Abstract
Enzymes in biology’s energy chains operate with low energy input distributed through multiple electron transfer steps between protein active sites. The general challenge of biological design is how to lower the activation barrier without sacrificing a large negative reaction free energy. We show that this goal is achieved through a large polarizability of the active site. It is polarized by allowing a large number of excited states, which are populated quantum mechanically by electrostatic fluctuations of the protein and hydration water shells. This perspective is achieved by extensive mixed quantum mechanical/molecular dynamics simulations of the half reaction of reduction of cytochrome c. The barrier for electron transfer is consistently lowered by increasing the number of excited states included in the Hamiltonian of the active site diagonalized along the classical trajectory. We suggest that molecular polarizability, in addition to much studied electrostatics of permanent charges, is a key parameter to consider in order to understand how enzymes work.
Similar content being viewed by others
Introduction
Cytochrome c is an essential redox protein in bacterial photosynthesis and respiratory energy chains of mitochondria. Its redox function is to shuttle electrons between membrane-bound energy complexes, such as between the bc1 complex and cytochrome c oxidase in respiration1. The redox activity occurs in heme c covalently bound to the polypeptide2 (Fig. 1A). The mechanism of transferring the electron, which alters the redox state of the heme, is generally understood within the Marcus theory of electron transfer3. It stipulates that the reaction is activated by nuclear fluctuations of the thermal bath, which in the case of protein electron transfer is a highly heterogeneous protein-water interface. The prevailing modes, frequencies and coupling strengths of those medium modes to the heme’s electronic states are the parameters establishing the overall activation barrier of the reaction4,5,6.
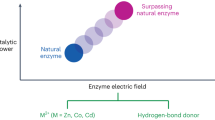
(A) Hydrated cytochrome c with the heme active site shown in green. (B) The free energy surfaces of a half reaction in the Marcus model (equation (1)) of fixed atomic charges (solid lines) and in the polarizable model with λSt < λ (equation (2)). (C) The active site representation by atomic charges qα in classical simulations and by a Hamiltonian matrix coupled to the classical bath in QM/MD simulations. (D) The mechanism of compensation of protein and water electrostatics through polarizing the interfacial water dipoles by the charged residues of the protein.
Hydration does not significantly affect vibrational cooling of the heme7 and THz absorption of well-hydrated samples is nearly insensitive to the oxidation state8. These observations suggest little direct contact of water with the heme7. Therefore, cytochrome c is a good model system to which basic assumptions of the Marcus theory apply9,10,11. Potential complications of water penetrating the active site12 and of conformational transitions upon changing the redox state2 are largely insignificant. Our present simulations support this general assessment when applied to the active site represented by fixed atomic charges. However, the main result of this study is the finding that polarizability of the active site extends the problem of protein electron transfer beyond the standard model13 by dramatic reduction of the activation barrier.
The established paradigm of the Marcus theory is based on the two-state description involving electronic energy levels of either the electron donor or the acceptor3. Nuclear fluctuations of the medium bring these two energy levels into resonance, allowing tunneling to occur14. The free energy (reversible work) required to create the resonance condition is determined by two parameters, the reaction free energy ΔG0 and the medium reorganization energy λ. The formulation further simplifies when ΔG0 is zero for either self-exchange electron transfer or for a half redox reaction occurring at the electrode. The activation free energy (activation barrier) is then fully determined by the reorganization energy3

Early calculations and numerical simulations of protein electron transfer produced values of the reorganization energy in the range of 0.7–0.8 eV (see ref. 5 for a review). More recent molecular dynamics (MD) simulations employing improved force fields and significantly longer trajectories resulted in an upward revision of these values toward those more traditional for redox chemistry, ~1.0–1.5 eV5,11,15,16 (or even higher17). The upward revision of the reorganization energy implies a higher activation barrier in equation (1) and a much slower rate. On the other hand, electrode kinetics measurements typically report much lower values, λ~0.4–0.6 eV18,19, when estimated from the Marcus relation (1). It implies that either the results of numerical simulations for λ are grossly incorrect or the relation between the activation barrier and the reorganization energy needs a revision. Here we present arguments that the latter is the case. The revision of the barrier height arises from introducing the polarizability of protein’s active site (Fig. 1B).
Equation (1) can be derived by considering two crossing parabolas Fi(X) = (X ± X0)2/(4λ) (i = Red, Ox) plotted against the energy gap reaction coordinate X as defined by Warshel20 (Fig. 1B). The crossing point F1(X) = F2(X) is the transition state of zero energy gap X = 0, where tunneling occurs21,22,23. The Marcus formulation follows from requiring X0 = λ as stipulated by the fluctuation-dissipation theorem24.
Recent simulations have shown that proteins are often unable to sample their entire phase space on the reaction time-scale. This ergodicity breaking6 eliminates the restriction on the reaction parameters imposed by the fluctuation-dissipation theorem25. In particular, X0 and λ become two separate parameters6,26 and finding the activation barrier requires three parameters (X0, λ and ΔG0), instead of two parameters of the Marcus theory. Sampling of the entire phase space is never realistically possible, but the problem is drastically elevated for proteins because of their rugged energy landscape, similar to those found for fragile glass formers27,28. The dynamics and statistics of proteins are characterized by many local minima, in which the protein-water system can be trapped, never reaching its true thermodynamic minimum29. Electron transfer reactions between non-equilibrium trapped states do not follow the strict restriction X0 = λ6.
The direct mechanistic consequence of this new perspective is an extended flexibility in fine-tuning the activation barrier of electron transfer6,30. Since X0 refers to the average of the vertical transition energy, it defines the position of the maximum of an optical spectroscopic line31 and can be associated with the Stokes shift of optical spectroscopy. One can therefore define the Stokes shift reorganization energy as λSt = X013. The three-parameter description leads to the following activation barrier when ΔG0 = 06

where the “reaction reorganization energy” λr can be identified with the Marcus reorganization energy in equation (1). λr = (λSt)2/λ is in fact consistent with the standard Marcus definition of the reorganization energy3 as the free energy invested to shift the system to the position of the products (thus moving 2λSt along X) while remaining on the free energy surface of the reactants characterized by the curvature (2λ)−1.
The reorganization energy in the denominator in equation (2) is defined as the variance of the reaction coordinate

where kB is the Boltzmann constant and T is the temperature. Note that long trajectories, >100 ns or longer32, are required to converge λ (see Supplementary Fig. S6). Because of this difficulty, most simulations, with few exceptions5, report λSt instead of λ.
It is clear that the activation barrier can be lowered compared to equation (1) of the Marcus model when λSt < λ. The parameter

quantifies the difference between two reorganization energies in the three-parameter model26. Note that electrochemical kinetic measurements report λr. The low values of such effective reorganization energies18,33,34 are therefore consistent with κG > 1 as schematically shown in Fig. 1B. It is also clear that the rate maximum, when the rate is plotted against the driving force −ΔG0 (the Marcus inverted parabola3), gives the value of λSt = X0 only and provides no access to λ.
Important for biological applications is that κG > 1 lowers the activation barrier without requiring more negative reaction free energy, which is a scarce commodity in biological energy chains1. It seems therefore possible that the evolutionary pressure has favored the glassy character of the protein fluctuations and their high fragility27, to promote electron transport consuming less free energy input for its operation.
The Marcus formulation of the electron-transfer theory can be viewed as the first-order quantum-mechanical perturbation of the electronic energy levels by the thermal bath. The perturbation Hamiltonian comes from integrating the electronic density ρe(r) with the electrostatic potential of the bath ϕ(r):  . When the electronic density is given by a set of atomic charges qα, one arrives at the force-field formulation often implemented in classical simulations. The solute-solvent Hamiltonian is obtained by summing up partial atomic charges with the bath potentials at their locations:
. When the electronic density is given by a set of atomic charges qα, one arrives at the force-field formulation often implemented in classical simulations. The solute-solvent Hamiltonian is obtained by summing up partial atomic charges with the bath potentials at their locations:  (Fig. 1C). However, fluctuations of the medium not only alter the donor-acceptor energy gap (between HOMO and LUMO), but also the entire manyfold of the electronic energy states. Each instantaneous nuclear configuration of the medium will produce a different extent of electronic delocalization between those available electronic states, or, alternatively, a different deformation of the electronic density.
(Fig. 1C). However, fluctuations of the medium not only alter the donor-acceptor energy gap (between HOMO and LUMO), but also the entire manyfold of the electronic energy states. Each instantaneous nuclear configuration of the medium will produce a different extent of electronic delocalization between those available electronic states, or, alternatively, a different deformation of the electronic density.
The ability of the electronic distribution to deform in an external field is associated with its electronic polarizability. In the dipolar approximation, it is given in terms of transition dipoles μkm linking different electronic sates of the molecule through the electric field of the bath Eb (Fig. 1C). The standard quantum-mechanical perturbation theory leads to the quadratic Stark effect35, shifting the energy level k by the amount  scaled with the polarizability of that state
scaled with the polarizability of that state

It is determined by a set of transition dipoles and energy gaps ΔEmk = Em − Ek of all possible virtual excitations.
Perturbation theory is not required to introduce polarizability into the description of electron transfer. A more accurate formalism is achieved by using the empirical valence-bond approximation introduced by Warshel and Weiss36,37,38. It produces the instantaneous energies of the donor and acceptor by diagonalizing the Hamiltonian matrix incorporating the coupling to the medium into the diagonal (electrostatics) and off-diagonal (transition dipoles) matrix elements. This approach has been widely used for a number of biologically relevant systems in the past39,40 and has recently been implemented in the form of the perturbed matrix algorithm11 in application to protein electron transfer. We follow this general formalism in the simulations presented in this paper. Our main goal is to explore the possibility of lowering the barrier for electron transfer by including mixing between the quantum states (polarizability). From the more fundamental perspective, our study asks the question of whether including polarizability of the enzyme’s active site, in contrast to the picture of fixed atomic charges, might reduce the barrier of an enzymetic reaction. In other words, the question is whether polarizability is one of the “tools” of biology’s catalytic capability41.
Results
QM/MD simulations
The goal of our simulation strategy is to go beyond the assumption of fixed atomic charges in the modeling of the redox active site. We introduce the ability of the electronic density of the heme in cytochrome c to redistribute in response to a thermal fluctuation of the bath. This goal is shared by essentially all QM/MM algorithms which all start by defining the quantum center, i.e., a part of the system which can be treated on the quantum-mechanical (QM) level5,30,40,42,43,44. The choice of the level of QM calculations is dictated by the physics of the problem and, to a large degree, by the time-scale required to capture the essential collective modes of the thermal bath contributing to the activation barrier45. Protein electron transfer is a difficult problem for QM algorithms because long time scales are very essential here. Classical simulations of electron transfer have shown that a broad range of bath time-scales affects the reorganization energy5,26. The time-scales of ~1 ns represent global elastic deformations of the protein shape, which have to be included for a realistic description of λ. These motions produce large fluctuations of electrostatic potential inside the protein by shifting charged surface residues and surface water polarized by them6 (Fig. 1D). As more elastic modes enter the observation window (the length of the simulation trajectory), the reorganization energy grows nearly continuously through the range of time-scales up to tens of microseconds currently reached by simulations32. Given these constraints imposed by the physics of the problem, a QM algorithm needs to capture the entire range of thermal motions sampled by classical simulations.
The method of perturbed matrix11 imposes essentially no QM overhead on the classical MD. It assumes that the forces acting on the atoms of the classical thermal bath can be well characterized by classical force fields. One therefore starts with a long classical MD simulation of the entire system producing the dynamics of the classical bath. These classical dynamics are then used to recalculate the parameters of the quantum center affected by the electrostatic interactions with the bath. Since long-range electrostatics is the main factor influencing the positions of the donor and acceptor energy levels involved in electron transfer4, this algorithm is particularly well suited for this problem.
The choice of the QM center (Fig. 1C and Supplementary Fig. S4) is explained in detail in the Supplementary Information. Briefly, several QM centers of increasing size have been tried. The largest QM center adopted for the analysis includes the heme, histidine, methionine and two cysteine amino acids. As we show in Supplementary Table S1, there is a fairly insignificant change in the spectrum of electronic states between this QM center and a somewhat smaller one, without two cysteine residues.
The QM component of the analysis was performed by expanding the electrostatic potential of the bath ϕ(r) around the potential ϕFe at the heme iron and truncating the expansion at the dipolar term. The matrix elements of the quantum center Hamiltonian then become

where Q is the total charge of the quantum center. The quantum states j = 0,…, M include the ground state of the quantum center, j = 0 and a number of its excited states produced here by ZINDO/S calculations for the oxidized (Ox, Q = −1) and reduced (Red, Q = −2) states. The polarizability is a slowly converging function of the number of excited states M46; the results presented here were obtained for M = 100. Reducing M makes the quantum center less polarizable and eventually brings the system back to the Marcus domain. This was the result of a recent calculation employing M = 1211.
The Hamiltonian matrix in equation (6) is diagonalized at each instantaneous value of the potential ϕFe and the electric field Eb along the simulation trajectory to produce the minimum eigenvalues  corresponding to the ground state in either oxidized or reduced states of the active site. The electron-transfer reaction coordinate, monitoring the transition to the activation state X = 0, is given as20
corresponding to the ground state in either oxidized or reduced states of the active site. The electron-transfer reaction coordinate, monitoring the transition to the activation state X = 0, is given as20

The limit of classical simulations is obtained by representing the quantum center by a set of atomic charges coupled to the bath through the corresponding electrostatic potentials ϕα (Fig. 1C). The reaction coordinate of electron transfer becomes in this case

where  . More details on the protocols of quantum calculations and of classical simulations are provided in Methods below and in the Supplementary Information. Here we discuss the results of our analysis.
. More details on the protocols of quantum calculations and of classical simulations are provided in Methods below and in the Supplementary Information. Here we discuss the results of our analysis.
Free energy surfaces of electron transfer
The free energy surfaces of electron transfer Fi(X) = −kBT ln[Pi(X)] (i = Ox, Red) follow from probabilities Pi(X) calculated from classical trajectories with the quantum (equation (7)) or classical (equation (8)) definitions of the reaction coordinate X. Since our focus is on a half reaction, we do not consider a more complex problem of calculating the redox potential5,43 and focus solely on the reorganization energies. We first note that the quantum and classical algorithms are consistent with each other when the polarizability of the active site is turned off (Fig. 2A). In order to study the effect of the active site polarizability, we introduced scaling of the transition dipoles with the scaling factor ξ: μmk → ξμmk. The non-polarizable active site corresponds to ξ = 0 when coupling between the quantum states is turned off. Even in this limit, the algorithms of calculating X are still somewhat different in the quantum and classical cases since we use an expansion of the potential in the quantum Hamiltonian in equation (6), in contrast to a full set of atomic charges in the classical MD. However, the free energy surfaces obtained in these two approaches are consistent (Fig. 2A), suggesting a little effect of re-distributing the charge within the active site on the electron-transfer barrier. One can further examine the effect of charge distribution in the active site by assuming complete localization of the charge on heme’s iron, ΔqFe = 1. This extreme case is compared to the calculation with distributed charge in Table 1. The results are consistent.
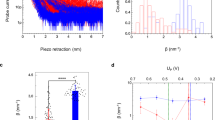
Free energy surfaces Fi(X), i = Red, Ox of cytochrome c in the Ox (left curves) and Red (right curves) states.
The blue points/lines refer to classical MD and the red points/lines refer to the QM/MD simulations. The solid lines are fits of the simulation data to parabolas. Panel (A) refers to a non-polarizable quantum center (ξ = 0). Panel (B) refers to a polarizable quantum center with ξ = 1 and Δα = −31 Å3. The lower panel demonstrates the depression of the barrier height upon allowing a non-zero αk (see Supplementary Fig. S7 for ξ = 2, Δα = −123 Å3).
Little sensitivity to electron delocalization might seem to be a trivial outcome since standard electrostatics suggests that the interaction of a point charge (localization) with the surrounding medium is equivalent to that of a charge uniformly spread over the conducting surface (delocalization). However, it is often suggested that delocalizing the electronic density of the active site is an optimization mechanism to reduce the reorganization energy43. While this mechanism is expected to lower the reorganization energy of localized skeletal vibrations2,15, we see little evidence for it altering the medium (protein and water) reorganization energy. It is also useful to keep in mind that most reactions relevant to biology’s energy chains occur at relatively small magnitudes of the driving force −ΔG0 and, therefore, proceed in the normal region of electron transfer when quantum vibrations have essentially no effect on the activation barrier47. We instead argue here that the reorganization energy λ is maximized and not minimized, for polarizable active sites. Since λSt remains nearly unaffected, the activation barrier in equation (2) can be reduced (Fig. 1B) due to a large value of the parameter κG (equation (4)).
Effect of polarizability on the reorganization energy
The main goal of our analysis is to establish whether λ can significantly exceed λSt when polarizability of the protein’s active site is turned on. We first note that  , in accord with the standard Marcus theory3, in the classical MD simulations (Table 1). This result is in agreement with previous simulations of this protein9,11, although we still find κG > 1. A reasonable agreement with the Marcus theory found here is not always shared by other redox proteins. We have found λ > λSt for a number of redox proteins (
, in accord with the standard Marcus theory3, in the classical MD simulations (Table 1). This result is in agreement with previous simulations of this protein9,11, although we still find κG > 1. A reasonable agreement with the Marcus theory found here is not always shared by other redox proteins. We have found λ > λSt for a number of redox proteins ( for electron transfer reactions in bacterial reaction centers6,26,32). The reasons why non-polarizable cytochrome c falls on the lower end of κG values are important to understand. We discuss below the mechanism of compensation between the protein and water fluctuations lowering κG. Here we first look at how altering the physical model from a set of fixed atomic charges to a fluctuating charge distribution affects the activation barrier.
for electron transfer reactions in bacterial reaction centers6,26,32). The reasons why non-polarizable cytochrome c falls on the lower end of κG values are important to understand. We discuss below the mechanism of compensation between the protein and water fluctuations lowering κG. Here we first look at how altering the physical model from a set of fixed atomic charges to a fluctuating charge distribution affects the activation barrier.
Table 1 and Fig. 3 summarize our findings. The reorganization energy λ is calculated according to equation (3), while 2λSt = 〈X〉Red − 〈X〉Ox is calculated from the average energy gaps in two redox states. The polarizability of the quantum center is continuously increased in Fig. 3 by scaling the ZINDO/S transition dipoles, μjk → ξμjk. The corresponding polarizabilities, calculated from equation (5), are listed in Table 1. As mentioned above, the polarizability significantly drops when fewer states are included and the statistics of the electron-transfer energy gap returns back to  of the Marcus theory (Supplementary Table S2). Even with M = 100 excited states, our calculations are likely to underestimate the polarizability of the active site46. The scaling parameter ξ > 1 allows us to estimate the potential effect of an increased polarizability on the reaction activation barrier.
of the Marcus theory (Supplementary Table S2). Even with M = 100 excited states, our calculations are likely to underestimate the polarizability of the active site46. The scaling parameter ξ > 1 allows us to estimate the potential effect of an increased polarizability on the reaction activation barrier.
Increasing the polarizability clearly separates λOx/Red from λSt (Fig. 3). According to equation (2), this should lower the activation energy, as is also seen from direct calculations shown in Fig. 2B. The main result of our calculations is that electron transfer involving polarizable active sites should proceed with lower activation barriers, without requiring more negative reaction free energy. Why this is the case can be understood from the following general arguments. The reversible work of creating a fluctuation of the bath field is a quadratic function of the field,  , in linear response. The negative free energy invested in polarizing the solute reduces this energy as
, in linear response. The negative free energy invested in polarizing the solute reduces this energy as  . One expects, therefore, a smaller activation barrier to reach X = 0, as we observe. There is also a possibility of breaking the harmonic stability at
. One expects, therefore, a smaller activation barrier to reach X = 0, as we observe. There is also a possibility of breaking the harmonic stability at  , when λ passes through a spike48. Since λ enters the denominator in Eq. (2) and λSt is not much affected by polarizability (Fig. 3), one can anticipate that an electron-transfer enzyme can reach its lowest activation barrier if operating in the regime
, when λ passes through a spike48. Since λ enters the denominator in Eq. (2) and λSt is not much affected by polarizability (Fig. 3), one can anticipate that an electron-transfer enzyme can reach its lowest activation barrier if operating in the regime  .
.
Electrostatics of protein and water
The overall reorganization energy is a gauge of the strength of thermal fluctuations affecting the active site (equation (3)), with water and protein being its two main components. It is therefore of great mechanistic interest to understand what are the relative contributions of protein and water to fluctuations experienced by the active site. In contrast to some early suggestions that soluble proteins can effectively screen water from the active site and thus produce an effectively nonpolar environment, a number of recent simulations have clearly shown that water can never be neglected5,49,50 and reorganization energies are comparable in magnitude to those traditionally reported for synthetic donor-acceptor complexes5,11,16,32. These new results are in line with the current shift of the view on the relation between proteins and water suggesting that water should be viewed as an integral part of the protein51. However, an upward revision of the reorganization energy for protein electron transfer raises an important mechanistic question of how high efficiency of biological energy chains is achieved.
One first needs to realize that there is a significant screening between the water and protein contributions to the electron-transfer energy gap. The water dipoles are oriented by the ionized surface residues of the protein to produce the electrostatic potential opposite in sign to the potential of the protein (Fig. 1D). As a result, the protein (p) and water (w) contributions to X0 = 〈Xw〉 + 〈Xp〉 are typically opposite in sign and similar in magnitude (Fig. 4A). The value of X0 is the result of their incomplete compensation. The same physics applies to the variance of X, that is to the reorganization energy in equation (3).
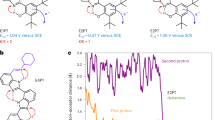
(A) Distribution of the electron-transfer coordinate (black) and its protein (green) and water (blue) parts (classical MD). The solid lines refer to the Ox state and the dashed lines refer to the Red state. The distance between the maxima of Red and Ox distributions is the Stokes shift, 2λSt. (B) Loss function χ′′(ν), 2πν = ω from protein (green), water (blue) and total (black) fluctuations of X. The solid black line shows the classical MD and the dashed line represents the QM/MD simulations at ξ = 1 (Table 1). The loss functions are normalized to give the corresponding reorganization component from  . The shaded areas represent separate Debye relaxation processes used to fit the time correlation function CX(t) from classical MD simulations.
. The shaded areas represent separate Debye relaxation processes used to fit the time correlation function CX(t) from classical MD simulations.
The reorganization energy obtained from equation (3) becomes the sum of three components: protein, λp, water, λw and a cross component, λpw = 〈δXpδXw〉/(kBT), produced by correlated protein and water fluctuations. Consistent with the opposite signs of 〈Xp〉 and 〈Xw〉, the cross component is negative and compensates much larger individual protein and water contributions52. For instance, for λOx = 1.67 eV listed in Table 1, one has λp = 2.28 eV, λw = 3.39 eV and λpw = −4.0 eV (Supplementary Table S3).
Dynamics
The compensation between the protein and water fluctuations, displayed in the overall value of λ, shows itself even more dramatically in the Stokes shift dynamics of the energy gap variable X(t). To study the dynamics, one has to turn to time correlation functions. The simplest one is the binary auto-correlation function CX(t) = 〈δX(t)δX(0)〉, where δX(t) = X(t) − X0. The t = 0 value of this correlation function is proportional to the reorganization energy and one can anticipate that the physics of protein-water electrostatic compensation should extend into the time domain. It does, but we also find new dynamics pertinent to each component, which loses its prominence in the overall Stokes-shift dynamics due to the compensation effect.
Figure 4B shows the loss spectrum of the Stokes-shift dynamics. The loss function χ′′(ω) characterizes the rate of energy exchange, at a given frequency, between the active site and the thermal bath. It can be thought of as the rate of energy dissipation, at a given frequency, of some energy (e.g., photon) absorbed by the active site. The overall energy dissipated into surrounding is the integral of χ′′(ω)/ω over all frequencies. In our calculations, 2kBTχ′′(ω) = ωCX(ω) is obtained from the frequency Fourier transform of the time correlation function24.
The peaks of χ′′(ω) show the characteristic relaxation times of the modes coupled to the electron-transfer coordinate and their intensities represent the coupling strengths. The main striking observation from the plot is the presence of slow dynamics in both the protein and water components characterized by nearly equal relaxation times. These common dynamics, in the nanosecond time domain (see SI), represent elastic modes altering the shape of the protein and simultaneously shifting the surface water molecules oriented by charged protein residues (Fig. 1D)6. The slow dynamics, however, nearly disappear in the overall χ′′(ω) due to a strong compensation (screening) between protein and water electrostatic contributions. It is this compensation that brings λ in a near accord with λSt in the case of non-polarizable heme of cytochrome c. The lack of this compensation makes the two reorganization energies deviate from each other, often significantly, for other proteins6.
Mechanistic aspects
Our QM/MD calculations produce the effective reorganization energy in equation (2)  eV, λ = (λOx + λRed)/2 not far from ~0.6 eV viewed to be the average number from solution-based measurements19. The error bars of the reorganization energy calculations are displayed in Fig. 3 and are additionally listed in Supplementary Table S3. The errors of reorganization energy calculations from QM/MD do not exceed 4%.
eV, λ = (λOx + λRed)/2 not far from ~0.6 eV viewed to be the average number from solution-based measurements19. The error bars of the reorganization energy calculations are displayed in Fig. 3 and are additionally listed in Supplementary Table S3. The errors of reorganization energy calculations from QM/MD do not exceed 4%.
One still has to be aware that the present simulations do not directly include polarizability of water and molecular groups of the protein in the production runs. A number of recent simulations employed polarizable force fields5,30,45,50. The polarizability is included a posteriori in our analysis of the trajectories by calculating the energy of induction interaction of the electric field of the QM center with the induced dipoles of the protein and water49,53 (see Supplementary Information for details). The induction (bath polarizability) component of X0 can be significant, but it mostly cancels out54 in λSt. The remaining nonzero contribution to the reorganization energy is caused by density fluctuations modulating the induction interaction energy.
The reorganization energies can potentially decrease if induced dipoles are included. The Pekar factor of dielectric models predicts a drop of λ by the factor  when switching from a nonpolarizable to a polarizable dielectric;
when switching from a nonpolarizable to a polarizable dielectric;  and
and  are, correspondingly, the static and high-frequency dielectric constants of the thermal bath. However, simulations of model systems55 show that this drop is an overestimate and the reorganization energy decreases only by ~20% upon the inclusion of induced dipoles. All these results apply, however, to the Marcus picture with λSt = λ. It is currently unclear how induced dipoles affect each distinct reorganization energy λSt and λ. In addition, a drop in the magnitude of the reorganization energy upon including induced dipoles is mostly off-set by the reorganization energy arising from translational motions of induced dipoles (induction reorganization energy, see Supplementary Information).
are, correspondingly, the static and high-frequency dielectric constants of the thermal bath. However, simulations of model systems55 show that this drop is an overestimate and the reorganization energy decreases only by ~20% upon the inclusion of induced dipoles. All these results apply, however, to the Marcus picture with λSt = λ. It is currently unclear how induced dipoles affect each distinct reorganization energy λSt and λ. In addition, a drop in the magnitude of the reorganization energy upon including induced dipoles is mostly off-set by the reorganization energy arising from translational motions of induced dipoles (induction reorganization energy, see Supplementary Information).
As mentioned above, κG > 1 requires either incomplete sampling (ergodicity breaking), when some configurations are not accessible, or the breakdown of the Gaussian statistics of the energy gap fluctuations and non-parabolic free energy surfaces. The latter scenario is indeed realized for donor-acceptor systems with polarizabilities different between the two electron-transfer states13. However, this scenario requires λOx ≠ λRed. This seems to be generally true for polarizable active sites (Table 1), but the extent of deviation is hard to estimate with limited sampling available from protein simulations. We also note that the dynamics of the energy gap X(t) follow the Gaussian approximation. It is tested by the ability to produce the fourth-order time correlation function of the energy gap in terms of the Stokes shift dynamics (Supplementary Fig. S8)56. Overall, we cannot clearly assign κG > 1 found in our simulations to the non-Gaussian character of the energy gap fluctuations.
Discussion
Energy chains of biology rely on a very short list of redox centers to transfer electrons1. They mostly include hemes of cytochromes, iron-sulfur clusters and cupredoxins. One wonders if they are used to allow distinctly different electron-transfer mechanisms or have been selected based on similar mechanistic properties. A partial answer comes from biology. Cytochrome c6, a heme protein, is used interchangeably with plastocyanin, a cupredoxin, in cyanobacteria to catalytically connect photosystems I and II2 (only plastocyanin is used in higher plants). Numerical simulations have shown that λ ≫ λSt, attributed in this study to a high polarizability of the active site, is achieved in plastocyanin through insufficient compensation between water and protein electrostatics, which does not require a polarizable active site26. Does it mean that evolutionary pressure chooses redox proteins with λ ≫ λSt, regardless of the mechanism producing the desired result? We do not have a definitive answer at this time. Studies of the effect of polarizability on electron transfer in all three classes of redox centers are required to address this question.
What our study convincingly shows is that increasing the polarizability of protein’s active site can significantly reduce the activation barrier of a catalytic reaction, electron transfer in this case. Interaction of atomic charges of the active site with the electrostatic potential of the surrounding medium is clearly an essential part of the enzyme’s catalytic action57. It might be true as well that not only the distribution of molecular charge, but also its ability to deform in the external field (polarizability) is an important “tool” employed by nature to catalyze biological reactions.
Polarizability of an active site is controlled by a set of transition dipoles and energy gaps between the state of the transferred electron and the excited electronic states (equation (5)). Minimizing the energy gaps and maximizing the transition dipoles enhances the polarizability. The primary pair of bacterial photosynthesis (a sandwich of closely spaced and parallel oriented bacteriochlorophylls58) is a dramatic example showing that polarizability can be effectively manipulated by design. The two bacteriochlorophylls form a partial charge transfer state with a strong electronic coupling and a low energy gap between the eigenstates49. The resulting polarizability of the photoexcited state59, ~103 Å3, is among the highest in the molecular world and it affects the energetics of primary charge separation49. The active site polarizability can be viewed as an additional “knob” turned by evolutionary pressure through chemical manipulation of the excited electronic states.
Methods
The NMR solution structure of horse heart cytochrome c (PDB 1GIW) was adopted as the starting configuration for classical MD simulations. The simulations were done with NAMD software suite60, with the trajectory length of 250 ns and 33231 TIP3P water molecules in the simulation box. Additional details of the simulation protocol and of the quantum calculations are given in the Supplementary Information.
Additional Information
How to cite this article: Dinpajooh, M. et al. Polarizability of the active site of cytochrome c reduces the activation barrier for electron transfer. Sci. Rep. 6, 28152; doi: 10.1038/srep28152 (2016).
References
Nicholls, D. G. & Ferguson, S. J. Bioenergetics 3 (Academic Press, London, 2002).
Liu, J. et al. Metalloproteins Containing Cytochrome, Iron-Sulfur, or Copper Redox Centers. Chem. Rev. 114, 4366–4469 (2014).
Marcus, R. A. & Sutin, N. Electron transfer in chemistry and biology. Biochim. Biophys. Acta 811, 265–322 (1985).
Simonson, T. Electrostatics and dynamics of proteins. Rep. Prog. Phys. 66, 737–787 (2003).
Blumberger, J. Recent Advances in the Theory and Molecular Simulation of Biological Electron Transfer Reactions. Chem. Rev. 115, 11191–11238 (2015).
Matyushov, D. V. Protein electron transfer: is biology (thermo)dynamic? J. Phys.: Condens. Matter 27, 473001 (2015).
Zhang, Y. & Straub, J. E. Diversity of solvent dependent energy transfer pathways in heme proteins. J. Phys. Chem. B 113, 825–830 (2009).
He, Y., Chen, J. Y., Knab, J. R., Zheng, W. & Markelz, A. G. Evidence of protein collective motions on the picosecond timescale. Biophys. J. 100, 1058–1065 (2011).
Muegge, I., Qi, P. X., Wand, A. J., Chu, Z. T. & Warshel, A. The reorganization energy of cytochrome c revisited. J. Phys. Chem. B 101, 825–836 (1997).
Simonson, T. Gaussian fluctuations and linear response in an electron transfer protein. Proc. Natl. Acad. Sci. 99, 6544–6549 (2002).
Bortolotti, C. A. et al. The Reversible Opening of Water Channels in Cytochrome c Modulates the Heme Iron Reduction Potential. J. Am. Chem. Soc. 134, 13670–13678 (2012).
Chakrabarty, S. & Warshel, A. Capturing the energetics of water insertion in biological systems: The water flooding approach. Proteins 81, 93–106 (2013).
Matyushov, D. V. & Voth, G. A. Modeling the free energy surfaces of electron transfer in condensed phases. J. Chem. Phys. 113, 5413 (2000).
Gray, H. B. & Winkler, J. R. Long-range electron transfer. Proc. Natl. Acad. Sci. 102, 3534–3539 (2005).
Blumberger, J. & Klein, M. L. Reorganization free energies of long-range electron transfer in porphyrin-binding four-helix bundle protein. J. Am. Chem. Soc. 128, 13854 (2006).
McCullagh, M. & Voth, G. A. Unraveling the Role of the Protein Environment for [FeFe]-Hydrogenase: A New Application of Coarse-Graining. J. Phys. Chem. B 117, 4062–4071 (2013).
Woiczikowski, P. B., Steinbrecher, T., Kubar, T. & Elstner, M. Nonadiabatic QM/MM Simulations of Fast Charge Transfer in Escherichia coli DNA Photolyase. J. Phys. Chem. B 115, 9846–9863 (2011).
Yue, H. et al. On the electron transfer mechanism between cytochrome c and metal electrodes. Evidence for dynamic control at short distances. J. Phys. Chem. B 110, 19906–19913 (2006).
Alvarez-Paggi, D., Zitare, U. & Murgida, D. H. The role of protein dynamics and thermal fluctuations in regulating cytochrome c/cytochrome c oxidase electron transfer. Biochim. Biophys. Acta - Bioenergetics 1837, 1196–1207 (2014).
Warshel, A. Dynamics of reactions in polar solvents. Semiclassical trajectory studies of electron-transfer and proton-transfer reactions. J. Phys. Chem. 86, 2218–2224 (1982).
Beratan, D. N., Betts, J. N. & Onuchic, J. N. Tunneling pathway and redox-state-dependent electronic couplings at nearly fixed distance in electron transfer proteins. J. Phys. Chem. 96, 2852–2855 (1992).
Hayashi, T., Stuchebrukhov, A. A. & Gray, H. B. Electron tunneling in respiratory complex I. Proc. Natl. Acad. Sci. USA 107, 19157–19162 (2010).
Zhang, Y., Liu, C., Balaeff, A., Skourtis, S. S. & Beratan, D. N. Biological charge transfer via flickering resonance. Proc. Natl. Acad. Sci. USA 111, 10049–10054 (2014).
Kubo, R. The fluctuation-dissipation theorem. Rep. Prog. Phys. 29, 255–284 (1966).
Crisanti, A. & Ritort, F. Violation of the fluctuation-dissipation theorem in glassy systems: basic notions and the numerical evidence. J. Phys. A: Math. Gen. 36, R181–290 (2003).
LeBard, D. N. & Matyushov, D. V. Protein-water electrostatics and principles of bioenergetics. Phys. Chem. Chem. Phys. 12, 15335–15348 (2010).
Angell, C. A. Formation of glasses from liquids and biopolymers. Science 267, 1924–1935 (1995).
Debenedetti, P. G. & Stillinger, F. H. Supercooled liquids and glass transitions. Nature 410, 259 (2001).
Frauenfelder, H. et al. A unified model of protein dynamics. Proc. Natl. Acad. Sci. USA 106, 5129–5134 (2009).
Cailliez, F., Müller, P., Firmino, T., Pernot, P. & de la Lande, A. Energetics of photoinduced charge migration within the tryptophan tetrad of an animal (6-4) photolyase. J. Am. Chem. Soc. 138, 1904–1915 (2016).
Marcus, R. A. Relation between charge transfer absorption and fluorescence spectra and the inverted region. J. Phys. Chem. 93, 3078–3086 (1989).
Martin, D. R. & Matyushov, D. V. Microsecond dynamics of the protein and water affect electron transfer in a bacterial bc1 complex. J. Chem. Phys. 142, 161101 (2015).
Monari, S. et al. Thermodynamic and kinetic aspect of electron transfer reaction of bovide cytochrome c immobilized on 4-mercaptopyridine and 11-mercapto-1-undecanoid acid films. J. Appl. Electrochem. 38, 885 (2008).
Monari, S. et al. Understanding the mechanism of short-range electron transfer using an immobilized cupredoxin. J. Am. Chem. Soc. 134, 11848–11851 (2012).
Lockhart, D. J. & Boxer, S. G. Stark effect spectroscopy of Rhodobacter sphaeroides and Rhodopseudomonas viridis reaction centers. Proc. Natl. Acad. Sci. USA 85, 107–111 (1988).
Warshel, A. & Weiss, R. M. An empirical valence bond approach for comparing reactions in solutions and in enzymes. J. Am. Chem. Soc. 102, 6218–6226 (1980).
Warshel, A. Computer modeling of chemical reactions in enzymes and solutions. (Wiley Interscience, New York, 1991).
Kamerlin, S. C. L. & Warshel, A. The empirical valence bond model: theory and applications. Wiley Interdisciplinary Reviews: Computational Molecular Science 1, 30–45 (2011).
Gao, J. & Truhlar, D. G. Quantum Mechanical Methods for Enzyme Kinetics. Annu. Rev. Phys. Chem. 53, 467–505 (2002).
Senn, H. M. & Thiel, W. QM/MM Methods for Biomolecular Systems. Angewandte Chemie International Edition 48, 1198–1229 (2009).
Garcia-Viloca, M., Gao, J., Karplus, M. & Truhlar, D. G. How enzymes work: Analysis by modern rate theory and computer simulations. Science 303, 186–195 (2004).
Truhlar, D. G. et al. The incorporation of quantum effects in enzyme kinetics modeling. Acc. Chem. Res. 35, 341–349 (2002).
Cascella, M., Magistrato, A., Tavernelli, I., Carloni, P. & Rothlisberger, U. Role of protein frame and solvent for the redox properties of azurin from Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. 103, 19641–19646 (2006).
de la Lande, A., Gillet, N., Chen, S. & Salahub, D. R. Progress and challenges in simulating and understanding electron transfer in proteins. Archives of Biochemistry and Biophysics 582, 28–41 (2015).
Kubař, T. & Elstner, M. A hybrid approach to simulation of electron transfer in complex molecular systems. J. R. Soc. Interface 10, 20130415 (2013).
Zheng, L., Polizzi, N. F., Dave, A. R., Migliore, A. & Beratan, D. N. Where Is the Electronic Oscillator Strength? Mapping Oscillator Strength across Molecular Absorption Spectra. J. Phys. Chem. A 120, 1933–1943 (2016).
Bixon, M. & Jortner, J. Electron transfer - from isolated molecules to biomolecules. Adv. Chem. Phys. 106, 35 (1999).
Dinpajooh, M. & Matyushov, D. V. Interfacial structural transition in hydration shells of a polarizable solute. Phys. Rev. Lett. 114, 207801 (2015).
LeBard, D. N., Kapko, V. & Matyushov, D. V. Energetics and kinetics of primary charge separation in bacterial photosynthesis. J. Phys. Chem. B 112, 10322–10342 (2008).
Tipmanee, V., Oberhofer, H., Park, M., Kim, K. S. & Blumberger, J. Prediction of Reorganization Free Energies for Biological Electron Transfer: A Comparative Study of Ru-Modified Cytochromes and a 4-Helix Bundle Protein. J. Am. Chem. Soc. 132, 17032–17040 (2010).
Linke, K. & Ho, F. M. Water in Photosystem II: Structural, functional and mechanistic considerations. Biochim. Biophys. Acta - Bioenergetics 1837, 14–32 (2014).
Nilsson, L. & Halle, B. Molecular origin of time-dependent fluorescence shift in proteins. Proc. Natl. Acad. Sci. 102, 13867 (2005).
Sterpone, F., Ceccarelli, M. & Marchi, M. Linear response and electron transfer in complex biomolecular systems and a reaction center protein. J. Phys. Chem. B 107, 11208–11215 (2003).
Matyushov, D. V. & Schmid, R. Optical and radiationless intramolecular electron transitions in nonpolar fluids: Relative effects of induction and dispersion interactions. J. Chem. Phys. 103, 2034–2049 (1995).
Gupta, S. & Matyushov, D. V. Solvent and solute polarizability effects on the reorganization energy of electron transfer. J. Phys. Chem. A 108, 2087 (2004).
Dinpajooh, M. & Matyushov, D. V. Non-Gaussian Lineshapes and Dynamics of Time-Resolved Linear and Nonlinear (Correlation) Spectra. J. Phys. Chem. B 118, 7925–7936 (2014).
Warshel, A. et al. Electrostatic basis for enzyme catalysis. Chem. Rev. 106, 3210–3235 (2006).
Hoff, A. J. & Deisenhofer, J. Photophysics of photosynthesis. Phys. Rep. 287, 1–247 (1997).
Middendorf, T. R., Mazzola, L. T., Lao, K. Q., Steffen, M. A. & Boxer, S. G. Stark effect (electroabsorption) spectroscopy of photosynthetic reaction centers at 1.5 K: evidence that the special pair has a large excited-state polarizability. Biochimica et Biophysica Acta 1143, 223 (1993).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Acknowledgements
This work was supported by the NSF (CHE-1464810 and TG-MCB080116N). We are grateful to Dr. Aschi for sharing with us the results of ref. 11 and to Prof. Beratan for useful discussions.
Author information
Authors and Affiliations
Contributions
M.D., D.R.M. and D.V.M. designed research, performed research, analyzed data and co-wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Dinpajooh, M., Martin, D. & Matyushov, D. Polarizability of the active site of cytochrome c reduces the activation barrier for electron transfer. Sci Rep 6, 28152 (2016). https://doi.org/10.1038/srep28152
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep28152
This article is cited by
-
Electron-transfer chain in respiratory complex I
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.