Abstract
Proteasome-catalyzed peptide splicing represents an additional catalytic activity of proteasomes contributing to the pool of MHC-class I-presented epitopes. We here biochemically and functionally characterized a new melanoma gp100 derived spliced epitope. We demonstrate that the gp100mel47–52/40–42 antigenic peptide is generated in vitro and in cellulo by a not yet described proteasomal condensation reaction. gp100mel47–52/40–42 generation is enhanced in the presence of the β5i/LMP7 proteasome-subunit and elicits a peptide-specific CD8+ T cell response. Importantly, we demonstrate that different gp100mel-derived spliced epitopes are generated and presented to CD8+ T cells with efficacies comparable to non-spliced canonical tumor epitopes and that gp100mel-derived spliced epitopes trigger activation of CD8+ T cells found in peripheral blood of half of the melanoma patients tested. Our data suggest that both transpeptidation and condensation reactions contribute to the frequent generation of spliced epitopes also in vivo and that their immune relevance may be comparable to non-spliced epitopes.
Similar content being viewed by others
Introduction
CD8+ T cell responses involve in most cases the proteasome-dependent processing of antigens and cell surface presentation of the resulting peptides by MHC-class I molecules to cytotoxic T lymphocytes (CTLs). The 20S standard proteasome (s-proteasome) with its active site β-subunits β1, β2 and β5, the immunoproteasome (i-proteasome) with the IFN-γ-induced catalytic β-subunits β1i/LMP2, β2i/MECL1 and β5i/LMP7 or intermediate proteasome types containing both standard and immuno-subunits1,2 are the catalytic cores of 30S proteasomes, which are formed by the association of two 19S regulator complexes with the 20S core3. The 30S proteasome isoforms execute the regulated degradation of ubiquitin (Ub)-tagged proteins, thereby generating peptides presented by MHC-class I proteins4,5. Exchange of standard catalytic subunits for the corresponding IFN-γ-induced β-subunits leads to variations of proteasome catalytic pocket structure and of the peptide transport along the proteasome inner channel6,7,8. Such variations may result in quantitative differences in the proteasome cleavage products, which in turn can strongly affect cell surface presentation of MHC-class I-bound virus- or tumor-specific antigenic peptides and in consequence also the efficacy of a peptide-specific CD8+ T cell response9,10.
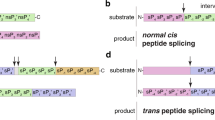
Canonical MHC-class I-bound peptides generated by proteasomes mirror the linear sequence of the parental protein10. In contrast, spliced peptides that are presented by MHC-class I molecules and generated by proteasome-catalyzed peptide splicing (PCPS) result from the ligation of two separate distant proteasomal cleavage products that are not contiguous in the parental protein. We previously provided experimental proofs for the hypothesis that PCPS proceeds via a transpeptidation reaction11,12,13,14. PCPS involves the formation of an O-acyl-enzyme intermediate consisting of a N-terminal peptide fragment linked to the Thr1 residue of one of the β-subunit active sites. In the course of the transpeptidation reaction, this acyl-enzyme intermediate is subjected to a nucleophilic attack by the amino-terminus of another peptide fragment leading to the creation of a new peptide containing two non-contiguous fragments of a protein11,12,15. Theoretically, spliced peptides may also be produced without the formation of a semi-stable O-acyl-enzyme intermediate by a so-called condensation reaction. Condensation reactions forming de novo peptide bonds have been described for some proteases to occur in vitro under conditions disfavoring diffusion of peptides from the active sites16,17. However, whether such a condensation reaction can take place in cellulo in the barrel-shaped proteasome controlling peptide diffusion and whether this can contribute to epitope production has yet to be shown.
Five spliced epitopes, derived from the fibroblast growth factor 5 (FGF-5172–176/217–220), melanocyte protein gp100mel (gp100mel40–42/47–52; gp100mel195–202/192), the SP100 nuclear phosphoprotein (SP100296–301/286–289) and tyrosinase (Tyr368–373/336–340) have been identified so far using cancer patient-derived CTLs for epitope identification11,12,18,19,20. Interestingly, two of these spliced peptide-specific CD8+ T cells have previously been shown to induce regression of tumors in a clinical setting20,21 or the engraftment of acute myeloid leukemia cells in non-obese diabetic/SCID mice11,22, indicating their potential immune relevance.
PCPS represents a genuine catalytic activity of proteasomes competing with the normal proteasomal hydrolysis event14. Initial estimations of epitope splicing efficacy were calculated to range between 0.0002% and 0.01% of the total proteasome-dependent epitope generation. Therefore, epitope production by PCPS was thought to be an extremely rare event12,13,16. However, performing in vitro experiments we showed that PCPS may be a relatively frequent process controlled by certain sequence requirements guaranteeing that PCPS is also a highly reproducible reaction14. In fact, in vitro experiments suggested that potential MHC-class I epitopes are relatively more frequent among spliced peptides than among the “canonical” non-spliced proteasome cleavage products14,23. The limited number of spliced tumor epitopes identified most likely resides in the fact that identification of immune-relevant spliced peptides has so far been limited by the serendipitous availability of patient-derived CTLs and the concomitant considerable experimental effort to identify the cognate epitopes.
To close this gap, we here report an algorithm-aided reverse immunology approach permitting the facilitated identification of spliced peptides from a given protein sequence thereby overcoming the need for patient-derived CTLs. By applying this method we here identified a new spliced peptide derived from melanocytic protein gp100mel. This peptide can be generated in vitro and in cellulo by a condensation reaction. This epitope is presented by patient-derived melanoma cells and triggers activation of CD8+ T cells found in the peripheral blood of melanoma patients. Importantly, we show that spliced epitopes are presented at the cell surface and trigger CTL responses with efficacies similar to non-spliced epitope underlining that PCPS may play an important role in anti-tumor immune responses.
Results
The gp100mel47–52/40–42 antigenic peptide is generated by a new proteasomal catalytic mechanism
For identification of new spliced peptides and/or epitopes we subjected the gp100mel-derived synthetic polypeptide substrate gp100mel40–52 [RTKAWNRQLYPEW] to in vitro digestion by purified human 20S proteasomes. By applying SpliceMet23 we identified three spliced peptides among the proteasomal processing products. These were the previously reported spliced antigenic peptide gp100mel40–42/47–52 [RTK][QLYPEW]12 and the novel gp100mel47–52/40–46 [QLYPEW][RTKAWNR] and gp10047–52/40–42 [QLYPEW][RTK] peptides (Fig. 1A, Fig. S1). Interestingly, a theoretical IC50 value of 86.1 μM for HLA-A*03:01 and HLA-A*011 binding was predicted for gp100mel47–52/40–42 [QLYPEW][RTK] by in silico analysis24. Therefore, we decided to analyze the formation of this potential new spliced tumor epitope in more detail.
Involvement of proteasome standard- and immuno-subunits in in vitro spliced peptide generation.
(A) B- and Y-ions of the spliced products gp100mel40–42/47–52 [RTK][QLYPEW], gp100mel47–52/40–42 [QLYPEW][RTK] and gp100mel47–52/40–46 [QLYPEW][RTKAWNR] as detected by MS/MS Orbitrap analysis (see also Fig. S1) and derived from gp100mel40–52 upon 20S proteasome-catalyzed digestion. (B) Erythrocyte 20S s-proteasomes generated larger amount of the spliced peptide gp100mel40–42/47–52 than of the gp100mel47–52/40–42 and gp100mel47–52/40–46 peptides. (C,D) Erythrocyte 20S s-proteasomes generated similar or larger amounts of the splice reactants gp100mel40–42 [RTK], gp100mel40–46 [RTKAWNR] and gp100mel47–52 [QLYPEW] (C) but less gp100mel47–52/40–42 and gp100mel47–52/40–46 spliced peptides than spleen 20S proteasomes (D). (B–D) The means of the amount measured after 3 hrs in vitro digestion and the SD between triplicates of two independent experiments are shown. Significant differences of the means are detected by student t test and marked by *. In particular, (B) gp100mel: 40–42/47–52 vs 47–52/40–42, p = 0.002; 40–42/47–52 vs 47–52/40–46, p < 0.001; (C) gp100mel40–46 s- vs i-proteasome, p = 0.036; gp100mel47–52 s- vs i-proteasome, p = 0.043; (D) gp100mel47–52/40–42: s- vs i-proteasome p < 0.001; gp100mel47–52/40–46: s- vs i-proteasome p < 0.001. (E) B- and Y-ions of peptide gp100mel47–52/40–52 -detected by MALDI-TOF/TOF-MS (Fig. S3) – that was produced by hydrolysis-independent ligation of the splice-reactants [RTK] and [QLYPEW] by 20S proteasome. (F) The peptide gp100mel47–52/40–42 amount, generated by hydrolysis-independent ligation by i- and s-proteasomes is shown. Amounts below 10 fmol could be detected but not quantified and are symbolized with ♦. (G) Absolute amounts of gp100mel47–52/40–42 generated in the hydrolysis-independent assay by T2 and LCL 20S proteasomes in the presence of the inhibitors YU-102 (2 μM; β1/β1i-specific), LU-102 (2 μM; β2/β2i-specific), PR-893 (0.5 μM; β5-specific) and PR-924 (0.5 μM; β5i-specific). Significant differences due to the effect of the inhibitors are detected by paired student t test and marked by * (T2 proteasome, control vs PR-893, p = 0.037; LCL proteasome, control vs PR-893, p = 0.050; control vs PR-924, p = 0.012). (F,G) The mean (fmol measured in 9 μl reaction) and the SD of independent experiments (n = 2–3), each measured in two duplicates are shown.
Monitoring the in vitro generation of the spliced peptides revealed that the spliced peptide gp100mel40–42/47–52 was produced approximately 6-fold more efficiently by erythrocytes s-proteasomes than the new spliced gp100mel47–52/40–42 peptide, being composed of the same peptide fragments but in reversed order (Fig. 1B). Interestingly, opposite to the generation kinetics of the individual splice-reactants [QLYPEW], [RTKAWNR] or [RTK] (Fig. 1C), spleen i-proteasomes generated the spliced peptides gp10047–52/40–46 and gp10047–52/40–42 more efficiently than erythrocytes s-proteasomes (Fig. 1D, Fig. S2).
Both the gp100mel47–52/40–42 and gp100mel47–52/40–46 peptides were the result of the linkage between the gp100 Trp52 and Arg40 residues of the gp100-derived substrate (Fig. 1E). However, the Trp52 residue of the N-terminal splice-reactant [QLYPEW] represents the C-terminal residue of the synthetic polypeptide substrate gp100mel40–52. This excluded the formation of an O-acyl-enzyme intermediate with the proteasomal active site Thr1 as a result of peptide-bond hydrolysis as demanded for the established transpeptidation reactions12,14. We therefore hypothesized that spliced peptides could also be produced in a proteasome-catalyzed condensation reaction.
To test this hypothesis we co-incubated purified 20S proteasomes with the two splice-reactant peptides gp100mel40–42 [RTK] and gp100mel47–52 [QLYPEW]. The resulting spliced peptide products were then analyzed by mass spectrometry. In support of our hypothesis, 20S proteasomes catalyzed the formation of the gp100mel47–52/40–42 peptide in vitro when the two peptides were added individually (Fig. 1E, Fig. S3). In accordance with the results shown in Fig. 1D i-proteasomes derived from different cellular sources catalyzed the condensation reaction with higher efficiency than s-proteasomes (Fig. 1F).
In principle, all active sites of both proteasome isoforms are capable of catalyzing a transpeptidation reaction14. However, because condensation reactions require specific favorable conditions16 we next asked whether any of the proteasome active sites were specifically involved in the condensation reaction. To do so, proteasome active site subunit specific inhibitors, whose active site specificity had been previously documented25 were added to 20S proteasomes derived from T2 cells (s-proteasome) or LCLs (i-proteasome) together with the two splice-reactants [RTK] and [QLYPEW]. PR-893 (0.5 μM) was used to block the activity of the β5 subunits of 20S s-proteasomes and PR-924 (0.5 μM) to inhibit the activity of the β5i immuno-subunit25. Blockage of either the β5 active site as in T2 cell-derived 20S proteasome or the β5 and β5i active sites as present in LCL-derived proteasomes led to a strong reduction of the condensation reaction (Fig. 1G). In contrast, blocking the β1/β1i and β2/β2i active sites with YU-102 (2 μM) and LU-102 (2 μM) proteasome inhibitors25, respectively, barely affected the production of the antigenic peptide, suggesting that the condensation reaction was mainly executed by the active sites of the β5 subunit isoforms (Fig. 1G).
Cellular condensation reaction results in the presentation of the gp100mel47–52/40–42 peptide
Although our in vitro experiments clearly demonstrated that proteasomes are able to generate spliced peptides in a condensation reaction, there still remained the possibility that the in vitro experiments as performed here may be artificially influenced by high enzyme and/or substrate concentrations. Therefore, to assess whether the gp100mel47–52/40–42 peptide is of any functional relevance and to study whether proteasomes could perform condensation reactions also in cellulo we established an immunological read-out system to monitor the generation and MHC class I presentation of gp100mel47–52/40–42 in HeLa cells. HLA-A binding affinity assays using HLA-A*03:01- transfected HeLa and T2 cells lines (Fig. 2A,B) as well as mature dendritic cells (data not shown) confirmed the predicted binding of the peptide to HLA-A*03:01 complex. We therefore generated a gp100mel47–52/40–42 peptide-specific CTL clone from peripheral blood mononuclear cells (PBMCs) of a HLA-A*03:01+ healthy donor (Fig. S4) allowing us to monitor the generation of this potential MHC class I-restricted epitope in cellulo.
The spliced gp100mel47–52/40–42 peptide is generated from the gp100mel40–42 peptide by a condensation reaction in cellulo.
(A,B) The spliced peptide gp100mel47–52/40–42 [QLYPEW][RTK] bound the HLA-A*03:01 complex on the cell surface of HeLa (A) and T2 (B) cell lines. In (A) the side / forward scatters charts are reported in the left panel. Cells included in the analysis are marked with a circle and their HLA-A*03:01 amount is shown in the right panel. (B) Shown is the binding affinity of the spliced peptide gp100mel47–52/40–42 compared to a known good HLA-A*03:01 – binder, i.e. [ILRGSVAHK] (NP265–273)35 and to the negative controls HLA-A*32:01-restricted gp100mel40–42/47–52 [RTK][QLYPEW] and HLA-A*02:01-restricted gp100mel209–217 [IMDQVPFSV] epitopes. Cells were transfected with HLA-A*03:01 protein encoding plasmids and pulsed with 100 μM (A) or different peptide concentrations (B). Binding affinity was measured by staining of HLA-A*03:01 and FACS analysis. Values are in mean fluorescence unit (MFU) and are means and SD of 2 replicates in a representative assay of 2–6 independent experiments. (C) Schematic representation of the HA-Ub/gp100mel40–52 fusion construct used in this study. The first amino acid of the gp100mel40–52 [RTKAWNRQLYPEW] sequence was directly fused in frame to the C-terminus of the Ub, which was N-terminally tagged with the YPYDVPDYA HA sequence. When expressed in cells, Ub hydrolases cleaved the fusion-polypeptide directly after the last amino acid (Gly76) of Ub, liberating the gp100mel40–52 peptide from the first residue. (D) CTL response towards HeLa 33/2 cells transfected with HLA-A*03:01 individually or together with HA-Ub/gp100mel40–52 for 24 hrs. Their ability to present gp10047–52/40–42 epitope was measured by using K631 1C CTL. After 16 hrs co-culture, the IFN-γ concentration in supernatants was determined by ELISA. Values are means and SD of 2 replicates in a representative assay of 6 independent experiments. Significant differences in the means of replicates are marked by * (student t-test; E:T ratio: 1:1, p = 0.003; 2:1, p = 0.050). (E) 10 μg of whole HeLa 33/2 cell extracts were resolved on a 15% SDS-PAGE for western blotting with anti-HA antibody to control the expression of the construct. The western blot is representative of 5 independent assays.
To monitor the condensation reaction in cellulo, HeLaHLA-A*03:01+ cells were transfected with a plasmid expressing a HA-Ub-gp100mel40–52 fusion protein (HelaA*03:01+Ub/gp100mel40–52). In the fusion protein the GG motif of the ubiquitin was directly linked to the first amino acid residue of the 13-mer gp10040–52 peptide allowing the release of the unchanged full-length gp100mel40–52 peptide substrate after cleavage behind the GG motif by de-ubiquitinating enzymes (Fig. 2C). CTL assays using the gp100mel47–52/40–42 specific CD8+ T cell clone demonstrated that proteasomes could indeed also perform a condensation reaction in cellulo (Fig. 2D,E), indicating that in addition to transpeptidation also condensation reactions may also contribute to the cellular pool of spliced epitopes.
The gp100mel47–52/40–42 peptide is a tumor epitope presented by melanoma cells
This data raised the question whether the proteasome-catalyzed generation of gp100mel47–52/40–42 was solely determined by the biochemical nature of the 13-mer gp100mel40–52 substrate and/or the specific assay conditions applied in vitro and/or in cellulo. To further validate our reverse immunology approach, we next investigated whether the gp100mel47–52/40–42 peptide was also naturally produced from full-length gp100mel antigen by patient-derived melanoma cells. Of the five different patient-derived melanoma cell lines tested for gp100mel protein and HLA-A*03:01 expression only Mel63a cells proved to be HLA-A*03:01+/gp100mel + (Fig. 3A,B). Immunoblotting revealed that Mel63a cells mostly expressed intermediate-type proteasomes, expressing all standard and IFN-γ inducible proteasome subunits except the β2i subunit (Fig. 3B). Interestingly, the Mel63a cells elicited a strong IFN-γ release by the gp100mel47–52/40–42 specific T-cell clone, suggesting that the spliced peptide gp100mel47–52/40–42 can indeed be produced from the full-length gp100mel protein (Fig. 3C). This proved that the spliced gp100mel47–52/40–42 peptide identified by applying SpliceMet in in vitro experiments represented a natural tumor epitope produced from the full-length gp100mel membrane protein and was presented on the surface of gp100mel/HLA-A*03:01+ patient-derived melanoma cells.
Melanoma Mel63a cells efficiently generate and present the gp100mel47–52/40–42 epitope.
(A) Overlay histograms obtained by flow cytometry showing the cell surface expression of the HLA-A*03:01 molecule (thick line) compared to isotype control (thin line) of the Mel63a melanoma cell line. Shown is a representative staining of 2 independent experiments. (B) The four melanoma cell lines Mel15a, Mel18, Mel21a and Mel63a were analyzed by western blotting for their content of gp100mel, the proteasome subunits β1, β2, β5, β1i, β2i, β5i and α6 (the latter as loading control). HeLa cells and HeLa 33/2 cells were used as controls for s- and i-proteasomes, respectively. (C) The four melanoma cell lines Mel15a (HLA-A*03:01−/gp100+); Mel18 (HLA-A*03:01−/gp100mel+), Mel21a (HLA-A*03:01−/gp100mel−) and Mel63a (HLA-A*03:01+/gp100mel+) were assessed for their ability to present the gp100mel47–52/40–42 epitope in a 16 hrs CTL assay to K631 1C CTL clone. Supernatants were collected and tested for their IFN-γ content by ELISA. Values are means and SD of 2 replicates in a representative assay of 2 independent experiments.
The gp100mel47–52/40–42 epitope is generated by the ERAD pathway
To exclude the possibility that generation of the gp100mel47–52/40–42 peptide was restricted to a specific cell type and to study the contribution of individual proteasome subunits in the production of the peptide, HeLa cells were transfected with plasmids encoding the HLA-A*03:01 protein and a full-length gp100mel-myc/His fusion protein. As shown in Fig. 4A, HeLa cells efficiently presented the gp100mel47–52/40–42 epitope in a proteasome-dependent manner providing evidence that epitope generation was not dependent on a specific cell type. The in vitro experiments had indicated a driving role for the two different β5 subunits in the generation of the gp100mel47–52/40–42 epitope (see Fig. 1F,G). To study the role of the β5-subunit isoforms in a cellular context, we monitored gp100mel47–52/40–42 epitope presentation in HeLa 33/2 cells, which constitutively express i-proteasomes, using a siRNA approach. The siRNA-induced silencing of β5i subunit expression resulted in its substitution by the standard β5 subunit within the 20S proteasome complex (Fig. 4B). This subunit substitution led to a remarkable reduction of gp100mel47–52/40–42 presentation underlining the importance of the β5i subunit for this splicing reaction (Fig. 4B). Corroborating this result, over-expression of the β5i subunit in HeLa cells and concomitant formation of β5i/ β1/β2 subunits containing intermediate type 20S proteasomes led to a strong increase in gp100mel47–52/40–42 presentation (Fig. 4C). Interestingly, the proteasome subunit composition found in Hela cells overexpressing β5i subunit was similar to that observed in Mel63a cells suggesting that the intermediate type proteasomes carrying β5i, β2 and β1 subunits are best equipped for the generation and presentation of this spliced epitope.
Ub-proteasome- and ERAD-dependent gp100mel47–52/40–42 epitope production is enhanced by β5i subunit.
(A) HeLa cells were transfected with HLA-A*03:01 and gp100mel and were subjected to epoxomicin treatment (250 nM; 2 hrs) or DMSO (control) and compared to HeLa cells transfected with HLA-A*03:01 treated with epoxomicin (250 nM; 2 hrs) and pulsed with the synthetic peptide gp100mel47–52/40–42 (50 μM). Values are means and SD of 2 replicates in a representative assay of 2 independent experiments. Significant differences (between the samples with or without epoxomicin) in the means of replicates are marked by * (student t-test; E:T ratio: 1:1, p = 0.002; 2:1, p = 0.001; 4:1, p < 0.001; 8:1, p < 0.001). (B) HeLa 33/2 cells were exposed to β5i subunit or control siRNAs (30 nM for 24 hrs) and subsequently transfected with gp100mel. Cell extracts were analyzed by western blotting (representative of 4 independent experiments). β5i subunit-depleted HeLa 33/2 cells were subjected to a 16 hr CTL assay. Values are means and SD of 2 replicates in a representative assay of 4 independent experiments. Significant differences in the means of replicates are marked by * (student t-test; p = 0.047). (C) HeLa cells were transfected with plasmids encoding HLA-A*03:01, gp100mel full-length protein, β5i subunit or the empty vector. The HeLa cell ability to process the gp100mel47–52/40–42 was assessed by CTL assay. Values are means and SD of 2 replicates in a representative assay of 4 independent experiments. Significant difference in the means of replicates is marked by * (student t-test; p = 0.001). (D) HeLa cells were exposed to 30 nM siRNA specific for the 19S Rpn10 subunit or p97/VCP (or control siRNA) prior to transfection with the HLA-A*03:01 and the full-length gp100mel protein plasmids and pulsed or not pulsed with the synthetic peptide gp100mel47–52/40–42 (30 μM). Values are means and SD of 2 replicates in a representative assay of 3 independent experiments. Significant differences in the means of replicates are marked by * (student t-test) for Rpn10 siRNA (E:T ratio: 1:1, p = 0.038; 2:1, p = 0.029; 4:1, p = 0.047; 8:1, p = 0.002) and for p97/VCP siRNA (E:T ratio: 1:1, p = 0.016; 2:1, p = 0.020; 4:1, p = 0.024; 8:1, p = 0.001). The knock-down efficiencies were monitored by determining the steady-state level of Rpn10 or p97/VCP proteins by western blotting.
Because gp100mel is a membrane protein we also asked whether generation of the gp100mel47–52/40–42 epitope from gp100mel protein resulted from the ER-associated ubiquitin-dependent protein degradation pathway (ERAD). To test this, expression of the ubiquitin-receptor subunit Rpn10 residing in the 19S complex and of the ERAD component p97/VCP were silenced by siRNA. As shown in (Fig. 4D) silencing of either Rpn10 or p97/VCP expression significantly impaired the endogenous gp100mel47–52/40–42 epitope presentation providing evidence that PCPS takes place as part of normal ubiquitin-dependent protein degradation and can also involve the ERAD pathway.
PBMCs of melanoma patients harbor spliced epitope-specific T cells
Because the gp100mel47–52/40–42 epitope had been identified using an open unbiased algorithm-aided reverse immunology approach, the possibility still remained that its identification and generation may have been the result of the experimental set ups. Hence, it was important to validate whether the gp100mel47–52/40–42 peptide was also generated in vivo and that melanoma patients possessed CD8+ T cells directed against this epitope.
For proof of principal, we therefore tested whether HLA-A*03:01+ melanoma patients possessed CD8+ T cells directed against the gp100mel47–52/40–42 peptide (Fig. 5A) and/or the HLA-A*03:01-restricted spliced peptide gp100mel195–202/192 [RSYVPLAH][R] (Fig. 5B), which was previously shown to trigger a T cell response in melanoma patients18. In total, PBMCs from 20 randomly chosen melanoma patients and 4 healthy controls were analyzed. Interestingly, 6/12 untreated patients and 2/8 vaccinated patients displayed T cells reactive against the gp100mel47–52/40–42 epitope (Fig. 5A). Only 1 out of 4 healthy controls had a measurable specific response toward the same spliced epitope (Fig. 5A), as further confirmed by FACS analysis (Fig. 5C). On the other hand, 5/6 untreated and 2/8 vaccinated melanoma patients contained T cells recognizing the gp100mel195–202/192 peptide (Fig. 5B). This data demonstrated that our algorithm-based reverse immunology approach is a potent strategy to identify new spliced epitopes. They also suggested that proteasome-catalyzed generation of spliced antigenic peptides is not an isolated singular event but is part of the physiological proteasome-dependent epitope generation process, which elicits specific CD8+ T cell responses in vivo in melanoma patients.
IFN-γ responses to the gp100mel47–52/40–42 and gp100mel195–202/192 spliced peptides in HLA-A*03:01+ healthy controls and melanoma patients.
PBMCs from healthy controls and patients with melanoma vaccinated or not with dendritic cells pulsed with the gp100mel209–217 peptide were cultured in 96-well plates and stimulated with 30 μM gp100mel47–52/40–42 (A) or gp100mel195–202/192 (B) synthetic peptides. After 10 days of culture, the presence of CTLs specific for gp100mel47–52/40–42 and gp100mel195–202/192 epitopes in each well was assessed by re-stimulating the cells with the gp100mel47–52/40–42 or gp100mel195–202/192 peptide in a 16 hr assay in the presence of CD28/CD49d co-stimulatory molecules. A negative control without peptide was included in all experiments. After 16 hrs, supernatants were collected and tested for their IFN-γ content by ELISA. Shown is the number of wells per plate in which the IFN-γ secretion upon peptide stimulation was larger than 100 pg/ml and at least twice as large as that detected without peptide for each patient and/or control. (C) Validation of the ELISA outcome on the representative healthy control well showing the larger IFN-γ response by CD8+ T cells in presence of the target spliced epitope. CTLs were double stained for CD8 and intracellular IFN-γ by FACS analysis. The side/forward scatters chart is reported in the left panel. Cells included in the analysis are marked with a circle and are shown in the two right panels, where CTLs were co-cultivated for 16 hrs with antigen presenting cells pulsed or not pulsed with the 30 μM gp100mel47–52/40–42 synthetic peptide prior the FACS analysis.
Spliced and non-spliced epitopes are presented at the cell surface in comparable amount
Despite being an important issue in judging the immune relevance of spliced epitopes, there exists no study comparing the relative presentation efficacy of different spliced epitopes with each other or with non-spliced epitopes. The gp100mel antigen seemed be ideally suited for such an analysis because the spliced peptides gp100mel47–52/40–42 (HLA-A*03:01), gp100mel40–42/47–52 (HLA-A*32:01), gp100mel195–202/192 (HLA-A*03:01) and the non-spliced gp100mel209–217 (HLA-A*02:01) were all generated from the same protein. To extend the analysis to a second and gp100mel-independent non-spliced epitope we also included the tumor-associated HLA-A*02:01-restricted epitope NY-ESO-1157–16526 into the comparison. However, to perform such comparison, different HLA-A peptide binding affinities, HLA expression levels as well as the different affinities of the CD8+ T cells have to be taken into consideration.
Therefore we first performed titration experiments with HeLa cells expressing the target HLA-A molecules, pulsed with the antigenic peptides and co-cultivated them with the respective peptide-specific CD8+ T-cell clones (Fig. 6A–E upper panels). By measuring the resulting IFN-γ release, informative titration curves where obtained for all antigenic peptides (Fig. 6B–E) but the non-spliced gp100mel209–217 epitope (data not shown). Because gp100mel209–217 is known be a poor HLA-A*02:01 binder, we consequently performed the titration experiments with the previously described M210 gp100mel209–217 peptide, in which the anchor residue T210 of gp100mel was exchanged with a Methionine for better HLA-A *02:01 binding. T cell assays were performed with transduced CD8+ T cells expressing the corresponding TCR. In this case satisfactory titration curves were obtained (Fig. 6A). We next transfected HeLaHLA-A*03:01+ cells with plasmids encoding the M210gp100mel variant, or the NY-ESO-1 proteins together with the three HLA-A*32:01, -A*02:01 and -A*03:01molecules. The transfected cells were then tested for their ability to activate CD8+ T-cells specific for each of the peptides as outlined above (Fig. 6A–E lower panels).
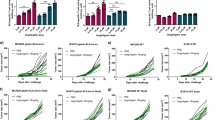
The processing efficacy of the gp100mel–derived spliced epitopes is comparable to that of the non-spliced M210gp100mel209–217 and NY-ESO-1157–165 epitopes.
(A–E) The IFN-γ-release by specific CD8+ T cell clones exposed to HLA-A*03:01/-A*02:01/-A*32:01+ HeLa cells (E:T = 1:1) pulsed with different concentrations of the synthetic peptides (upper panels) or HLA-A*03:01/-A*02:01/-A*32:01+ HeLa cells transfected with the target antigen (lower panels) is shown for the epitopes M210gp100mel209–217 (A), NY-ESO-1157–165 (B), gp100mel40–42/47–52 (C), gp100mel47–52/40–42 (D) or gp100mel195–202/192 (E). The data are the mean and bars the SD of three replicates of a representative assay of independent experiments (n = 2–6). (F) We here report a marker of the amount of epitope presented on the cell surface of HLA-A*03:01/-A*02:01/-A*32:01+ HeLa cells transfected with the target antigen. By titrations (upper panels in A–E) we calculated the amount of synthetic peptide pulsed on to HeLa cells, which triggered the release of IFN-γ amount corresponding to the IFN-γ released by the specific CD8+ T cell clones exposed to HLA-A*03:01/-A*02:01/-A*32:01+ HeLa cells transfected with the target antigen and cultivated in an E:T ratio = 4:1 (see lower panels in A–E). Values are the means and bars the SEM of independent experiments (n = 2–7; each experiment with three independent replicates). Significant differences in the means are marked and the corresponding by p values are reported (we applied the Mann-Whitney test with Bonferroni correction for multiple comparison). The IFN-γ-release in the medium was measured by ELISA after 16 hrs co-culture.
We considered the measured IFN-γ release obtained with an 4:1 E:T ratio and determined the corresponding peptide concentrations on the basis of the titration experiments. We chose the 4:1 E:T ratio to obtain IFN-γ release values in the log-phase of the titration curves. By applying this method we obtained a semi-quantitative comparison of the amount of non-spliced vs spliced epitopes generated by proteasomes of HeLa cells and presented by MHC class I molecules and. As shown in Fig. 6F the amounts of the spliced gp100mel40–42/47–52 epitope and of the non-spliced epitope M210gp100mel209–217 were comparable and the amounts of the spliced gp100mel40–42/47–52 epitope exceeded those of all other epitopes analyzed. In contrast, cell surface expression of the new spliced gp100mel47–52/40–42 epitope was significantly lower, which is in agreement with our observation that s-proteasomes produce only relatively low amounts of the gp100mel47–52/40–42 epitope. Interestingly, also the expression of the non-spliced NY-ESO-1157–165 epitope was lower than that of the gp100mel40–42/47–52 and T210Mgp100mel209–217 epitopes and comparable with the spliced gp100mel195–202/192 epitope, requiring C-terminal trimming. As a control, we compared the CD8+ T-cell response towards gp100mel40–42/47–52 and gp100mel47–52/40–42 epitopes in HeLa cells expressing the wild-type gp100mel and the mutated T210Mgp100mel proteins in two independent experiments. No difference emerged confirming that the use of the T210Mgp100mel variant did not affect the generation of the distant spliced epitopes (Fig. S5).
In summary, these experiments demonstrate that generation and presentation of spliced epitopes occurs with efficacies similar to what is observed for canonical non-spliced epitopes and do not represent exceptions with respect to their immune relevance.
Discussion
Generation of spliced epitopes is an additional proteasome-dependent activity, contributing to the pool of MHC-class I presented antigenic peptides. A number of features of the proteasome-dependent splicing events indicate that peptide splicing is not an accidental side reaction of proteasome activity but rather an authentic post-translational modification. One important aspect of PCPS is the fidelity of the process because to trigger a CD8+ T cell response a spliced peptide must be repeatedly generated from a given substrate protein. Thus, for a functional purpose of the resulting splicing products PCPS must be a finely tuned event27. However, a proteasome-catalyzed splicing event is difficult to detect because it requires the complementary information on gene sequence and processing events. Because of that, spliced epitopes have so far escaped wide spread identification using available databases in combination with mass spectrometric analyses. We here applied an algorithm aided reverse immunology approach, which a priori does not rely on available patient-derived CD8+ T cells and therefore greatly facilitates the identification of new spliced epitopes.
As part of our analyses the spliced gp100mel47–52/40–42 peptide was identified as a new HLA-A*03:01-restricted tumor epitope expressed on patient-derived gp100mel+ melanoma cells. Interestingly, the gp100mel47–52/40–42 peptide was generated in vitro from the same gp100mel40–52 13-mer polypeptide substrate as the previously reported spliced gp100mel40–42/47–52 epitope12 but with the same peptide fragments being arranged in reverse order. This disclosed that in addition to the well-established transpeptidation reaction11,12,14 PCPS could also involve condensation reactions that in vitro are independent of prior proteasomal peptide-bond hydrolysis. Production of the spliced antigenic peptide in a condensation reaction could also be observed in cellulo when the 13-mer gp100mel40–52 polypeptide substrate was expressed in HeLa cells as part of an ubiquitin fusion protein. Condensation reactions performed by proteinases have only been reported to occur in vitro under defined diffusion-limiting conditions. The gated barrel-shaped catalytic cavity of the 20S proteasome hindering rapid diffusion of the generated peptide as well as the catalytic activity of Thr1 via the formation of an ester-bond intermediate may however explain why proteasomes are able to perform condensation reactions also in a cellular environment with relative efficacy. In fact, in case the peptide ligands are held in the right orientation in close proximity, this does not even require catalysis. It is however not possible to determine the extent to which condensations reactions participate in cellulo to the overall production of the gp100mel47–52/40–42 peptide from the full-length gp100mel protein.
Theoretically, only relatively few MHC-class I-peptide complexes have to be exposed at the cell surface to trigger an immune response28 and T cells directed against spliced epitopes have been identified among a subset of tumor infiltrating lymphocytes10. Nevertheless, the notion that spliced epitopes represented a curiosity still prevailed and doubts remained whether PCPS was a sufficiently frequent event to play any significant role in MHC class I-restricted immune responses. Importantly, PCPS does not randomly produce an unlimited number of all theoretically possible spliced peptides14,23. Rather, PCPS seemed to be restricted by certain sequence characteristics of the substrate protein, guaranteeing a high reproducibility in the generation of spliced epitopes14. But similar to the well-characterized proteasome-generated non-spliced epitopes for which reliable rules are still missing, general sequence rules governing spliced epitope generation are so far unresolved.
An important issue with respect to their immune relevance and a still unresolved question was whether spliced epitope presentation is at all comparable with that of non-spliced conventional tumor epitopes. Therefore, in a semi-quantitative analysis we also compared for the first time the relative efficacy of endogenous presentation of three different gp100mel derived spliced epitopes with that of two non-spliced tumor epitopes. Relative cell surface presentation efficacy of the three gp100mel derived spliced epitopes analyzed here differed considerably among each other. In particular, cell surface expression of the spliced gp100mel40–42/47–52 epitope was significantly superior to the presentation of the newly identified gp100mel47–52/40–42 epitope, which is composed of the same splice-reactants but in reverse order. This difference may be explained in that the reverse order peptide-ligation by either transpeptidation or condensation is less efficient and/or that the generation of the gp100mel47–52/40–42 epitope is not favored by s-proteasomes. Importantly, our data show for the first time that the overall relative cell surface expression levels of the spliced epitopes is comparable to that of the non-spliced gp100mel209–217 and NY-ESO-1157–165 epitopes. This suggests that with regard to their relative presentation efficacy proteasome-generated spliced epitopes exert features almost identical to those known for non-spliced epitopes and do not greatly differ from what is known for canonical epitopes.
Considering that the spliced gp100mel47–52/40–42 epitope was identified in an unbiased reverse immunology approach and the fact that the PBMCs from approximately 50% of randomly chosen untreated melanoma patients contained CD8+ T cells that were stimulated by the gp100mel47–52/40–42 peptide suggested that generation of spliced epitopes is indeed a rather frequent event. Corroborating this result, five out of six untreated patient samples turned out to contain memory CD8+ T cells responsive to the spliced HLA-A*03:01-restricted epitope gp100mel195–202/191 [RSYVPLAH][R] that had initially been identified through the analysis of tumor infiltrating lymphocytes18. That a fewer number of samples obtained from melanoma patients having undergone vaccination with dendritic cells were responsive to the spliced antigenic peptides might be owed to the fact that they contained a relatively larger percentage of T cells directed against the non-spliced peptides, which were used for vaccination. Although our assay conditions are not designed to prime naïve T-cells in that dendritic cells were in low number and not activated, we can not completely exclude that the immune response towards the spliced epitopes may in part be due to priming and the activation of naïve CD8+ T cells during the in vitro assays
Supporting the conclusion that generation of MHC-class I ligands by PCPS was not a rare event, we also identified, by applying our SpliceMet-supported reverse immunology approach, a significant number of potential spliced epitopes derived from human gp100mel or Listeria monocytogenes LLO antigens and with high HLA-A and/or -B binding affinity, which warrant further immunological characterization (Table 1).
Our data also demonstrated that PCPS is intrinsically linked to ubiquitin-dependent proteasomal protein turnover and proteasomal hydrolysis events and that it was not a rare side reaction. Since peptide exhibiting features suitable for binding to MHC class I molecules seem to occur in among spliced peptides with a certain preference14, the relative probabilities for generation of a spliced epitope or a non-spliced epitope from a given substrate are most likely not very different. Therefore, the pool of spliced antigenic peptides available for selection by the antigen presentation machinery may be significantly larger than previously estimated. In fact, as shown here for the gp100mel protein PCPS also seems to enlarge the HLA haplotype-restriction of antigenic peptides that can be produced from a single substrate protein thereby broadening the cellular immune response.
Materials and Methods
A detailed Material and Method section is disclosed in the Supplementary information.
Peptide synthesis and quantification
The sequence enumeration for the polypeptides gp10040–52 [RTKAWNRQLYPEW], gp100mel209–217 [IMDQVPFSV], gp100mel195–202/92 [RSYVPLAH][R] are referred to the human protein gp100PMEL17, for the peptide [SLLMWITQC] to the NY-ESO-1 human protein. All peptides were synthesized as described elsewhere23. LC-MS analyses were performed as previously described14. Database searching was performed using the SpliceMet’s ProteaJ algorithm23. Quantification of proteasome-generated non-spliced and spliced peptides (Fig. 1, Fig. S2) was carried out by applying the QME method to LC-MS analyses as described elsewhere14. For semi-quantitative studies of the amount of the two spliced peptides [QLYPEW][RTK] and [RTK][QLYPEW] (Fig. 1F,G), the heavy peptides QLYPE+6WRTK and RTK+6QLYPEW were used as internal standards. The limit of quantification was set to 10 fmol upon preliminary experiments with synthetic peptide titration
In vitro processing of synthetic peptides
The synthetic peptide gp100mel40–52 (40 μM) was digested by 3 μg 20S proteasomes in 100 μl TEAD buffer over time at 37 °C. In the in vitro hydrolysis-independent proteasome-mediated experiments (Fig. 1F,G) 5 μg 20S proteasomes were incubated in 100 μl TEAD buffer in presence of 100 μM peptides for 20 hours. For the experiments with the inhibitor PR-893, PR-924 (Onyx Pharmaceuticals, US), YU-102 and LU-10225 the specific binding to the proteasome subunits was monitored.
20S proteasome purification
20S proteasomes were purified from LCLs and T2 cell line as previously described29. 20S proteasome purified from human erythrocytes and spleen were purchased from BioMol.
Cell culture
Cells used in this study include LCLs, T2 cells, HeLa cells, HeLa 33/2 cells which were all previously described9,30,31. The melanoma cell lines Ma-Mel15a, Ma-Mel18, Ma-Mel21a and Ma-Mel63a were established from tumor metastasis and grown in RPMI 1640 with 10% FCS, 1% L-glutamine and 1% penicillin. The CTL clones K631 1C specific for the HLA-A*03:01-restricted gp100mel47–52/40–42 epitope and RG39 specific for the HLA*02:01-restricted NY-ESO-1157–165 epitope were prepared according to the procedure of Fonteneau et al.32. The LG2-37.7.14 and M45-3B CD8+ T cell clones specific for the HLA-A*32:01-restricted gp100mel40–42/47–52 and HLA-A*03:01-restricted gp100mel195–202/192 spliced peptides, respectively were described elsewhere12,18. The gp100mel-specific TCR-transduced T cells were generated as previously described33,34.
HLA-A03:01-peptide binding affinity
Binding affinity of synthetic peptides were computed with different peptide concentration in a standard binding affinity assay measured by flow cytometry as described elsewhere29, using HeLa or T2 cells transfected with the HLA-A*03:01-expressing plasmid.
Plasmid and siRNA transfection
Expression vectors used in thy study include the pcDNA3.1 (HLA-A*02:01, HLA-A*32, β5i), pcDNA3.1/myc-HIS (gp100mel, NY-ESO-1) and the pcDNA3.1(+)/Zeo (HLA-A*03:01, HA-Ub/gp100mel40–52) plasmids. Transfection of HeLa and 33/2 cells or T2 cells was performed using Lipofectamin2000 or Amaxa-Nucleofactor technology, respectively, according to the manufacturer’s instructions. All siRNA (control, Rpn10 and p97/VCP) used in this study were purchased from Dharmacon and introduced into HeLa or 33/2 cells as previously described31.
Antibodies and western blotting
Cell extracts were prepared as previously described31. Proteins were resolved by SDS-PAGE and blotted with β1i, β2i, β5i, β1, β2, β5, α6 proteasome subunits, Rpn10, HA, gp100mel, p97/VCP and β-actin (as a loading control).
CTL assays and screening of gp10047–52/40–42-specific CD8+ T cells in HLA-A*03:01 healthy donors’ and melanoma patients’ PBMCs
Targets cells were serially diluted and co-cultured with a fixed amount of T cells, resulting in graded effector-to-target (E:T) ratio for 12 h prior to the measurement of IFN-γ release by ELISA. PBMCs from healthy donor or patients with melanoma were pulsed with 30 μM of the gp100mel47–52/40–42 synthetic peptide in RPMI medium supplemented with 8% human AB serum, 1% penicillin and IL-2 (50 U/ml). After 10 days of culture, their capacity to produce IFN-γ in response to the cognate peptide was measured by ELISA.
Statistical Analysis
Data were tested for normality distribution and homoscedasticity by Kolmogorov-Smirnov, Shapiro-Wilk and Levene tests. To identify significant difference between groups reported in Fig. 6, Kruskal-Wallis followed by Mann-Whitney tests with Bonferroni Post Hoc correction for multiple comparisons were applied. Otherwise, paired and unpaired Student t-tests were carried out. Descriptive statistics were carried out with SPSS (version 17) and R; a p-value <= 0.05 was considered statistically significant.
Additional Information
How to cite this article: Ebstein, F. et al. Proteasomes generate spliced epitopes by two different mechanisms and as efficiently as non-spliced epitopes. Sci. Rep. 6, 24032; doi: 10.1038/srep24032 (2016).
References
Dahlmann, B., Ruppert, T., Kuehn, L., Merforth, S. & Kloetzel, P. M. Different proteasome subtypes in a single tissue exhibit different enzymatic properties. J Mol Biol 303, 643–653 (2000).
Guillaume, B. et al. Two abundant proteasome subtypes that uniquely process some antigens presented by HLA class I molecules. Proc Natl Acad Sci USA 107, 18599–18604 (2010).
Groll, M. et al. Structure of 20S proteasome from yeast at 2.4 A resolution. Nature 386, 463–471 (1997).
Navon, A. & Ciechanover, A. The 26 S proteasome: from basic mechanisms to drug targeting. J Biol Chem 284, 33713–33718 (2009).
Kloetzel, P. M. Antigen processing by the proteasome. Nat Rev Mol Cell Biol 2, 179–187 (2001).
Liepe, J. et al. Quantitative time-resolved analysis reveals intricate, differential regulation of standard- and immuno-proteasomes. Elife doi: 10.7554/eLife.07545 (2015).
Huber, E. M. et al. Immuno- and constitutive proteasome crystal structures reveal differences in substrate and inhibitor specificity. Cell 148, 727–738 (2012).
Arciniega, M., Beck, P., Lange, O. F., Groll, M. & Huber, R. Differential global structural changes in the core particle of yeast and mouse proteasome induced by ligand binding. Proc Natl Acad Sci USA 111, 9479–9484 (2014).
Mishto, M. et al. Proteasome isoforms exhibit only quantitative differences in cleavage and epitope generation. Eur J Immunol 44, 3508–3521 (2014).
Vigneron, N. & Van den Eynde, B. J. Proteasome subtypes and the processing of tumor antigens: increasing antigenic diversity. Curr Opin Immunol 24, 84–91 (2011).
Warren, E. H. et al. An antigen produced by splicing of noncontiguous peptides in the reverse order. Science 313, 1444–1447 (2006).
Vigneron, N. et al. An antigenic peptide produced by peptide splicing in the proteasome. Science 304, 587–590 (2004).
Dalet, A., Vigneron, N., Stroobant, V., Hanada, K. & Van den Eynde, B. J. Splicing of distant Peptide fragments occurs in the proteasome by transpeptidation and produces the spliced antigenic peptide derived from fibroblast growth factor-5. J Immunol 184, 3016–3024 (2010).
Mishto, M. et al. Driving Forces of Proteasome-catalyzed Peptide Splicing in Yeast and Humans. Mol Cell Proteomics 11, 1008–1023 (2012).
Borissenko, L. & Groll, M. Diversity of proteasomal missions: fine tuning of the immune response. Biol Chem 388, 947–955 (2007).
Berkers, C. R., de Jong, A., Ovaa, H. & Rodenko, B. Transpeptidation and reverse proteolysis and their consequences for immunity. Int J Biochem Cell Biol 41, 66–71 (2009).
Somalinga, B. R. & Roy, R. P. Volume exclusion effect as a driving force for reverse proteolysis. Implications for polypeptide assemblage in a macromolecular crowded milieu. J Biol Chem 277, 43253–43261 (2002).
Michaux, A. et al. A Spliced Antigenic Peptide Comprising a Single Spliced Amino Acid is Produced in the Proteasome by Reverse Splicing of a Longer Peptide Fragment followed by Trimming. J Immunol 192, 1962–1971 (2014).
Hanada, K., Yewdell, J. W. & Yang, J. C. Immune recognition of a human renal cancer antigen through post-translational protein splicing. Nature 427, 252–256 (2004).
Dalet, A. et al. An antigenic peptide produced by reverse splicing and double asparagine deamidation. Proc Natl Acad Sci USA 108, 323–331 (2011).
Robbins, P. F. et al. Recognition of tyrosinase by tumor-infiltrating lymphocytes from a patient responding to immunotherapy. Cancer Res 54, 3124–3126 (1994).
Bonnet, D., Warren, E. H., Greenberg, P. D., Dick, J. E. & Riddell, S. R. CD8(+) minor histocompatibility antigen-specific cytotoxic T lymphocyte clones eliminate human acute myeloid leukemia stem cells. Proc Natl Acad Sci USA 96, 8639–8644 (1999).
Liepe, J. et al. The 20S Proteasome Splicing Activity Discovered by SpliceMet. Plos Computational Biology 6, e1000830 (2010).
Peters, B. & Sette, A. Integrating epitope data into the emerging web of biomedical knowledge resources. Nat Rev Immunol 7, 485–490 (2007).
Bellavista, E. et al. Immunoproteasome in cancer and neuropathologies: a new therapeutic target? Curr Pharm Des 19, 702–718 (2013).
Jager, E. et al. Induction of primary NY-ESO-1 immunity: CD8+ T lymphocyte and antibody responses in peptide-vaccinated patients with NY-ESO-1+ cancers. Proc Natl Acad Sci USA 97, 12198–12203 (2000).
Saska, I. & Craik, D. J. Protease-catalysed protein splicing: a new post-translational modification? Trends Biochem Sci 33, 363–368 (2008).
Cresswell, P. Antigen processing and presentation. Immunol Rev 207, 5–7 (2005).
Mishto, M. et al. Immunoproteasome LMP2 60HH variant alters MBP epitope generation and reduces the risk to develop multiple sclerosis in Italian female population. Plos One 5, e9287 (2010).
Mishto, M. et al. A structural model of 20S immunoproteasomes: effect of LMP2 codon 60 polymorphism on expression, activity, intracellular localisation and insight into the regulatory mechanisms. Biol Chem 387, 417–429 (2006).
Ebstein, F., Lehmann, A. & Kloetzel, P. M. The FAT10- and ubiquitin-dependent degradation machineries exhibit common and distinct requirements for MHC class I antigen presentation. Cell Mol Life Sci 69, 2443–2454 (2012).
Fonteneau, J. F. et al. Generation of high quantities of viral and tumor-specific human CD4+ and CD8+ T-cell clones using peptide pulsed mature dendritic cells. J Immunol Methods 258, 111–126 (2001).
Sommermeyer, D. et al. Designer T cells by T cell receptor replacement. Eur J Immunol 36, 3052–3059 (2006).
Morgan, R. A. et al. High efficiency TCR gene transfer into primary human lymphocytes affords avid recognition of melanoma tumor antigen glycoprotein 100 and does not alter the recognition of autologous melanoma antigens. J Immunol 171, 3287–3295 (2003).
Grant, E. et al. Nucleoprotein of influenza A virus is a major target of immunodominant CD8+ T-cell responses. Immunol Cell Biol 91, 184–194 (2013).
Acknowledgements
Work described here was supported by a Collaborative Research Grant of the Berlin Institute of Health (P.M.K. & W.U.), the Einstein-Stiftung Berlin (P.M.K.) and the Berlin Krebsgesellschaft (P.M.K.). N.V. is supported by Belgian Program of Interuniversity Poles of Attraction initiated by the Belgian State (Prime Minster´s Office, Science Policy Programming). We thank K. Hummel and E. Bellavista for excellent technical assistance, P. Kunert and B. Brecht-Jachan for the peptide synthesis, T. Ruppert (ZMBH) for Orbitrap support, H.S. Overkleeft, S. Stefanović, K.B. Kim, P. Coulie and Onyx Pharmaceuticals-Amgen subsidiary for providing biotechnological tools to carry out the experiments.
Author information
Authors and Affiliations
Contributions
F.E. and M.M. designed the project, performed experiments, analyzed data and contributed to write the manuscript; P.M.K. designed the project, analyzed data and contributed to write the manuscript; K.T.-T., C.K. and V.N. performed experiments and analyzed data; F.K.M.L., S.U., N.A.-K., P.H., A.N., A.L. and R.G. performed the experiments; K.J. analyzed the data; B.J.V.d.E. and W.U. contributed to write the manuscript; B.S.-T. and D.S. provided clinical samples.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ebstein, F., Textoris-Taube, K., Keller, C. et al. Proteasomes generate spliced epitopes by two different mechanisms and as efficiently as non-spliced epitopes. Sci Rep 6, 24032 (2016). https://doi.org/10.1038/srep24032
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24032
This article is cited by
-
Protein degradation by human 20S proteasomes elucidates the interplay between peptide hydrolysis and splicing
Nature Communications (2024)
-
InvitroSPI and a large database of proteasome-generated spliced and non-spliced peptides
Scientific Data (2023)
-
One-step artificial antigen presenting cell-based vaccines induce potent effector CD8 T cell responses
Scientific Reports (2019)
-
Novel insights into the HLA class I immunopeptidome and T-cell immunosurveillance
Genome Medicine (2017)
-
Extracellular proteasome-osteopontin circuit regulates cell migration with implications in multiple sclerosis
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.