Abstract
Protocadherin 19 (Pcdh19) is an X-linked gene belonging to the protocadherin superfamily, whose members are predominantly expressed in the central nervous system and have been implicated in cell-cell adhesion, axon guidance and dendrite self-avoidance. Heterozygous loss-of-function mutations in humans result in the childhood epilepsy disorder PCDH19 Girls Clustering Epilepsy (PCDH19 GCE) indicating that PCDH19 is required for brain development. However, understanding PCDH19 function in vivo has proven challenging and has not been studied in mammalian models. Here, we validate a murine Pcdh19 null allele in which a β-Geo reporter cassette is expressed under the control of the endogenous promoter. Analysis of β-Geo reporter activity revealed widespread but restricted expression of PCDH19 in embryonic, postnatal and adult brains. No gross morphological defects were identified in Pcdh19+/β-Geo and Pcdh19Y/β-Geo brains and the location of Pcdh19 null cells was normal. However, in vitro migration assays revealed that the motility of Pcdh19 null neurons was significantly elevated, potentially contributing to pathogenesis in patients with PCDH19 mutations. Overall our initial characterization of Pcdh19+/β-Geo, Pcdh19β-Geo/β-Geo and Pcdh19Y/β-Geomice reveals that despite widespread expression of Pcdh19 in the CNS and its role in human epilepsy, its function in mice is not essential for brain development.
Similar content being viewed by others
Introduction
Protocadherins comprise a large family of single pass transmembrane glycoproteins that have important roles in cell-cell adhesion, dendrite self-avoidance and axon guidance. In mammals, ~70 protocadherin genes have been identified, the majority of which are located in three genomic “clusters” termed α, β and γ. The remainder, termed “non-clustered protocadherins” are scattered throughout the genome and are classified as δ1, δ2 and “other”. Non-clustered protocadherins are widely expressed in the central nervous system and have been implicated in homotypic cell adhesion, neuronal migration and synaptic plasticity1,2,3,4.
Pcdh19 is an X-chromosome linked member of the δ2 protocadherin subfamily and is highly expressed in the developing mouse cortex and hippocampus5. PCDH19 is highly conserved across human, mouse, zebrafish and chicken and has weak homotypic cell adhesion properties. It has been shown to interact with N-cadherin and members of the WAVE complex (Nap1 and Cyfip2) suggesting a role in actin cytoskeletal dynamics6,7. Genetic disruption of the closely related Pcdh10 or Pcdh17 genes (members of the δ2 subfamily) in mice have revealed roles in axon extension, regulation of presynaptic assembly and growth of striatal axons and thalamocortical projections8,9,10. These studies raise the possibility Pcdh19 functions similarly during mouse brain development; however this remains unexplored due to the lack of a validated Pcdh19 null mouse.
The majority of studies investigating the endogenous role of Pcdh19 have been performed in zebrafish11,12,13. These studies have utilized either morpholino knockdown or mutation of Pcdh19 with each approach producing different phenotypes. Morpholino knockdown results in a severe phenotype where cell motility in the neural plate is compromised resulting in a severe disruption to early brain morphogenesis13. In contrast, pcdh19 null zebrafish generated by TALEN-induced frame-shift mutation are viable and fertile and exhibit disruption of columnar architecture in the optic tectum resulting in impaired visually guided behaviours12. While these data support a role for pcdh19 in cell-cell recognition during zebrafish brain development, the discordance between the mutant and morphant phenotypes, which is commonly observed in zebrafish14, makes interpretation of these data difficult.
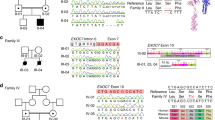
In humans, heterozygous PCDH19 mutations cause PCDH19 GCE5,15 which is now recognized as the second most common cause of monogenic epilepsy16. Mutations of PCDH19 that cause PCDH19 GCE are mainly missense, however as some cases have complete gene deletions, all pathogenic mutations are almost certainly loss-of-function. The phenotype of PCDH19 GCE is variable – affected girls present with symptoms ranging from benign focal epilepsy with normal cognitive function to severe seizures and intellectual disability that resemble Dravet syndrome16,17. The inheritance pattern of PCDH19 GCE is highly unusual for an X-linked gene and is described as X-linked dominant with male sparing i.e. heterozygous females are affected whereas hemizygous males are not. Due to random X-chromosome inactivation affected females are composed of a mosaic population of normal and PCDH19 mutant cells. Intriguingly, rare cases of affected males have also been described, which arise from somatic mutation and also display mosaicism5,18. It has been proposed that the mosaicism of normal PCDH19 and mutant expressing cells leads to ‘scrambling’ of the neuronal circuitry in the brain of affected individuals. However, experimental support for this ‘cellular interference’ model remains limited.
Here we investigate the in vivo expression and function of Pcdh19 using mutant mice where the Pcdh19 gene is disrupted by a β-galactosidase-neomycin (β-Geo) reporter cassette. At this allele, β-Geo was expressed under the control of the endogenous Pcdh19 promoter, whilst Pcdh19 expression itself was ablated (i.e. a knock-out allele). Using β-Geo expression and X-Gal staining we show that Pcdh19 expression is widely expressed in the developing CNS through to adulthood. Although brain morphology in Pcdh19Y/β-Geo, Pcdh19β-Geo/β-Geo and Pcdh19+/β-Geo mutants appears grossly normal, in vitro analysis indicates that Pcdh19 null neurons have a slight but significant increase in motility. These data suggest more subtle roles for Pcdh19 beyond establishment of gross brain architecture.
Results
Generation and validation of a Pcdh19 null mouse model
The Pcdh19 gene contains six exons, with exon 1 encoding the majority of the protein including the entire extracellular and transmembrane domains (Fig. 1A). To investigate the physiological role of Pcdh19 and the molecular pathology of PCDH19 GCE, we acquired a Pcdh19 null mouse model in which exons 1–3 were replaced with a β-geo cassette (Fig. 1B). Removal of the first 3 exons leads to deletion of both the extracellular and transmembrane domains of PCDH19. The genetic disruption of Pcdh19 gene was validated by PCR of genomic DNA (Fig. 1C) and ablation of Pcdh19 expression confirmed by qPCR of 2 week old hippocampal cDNA (Supplementary Fig. 1).
Generation of Pcdh19 Null Mice.
(A) Pcdh19 contains 6 exons. Exon 1 encodes more than half of the protein including the entire extracellular domain and transmembrane domain. (B) Pcdh19 null mice were generated by Taconic through targeted homologous recombination. Exon 1, 2 and 3 were removed and replaced with a β-Galactosidase-Neomycin fusion cassette. (C) PCR was used to confirm the presence of the β-Galactosidase/Neomycin fusion cassette with primers P1, P2 and P3 (black arrowheads in B). The β-Galactosidase-Neomycin fusion cassette was only present in Pcdh19+/β-Geo and Pcdh19Y/β-Geo mice.
Previous in situ hybridisation analysis has demonstrated Pcdh19 expression in the hippocampus and cortex of postnatal mouse brains5. However, expression of endogenous PCDH19 protein has not been shown. To investigate this issue, we performed western blot analysis using a commercially available PCDH19 antibody. Consistent with mRNA expression data, robust expression was observed in the hippocampus and cortex (Fig. 2A).
Validation of Pcdh19 Null mice.
(A) Protein extracts from hippocampus, cortex and cerebellum brain regions were immunoblotted with PCDH19 antibody. PCDH19 was detected in wild type extracts but was absent from Pcdh19Y/β-Geo extracts. (B) Frozen sections from the same regions as A) were stained for β-Galactosidase activity using X-Gal. Staining was present in both Pcdh19+/β-Geo and Pcdh19Y/β-Geo brain sections but absent from wild type brain sections. Scale bar represents 250 μm.
Integration of the β-geo cassette into the Pcdh19 locus potentially provides a reporter allele for null cells that would normally express Pcdh19. To confirm the activity of the β-geo cassette, X-Gal staining was performed on brain tissue from Pcdh19Y/β-Geo, Pcdh19+/β-Geo and Pcdh19+/Y mice. Pcdh19Y/β-Geo brains exhibit positive staining in multiple brain regions including the hippocampus, cortex and cerebellum (Fig. 2B) which is consistent with western blot analysis (Fig. 2A) and previously published RNA expression data5. Pcdh19+/β-Geo brains showed staining in the same brain regions but exhibited a mosaic staining pattern (Fig. 2B). Pcdh19+/Y brains showed no staining. This confirms the β-Geo cassette is active and that Pcdh19 undergoes random X-inactivation across multiple tissues similar to results in humans19.
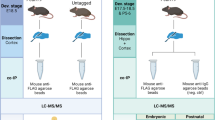
Chemical fractionation of adult hippocampal samples revealed that PCDH19 was present within the synaptosome, providing evidence for a role in synaptic function (Fig. 3A). Importantly, no signal was detected in Pcdh19Y/β-Geo samples, further confirming PCDH19 loss-of-function in this KO mouse model. Furthermore overexpression of FLAG tagged human PCDH19 (PCDH19-FLAG) in primary hippocampal mouse neurons revealed PCDH19 localisation with SYNAPSIN further supporting PCDH19 expression in synapses (Fig. 3B).
PCDH19 is expressed in the synapses of hippocampal neurons.
(A) Synaptosome fractionation of P14 mouse hippocampus was performed and immunoblotted for PCDH19. PCDH19 protein was present in the synaptosome fraction which was also enriched for the synapse proteins PSD95 and SYNAPTOPHYSIN. PCDH19 was absent from all Pcdh19Y/β-Geo fractions. (B) Primary neurons were harvested from 18.5 dpc hippocampi and nucleofected with PCDH19-FLAG ORF. Immunostaning at DIV12 shows PCDH19 is colocalised with SYNAPSIN positive synapses. Scale bar represents 20 μm.
Pcdh19+/β-Geo, Pcdh19β-Geo/β-Geo and Pcdh19Y/β-GeoMice have No Gross Phenotypic Abnormalities
Breeding of the Pcdh19 mouse model generated Pcdh19+/β-Geo, Pcdh19β-Geo/β-Geo and Pcdh19Y/β-Geo mice at the expected frequencies (Supplementary Fig. 2). Adult mice with these genotypes had normal body size and did not exhibit any overt health issues (data not shown). To date, spontaneous seizures have not been detected in Pcdh19+/β-Geo, Pcdh19β-Geo/β-Geo or Pcdh19Y/β-Geo mice after weaning (ie. from 3 weeks of age).
Morpholino knock-down of pcdh19 in zebrafish embryos results in profound CNS abnormalities. We therefore examined gross brain morphology in Pcdh19 null mice. Histological analysis of Pcdh19+/β-Geo, Pcdh19β-Geo/β-Geo and Pcdh19Y/β-Geo brains showed no gross morphological abnormalities (Fig. 4A). As Pcdh19 is highly expressed in the hippocampus and cortex, we performed morphometric analysis of these regions to determine if more subtle defects may be present. Measurements were made of the cortical thickness and CA1 region of the hippocampus with no significant difference observed in Pcdh19+/β-Geo, Pcdh19β-Geo/β-Geo and Pcdh19Y/β-Geo adult brains compared with Pcdh19+/+ and Pcdh19Y/+controls (Fig. 4B). We therefore conclude that mutation of Pcdh19 in mice does not overtly affect brain structure.
Histological and morphometric analysis of adult Pcdh19 mutant mouse brains.
(A) Haematoxylin and Eosin staining was performed on coronal P42 brain sections. Scale bar represents 1 mm. (B) Thickness measurement of the lateral parietal association cortex and hippocampus CA1 region in WT and Pcdh19 mutant brains. Three animals were analysed for each genotype. Error bars = SEM.
Pcdh19 Null Cells are Located Normally in the Developing and Adult Brain
Previous studies in zebrafish have shown that morpholino knockdown of Pcdh19 results in severe disruption of brain morphogenesis due to an arrest of cell convergence in the anterior neural plate as well as abnormal cell migration during neurulation11,13, although this was not seen in pcdh19 null zebrafish12. To investigate whether mutation of Pcdh19 in mice affects neuronal migration during brain development we looked for ectopic localisation of β-Geo positive cells in developing (15.5 dpc) brains of Pcdh19+/β-Geo and Pcdh19Y/β-Geo mice using X-gal staining. As the Pcdh19-specific antibody used for western blot analysis was not compatible with immunohistochemistry (data not shown), wild type Pcdh19-expressing cells were identified by in situ hybridisation. At 15.5 dpc, robust Pcdh19 expression was detected in the developing cortex and hippocampus of Pcdh19Y/+brains (Fig. 5A). Wild type Pcdh19-expressing cells were also present in the cortex and hippocampus of Pcdh19+/β-Geo brains, although these were fewer in number consistent with X-inactivation (Fig. 5B). Importantly, analysis of Pcdh19Y/β-Geo and Pcdh19+/β-Geo embryonic brains revealed β-Geo-positive cells throughout the cortex and hippocampus in the same regions as wild type Pcdh19 expressing cells (Fig. 5E,F). To confirm that the different methods used to detect Pcdh19 expressing and mutant cells did not confound the analysis we performed in situ hybridisation of 15.5 dpc brains with a β-Geo probe (Supplementary Fig. 3). Cells expressing β-Geo were identified in the cortex and hippocampus as expected including the marginal zone, intermediate zone and ventricular/sub-ventricular zones (Fig. 5G). Taken together, these data suggest that mutant cells are not ectopically positioned in Pcdh19+/β-Geo and Pcdh19Y/β-Geoembryonic brains.
Pcdh19 null cells are not ectopically located in the developing brain (15.5 dpc).
(A–C) Pcdh19 WT cells were detected by in situ hybridisation and were located in the cortex and hippocampus of Pcdh19Y/+(A) and Pcdh19+/β-Geo (B) developing brains. Pcdh19+/β-Geo brains exhibited less extensive staining consistent with X-inactivation. No specific signal was detected in Pcdh19Y/β-Geo brains (C). (D–F) Pcdh19 null cells were detected by X-Gal staining and were located in the cortex and hippocampus of Pcdh19+/β-Geo (E) and Pcdh19Y/β-Geo (F) developing brains. Pcdh19+/β-Geo brains showed reduced staining consistent with X-inactivation. No signal was detected in Pcdh19Y/+brains (D). Scale bar represents 500 μm. Three animals were analysed for each genotype and age. (G) Pcdh19 null cells are predominantly located in the ventricular zone (VZ)/subventricular zone (SVZ), the intermediate zone (IZ) and the marginal zone (MZ). Scale bar represents 100 μm.
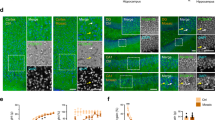
Given that Pcdh19 expression is maintained after birth5, we next investigated whether β-Geo expressing cells were ectopically expressed in Pcdh19+/β-Geo and Pcdh19Y/β-Geobrains at postnatal day 42 (P42). Consistent with our western blot analysis (Fig. 2A), robust expression of Pcdh19 was detected in the hippocampus and cortex of Pcdh19 WT brains (Fig. 6A). Pcdh19 expression was strongest in the CA1 and dentate gyrus of the hippocampus and in layers 2/3 and 5 of the cortex. Mosaic expression was identified in the same regions of Pcdh19+/β-Geo brains (Fig. 6B).
Pcdh19 null cells are not ectopically located in the adult brain.
(A–C) Pcdh19 WT cells were detected by in situ hybridisation and were located in the cortex and hippocampus of Pcdh19Y/+(A) and Pcdh19+/β-Geo (B) adult (P42) brains. Pcdh19+/β-Geo brains exhibited less staining consistent with X-inactivation. No specific signal was detected in Pcdh19Y/β-Geo brains (C). (D–F) Pcdh19 null cells were detected by X-Gal staining and were located in the cortex and hippocampus of Pcdh19+/β-Geo (E) and Pcdh19Y/β-Geo (F) adult brains. Pcdh19+/β-Geo brains showed less staining consistent with X-inactivation. No signal was detected in Pcdh19Y/+brains (D). Scale bar represents 500 μm. Three animals were analysed for each genotype and age. (G) Pcdh19 null cells are predominantly expressed in layer II/III and V of the adult cortex. Scale bar represents 100 μm.
β-Geo staining of Pcdh19Y/β-Geo brains with X-Gal revealed robust expression in the cortex and hippocampus that resembled the pattern of Pcdh19 expression in Pcdh19+/Y brains (Fig. 6). Mosaic expression of β-Geo was observed in these regions in Pcdh19+/β-Geo brains (Fig. 6E). No evidence of ectopic β-Geo-positive cells was detected. Thus, it appears that the mutation of Pcdh19 does not perturb positioning of neurons in postnatal brain development.
The Migration Potential of Pcdh19 Null Neurons is Increased In Vitro
As subtle defects in neuronal migration can be difficult to detect in vivo, we next performed in vitro migrations assays using neurospheres generated from Pcdh19 WT and Pcdh19 null embryos. Neural progenitor cells were isolated from the developing cortex (14.5 dpc) and cultured as non-adherent neurospheres. To model neuronal migration, whole neurospheres were adhered to a poly-L-lysine substrate and the migration of neurons outward from the neurosphere boundary measured20. Increased neuronal migration in the Pcdh19 null neurons was observed, as shown by an increase in the percentage of migrating neurons in the outer migration regions (bins) (Fig. 7A,B). However, this only reached significance in 4/9 of the bins. This data suggests that the mutant cells may have a subtle change in neuronal migration potential that does not translate into overt positional changes of neurons in the developing and adult mouse brain.
Loss of Pcdh19 leads to an increase in neuronal migration.
Neural progenitor cells (NPCs) were isolated from the telencephalic vesicles of Pcdh19 WT or Pcdh19 null E14.5 embryos and grown as non-adherent neurospheres. Neurospheres were plated on a Poly-L-Lysine substrate and cells were allowed to attach and migrate. (A) Neuronal migration was scored at 48 hrs by co-staining with a neuronal specific antibody β-III Tubulin (Cyan) and DAPI. (B) Neuronal migration was determined by drawing concentric circles around the contiguous boundary of the neurosphere and counting the number of neurons within each circle. Differences in the number of migrating neurons was determined by the percentage of neurons within each circle (bin). The percentage of neurons is greater in the more distal circles in the KO Pcdh19 neurospheres compared to the WT Pcdh19 neurospheres. Each data point represents 7 embryos collected from 3 litters with at least 20 neurospheres scored for each embryo. Error bars = SEM, *differs from WT; P < 0.05, Student’s T-test.
Discussion
Mutations of protocadherin genes in humans have been implicated in several neurological diseases and in mice have revealed important functions in neuronal development including axon guidance, neural circuit formation and synaptogenesis. Here we performed validation and gross morphological characterization of Pcdh19 null mice to determine the function of Pcdh19 in vivo.
Previous studies indicate that Pcdh19 expression is temporally and spatially restricted within the embryonic and neonatal CNS5,21. Using in situ hybridization and β-Geo reporter activity, we have shown that Pcdh19 expression is maintained in the adult brain. Expression was particularly robust in regions of the brain that are implicated in epilepsy including the hippocampus and in the cortex, where it was restricted to layers 2/3 and 5, although expression in other areas such as the cerebellum and the hypothalamus was also evident. We have attempted to further characterize endogenous PCDH19 CNS protein expression by immunohistochemistry using several commercially available PCDH19 antibodies, however none bind specifically to PCDH19 as evident by the immunostaining patterns that were indistinguishable between Pcdh19 WT and Pcdh19 null brain sections (our unpublished data). The β-Geo reporter gene in our Pcdh19 null mouse therefore provides a useful alternative method for identification of cells that normally express Pcdh19, especially given that loss of Pcdh19 function does not overtly affect neuron localization in vivo (see below).
Expression of Pcdh19 in the adult brain also raises the question of its subcellular localization. Using chemical fractionation and primary hippocampal neuron culture, we have shown that PCDH19 is located within both the synaptosome fraction in vivo and synapses in vitro suggesting it has synaptic function. Consistent with these results, analysis of chick PCDH19 in the optic tectum revealed substantial overlap with the synaptic marker Syntaxin7. Interestingly, Pcdh17, the closest related family member to Pcdh19, has recently been shown to play a role in synapse formation and synaptic vesicle assembly in murine corticobasal ganglia9. These data raise the possibility that PCDH19 may perform a similar function in vivo and supports the hypothesis that abnormal synaptic communication between PCDH19 WT and PCDH19 mutant cells contributes to the pathobiology of PCDH19 GCE5.
Previous reports have shown that morpholino knockdown or genetic mutation of pcdh19 in zebrafish resulted in contrasting phenotypes11,12,13. In agreement with mutation of pcdh19 in zebrafish by Cooper et al. we show here that Pcdh19Y/β-Geo, Pcdh19β-Geo/β-Geo and Pcdh19+/β-Geo mice are healthy, fertile and do not exhibit gross defects in brain morphology. These striking phenotypic differences in zebrafish may reflect lack of genetic compensation in response to acute morpholino Pcdh19 knockdown in zebrafish which has been shown to account for similar differences in other studies22. In humans, hemizygous males do not exhibit intellectual disability or seizures indicating that brain development occurs relatively normally without PCDH19, perhaps due to functional compensation by other PCDH19 family members23. The absence of morphological defects in Pcdh19Y/β-Geoand Pcdh19β-Geo/β-Geomice is therefore not unexpected and suggests that Pcdh19 functional redundancy also occurs in mice. In contrast, the majority of PCDH19 heterozygous human females develop infantile seizures with variable cognitive defects, although there is no evidence of gross morphological defects affected individuals. It has been proposed that this phenotype results from “cellular interference” i.e. segregation of PCDH19 expressing neurons and PCDH19 mutant neurons in the developing brain development resulting in abnormal neural circuitry5,23. While there is little in vivo data to support this model, it is interesting to note that cortical dysplasia and clustering of dysplastic pyramidal neurons has been detected in a single female with PCDH19 GCE15. Although we did not detect any evidence of cortical dysplasia or gross abnormalities in Pcdh19+/β-Geo mice, we cannot exclude the possibility that subtle regionally-restricted defects may be present. Ryan et al. also detected dysplastic pyramidal cells ectopically positioned in the deep white layer, suggesting defective neuronal migration. To investigate this phenotype in Pcdh19+/β-Geo mice, we determined the location of Pcdh19 null cells using X-Gal staining in both developing and postnatal Pcdh19+/β-Geo brains. However, no evidence of ectopic Pcdh19 null cells was detected indicating that PCDH19 is not critical for neuronal cell migration in vivo. In contrast, a modest but significant increase in migration was detected in Pcdh19 null neurons generated from neurosphere explants. It is therefore possible that loss of Pcdh19 function compromises the migration potential of some neurons but that this defect is subtle and not associated with overt morphological defects in vivo.
In summary, our characterization of Pcdh19 null mice reveals that Pcdh19 is widely expressed in the CNS but not required for brain development per se. However, given the significantly increased motility of Pcdh19 null neurons in vitro and the identification of Pcdh19 mutant phenotypes in other species, further detailed analysis of this mouse model may reveal subtle abnormalities in Pcdh19 null brains.
Methods
Animals
Pcdh19 null mice (TF2108) were purchased from Taconic Biosciences. Both male and female animals were used in this study. The day the vaginal plug was detected was designated as embryonic day 0.5 (0.5 dpc). The day of birth was designated postnatal day 0 (P0). Each group subjected to analysis contained at least three mice. All of the experiments were approved by The University of Adelaide Animal Ethics Committee and performed according to their ethical guidelines.
Genotyping
Genotyping was performed by PCR amplification using three primers: one primer common to both Pcdh19 WT and Pcdh19 null alleles F-5′-AGTCCACTACCGACTCTGCTG -3′, one specific to the Pcdh19 WT allele R-5′-CAAAGTTAGCCAGGCGGGAC-3′ and one specific to the Pcdh19 null allele R-5′-AACTCACAACGTGGCACTGG-3′. PCR products were separated on a 1% agarose gel. Pcdh19 WT and null allele products were 102 bp and 471 bp, respectively.
Protein extraction, synaptosome fractionation and western blotting
Cortex, hippocampi and cerebellum for protein extraction were minced in extraction buffer (50 mM Tris, 150 mM NaCl, 1% NP40 (Roche) and 1% Triton X-100 (Sigma) and incubated at 4 degrees C for 30 minutes. Synaptosome fractions were isolated using Syn-PER synaptic protein extraction reagent (ThermoFisher Scientific) according to the manufacturer’s instructions. Lysates and fractions were separated on Invitrogen Bolt™ precast 4–12% polyacrylamide gels and transferred to PVDF membrane before being blotted. Antibodies and their corresponding dilutions were: mouse anti-PCDH19 1:100 (Abcam, ab57510) and rabbit anti-β-ACTIN 1:1000 (Cell Signalling Technology, #4967), mouse anti-PSD95 1:2000 (ThermoScientific, 7E3-1138) and rabbit anti-SYNAPTOPHYSIN 1:5000 (Abcam, ab32127). Membranes were blocked in 5% BSA+5% Skim milk in Tris-buffered saline+0.1% Tween 20 (TBST) and antibodies were incubated with the membrane in 5% BSA in TBST. Membranes were developed using Bio-Rad Clarity Western ECL substrate and imaged using a Bio-Rad ChemiDoc.
Primary Hippocampal Neuron Culture, Nucleofection and Immunohistochemistry
18.5 dpc hippocampi were dissected from wild type embryos and neurons dissociated with 0.5% Trypsin. 1 × 105 neurons were nucleofected with 1 μg PCDH19-FLAG using the Neon Transfection System (Invitrogen) and seeded onto poly-D-lysine coated coverslips. Primary neurons were maintained in Neurobasal medium+B27 supplement (Life Technologies). Primary neurons were cultured for 12 days then fixed in 4% PFA for 15-30 minutes. For immunofluorescence, cells were block permeablised with 0.1% Tween20 (PBST) and 10% foetal calf serum for 1 hour at room temperature. Primary neurons were then incubated overnight with anti-FLAG (1:1000, Sigma Aldrich) and anti-SYNAPSIN (1:500, Millipore) at 4˚C and then with secondary antibody (donkey anti-mouse Cy3 and donkey anti-rabbit Cy5; JacksonImmunoResearch). Images were acquired on a Nikon Eclipse Ti microscope.
X-gal staining
15.5 dpc embryonic heads and dissected P42 brains were fixed in 4% paraformaldehyde at 4 °C, cryoprotected in 30% sucrose and frozen in OCT embedding medium. Embryos were fixed in 4% paraformaldehyde in PBS. Whole embryos or cryosections were incubated in staining solution: 19 mM Sodium dihydrogen phosphate, 81 mM Disodium hydrogen phosphate, 2 mM MgCl2, 5 mM EGTA, 0.01% Sodium deoxycholate, 0.02% NP-40, 5 mM Potassium ferricyanide, 5 mM Potassium ferrocyanide and 1 mg/ml X-gal substrate, at 37 °C until blue staining was sufficient. Images were acquired on a Nikon Eclipse Ti microscope, compiled and minimally processed (adjusted for color and light/dark) using Adobe Photoshop CS.
Histological analysis
Embryos for histological examination were fixed in 4% PFA, embedded in paraffin, cut at 5μm and stained with Mayer’s haematoxylin and Eosin using standard approaches. Images were acquired on a Nikon Eclipse Ti Microscope and measurements taken using Nikon Imaging Software (NIS Elements). Three animals were analysed for each genotype with cortical measurements being made in the visual cortex and hippocampal measurements in the CA1 region. A total of ten measurements for each region was recorded for each individual animal and averaged. The mean was then determined from the average of the three animals in each group.
In situ hybridisation
15.5 dpc embryonic heads and dissected P42 brains were fixed in 4% paraformaldehyde at 4 °C, cryoprotected in 30% sucrose and frozen in OCT embedding medium. In situ hybridization of 16-μM sections was performed as described previously24. The Pcdh19 probe was a digoxigenin-labeled antisense RNA probe prepared as described21. The LacZ probe sequence (500 bp) was generated by PCR using the following primers F-5′-ATGTGCGGCGAGTTGCGTGA-3′ R-5′-CGCTCATCGCCGGAGCCAGC-3′. At least three independent samples of each group were analyzed and representative sections are shown. No signal was detected using a sense control probe. Images were acquired on a Nikon Eclipse Ti microscope, compiled and minimally processed (adjusted for color and light/dark) using Adobe Photoshop CS.
Neurosphere Migration Assay
NPCs were isolated from the 14.5 dpc embryonic mouse cortex (telencephalic vesicles) and grown as non-adherent neurospheres in the presence of 20 ng/mL bFGF and EGF as previously described25. Passage 4 neurospheres were cultured for 5 days and seeded onto a poly-L-lysine substrate at a density of 100 neurospheres/9.5 cm2 with the removal of growth factors. Cells were allowed to migrate for 48 hrs before being fixed in 4% paraformaldehyde (PFA) in phosphate buffered saline (PBS) for 15 minutes at room temperature. For immunofluorescence analysis, cells were block permeablised in 0.2% tween20 (PBST) and 5% horse serum for 1 hour at room temperature. To identify neurons, cells were stained overnight with β-III tubulin primary antibody (1:300, Sigma Aldrich) in PBST with 0.5% horse serum at 4 °C and then with secondary antibody, donkey anti-mouse AlexaFluor647 (1:700, Invitrogen) for 1 hr at room temperature. Cells were counterstained with 300 nM 4′,6-diamidino-2-phenylindole (DAPI) to identify single cells and mounted with Slow-fade mounting media (Invitrogen) for microscopy. Fluorescence was viewed using the Zeiss AxioImager M2 fluorescent microscope (Carl Zeiss, Germany). Images of isolated single neurospheres and migrating cells were captured at 20× magnification using an Axiocam Mrm camera and Vs4.9.1.0 software (Axiovision, Carl Zeiss). Neurospheres were reconstructed using the Image J Stitching plugin26 and migration analysis preformed using the ImageJ2 software27. Briefly, the neurosphere boundary was identified and the radius of the neurosphere determined. The concentric circles plugin was used to draw 10 concentric circles (ie. migration zones) around the boundary of the sphere, with an outer radius calculated by the radius of the neurosphere+600 pixels (307 μM). Using the cell counter plugin the number of migrating neurons within each concentric circle was determined by counting co-stained β-III tubulin and DAPI cells. The migration distance was then calculated as a percentage of total migrating cells within each circle (bin). Each data point represents 7 embryos collected from 3 litters with at least 20 neurospheres scored for each embryo. Analysis conducted by Student’s two-tailed, unpaired t test, assuming equal variance.
Real time PCR
RNA was extracted from P14 hippocampi by using an RNeasy mini column (Qiagen) according to the manufacturer’s instructions. Reverse transcription was performed on 1μg of RNA using a High Capacity RNA to cDNA synthesis kit (Life Technologies). Real time PCR was performed on a Step One Plus thermocycler (Life Technologies) using Fast Sybr green master mix (Life Technologies) according to manufacturer’s instructions on a two-step program. Primers and product sizes were as follows; Pcdh19 (130 bp) F-5′ -TGGCAATCAAATGCAAGCGT-3′ R-5′ ACCGAGATGCAATGCAGACA-3′, β-Galactosidase (85 bp) F-5′ TACGATGCGCCCATCTACAC-3′ R5′ AACAACCCGTCGGATTCTCC-3′ β-actin (89 bp) F-5′-CTGCCTGACGGCCAGG-3′ R-5′-GATTCCATACCCAAGAAGGAAGG-3′.
Additional Information
How to cite this article: Pederick, D. T. et al. Pcdh19 Loss-of-Function Increases Neuronal Migration In Vitro but is Dispensable for Brain Development in Mice. Sci. Rep. 6, 26765; doi: 10.1038/srep26765 (2016).
References
Frank, M. & Kemler, R. Protocadherins. Curr. Opin. Cell Biol. 14, 557–562 (2002).
Junghans, D., Haas, I. G. & Kemler, R. Mammalian cadherins and protocadherins: about cell death, synapses and processing. Curr. Opin. Cell Biol. 17, 446–452 (2005).
Morishita, H. & Yagi, T. Protocadherin family: diversity, structure and function. Curr. Opin. Cell Biol. 19, 584–592 (2007).
Redies, C., Vanhalst, K. & Roy, F. van. delta-Protocadherins: unique structures and functions. Cell. Mol. Life Sci. CMLS 62, 2840–2852 (2005).
Dibbens, L. M. et al. X-linked protocadherin 19 mutations cause female-limited epilepsy and cognitive impairment. Nat. Genet. 40, 776–781 (2008).
Emond, M. R., Biswas, S., Blevins, C. J. & Jontes, J. D. A complex of Protocadherin-19 and N-cadherin mediates a novel mechanism of cell adhesion. J. Cell Biol. 195, 1115–1121 (2011).
Tai, K., Kubota, M., Shiono, K., Tokutsu, H. & Suzuki, S. T. Adhesion properties and retinofugal expression of chicken protocadherin-19. Brain Res. 1344, 13–24 (2010).
Hayashi, S. et al. Protocadherin-17 mediates collective axon extension by recruiting actin regulator complexes to interaxonal contacts. Dev. Cell 30, 673–687 (2014).
Hoshina, N. et al. Protocadherin 17 regulates presynaptic assembly in topographic corticobasal Ganglia circuits. Neuron 78, 839–854 (2013).
Uemura, M., Nakao, S., Suzuki, S. T., Takeichi, M. & Hirano, S. OL-protocadherin is essential for growth of striatal axons and thalamocortical projections. Nat. Neurosci. 10, 1151–1159 (2007).
Biswas, S., Emond, M. R. & Jontes, J. D. Protocadherin-19 and N-cadherin interact to control cell movements during anterior neurulation. J. Cell Biol. 191, 1029–1041 (2010).
Cooper, S. R. et al. Protocadherins control the modular assembly of neuronal columns in the zebrafish optic tectum. J. Cell Biol. 211, 807–814 (2015).
Emond, M. R., Biswas, S. & Jontes, J. D. Protocadherin-19 is essential for early steps in brain morphogenesis. Dev. Biol. 334, 72–83 (2009).
Kok, F. O. et al. Reverse genetic screening reveals poor correlation between morpholino-induced and mutant phenotypes in zebrafish. Dev. Cell 32, 97–108 (2015).
Ryan, S. G. et al. Epilepsy and mental retardation limited to females: an X-linked dominant disorder with male sparing. Nat. Genet. 17, 92–95 (1997).
Depienne, C. & LeGuern, E. PCDH19-related infantile epileptic encephalopathy: an unusual X-linked inheritance disorder. Hum. Mutat. 33, 627–634 (2012).
Scheffer, I. E. et al. Epilepsy and mental retardation limited to females: an under-recognized disorder. Brain 131, 918–927 (2008).
Terracciano, A. et al. PCDH19-related epilepsy in two mosaic male patients. Epilepsia 57, e51–e55 (2016).
Cotton, A. M. et al. Landscape of DNA methylation on the X chromosome reflects CpG density, functional chromatin state and X-chromosome inactivation. Hum. Mol. Genet. 24, 1528–1539 (2015).
Homan, C. C. et al. Mutations in USP9X are associated with X-linked intellectual disability and disrupt neuronal cell migration and growth. Am. J. Hum. Genet. 94, 470–478 (2014).
Gaitan, Y. & Bouchard, M. Expression of the δ-protocadherin gene Pcdh19 in the developing mouse embryo. Gene Expr. Patterns 6, 893–899 (2006).
Rossi, A. et al. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature 524, 230–233 (2015).
Depienne, C. et al. Sporadic Infantile Epileptic Encephalopathy Caused by Mutations in PCDH19 Resembles Dravet Syndrome but Mainly Affects Females. PLoS Genet 5, e1000381 (2009).
Wilson, L. D. et al. Developmentally regulated expression of the regulator of G-protein signaling gene 2 (Rgs2) in the embryonic mouse pituitary. Gene Expr. Patterns 5, 305–311 (2005).
Basak, O. & Taylor, V. Identification of self-replicating multipotent progenitors in the embryonic nervous system by high Notch activity and Hes5 expression. Eur. J. Neurosci. 25, 1006–1022 (2007).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinforma. Oxf. Engl. 25, 1463–1465 (2009).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Acknowledgements
This study was supported by the Australian National Health and Medical Research Council and the PCDH19 Alliance.
Author information
Authors and Affiliations
Contributions
P.Q.T., D.T.P., J.G., J.N.H., L.A.J., B.T.B., E.J.J. and C.C.H. conceived and designed the experiments. P.Q.T., J.N.H., L.A.J., B.P.H., D.T.P. and C.C.H. interpreted the data. D.T.P., C.C.H. and S.G.P. performed the experiments. D.T.P., P.Q.T. and C.C.H. wrote the manuscript. P.Q.T. and J.G. obtained the funding.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Pederick, D., Homan, C., Jaehne, E. et al. Pcdh19 Loss-of-Function Increases Neuronal Migration In Vitro but is Dispensable for Brain Development in Mice. Sci Rep 6, 26765 (2016). https://doi.org/10.1038/srep26765
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep26765
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.