Abstract
Molecular diagnostics in cancer pharmacogenomics is indispensable for making targeted therapy decisions especially in lung cancer. For routine clinical practice, the flexible testing platform and implemented quality system are important for failure rate and turnaround time (TAT) reduction. We established and validated the multiplex EGFR testing by MALDI-TOF MS according to ISO15189 regulation and CLIA recommendation in Taiwan. Totally 8,147 cases from Aug-2011 to Jul-2015 were assayed and statistical characteristics were reported. The intra-run precision of EGFR mutation frequency was CV 2.15% (L858R) and 2.77% (T790M); the inter-run precision was CV 3.50% (L858R) and 2.84% (T790M). Accuracy tests by consensus reference biomaterials showed 100% consistence with datasheet (public database). Both analytical sensitivity and specificity were 100% while taking Sanger sequencing as the gold-standard method for comparison. EGFR mutation frequency of peripheral blood mononuclear cell for reference range determination was 0.002 ± 0.016% (95% CI: 0.000–0.036) (L858R) and 0.292 ± 0.289% (95% CI: 0.000–0.871) (T790M). The average TAT was 4.5 working days and the failure rate was less than 0.1%. In conclusion, this study provides a comprehensive report of lung cancer EGFR mutation detection from platform establishment, method validation to clinical routine practice. It may be a reference model for molecular diagnostics in cancer pharmacogenomics.
Similar content being viewed by others
Introduction
The paradigm of lung cancer therapy has shifted from the histopathology-based to the companion diagnosis-based guidance since the emergence of TKIs (tyrosine kinase inhibitors) and immune check point therapy1. One of the important landmarks in this shift is the advance in discovery of tumor driver mutations. Tumor onset, progression and drug resistance are involved in altered signaling pathways that modulate the cancer hallmarks including tumor cell proliferation, motility, adhesion, angiogenesis and apoptosis and immune escape2. Molecules that can reverse or compensate the effects caused by these alterations have the potential to develop as anti-tumor drugs for molecular targeting therapy (MTT) or immune therapy. MTT has been proved as an efficient and better strategy to benefit cancer patients with actionable mutations in various cancers, especially in lung cancer3,4,5. Since the mutation burden was diverse in different cancer types, certain genetic aberrations, so-called driver mutations, led tumor cells to depend on or addict to a mutation-dependent signaling pathway6,7. Hence, the identification of these driver mutations is necessary for the treatment decision. To identify patients who benefit from MTT by molecular testing is one of the most important issues of precision medicine. In lung cancer, the leading cause of cancer-related death worldwide8, the molecular testing-based target therapy has been routinely practiced. Recently, Kris et al. showed that patients with driver mutations who received the corresponding drugs had a prolonged progress-free survival than those with a driver mutation who did not receive the drugs and those without driver mutations9. This suggested that the companion molecular diagnostics-guided therapy is the trend in cancer management to improve patients’ survival10.
However, several critical issues that should be concerned including reliability, reproducibility, specimen amount, sample quality and turnaround time before cancer molecular testing become routine assays. Until now, many home-made and commercialized methods have been utilized for detecting the specific cancer associated gene mutations, such as EGFR mutations in lung cancer measured by Sanger sequencing, PCR-SSCP (single-strand conformation polymorphism), TaqMan PCR, Loop-hybrid mobility shift assay, cycleave PCR and PCR-RFLP11,12,13,14. To meet the more and more stringent clinical requests, high- or ultra-sensitive methods were enthusiastically developed including MALDI-TOF MS (matrix-assisted-laser-desorption-ionization time-of-flight mass spectrometry)15, PNA-LNA (peptide nucleic acid-locked nucleic acid) PCR clamp, Scorpions ARMS (amplified refractory mutation system), dHPLC (denaturing high performance liquid chromatography), single-molecule sequencing and digital PCR-based or next generation sequencing (NGS)-based strategies16,17,18,19. CE-marked or FDA-approved assays are validated in reliability, traceability, procedure standardization, easily and popularly used in routine clinical service as companion diagnoses especially EGFR assays in international multicenter clinical trials. However, with the rapid growing of novel actionable and druggable candidates, the laboratory developed tests (LDT) have higher flexibility to meet the immediate clinical requests. Even the issue of well characterized quality assurance has come to a consensus, the guideline or the regulation is still debated. This was because of a lot of high-sensitive and high throughput platforms were developed and the difficulties in method validation needed to be solved. This study aimed to establish a customized EGFR mutation molecular testing by MALDI-TOF MS and to validate the characteristics of this platform for routine clinical practice. The first-line EGFR TKI was reimbursed by National Health Insurance (NHI) in Taiwan since June 2011. In this report, we have conducted the EGFR mutation companion diagnostics from Aug-2011 to Jul-2015 in Taiwan. We focused on the quality issues including method validation, procedure, turnaround time and statistical characteristics. This can be a reference for cancer molecular diagnostics.
Results
Procedure of Mutation Testing by MALDI-TOF MS
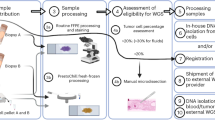
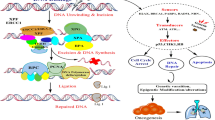
To establish routine clinical EGFR genetic testing in lung cancer patients, the pipeline of testing was firstly constructed (Fig. 1A). The experimental procedure was started from genomic DNA extraction from samples followed by PCR-based target amplification. After inactivation of dNTP by shrimp alkaline phosphatase (SAP) treatment, the single nucleotide extension reaction was performed by the specific probe annealed to one nucleotide before the mutation site. The incorporated ddNTP was different in the wild-type and mutant allele and the final products were further analyzed by MALDI-TOF MS. The mutation specific products can be distinguished from the wild-type ones in the spectrum due to the different molecular weights. The EGFR genetic testing was performed by the Pharmacogenomics Lab funded by National Research Program for Biopharmaceuticals (NRPB) Taiwan. All standard operation procedures were certified by ISO15189 regulation (Medical Laboratory 2695, No. L2695-140527). The testing process including sample receiving, testing and reporting was taken about four working days (Fig. 1B). Day 1 was initiated from sample receiving and unique barcode tabbing followed by nucleic acid extraction. Most of cases were continued to parts of biochemistry reactions. The biochemistry reactions included PCR, SAP reaction and single nucleotide extension. The reactions were ended at day 2 followed by MALDI-TOF MS analysis and data interpretation. At Day 3, the result was primary checked by laboratory scientists including quality control. After that, the final report was signed by two medical technologists at day 4. All cases adjudged to be needed confirmation at primary check will be repeated the testing procedure from biochemistry reaction.
Schematic presentation of the principle of mutation detection by MALDI-TOF MS and the molecular testing process.
(A) Genomic DNA extracted from samples was amplified by PCR primers. After inactivation of dNTP by SAP treatment, the target site-containing amplicons were further performed single nucleotide extension by the probe annealing to the nucleotide before the mutation site and ddNTP. The mutation specific product can be distinguished from the wild-type one in the mass spectrometry due to the incorporated nucleotide. (B) The procedure of molecular diagnostics can be completed within four working days starting with the sample receiving until data reported. dNTP, deoxynucleotide; ddNTP, dideoxynucleotide; SAP, shrimp alkaline phosphatase; PCR, polymerase chain reaction.
Quantification of EGFR Mutations Determined by MALDI-TOF MS
MALDI-TOF MS is a multi-function and flexible platform for gene testing. The major advantages are high sensitivity, low DNA quality requirement, capable of multiplex gene testing and quantification of mutation frequency. The principle of mutation quantification was shown in Supplementary Fig. 1A. The mutant allele competes with the wild-type allele for binding to detection probes. The ratio of mutant to wild-type signal height was calculated and reflected the percentage of mutant alleles among all alleles in the tested samples. To optimize the operation procedures of MALDI-TOF MS, the gDNAs from PBMC of healthy individuals and the DNAs from two well-established lung adenocarcinoma cell lines in which H1975 harbors both EGFR L858R and T790M mutations and PC9 harbors Del19 mutation were subjected to test as reference materials. The clear reproducible signals were obtained from MALDI-TOF MS for all control samples (Supplementary Fig. 1B). In EGFR L858R, Del19 and T790M detection, PBMC had no mutation signal while H1975 and PC9 showed the L858R/T790M mutation signals and the Del19 mutation signal, respectively. To test the repeatability and reproducibility, the calculated EGFR mutation frequency by MALDI-TOF MS was used as an index. Each control sample from 30 independent inspections was collected for variation analysis (Supplementary Fig. 1C). Among these mutations, PBMC had a background mutation frequency in L858R (0.0 ± 0.0), T790M (0.3 ± 0.3) and Del19 (1.4 ± 0.5). H1975 had high mutation frequency in L858R (67.4 ± 2.0), T790M (73.0 ± 1.2) but low mutation frequency in Del19 (1.3 ± 0.4). PC9 only had high mutation frequency in Del19 (87.1 ± 2.2) and low mutation frequency in L858R (0.0 ± 0.0) and T790M (0.3 ± 0.5). The coefficient of variation (CV) of mutation frequency were 2.98% for L858R (in H1975), 1.66% for T790M (in H1975) and 2.56% for Del19 (in PC9) respectively. These results suggested that MALDI-TOF MS can quantitatively and reproducibly detect EGFR mutations for clinical practice.
Precision, Reference Range and Limit of Detection
To verify the analytic validity of MALDI-TOF MS platform we first performed the intra-run and inter-run precision test (Fig. 2A,B). In the intra-run test, the EGFR L858R and T790M mutations of H1975 cells were assayed by independent four technicians in 20 replicates independently. In total 80 replicates, the averaged mutation frequency of L858R was 67.66 ± 1.46% with 2.15% CV while T790M was 73.94 ± 2.05% with 2.77% CV (Fig. 2A and Supplementary Table 1). In the mention of technical variation, the CVs of 20 replicates by each technician were ranged from 1.65% to 2.68% in L858R and 1.67% to 3.69% in T790M. In the inter-run test, the averaged mutation frequency of L858R was 67.75 ± 2.37% with 3.50% CV while T790M was 73.86 ± 2.10% with 2.84% CV in total 80 replicates (Fig. 2B and Supplementary Table 1). For each technician, the CVs of 20 replicates were ranged from 1.91% to 5.87% in L858R and 1.76% to 3.29% in T790M. The scatter plot of L858R vs T790M mutation frequency in total 160 replicates from the intra-run and inter-run showed high performance of MALDI-TOF MS in precision (Supplementary Fig. 2). The evaluation of precision for Del19 was also performed by using PC9 cells (Del E746-A750 mutation) as a reference material (Supplementary Fig. 3). The CV for Del19 was 0.79% in the intra-run test while 1.50% in the inter-run test. In the mention of reference range determination, 60 genomic DNAs from PBMC of healthy individuals were utilized as normal samples (Fig. 2C). The result indicated that the EGFR mutation frequency in PBMC was 0.002 ± 0.016% (95% CI: 0.000–0.036) in L858R and 0.292 ± 0.289% (95% CI: 0.000–0.871) in T790M and 1.658 ± 0.625% in Del19 (95% CI: 0.000–2.961) (Supplementary Table 2). Limit of detection (LOD) for EGFR mutation was defined as the lowest percentage of mutant allele content among wild-type allele background. It was determined by the serial dilutions made by mixing the mutant EGFR plasmids with wild-type ones (Fig. 2D). Among totally constant 1000 plasmid copies, the correlation between theoretical diluted mutation ratio and MALDI-TOF MS calculated mutation frequency was plotted. The R2 of diluted mutation ratio versus mutation frequency was 0.9837 in L858R and 0.9735 in T790M. However, the confident quantification of mutation frequency was around 1% (Fig. 2D, inserted box).
Precision, reference range and LOD of MALDI-TOF MS for EGFR mutation detection.
(A) Intra-run precision test. Four independent technicians (No. 1~4) performed testing in 20 replicates by using the EGFR L858R/T790M harboring cells, H1975. The mutation frequencies were plotted in the box chart. (B) Inter-run precision test. Four independent technicians (No. 1~4) performed testing in 20 replicates in independent runs by using H1975 cells. The mutation frequencies were plotted in the box chart. (C) Reference range identification. Sixty PBMC samples were assessed for the EGFR mutation testing. The mutation frequencies were plotted in the box chart. (D) LOD of MALDI-TOF MS in the EGFR mutation detection. Correlation of theoretic dilution ratios and calculated mutation frequencies was calculated by linear regression analysis in EGFR L858R and T790M detections. Each dilution was assayed in triplicate. LOD, limit of detection.
Accuracy Test, Analytical Sensitivity and Analytical Specificity
To address the accuracy of MALDI-TOF MS in EGFR mutation detection, we utilized the reference immortalized cell lines with naturally occurring disease-associated sequence variations or synthetic cloned DNA for testing according to the suggestion guideline20. All materials can be traced according to the information from quality documents, literatures, reference articles as well as database from bioresources (Table 1). In the double blind test, the EGFR mutation statuses including L858R, T790M and Del19 determined by MALDI-TOF MS were totally consistent with the statements in the public database. The artificial 50% mutant allele DNAs made up of the EGFR wild-type and L858R/T790M expression plasmids also exhibited the anticipated mutation frequency (Table 1 and Supplementary Fig. 4).
Analytical sensitivity and analytical specificity were tested by another set of 45 clinical FFPE samples (with sufficient amounts for quantitative DNA extraction, Supplementary Methods) from lung cancer patients and three PBMC samples. These samples were assessed for double blind EGFR mutation testing by traditional Sanger sequencing and MALDI-TOF MS methods in parallel (Table 2). None of EGFR L858R, T790M and Del19 was detected by both methods in three PBMC samples. Among the 45 FFPE samples, 14 had the L858R mutation and 9 had the Del19 mutation and one had L858R/T790M double mutations and 21 had no mutation. The results of MALDI-TOF MS were consistent with those of Sanger sequencing with 100% analytical sensitivity and 100% analytical specificity.
Routine Testing Characteristics and Quality Monitoring
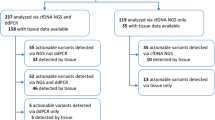
Since MALDI-TOF MS was established as the routine lung cancer molecular testing in Pharmacogenomics Lab, we analyzed totally 8,147 lung adenocarcinoma cases from Aug 2011 to Jul 2015 under ISO15189 regulation (Fig. 3 and Table 3). Among these cases, 4,299 cases were tested from Aug 2011 to Nov 2013 and parts of these (n = 1,772) had been reported in our previous study21. Additional 3,848 cases were tested from Dec 2013 to Jul 2015 by the same platform (additional 6,375 cases were included in this study). According to the statistical result, we analyzed 170 cases per month in average and 74.7% (n = 6,089) were FFPE samples (Table 3). Regarding to the testing fail rate, only 0.1% (n = 5) samples were fail in testing due to the poor DNA quality or reaction. Up to 94.6% (n = 7,708) of cases were reported at the first testing process while 5.3% samples (n = 434) were reported by further confirmation due to the inconsistence of replicates within one run (Table 3). Given EGFR L858R, T790M and Del19 mutations, the mutation prevalence were 24.7%, 3.8% and 23.1% respectively in tested cases similar with our previous study. The DNA concentrations from different sample types showed that all were various with a wide range. Pleura effusion and other sample types yielded related higher DNA concentration compared with other types (Supplementary Fig. 5). In each testing run, control materials including H1975 cell line harboring L858R/T790M and PBMC gDNA were assayed in parallel as a quality monitor of system. According to previous results, the H1975 cell line had stable mutation frequency and was suitable for systematic monitoring in the routine practice. Taking the mutation frequency of L858R or T790M in H1975 cells for Levey-Jennings quality graph, there were three L858R and one T790M tests out of 652 runs fail in quality monitoring (Fig. 3A,B). Starting from the sample receiving, we were in principle to report the data for clinical applicants in averaged 4.5 turnaround days (Fig. 3C).
Quality monitor and turnaround time of MALDI-TOF MS.
Levey-Jennings quality graph was used to monitor quality of MALDI-TOF MS by using DNA of H1975 cells from each run for the EGFR L858R mutation frequency (A) and the EGFR T790M mutation frequency (B). (C) The turnaround time of the EGFR mutation detection in 8,147 cases.
Discussion
Precision medicine points out that the treatment for individual cancer patient should consider their genetic information. Taking the advantage of new sequencing techniques and vast databases of information, the identification of potential actionable genetic aberrations is dramatically growing. On the other hand, this advance introduces an unprecedented revolutionary progress in laboratory practice. However, it has difficulties in the establishment of the standard operation procedure even consensus guidelines. According to the results from clinical trials, the prediction power of the molecular testing for therapeutic response was better than the traditional laboratory testing. The success of EGFR target therapy in lung cancer patients with EGFR mutations initiated the era of molecular diagnostics in cancer management4. In addition, the experience of prospective testing in Taiwan by Pharmacogenomics Lab can be a reference of cancer molecular testing in the future.
Even though we consider the precision issues including reproducibility (inter-run precision) and repeatability (intra-run precision) as well as the performance variations of technicians in this study, the accuracy is still a trouble due to the availability of reference materials. Herein, we utilized the traceable cell lines or synthetic DNAs as biomaterials to perform the accuracy testing according to recommendations20. It was noticed that occasionally EGFR mutations were detected in normal PBMCs with low mutation frequency (Fig. 2 and Supplementary Table 2). Based on the LOD established by the synthetic DNAs, the EGFR mutation frequency of PBMCs was lower than LOD. The data suggested that the low frequency found in PBMCs should be derived from the background noise of the assay. Characterization of the background is necessary for defining the cutoff value in routine practice. Furthermore, our results showed that the L858R mutation rate was 24.7% (2,012/8,147), Del19 mutation rate was 23.1% (1,884/8,147). The rate of overall EGFR activating mutations is consistent with the epidemiological statistics in Asian population4,22,23,24,25,26, this fact provides a robust clinical validation to prove the clinical utility of our system. To determine the analytical sensitivity and specificity of MALDI-TOF MS Sanger sequencing was acted as the gold standard method although the performance and successful rate of Sanger sequencing is largely limited in the poor DNAs or specimens. The main purpose of comparison between MALDI-TOF MS data and Sanger sequencing data is to perfect the analytic validity of MALDI-TOP MS not to investigate the limit of clinical specimen quality between both assays. Although the basic performance characteristics for Sanger sequencing had been mentioned, it still needed to consider whether the item of these characteristics should be concerned in the different mutation testing27. Furthermore, in our previous study have shown that some EGFR mutations of clinical specimens detected by a highly sensitive method cannot be identified by Sanger sequencing15.
Next, the TAT of molecular diagnostics was largely dependent on the methodology used and the average TAT was around two weeks (10 working days)28. The averaged working time for sequencing-based assays particularly for the case of NGS was four to five working days indicated that the post-analytical data processing is time-intensive and complex28. Our system exhibited a relative short turnaround time (4.5 working days) (Fig. 3 and Table 3). Finally, the DNA concentration of extracted samples is an issue. In this study, the DNA concentration was varied with a wide range (Supplementary Fig. 5) which may be attributed by several confounding factors including the handling process of samples, the size of biopsy, the basic property of sample type and the technical variation of extraction. In our routine practice, three 10μm thickness FFPE slices with over 0.5 cm-square tumor biopsy were recommended while the size with 2 mm cubic was recommended for fresh tissues.
In spite of growing up in sequencing-based or quantitative PCR-based detection platforms, more than 10 well-documented methods were used in EGFR mutation identification29. Recently, the emergence of NGS facilitated the high throughput and multiplex genetic testing in personalized medicine of cancers. Although the trend of NGS used in clinical molecular diagnostics was a consensus and authorized by US FDA, the risk-based regulatory framework was still a critical issue for quality assurance30.
Although the quality assurance of molecular diagnostics still had a lot of gray zone due to objective difficulties such as method validation, independent proficiency testing and reference material availability, many guide lines and consensus agreements from the expert workgroups consist of experts were established20. According to CLIA (Clinical laboratory improvement amendment) regulations, the analytical validation should consider several characteristics such as precision, accuracy, analytical sensitivity, analytical specificity, reference range and reportable range as well as other relevant performance metrics31. In house or LDT assays used in cancer pharmacogenomics testing should follow such kinds of regulations. Although there was still a gray zone in method validation and quality system of molecular diagnostics, more and more consensuses from experts will form mature regulations32,33,34. In this study we demonstrated that our system including MALDI-TOF MS and the entire validation process is a convincing system and adheres to the consensus guidelines of CLIA. The clinical utility of our system is confirmed by more than 8,000 patients with lung adenocarcinoma since 2011 to 2015. The first-line TKIs for the EGFR mutation patients identified by our system were reimbursed by Taiwan NHI. The goal of this study is to provide an update on recent developments for advanced NSCLC patients with EGFR mutations characterized by actionable molecular or histological alterations. Taken together, the molecular diagnostics of cancer pharmacogenomics aimed to understand and identify the genetic aberrations that influence drug efficacy and cytotoxicity in cancer patients. The pipeline of molecular diagnostics in cancer pharmacogenomics has been widely executed in worldwide such as United States, France, Japan, China, Germany and Taiwan9,21,35,36,37. Each step of cancer pharmacogenomics study to prepare for the clinical routine practice including testing cohort selection, sample size optimization, phenotype consideration, statistical analysis and validation needed to be carefully conducted38. The implementation required the cooperation between clinical physicians, pathologists, laboratory scientists and executive support. In conclusion, this study firstly provides the experience of an in-house molecular diagnostics system in cancer pharmacogenomics, especially EGFR mutations in lung cancer, from setup to routine practice and quality control in Taiwan.
Methods
Study cases
The 8,147 study cases were from a multicenter prospective observational trial approved by the Institutional Review Board of the participating institutes including IRB No. 201111039RIC (National Taiwan University Hospital Research Ethics Committee), IRB No. C08197 (Institutional Review Board of Taichung Veterans General Hospital), IRB No. DMR100-IRB-284(CR-2) (China Medical University and Hospital Research Ethics Committee), IRB No.CS12022 (Institutional Review Board of Chung Shan Medical University Hospital) and IRB No. REC102-7 (Taichung Tzu Chi Hospital Research Ethics Committee). Written informed consents for the genetic testing and clinical data records were obtained from all patients.
Genomic DNA Extraction, EGFR Mutation Detection by Sanger Sequencing and MALDI-TOF MS
Genomic DNAs were extracted from the tumor samples by using QIAmp DNA Minikit (QIAGEN, CA) according to the manufacturer’s instruction. The mutation analysis of EGFR by Sanger sequencing has been described previously39. Detection and quantification of EGFR mutations by MALDI-TOF MS was described in our previous studies15,40. The method and the procedure were detailed in the supplementary material (see the supplementary material for additional details).
Quality System
The testing performed in Pharmacogenomics Lab was under the regulation guideline of International Organization for Standardization (ISO) 15189. Pharmacogenomics Lab obtained ISO15189 certification from Taiwan Accreditation Foundation (TAF) since April-2013 (No. 2695). For external quality control, we participated proficiency test (PT) programs from College of American Pathologists (CAP) (Molecular Oncology, Program Code: EGFR) and European Molecular Genetics Quality Network (EMQN) (Program: lung cancer) twice a year since 2011.
Method Validation
Materials used for MALDI-TOF MS method validation including cell lines and control DNAs were purchased or obtained from ATCC, NCI or other institutes (Table 1). The validation items included precision, accuracy, analytical sensitivity, analytical specificity and reference range. The strategy and the procedure were detailed in the supplementary material (see the supplementary material for additional details).
Additional Information
How to cite this article: Su, K.-Y. et al. Implementation and Quality Control of Lung Cancer EGFR Genetic Testing by MALDI-TOF Mass Spectrometry in Taiwan Clinical Practice. Sci. Rep. 6, 30944; doi: 10.1038/srep30944 (2016).
References
Reck, M., Heigener, D. F., Mok, T., Soria, J. C. & Rabe, K. F. Management of non-small-cell lung cancer: recent developments. Lancet 382, 709–719, doi: 10.1016/S0140-6736(13)61502-0 (2013).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674, doi: 10.1016/j.cell.2011.02.013 (2011).
Rini, B. I. et al. Randomized phase III trial of temsirolimus and bevacizumab versus interferon alfa and bevacizumab in metastatic renal cell carcinoma: INTORACT trial. J Clin Oncol 32, 752–759, doi: 10.1200/JCO.2013.50.5305 (2014).
Mok, T. S. et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med 361, 947–957, doi: 10.1056/NEJMoa0810699 (2009).
Ismael, G. et al. Subcutaneous versus intravenous administration of (neo)adjuvant trastuzumab in patients with HER2-positive, clinical stage I-III breast cancer (HannaH study): a phase 3, open-label, multicentre, randomised trial. Lancet Oncol 13, 869–878, doi: 10.1016/S1470-2045(12)70329-7 (2012).
Weinstein, I. B. Cancer. Addiction to oncogenes–the Achilles heal of cancer. Science 297, 63–64, doi: 10.1126/science.1073096 (2002).
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489, doi: 10.1126/science.aab4082 (2015).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2015. CA: a cancer journal for clinicians 65, 5–29, doi: 10.3322/caac.21254 (2015).
Kris, M. G. et al. Using multiplexed assays of oncogenic drivers in lung cancers to select targeted drugs. Jama 311, 1998–2006, doi: 10.1001/jama.2014.3741 (2014).
McDermott, U., Downing, J. R. & Stratton, M. R. Genomics and the continuum of cancer care. N Engl J Med 364, 340–350, doi: 10.1056/NEJMra0907178 (2011).
Zhou, C., Ni, J., Zhao, Y. & Su, B. Rapid detection of epidermal growth factor receptor mutations in non-small cell lung cancer using real-time polymerase chain reaction with TaqMan-MGB probes. Cancer J 12, 33–39 (2006).
Yatabe, Y. et al. A rapid, sensitive assay to detect EGFR mutation in small biopsy specimens from lung cancer. J Mol Diagn 8, 335–341, doi: 10.2353/jmoldx.2006.050104 (2006).
Matsukuma, S. et al. Rapid and simple detection of hot spot point mutations of epidermal growth factor receptor, BRAF and NRAS in cancers using the loop-hybrid mobility shift assay. J Mol Diagn 8, 504–512, doi: 10.2353/jmoldx.2006.060030 (2006).
Marchetti, A. et al. EGFR mutations in non-small-cell lung cancer: analysis of a large series of cases and development of a rapid and sensitive method for diagnostic screening with potential implications on pharmacologic treatment. J Clin Oncol 23, 857–865, doi: 10.1200/JCO.2005.08.043 (2005).
Su, K. Y. et al. Pretreatment epidermal growth factor receptor (EGFR) T790M mutation predicts shorter EGFR tyrosine kinase inhibitor response duration in patients with non-small-cell lung cancer. J Clin Oncol 30, 433–440, doi: 10.1200/JCO.2011.38.3224 (2012).
Watanabe, M. et al. Ultra-Sensitive Detection of the Pretreatment EGFR T790M Mutation in Non-Small Cell Lung Cancer Patients with an EGFR-Activating Mutation Using Droplet Digital PCR. Clin Cancer Res, doi: 10.1158/1078-0432.CCR-14-2151 (2015).
Thomas, R. K. et al. Sensitive mutation detection in heterogeneous cancer specimens by massively parallel picoliter reactor sequencing. Nat Med 12, 852–855, doi: 10.1038/nm1437 (2006).
Kimura, H. et al. Detection of epidermal growth factor receptor mutations in serum as a predictor of the response to gefitinib in patients with non-small-cell lung cancer. Clin Cancer Res 12, 3915–3921, doi: 10.1158/1078-0432.CCR-05-2324 (2006).
Chin, T. M. et al. Detection of epidermal growth factor receptor variations by partially denaturing HPLC. Clin Chem 53, 62–70, doi: 10.1373/clinchem.2006.074831 (2007).
Gargis, A. S. et al. Assuring the quality of next-generation sequencing in clinical laboratory practice. Nat Biotechnol 30, 1033–1036, doi: 10.1038/nbt.2403 (2012).
Hsu, K. H. et al. Identification of five driver gene mutations in patients with treatment-naive lung adenocarcinoma in Taiwan. Plos one 10, e0120852, doi: 10.1371/journal.pone.0120852 (2015).
Girard, N. et al. Nomogram to predict the presence of EGFR activating mutation in lung adenocarcinoma. The European respiratory journal 39, 366–372, doi: 10.1183/09031936.00010111 (2012).
Sahoo, R. et al. Screening for EGFR mutations in lung cancer, a report from India. Lung cancer 73, 316–319, doi: 10.1016/j.lungcan.2011.01.004 (2011).
Taron, M. et al. Activating mutations in the tyrosine kinase domain of the epidermal growth factor receptor are associated with improved survival in gefitinib-treated chemorefractory lung adenocarcinomas. Clin Cancer Res 11, 5878–5885, doi: 10.1158/1078-0432.CCR-04-2618 (2005).
Wu, J. Y. et al. Gefitinib therapy in patients with advanced non-small cell lung cancer with or without testing for epidermal growth factor receptor (EGFR) mutations. Medicine 90, 159–167, doi: 10.1097/MD.0b013e31821a16f4 (2011).
Otani, H. et al. Detection of EGFR gene mutations using the wash fluid of CT-guided biopsy needle in NSCLC patients. J Thorac Oncol 3, 472–476, doi: 10.1097/JTO.0b013e31816de2cd (2008).
Pont-Kingdon, G. et al. Design and analytical validation of clinical DNA sequencing assays. Archives of pathology & laboratory medicine 136, 41–46, doi: 10.5858/arpa.2010-0623-OA (2012).
Konig, K. et al. Implementation of Amplicon Parallel Sequencing Leads to Improvement of Diagnosis and Therapy of Lung Cancer Patients. J Thorac Oncol 10, 1049–1057, doi: 10.1097/JTO.0000000000000570 (2015).
Pao, W. & Ladanyi, M. Epidermal growth factor receptor mutation testing in lung cancer: searching for the ideal method. Clin Cancer Res 13, 4954–4955, doi: 10.1158/1078-0432.CCR-07-1387 (2007).
Collins, F. S. & Hamburg, M. A. First FDA authorization for next-generation sequencer. N Engl J Med 369, 2369–2371, doi: 10.1056/NEJMp1314561 (2013).
Chen, B. et al. Good laboratory practices for molecular genetic testing for heritable diseases and conditions. MMWR. Recommendations and reports: Morbidity and mortality weekly report. Recommendations and reports/Centers for Disease Control 58, 1–37, quiz CE-31-34 (2009).
Leighl, N. B. et al. Molecular testing for selection of patients with lung cancer for epidermal growth factor receptor and anaplastic lymphoma kinase tyrosine kinase inhibitors: American Society of Clinical Oncology endorsement of the College of American Pathologists/International Association for the study of lung cancer/association for molecular pathology guideline. J Clin Oncol 32, 3673–3679, doi: 10.1200/JCO.2014.57.3055 (2014).
Rekhtman, N., Leighl, N. B. & Somerfield, M. R. Molecular testing for selection of patients with lung cancer for epidermal growth factor receptor and anaplastic lymphoma kinase tyrosine kinase inhibitors: american society of clinical oncology endorsement of the college of american pathologists/international association for the study of lung cancer/association for molecular pathology guideline. J Oncol Pract 11, 135–136, doi: 10.1200/JOP.2014.002303 (2015).
Lindeman, N. I. et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer and Association for Molecular Pathology. J Thorac Oncol 8, 823–859, doi: 10.1097/JTO.0b013e318290868f (2013).
Wang, R. et al. Analysis of major known driver mutations and prognosis in resected adenosquamous lung carcinomas. J Thorac Oncol 9, 760–768, doi: 10.1097/JTO.0b013e3182a406d1 (2014).
Serizawa, M. et al. Assessment of mutational profile of Japanese lung adenocarcinoma patients by multitarget assays: a prospective, single-institute study. Cancer 120, 1471–1481, doi: 10.1002/cncr.28604 (2014).
Nowak, F., Soria, J. C. & Calvo, F. Tumour molecular profiling for deciding therapy-the French initiative. Nature reviews. Clinical oncology 9, 479–486, doi: 10.1038/nrclinonc.2012.42 (2012).
Wheeler, H. E., Maitland, M. L., Dolan, M. E., Cox, N. J. & Ratain, M. J. Cancer pharmacogenomics: strategies and challenges. Nature reviews. Genetics 14, 23–34, doi: 10.1038/nrg3352 (2013).
Shih, J. Y. et al. Epidermal growth factor receptor mutations in needle biopsy/aspiration samples predict response to gefitinib therapy and survival of patients with advanced nonsmall cell lung cancer. Int J Cancer 118, 963–969, doi: 10.1002/ijc.21458 (2006).
Tsai, T. H. et al. RNA is favourable for analysing EGFR mutations in malignant pleural effusion of lung cancer. The European respiratory journal 39, 677–684, doi: 10.1183/09031936.00043511 (2012).
Acknowledgements
We thank the supported grants from MOST102-2325-B-002-078, MOST103-2325-B-002-026 (S.L.Y.), MOHW-TDU-B-211-113001, MOST103-2320-B-002-052 and MOST104-2320-B-002-030 (K.Y.S.) from Ministry of Science and Technology, Ministry of Education and Ministry of Health and Welfare and Mathematics in Biology Groups. The study was conducted by the members of Taiwan Clinical Trial Consortium for Lung Cancer.
Author information
Authors and Affiliations
Contributions
G.-C.H., C.-C.H. and S.-L.Y. initiated the project. K.-Y.S. performed the experiments and analyzed data. J.-T.K., H.-Y.C. and B.-C.H. analyzed data. K.-Y.S., J.-T.K. and S.-L.Y. contributed to manuscript preparation.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Su, KY., Kao, JT., Ho, BC. et al. Implementation and Quality Control of Lung Cancer EGFR Genetic Testing by MALDI-TOF Mass Spectrometry in Taiwan Clinical Practice. Sci Rep 6, 30944 (2016). https://doi.org/10.1038/srep30944
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep30944
This article is cited by
-
Molecular Genetic Techniques in Biomarker Analysis Relevant for Drugs Centrally Approved in Europe
Molecular Diagnosis & Therapy (2022)
-
Rapid Sputum Multiplex Detection of the M. tuberculosis Complex (MTBC) and Resistance Mutations for Eight Antibiotics by Nucleotide MALDI-TOF MS
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.