animals fed L. acidophilus 1244 stopped growing on Day 3 of adulthood, while animals fed OP50 or L. paracasei subsp. paracasei 2004 continued growing on Day 4, although the size of animals fed L. paracasei subsp. paracasei 2004 was smaller than that of the animals fed E. coli OP50 (Fig. 2D and Table S6). This result implies that L. acidophilus 1244 feeding causes malnutrition that makes it difficult for animals to grow, while L. paracasei subsp. paracasei 2004 is nutritious enough to allow the animals to grow steadily.
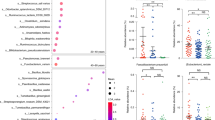
Our results show that L. paracasei subsp. paracasei 2004 feeding, compared with L. acidophilus 1244 feeding, has a more prominent effect on lifespan (Fig. 1C and Table S2) and a weaker food restriction effect on body size (Fig. 2D and Table S6). Transcription factors, such as DAF-16 and SKN-1, regulate longevity in C. elegans, and transcriptome alterations are vital for lifespan regulation29,30,31. Therefore, we then explored the effect of L. paracasei subsp. paracasei 2004 feeding on the transcriptome profile to understand the mechanisms of the longevity effect of lactic acid bacteria. We explored the transcriptome alterations after 3 days of L. paracasei subsp. paracasei 2004 feeding by RNA-seq analysis. We defined the differentially expressed genes (DEGs) whose fold changes were more than twofold with an FDR cutoff (less than 0.1). There were 1102 upregulated DEGs and 1817 downregulated DEGs in response to L. paracasei subsp. paracasei 2004 feeding (Fig. 3A). Gene Ontology analysis with PANTHER32 revealed that genes related to glutathione metabolic process, cellular protein modification process, multicellular organism development, and response to stress were overrepresented among the upregulated DEGs (Fig. 3B); genes related to innate immune response, alpha-amino acid metabolic process, transmembrane transport, fatty acid process, and carboxylic acid catabolic process were overrepresented among the downregulated DEGs (Fig. 3B–E). Glutathione plays a vital role in the antioxidative process33, which is consistent with our observation that L. paracasei subsp. paracasei 2004 feeding improved oxidative stress resistance (Fig. 2C and Table S5).
L. paracasei subsp. paracasei 2004 feeding upregulates the expression of genes related to stress response. (A) Heatmap of differentially expressed genes (DEGs) in response to L. paracasei subsp. paracasei 2004 feeding (FDR cutoff < 0.1, minimum fold change = 2.0). L. paracasei subsp. paracasei 2004 feeding upregulates 1102 genes and downregulates 1817 genes. (B, C) Gene Ontology (GO) analysis of biological processes: the 1102 (> 2.0-fold, FDR < 0.1) upregulated gene list (B) and the 1817 (< 0.5-fold, FDR < 0.1) downregulated gene list were analyzed by PANTHER32. GO terms corresponding to biological processes whose p-value was less than 0.0001 were extracted. (D, E) Bar plots show the expression level of glutathione genes (D) and transcription factors, daf-16 and skn-1 (E).
The upregulated DEGs included daf-16 h, one of the daf-16 transcript variants, and skn-1 (Fig. 3E and Table S6), both of which are transcription factors promoting stress resistance and longevity in C. elegans29,30. This suggests that lactic acid bacterial strain L. paracasei subsp. paracasei 2004 feeding may induce these transcription factors and play an essential role in longevity. Thus, we validated the upregulation of the daf-16 h and skn-1 genes by lactic acid bacterial strain L. paracasei subsp. paracasei 2004 feeding. Our analyses indicated that the expression of daf-16 and daf-16 h was slightly upregulated with statistical significance while that of skn-1 was not (Fig. 4A). These results suggest that DAF-16 might be involved in L. paracasei subsp. paracasei 2004 feeding-induced longevity. To examine this possibility, we measured the lifespan of wild-type N2 and skn-1(zu135) and daf-16(mu86) deletion mutants fed L. paracasei subsp. paracasei 2004 (Fig. 4B). While L. paracasei subsp. paracasei 2004 feeding significantly increased the lifespan of wild-type animals, and that of the skn-1 deletion mutant, daf-16 deletion suppressed the longevity effect induced by L. paracasei subsp. paracasei 2004 feeding. These results suggest that DAF-16 plays a vital role in lifespan extension by L. paracasei subsp. paracasei 2004 feeding.
Longevity increase induced by L. paracasei subsp. paracasei 2004 feeding requires the transcription factor DAF-16. (A) Box plot shows the relative mRNA levels of daf-16 h (left), skn-1 (middle), and daf-16 (right) in animals fed the indicated bacterium. L. paracasei subsp. paracasei 2004 feeding upregulated the expression of daf-16 mRNA. P-values were calculated by an unpaired t-test. *, p < 0.05. (B) Survival curves of the wild-type N2 (left) and two mutants, EU31[skn-1(zu135)] and CF1038[daf-16(mu86)], fed the indicated bacterium. L. paracasei subsp. paracasei 2004 feeding extended the C. elegans lifespan in a DAF-16-dependent manner. P-values were calculated by a log-rank test with Bonferroni correction. **, p < 0.01. See Table S8 for detailed statistics.
Discussion
Our study aimed to investigate the potential effect of various lactic acid bacteria on health and longevity. We found that administration of L. acidophilus 1244 and L. paracasei subsp. paracasei 2004 has a longevity effect. Interestingly, short-term supplementation of these bacteria in early life (for 3 days from Day 1 of adulthood) significantly improved health, including food consumption, motility rate, and resistance to oxidative stress in middle-aged animals. These results suggest that supplementation with these bacteria ameliorates the fitness decline with age. Animals fed L. paracasei subsp. paracasei 2004 continued to grow, indicating that supplementation with this bacterial strain could have a beneficial probiotic effect on the host organism, C. elegans. On the other hand, we observed that feeding L. acidophilus 1244 resulted in stunted growth. Because some probiotics improve health34 and lifespan35 through mimicking dietary restriction, L. acidophilus 1244 administration might also improve health and lifespan by mimicking the dietary restriction. Further study should be done to examine mechanisms underlying improved health and lifespan by L. acidophilus 1244 administration.
The proposed pro-longevity mechanisms of probiotics include modulating immune responses and stress responses20. Supplementation of some lactic acid bacterial strains has been shown to enhance longevity in C. elegans through the transcription factors DAF-16 and SKN-1, both of which play a role in pro-longevity effects through modulating immune responses and stress responses23,25,36,37,38. Our study demonstrated that the longevity increase induced by L. paracasei subsp. paracasei 2004 supplementation requires DAF-16, but not SKN-1, and that administration with this bacterial strain increases the expression of genes related to stress responses but decreases the expression of genes related to the innate immune system. These results suggest that L. paracasei subsp. paracasei 2004 might increase lifespan by enhancing stress responses through the activity of DAF-16.
Overall, we found that the lactic acid bacterial strain L. paracasei subsp. paracasei 2004 can potentially improve health and lifespan with one-time supplementation for 3 days in C. elegans. The mechanism underlying this effect of the bacterial strain on healthy aging is mediated by DAF-16-induced stress response modulation, a highly conserved pro-longevity transcription factor. The involvement of DAF-16 in the beneficial effects of L. paracasei subsp. paracasei 2004 suggests potential applications in other organisms. Further research is needed to fully understand these mechanisms and assess their relevance in other organisms.
Methods
Bacterial strains and culture conditions
E. coli OP50 was provided by the Caenorhabditis Genetics Center, University of Minnesota (CGC), and used as a control food source. E. coli OP50 was grown in LB medium at 37 °C for 10–12 h with shaking. Lactic acid bacterial strains were grown at 37 °C in a modified GAM medium (29.5 g of Nissui modified GAM broth, 3.5 g of glucose, 0.5 g of tween80 in 500 mL of H2O) without shaking for one or two days until the lactic acid bacteria grew. Bacteria were harvested by centrifugation at 3,000 × g for 10 min and washed three times: twice in M9 buffer and then once in S basal buffer. Then, the bacteria were adjusted to a final concentration of 5 × 109 CFU (colony-forming units).
Nematodes and growth conditions
The C. elegans Bristol strain N2 and mutant strains EU31[skn-1(zu135)] and CF1038[daf-16(mu86)] were provided by the CGC. The Bristol N2 strain was used for all measurements except those in the longevity assay using the EU31[skn-1(zu135)] and CF1038[daf-16(mu86)] mutants. Animals were maintained on nematode growth medium (NGM) plates seeded with E. coli OP50, as explained before39 or lactic acid bacterial strains. We defined animals as Day 1 adult animals 3 days after hatching. Synchronized germ-free animals were obtained by the bleaching method40.
Body size measurements
Synchronized eggs were grown on NGM plates seeded with E. coli OP50 until animals reached adulthood. Then, the animals were transferred to M9 buffer containing E. coli OP50 or lactic acid bacterial strains. The animals were incubated with gentle shaking. After 3 days, images of the animals were taken on Day 4 of adulthood with a stereomicroscope (Olympus SZX16). Images were analyzed using ImageJ software.
Longevity assay
For Fig. 1C, trials were conducted on NGM plates containing 5-fluoro-2′-deoxyuridine (FUdR) at 100 μg/mL seeded with E. coli OP50 or lactic acid strains from Day 1 of adulthood until all animals died. For Fig. 4B, young adult (Day 1) animals (3 days after synchronization) were cultured in E. coli OP50 or L. paracasei subsp. paracasei 2004 suspended in S basal buffer with FUdR (100 μg/mL). After 3 days, the animals were transferred, and trials were conducted on NGM plates with FUdR (100 μg/mL) seeded with E. coli OP50. Mortality was scored every 2 or 3 days when the animals were transferred to a new plate. Animals were scored as dead if they failed to respond to being touched by a pick. Survival plots were generated by using single lifespan data.
Pharyngeal pumping rate measurements
For pharyngeal pumping rate measurements, Day 1 adulthood animals were cultured on NGM plates containing FUdR seeded with E. coli OP50, L. acidophilus 1244, or L. paracasei subsp. paracasei 2004 for 3 days. Animals’ pumping rate on food was recorded. The number of pumping events was measured under an optical microscope for 30 s. One pharyngeal pump was defined as a complete forward and backward movement of the grinder in the pharynx.
Bending rate measurements
For bending measurements, Day 1 adult animals (3 days after synchronization) were transferred to NGM plates containing FUdR seeded with E. coli OP50, L. acidophilus 1244, or L. paracasei subsp. paracasei 2004 for 3 days. After Day 4 of adulthood, animals were transferred and cultured on NGM plates seeded with E. coli OP50 until Day 10 of adulthood. Then, the animals’ bending rate in M9 buffer was recorded under an optical microscope (Olympus SZX16 with CellSens system) for 30 s. The number of bends was counted manually afterward.
Oxidative stress assay
For the oxidative stress assay, Day 1 adult animals were transferred to NGM plates containing FUdR seeded with E. coli OP50, L. acidophilus 1244, or L. paracasei subsp. paracasei 2004 for 3 days. After Day 4 of adulthood, animals were transferred and cultured on NGM plates seeded with E. coli OP50 until Day 10 of adulthood, and they were cultured for nine days on NGM plates seeded with E. coli OP50. Then, the animals were soaked in M9 buffer with 2 mM hydrogen peroxide. The animals’ movements in response to blue light were recorded hourly under the camera system (DMK23GP031, ImageingSource) for one minute. The movement of each animal was measured using Python (code used for analyses is provided in the Supplementary manuscript). Animals were scored as dead if they failed to move in response to blue light for two hours in a row. The oxidative stress assays were repeated twice, and survival plots were generated using single lifespan data.
RNA-seq analysis
Total RNA was isolated using TRIzol reagent (Invitrogen) from around 200 C. elegans Bristol strain N2 cultured on NGM plates with E. coli OP50 or L. paracasei subsp. paracasei 2004 for 3 days from Day 1 of adulthood. RNA-seq library preparation and RNA sequencing were performed at Macrogen Inc. using the Illumina HiSeq 2000 platform. FASTQC41 was used to inspect the quality scores of the raw sequence data and to look for biases. The reads were trimmed using the Cutadapt42 wrapper, TrimGalore43. The reads were mapped by aligning the software HISAT244 to the reference genome (WB235). The mapped reads were sorted by SAMtools45. Read counts per gene were obtained using Stringtie46. The DEGs obtained from RNA-seq-based expression profiling were analyzed by using iDEP0.96 (Integrated Differential Expression and Pathway analysis) online tools47. GO analysis on DEGs was performed using PANTHER32: statistical overrepresentation test of up-regulated or down-regulated DEGs was conducted with an option, biological GO terms. Expression level is log2(CPM: counts per million + 2).
Quantitative RT‒PCR
Total RNA was isolated using TRIzol reagent (Invitrogen) from around 200 C. elegans Bristol strain N2 cultured on NGM plates with E. coli OP50 or L. paracasei subsp. paracasei 2004 with FUDR supplementation for 3 days from Day 1 of adulthood. The isolated total RNA was reverse transcribed into single-stranded cDNA using ReverTra Ace qPCR RT Master Mix with gDNA remover (TOYOBO) according to the manufacturer’s protocol. Quantitative RT‒PCR was performed with an ABI 7300 Real-Time PCR system (Applied Biosystem) using Power SYBR® Green Master Mix (Thermo Fisher Scientific). Relative mRNA quantification was performed with the standard curve method. The relative mRNA levels were normalized to the expression of act-1, a C. elegans housekeeping gene. Primer sequences were determined using Primer3Plus (https://www.primer3plus.com/), and we used the primers that did not generate non-specific products or primer dimers, which was verified with the melt curve analysis.
-
act-1 Fw: 5′- CCCATCAACCATGAAGATCAA-3′
-
act-1 Rv: 5′-CACATCTGTTGGAAGGTGGA-3′
-
daf-16 h Fw: 5′-TTCTCACAGGACATGCAAGC-3′
-
daf-16 h Rv: 5′-ACGCTCTTGTTGATGGAGGT-3′
-
daf-16 Fw: 5′-TGGAATTCAATCGTGTGGAA-3′
-
daf-16 Rv: 5′-ATGAATATGCTGCCCTCCAG-3′
-
skn-1 Fw: 5′-CTCCATTCGGTAGAGGACCA-3′
-
skn-1 Rv: 5′-ACTGATCAGCAGGAGCCACT-3′
Data availability
The RNA sequencing data generated during the current study are available in the NCBI Sequence Read Archive (Accession#: GSE241495). All the other datasets are available from the corresponding author upon reasonable request.
References
Ghosh, T. S., Shanahan, F. & O’Toole, P. W. The gut microbiome as a modulator of healthy ageing. Nat. Rev. Gastroenterol. Hepatol. 19, 565–584. https://doi.org/10.1038/s41575-022-00605-x (2022).
Bairamian, D. et al. Microbiota in neuroinflammation and synaptic dysfunction: A focus on Alzheimer’s disease. Mol. Neurodegener. 17, 19. https://doi.org/10.1186/s13024-022-00522-2 (2022).
Cukrowska, B., Bierla, J. B., Zakrzewska, M., Klukowski, M. & Maciorkowska, E. The relationship between the infant gut microbiota and allergy the role of bifidobacterium breve and prebiotic oligosaccharides in the activation of anti-allergic mechanisms in early life. Nutrients 12, 946. https://doi.org/10.3390/nu12040946 (2020).
Ford, A. C., Sperber, A. D., Corsetti, M. & Camilleri, M. Irritable bowel syndrome. Lancet 396, 1675–1688. https://doi.org/10.1016/S0140-6736(20)31548-8 (2020).
Gomes, A. C., Hoffmann, C. & Mota, J. F. The human gut microbiota: Metabolism and perspective in obesity. Gut Microbes 9, 308–325. https://doi.org/10.1080/19490976.2018.1465157 (2018).
Gurung, M. et al. Role of gut microbiota in type 2 diabetes pathophysiology. EBioMedicine 51, 102590. https://doi.org/10.1016/j.ebiom.2019.11.051 (2020).
Safari, Z. & Gerard, P. The links between the gut microbiome and non-alcoholic fatty liver disease (NAFLD). Cell. Mol. Life Sci. 76, 1541–1558. https://doi.org/10.1007/s00018-019-03011-w (2019).
Witkowski, M., Weeks, T. L. & Hazen, S. L. Gut microbiota and cardiovascular disease. Circ. Res. 127, 553–570. https://doi.org/10.1161/CIRCRESAHA.120.316242 (2020).
Xu, Y. et al. The roles of the gut microbiota and chronic low-grade inflammation in older adults with frailty. Front. Cell. Infect. Microbiol 11, 675414. https://doi.org/10.3389/fcimb.2021.675414 (2021).
Wong, S. H. & Yu, J. Gut microbiota in colorectal cancer: Mechanisms of action and clinical applications. Nat. Rev. Gastroenterol. Hepatol. 16, 690–704. https://doi.org/10.1038/s41575-019-0209-8 (2019).
Salazar, J. et al. Microbiota and diabetes mellitus: Role of lipid mediators. Nutrients 12, 339. https://doi.org/10.3390/nu12103039 (2020).
Odamaki, T. et al. Age-related changes in gut microbiota composition from newborn to centenarian: A cross-sectional study. BMC Microbiol. 16, 90. https://doi.org/10.1186/s12866-016-0708-5 (2016).
Biagi, E. et al. Gut microbiota and extreme longevity. Curr. Biol. 26, 1480–1485. https://doi.org/10.1016/j.cub.2016.04.016 (2016).
Claesson, M. J. et al. Gut microbiota composition correlates with diet and health in the elderly. Nature 488, 178–184. https://doi.org/10.1038/nature11319 (2012).
Claesson, M. J. et al. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc. Natl. Acad. Sci. U S A 108(Suppl 1), 4586–4591. https://doi.org/10.1073/pnas.1000097107 (2011).
Clark, R. I. et al. distinct shifts in microbiota composition during drosophila aging impair intestinal function and drive mortality. Cell. Rep. 12, 1656–1667. https://doi.org/10.1016/j.celrep.2015.08.004 (2015).
Smith, P. et al. Regulation of life span by the gut microbiota in the short-lived African turquoise killifish. Elife https://doi.org/10.7554/eLife.27014 (2017).
Barcena, C. et al. Healthspan and lifespan extension by fecal microbiota transplantation into progeroid mice. Nat. Med. 25, 1234–1242. https://doi.org/10.1038/s41591-019-0504-5 (2019).
Hill, C. et al. The international scientific association for probiotics and prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 11, 506–514. https://doi.org/10.1038/nrgastro.2014.66 (2014).
Miller, B. C., Mathai, M., Yadav, H. & Jain, S. Geroprotective potential of microbiome modulators in the Caenorhabditis elegans model. Geroscience 46, 129–151. https://doi.org/10.1007/s11357-023-00901-7 (2024).
Lin, S. W. et al. Lactobacillus plantarum GKM3 promotes longevity, memory retention, and reduces brain oxidation stress in SAMP8 mice. Nutrients 13, 2860. https://doi.org/10.3390/nu13082860 (2021).
Depommier, C. et al. Supplementation with Akkermansia muciniphila in overweight and obese human volunteers: A proof-of-concept exploratory study. Nat. Med. 25, 1096–1103. https://doi.org/10.1038/s41591-019-0495-2 (2019).
Grompone, G. et al. Anti-inflammatory Lactobacillus rhamnosus CNCM I-3690 strain protects against oxidative stress and increases lifespan in Caenorhabditis elegans. PLoS One 7, e52493. https://doi.org/10.1371/journal.pone.0052493 (2012).
Kwon, G., Lee, J. & Lim, Y. H. Dairy propionibacterium extends the mean lifespan of Caenorhabditis elegans via activation of the innate immune system. Sci. Rep. 6, 31713. https://doi.org/10.1038/srep31713 (2016).
Nakagawa, H. et al. Effects and mechanisms of prolongevity induced by Lactobacillus gasseri SBT2055 in Caenorhabditis elegans. Aging Cell 15, 227–236. https://doi.org/10.1111/acel.12431 (2016).
Trent, C., Tsung, N. & Horvitz, H. R. Egg-laying defective mutants of the nematode Caenorhabditis elegans. Genetics 104, 619–647. https://doi.org/10.1093/genetics/104.4.619 (1983).
Chen, J. & Caswell-Chen, E. P. Facultative vivipary is a life-history trait in Caenorhabditis elegans. J. Nematol. 36, 107–113 (2004).
Bansal, A., Zhu, L. J., Yen, K. & Tissenbaum, H. A. Uncoupling lifespan and healthspan in Caenorhabditis elegans longevity mutants. Proc. Natl. Acad. Sci. U S A 112, E277-286. https://doi.org/10.1073/pnas.1412192112 (2015).
An, J. H. & Blackwell, T. K. SKN-1 links C. elegans mesendodermal specification to a conserved oxidative stress response. Genes Dev. 17, 1882–1893. https://doi.org/10.1101/gad.1107803 (2003).
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R. A C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464. https://doi.org/10.1038/366461a0 (1993).
Kenyon, C. J. The genetics of ageing. Nature 464, 504–512. https://doi.org/10.1038/nature08980 (2010).
Thomas, P. D. et al. PANTHER: A library of protein families and subfamilies indexed by function. Genome Res. 13, 2129–2141. https://doi.org/10.1101/gr.772403 (2003).
Monostori, P., Wittmann, G., Karg, E. & Turi, S. Determination of glutathione and glutathione disulfide in biological samples: An in-depth review. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 877, 3331–3346. https://doi.org/10.1016/j.jchromb.2009.06.016 (2009).
Goya, M. E. et al. Probiotic Bacillus subtilis protects against alpha-synuclein aggregation in C. elegans. Cell. Rep. 30, 367–380. https://doi.org/10.1016/j.celrep.2019.12.078 (2020).
Barathikannan, K. et al. Anti-obesity efficacy of Pediococcus acidilactici MNL5 in Canorhabditis elegans gut model. Int. J. Mol. Sci. 23, 1276. https://doi.org/10.3390/ijms23031276 (2022).
Lee, J., Kwon, G. & Lim, Y. H. Elucidating the mechanism of Weissella-dependent lifespan extension in Caenorhabditis elegans. Sci. Rep. 5, 17128. https://doi.org/10.1038/srep17128 (2015).
Shmookler Reis, R. J. et al. Modulation of lipid biosynthesis contributes to stress resistance and longevity of C. elegans mutants. Aging 3, 125–147. https://doi.org/10.18632/aging.100275 (2011).
Komura, T., Takemoto, A., Kosaka, H., Suzuki, T. & Nishikawa, Y. Prolonged lifespan, improved perception, and enhanced host defense of Caenorhabditis elegans by Lactococcus cremoris subsp. cremoris. Microbiol. Spectr. 10, e0045421. https://doi.org/10.1128/spectrum.00454-21 (2022).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94. https://doi.org/10.1093/genetics/77.1.71 (1974).
Stiernagle, T. Maintenance of C. elegans. WormBooK https://doi.org/10.1895/wormbook.1.101.1 (2006).
S., A. (http://www.bioinformatics.babraham.ac.uk/projects/fastqc, 2010).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Felix Krueger, F. J., Phil Ewels, Ebrahim Afyounian, Michael Weinstein, Benjamin Schuster-Boeckler, Gert Hulselmans, sclamons. FelixKrueger/TrimGalore: v0.6.10 - add default decompression path (0.6.10). (2023).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915. https://doi.org/10.1038/s41587-019-0201-4 (2019).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079. https://doi.org/10.1093/bioinformatics/btp352 (2009).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295. https://doi.org/10.1038/nbt.3122 (2015).
Ge, S. X., Son, E. W. & Yao, R. iDEP: An integrated web application for differential expression and pathway analysis of RNA-Seq data. BMC Bioinform. 19, 534. https://doi.org/10.1186/s12859-018-2486-6 (2018).
Acknowledgements
We thank the Caenorhabditis Genetic Center (USA), which is funded by the NIH Office of Research Infrastructure Programs, for providing the wild-type N2, EU31[skn-1(zu135)], and CF1038[daf-16(mu86)] strains.
Author information
Authors and Affiliations
Contributions
S.K., M.N., and M. U. performed the experiments with the help of Y.M., Y.T., and H.O. S.K. and M.U. designed the research under supervision by Y.M., Y.T., H.O., and E.N. M.U. and S.K. wrote the manuscript under the supervision of E.N.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kishimoto, S., Nono, M., Makizaki, Y. et al. Lactobacillus paracasei subsp. paracasei 2004 improves health and lifespan in Caenorhabditis elegans. Sci Rep 14, 10453 (2024). https://doi.org/10.1038/s41598-024-60580-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60580-y
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.