Abstract
Cellular RNA-dependent RNA polymerases (RDRs) catalyze synthesis of double-stranded RNAs that can serve to initiate or amplify RNA silencing. Arabidopsis thaliana has six RDR genes; RDRs 1, 2 and 6 have roles in anti-viral RNA silencing. RDR6 is constitutively expressed but RDR1 expression is elevated following plant treatment with defensive phytohormones. RDR1 also contributes to basal virus resistance. RDR1 has been studied in several species including A. thaliana, tobacco (Nicotiana tabacum), N. benthamiana, N. attenuata and tomato (Solanum lycopersicum) but not to our knowledge in potato (S. tuberosum). StRDR1 was identified and shown to be salicylic acid-responsive. StRDR1 transcript accumulation decreased in transgenic potato plants constitutively expressing a hairpin construct and these plants were challenged with three viruses: potato virus Y, potato virus X, and tobacco mosaic virus. Suppression of StRDR1 gene expression did not increase the susceptibility of potato to these viruses. Phylogenetic analysis of RDR genes present in potato and in a range of other plant species identified a new RDR gene family, not present in potato and found only in Rosids (but apparently lost in the Rosid A. thaliana) for which we propose the name RDR7.
Similar content being viewed by others
Introduction
RNA silencing is a set of mechanisms whereby gene expression is suppressed in a sequence-specific manner. In plants RNA silencing is initiated by cleavage of double-stranded (ds) RNA precursors into 21–24 nucleotide small RNA (sRNA) by DICER-LIKE (DCL) endonuclease1. Depending on the nature of the precursor molecules sRNAs are classified as microRNAs [miRNAs: imperfectly ds molecules derived from stem-loop structures in endogenous transcripts] or short-interfering (si)RNAs, which are perfectly complementary dsRNA molecules derived from ds structures occurring in a variety of cellular and foreign substrates, including virus-derived RNA molecules. Following methylation catalyzed by HUA ENHANCER1 (HEN1) and unwinding, single-stranded (ss) sRNAs are incorporated into RNA-induced silencing complexes (RISCs) in which they direct the sequence-specific activity of an ARGONAUTE (AGO) protein against nucleic acid sequences that are complementary to the sRNA. RISC activity may include RNA cleavage or translational inhibition of mRNA1,2,3,4,5,6,7,8, or the methylation of specific DNA sequences9,10,11.
In plants and certain other eukaryotes, RNA silencing can be amplified by the activity of cellular RNA-dependent RNA polymerases (RDRs)12,13,14,15. RDRs synthesize dsRNA from ssRNA templates by primer-independent and primer-dependent mechanisms16. Primer-dependent production of dsRNA requires primary siRNA in order to recruit the RDR and initiate synthesis. The dsRNA synthesized by RDR activity is in turn processed by DCLs into secondary siRNAs that feed back into the silencing pathway17,18,19.
RDRs are encoded by a small gene family and share a putative catalytic domain, the DLDGD motif1,19,20. Varying numbers of RDR genes with various biological effects have been identified in different plant species. For example, Arabidopsis thaliana (hereafter referred to as Arabidopsis) has six RDR genes (RDR1, 2, 3a, 3b, 3c and 6), rice has five and Nicotiana benthamiana has three19. In tomato, the resistance genes Ty-1 and Ty-3 encode RDRs that direct methylation-mediated inactivation of the DNA genomes of geminiviruses and these genes possess some sequence similarities to AtRDRs 3a, 3b and 3c21. This suggests there is a degree of specialization or redundancy between different RDRs.
In Arabidopsis and a number of other plants, RDRs 1 and 6 have been shown to contribute to resistance against RNA viruses. RDR2, which is involved in establishment of transcriptional gene silencing, has also been implicated in anti-viral defence12,17,22,23,24,25,26,27,28,29,30,31. RDR6 is the best studied RDR with respect to its antiviral activities and it likely provides protection against the widest range of viruses. The other main ‘anti-viral’ RDR, RDR1, has stimulated considerable research because its expression is increased transiently by the defence-related signal molecules salicylic acid (SA), jasmonic acid (JA), ethylene, nitric oxide, as well as several other phytohormones, leading to suggestions that is involved not only in basal defence mechanisms but also in induced resistance mechanisms, such as systemic acquired resistance22,23,25,32,33,34,35. The importance of RDR1 in plant defence extends beyond resistance to viruses; RDR1 regulates JA-mediated changes in gene expression and defence against herbivory in Nicotiana attenuata32, and it influences expression of transcripts associated with pathogen resistance in tobacco (N. tabacum)36.
Potato (Solanum tuberosum) is the third most important food crop with an annual global production exceeding 374 million tonnes37. At least 40 viruses and two viroids are known to infect potato plants in the field, while a further 22 viruses and four viroids can infect potato plants experimentally38,39. However, the importance or otherwise of RDRs in potato-virus interactions is not well understood. In this paper we sought to explore the extent of the potato RDR gene family and investigate the role of StRDR1 in basal resistance to potato virus Y (PVY), potato virus X (PVX) and tobacco mosaic virus (TMV), which are three potato-infecting viruses for which RDR1 provides some level of protection in other hosts.
Results
Bioinformatic and phylogenetic analysis of plant RNA-dependent RNA polymerase genes
RDR genes with various biological effects have been identified in different plant species (summarized in Supplementary Table S1). Eight potential StRDR genes were identified by analysis of the Potato Genome Sequencing Consortium (PGSC) data. These include two RDR1 genes, referred to as StRDR1a and b and three RDR6 genes, referred to as StRDR6a, b and c. The Solanum RDR3 genes are the product of recent duplications peculiar to the Solanaceae and the Arabidopsis RDR3 genes are similarly recent and due to duplications in the Brassicaceae. As such there is no one-to-one orthology between the Arabidopsis RDR3a, b, and c genes (in earlier papers sometimes abbreviated as AtRDR3, 4, and 5, respectively: ref. 19) and the Solanum RDR3 genes, although they do belong to the same orthogroup. To recognize this we refer to them here as StRDR3e, d, and f (Supplementary Fig. S1).
The PGSC reference numbers identified for each StRDR gene locus, transcript, and coding sequence, as well as potential redundant and non-redundant RDR genes, are presented in Supplementary Fig. S1). Phylogenetic analysis of potato RDR sequences was performed to ensure that annotation of the potato RDRs by orthology assignment was to authenticated plant RDRs. It also allowed crosschecking for the presence or absence of a potato RDR2, for which no candidate sequence was initially apparent in the PGSC data. In Fig. 1a, the plant RDR phylogenetic tree that was produced using all published embryophyte plant genomes available in Phytozome v9 (http://www.phytozome.net/; date of access 17/09/2015) shows that most Arabidopsis and potato RDRs grouped according to the standard annotation19. Incorporating data from additional genetic resources such as the Solanum cDNA EST databases [NCBI BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi#alnHdr_56775579; date of access 17/09/2015) and Phytozome (http://www.phytozome.net/; date of access 17/09/2015)] revealed the existence of a potato RDR2 (Fig. 1a).
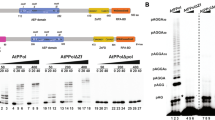
(a) Phylogeny of the potato RDR genes with other plant RDRs. White open circles are clade names, black circles are inferred gene duplications, red boxes are Solanum tuberosum sequences and grey boxes are Arabidopsis thaliana sequences. The S. tuberosum RDR sequences found on the Potato Genome Sequencing Consortium (PGSC) database are accompanied by their PGSC coding-sequence number. (b) Rosid-specific RDR7 phylogeny. This enlarged section of (a) shows the Rosid species that possess RDR7 gene sequences.
Figure 1a shows two separate groups of RDR genes, indicated by γ and α. This tree was constructed by separating these two groups because the Arabidopsis and potato RDR3 genes (shown in γ) bear little sequence resemblance to each other or to other RDR genes and were essentially non-alignable. They were therefore analysed separately and, as referred to above, the three potato RDR3 genes were referred to as StRDR3e, d, and f, to distinguish them from the Arabidopsis genes AtRDR3 a, b and c (AtRDR3, 4, and 5: ref. 19). Although these three Arabidopsis and potato RDR3 genes fall into the γ lineage they are each the product of recent duplications. Thus, the potato RDR3 genes are more closely related to each other than to the Arabidopsis RDR genes. Consequently, the potato RDR3 genes have not been named according to an equivalent Arabidopsis RDR in this clade.
Analysis of the available genome data for potato showed that there are two StRDR1 genes, which will be subsequently referred to as StRDR1a and StRDR1b. Alignment of the two transcribed StRDR1 sequences demonstrated that both are 3,348 nt in length and differ only by 17 nt, which are spread throughout the length of the sequences. These nucleotide differences result in 11 amino acid differences at the protein level. An alignment between the two predicted protein sequences demonstrated that the differences between the two StRDR1 proteins do not affect regions of amino acid sequence conserved among RDRs or those characteristic of RDR1 proteins19,40. Therefore, although there are two StRDR1 genes, comparison of the nucleotide and protein sequence alignments indicates they are equally likely to be functional in vivo. There is no evidence suggesting that either have been rendered non-functional due to a change in nucleotide sequence, such as, for example, the RDR1m gene that occurs in the widely used laboratory strain of N. benthamiana41, and which is a natural mutant due to a 72 nt insert within the open reading frame, containing tandem, in-frame stop codons25.
The apparent existence of three RDR6 genes in potato (indicated in the PGSC data) was puzzling as in most other plant species examined to date only one RDR6 gene sequence has been identified; barley (Hordeum vulgare) being the only known exception, with two RDR6 genes42. However, our alignment of the three potato RDR6 protein sequences (data not shown) indicates that a sequencing error was included in the PGSC data and that the putative StRDR6b and StRDR6c sequences are two parts of the same RDR6 locus. The alignment indicates that only one of these three RDR6 proteins (StRDR6a) is of full length (1,199 amino acids) and the other two proteins align roughly to either the N or the C termini of the full-length StRDR6 proteins (StRDR6b and StRDR6c, respectively). Furthermore, using the PGSC genome browser (http://solanaceae.plantbiology.msu.edu/cgi-bin/gbrowse/potato/; date of access 17/09/2015) it was found that the StRDR6b and StRDR6c loci are immediately adjacent to each other in the genome. Thus, potato has two RDR6 genes, here designated as RDR6a, and RDR6b. Figure 1a shows that tomato also has two RDR6 genes. This was confirmed also when checking the second available gene model produced by the International Tomato Genome Sequencing Project consortium: https://solgenomics.net/organism/Solanum_lycopersicum/genome; date of access 17/09/2015. This implies that RDR6 gene duplication occurred during the evolution of the genus Solanum and hence this is not specific to potato.
Identification of a novel Rosid-specific member of the RDR gene family: RDR7
Phylogenetic analyses were extended to include genomic data from a number of species in which RDR genes had not been examined previously. This resulted in the discovery of a previously unknown RDR clade, which we named RDR7 (Fig. 1b). RDR7 is Rosid-specific and although Arabidopsis is a Rosid43, RDR7 does not appear to exist in Arabidopsis, suggesting that RDR7 has been lost in this lineage (Fig. 1a). Potato is an Asterid and does not have an RDR7. Supplementary Table S1 shows in which plant species RDR genes have been studied and illustrates that the majority of research has been done on Arabidopsis as well as Nicotiana species, which are Asterids. Therefore, it is perhaps not surprising that RDR7 has remained unnoticed. Rosid species that were found to have RDR7 are shown in Fig. 1b, which presents the RDR7 region from the RDR phylogeny. These include a number of well-known food crops such as cassava (Manihot esculenta), common bean (Phaseolus vulgaris), soybean (Glycine max), cucumber (Cucumis sativus), cocoa (Theobroma cacao), papaya (Carica papaya), apple (Malus domestica), strawberry (Fragaria vesca), grape (Vitis vinifera), and orange (Citrus sinensis), as well as tree species such as Populus trichocarpa and eucalyptus (Eucalyptus grandis) (Fig. 1b).
Two Rosid species identified as having RDR7 genes, peach (Prunus persica) and cotton (Supplementary Table S1), have been used in previous research on RDRs by Di Serio et al.44 and Wang and co-workers45, respectively, but neither earlier study noted the existence of a Rosid-specific RDR gene. However, characterization of peach RDR genes was not the focus of the paper by Di Serio and colleagues44 and the cotton species (Gossypium hirsutum) studied by Wang and co-workers45 was different to the cotton species (G. raimondii) identified in our phylogenetic analysis (Fig. 1b).
Salicylic acid enhances accumulation of potato RDR1 transcripts
SA increases RDR1 gene expression in Nicotiana species and Arabidopsis22,25,33. To investigate whether potato RDR1 expression also responds to this phytohormone, potato plants (cv. Pentland Dell) were treated with 1 mM SA and StRDR1 expression was analysed by a reverse transcription coupled to the quantitative polymerase reaction (RTqPCR) using appropriate primers designed to allow detection of both StRDR1a and StRDR1b. Figure 2 shows relative StRDR1 transcript accumulation in control (water-treated) and SA-treated leaves at 0 (immediately before treatment), 24 and 48 hours after treatment. SA-treated leaves show a significant two-fold increase in StRDR1 expression at 24 hours post treatment in comparison to the control treatment at 24 hours. By 48 hours post treatment, StRDR1 expression had declined but was still significantly higher than in control-treated plants at that time point. This experiment was done three times with similar results.
Samples were taken at 0 hours (immediately before treatment), 24 hours and 48 hours after SA or control (water) treatment. Control-treated Pentland Dell plants were sprayed with water and the control sample at 0 hours was used as the reference. Cyclophilin was used as the reference gene for the RTqPCR analysis. Significant increases in StRDR1 transcript accumulation were seen at 24 hours and 48 hours after SA treatment (p < 0.005; *). Error bars represent standard error of the mean.
Depletion of StRDR1 transcript accumulation in transgenic potato
Two DNA constructs encoding dsRNA hairpin transcripts were designed to silence the transcripts of StRDR1a and StRDR1b (Supplementary Table S2). Construct 1 corresponded to a sequence in the 5′ region of the StRDR1 transcripts and the other (Construct 2) to a sequence in the 3′ region (Supplementary Fig. S2). A BLAST search using the two chosen RNA interference (RNAi) inducing sequences was done to ensure they did not match other plant gene sequences. Transgenic potato (cv. Pentland Dell) lines expressing either of the two hairpin constructs (Supplementary Fig. S2) were made and lines showing decreased accumulation of StRDR1 transcripts were identified (Fig. 3 and Supplementary Fig. S3). Both hairpin constructs appeared to be equally effective at diminishing accumulation of StRDR1 transcripts. Therefore, in most subsequent experiments to assess the effect of silencing StRDR1 on the susceptibility of potato plants to viral disease, line 1.3 (expressing Construct 1) was used. Figure 3c shows the relative RDR1 expression for lines 1.3 and 2.5 in comparison to non-transformed plants. To ensure that there were no off-target effects of either of the RNAi constructs, the accumulation of StRDR2 (Fig. 3d) and StRDR6 (Fig. 3e) transcripts were compared in non-transformed plants and transgenic lines 1.3 and 2.5. It was found that expression of silencing constructs for StRDR1 did not affect the expression of the StRDR2 or the StRDR6 genes.
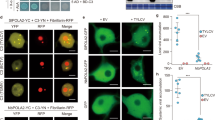
PCR of plant genomic DNA to verify the presence of the StRDR1-specific hairpin constructs RNAi1 (a) or RNAi2 (b) in the transformed potato lines. PCR used a primer complementary to a sequence located in the 35S promoter of the T-DNA insert and primers specific to either RNAi 1 and 2, to amplify regions of 535 bp and 550 bp, respectively. DNA from non-transformed (NT) plants and a no template PCR control (negative: -ve) were included in the analysis and control PCR reactions were carried out using primers specific for β-tubulin (expected product of 480 bp). (c) StRDR1 accumulation is efficiently decreased in a potato lines expressing the RNAi1 (Line 1.3) and RNAi2 construct (Line 2.5) (for other examples see Supplementary Fig. S3). Accumulation levels of transcripts of StRDR2 (d) and StRDR6 (e) were not diminished in plants of transgenic lines expressing either RNAi1 or RNAi2 to inhibit StRDR1 expression. Error bars represent standard error of the mean.
Down-regulation of StRDR1 expression did not enhance susceptibility to tobacco mosaic virus
TMV has been used as the model virus in several previous studies of RDR1 in solanaceous hosts (see introduction). Non-transgenic plants of the potato cultivar Pentland Dell were susceptible to TMV infection in directly-inoculated leaves but the virus did not spread systemically. Figure 4a shows relative TMV RNA accumulation at 7 days post inoculation (dpi) in inoculated leaves of non-transformed and StRDR1-silenced line 1.3 in three independent experiments. TMV RNA accumulated to similar levels in the inoculated leaves of non-transformed plants and transgenic plants with diminished StRDR1 transcript accumulation. In only one out of three experiments was any increase noted in the accumulation of TMV RNA in transgenic plant tissue compared to non-transformed (Fig. 4a left panel: p = 0.007). TMV RNA could not be detected in the upper, non-inoculated leaves of either non-transformed or transgenic plants and infection with TMV did not appear to result in the appearance of disease symptoms in either type of plant. Increased RDR1 expression has been proposed to be associated with TMV infection in susceptible tobacco22. However, we did not see any increase in StRDR1 transcript accumulation following infection of non-transformed or transgenic potato plants with this virus (Fig. 4b).
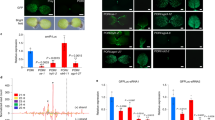
(a) RTqPCR was used to determine the relative level of TMV coat protein (CP) RNA (a) in inoculated leaves of non-transformed (NT) and transgenic Line 1.3 plants at 7 days post-inoculation (dpi). RNA was also extracted from mock-inoculated leaves. In each of three independent experiments, TMV CP RNA accumulation in transgenic samples was normalized to that in the non-transformed control. In only one experiment was any significant increase in CP accumulation noted (p = 0.007: *) (b) RTqPCR was used to measure accumulation, relative to that mock-inoculated non-transformed plant, of RDR1 in mock-inoculated and TMV-inoculated plants at 7 dpi in inoculated, mock-inoculated leaves and upper non-inoculated leaves. Error bars represent standard error of the mean.
Down-regulation of StRDR1 expression did not diminish resistance to PVX or PVYO
We examined the effect of depletion of StRDR1 transcript accumulation on the ability of resistant potato plants to inhibit virus spread. PVX induces visible necrotic hypersensitive response (HR) lesions at inoculation sites in the cultivar Pentland Dell due to the presence of the resistance genes, Nx and Nb39,46. PVYO causes severe disease symptoms in susceptible potato cultivars but in Pentland Dell the virus triggers visible HR lesions but does not normally spread systemically. We examined, in plants with diminished StRDR1 transcript accumulation, if either virus was able to escape the confinement imposed by HR-type resistance or replicate to higher levels than in non-transformed plants.
Using primers specific for either PVX or PVYO viral RNA accumulation was assessed by RTqPCR in inoculated leaves of non-transformed plants and plants of the StRDR1-depleted line 1.3 (Fig. 5). Three independent experiments were carried out for each virus but neither PVX nor PVYO showed any consistent increase in accumulation in the inoculated leaves of transgenic line 1.3 versus the leaves of the non-transformed controls (Fig. 5). Although there was variation in the relative viral RNA accumulation in inoculated leaves of plants of transgenic line 1.3 (see Fig. 5a, right panel and Fig. 5b, middle panel), it is likely to have resulted from variation in the numbers of inoculation sites per leaf. There were no obvious changes in HR lesion morphology on leaves of transgenic line 1.3 inoculated with PVX or PVYO (Supplementary Fig. S4) and no symptoms were apparent on the upper, non-inoculated leaves of these plants (Supplementary Fig. S5). Attempts to detect RNA of either virus in non-inoculated leaves of transgenic line 1.3 plants were unsuccessful. Therefore, silencing StRDR1 did not result in any breakdown in HR-type resistance to PVX or PVYO.
RTqPCR was used to determine the relative levels of PVX (a) and PVYO (b) RNA in inoculated leaves of non-transformed (NT) and transgenic Line 1.3 plants at 7 and 10 days post-inoculation, respectively. RNA was also extracted from mock-inoculated leaves. In each of three independent experiments, viral RNA accumulation in transgenic samples was normalized to that in the non-transformed control. Error bars represent standard error of the mean.
Since StRDR1 is SA-responsive and since SA is produced at increased levels following an HR in other solanaceous hosts, particularly in the vicinity of the lesion47,48, we investigated whether StRDR1 transcript accumulation was increased in leaves of non-transformed or StRDR1-depleted transgenic plants following inoculation with either PVX or PVYO. However, no increase in StRDR1 expression was detectable in these tissues (Supplementary Fig. S6).
Discussion
Prior to designing constructs to down-regulate StRDR1 expression in potato we carried out bioinformatic and phylogenetic analyses of the RDR gene family in this plant. Initially, this was carried out to aid the design of silencing constructs. However, we extended our analyses to encompass a range of other plant species to obtain a broader insight into the evolution of RDR genes. We found that potato has eight RDR genes in total, which were named StRDR1a, StRDR1b, StRDR2, StRDR3d, StRDR3e, StRDR3f, StRDR6a and StRDR6b (Fig. 1a). The naming of StRDR3 family members as StRDR3d, StRDR3e, StRDR3f was used since there is no direct orthology with the Arabidopsis genes AtRDR3a, b, or c. Construction of a phylogenetic tree indicated that the duplication of StRDR6 is not specific to S. tuberosum, as S. lycopersicum also has two RDR6 genes. Eight RDR genes is the largest complement of RDR genes possessed by any plant species examined to date. Until now, Arabidopsis was thought to have the highest number of RDR genes – six19. Our wider phylogenetic analysis identified a novel RDR clade. We have tentatively named this RDR7 to distinguish it from RDR1 since the RDR1 and RDR7 lineages split from each other approximately 108 years ago and because RDR7 is widely conserved across Rosid species but, does not occur in non-Rosid species (Fig. 1b). RDR7 appears to have been lost during the evolution of Arabidopsis, the best studied Rosid species (Supplementary Table S1), and as a consequence it appears that RDR7 has been overlooked in previous studies of RDR genes. To comment further on the biological function(s) of RDR7 or the evolution of the RDR7 clade would be speculative at this stage since further research needs to be done to characterize the properties of this gene and its product.
A key characteristic shared by RDR1 genes studied to date has been their transcriptional response to SA, as well as to a number of other phytohormones22,23,25,32,33,34. Consistent with previous studies, we found that SA treatment increased StRDR1 transcript accumulation, which indicates that regulation of RDR1 by this phytohormone is conserved in potato. RDR1 expression has been reported to increase following tobamovirus infection in susceptible tobacco and Arabidopsis plants22,23 although this was likely an artefact of the wounding associated with mechanical inoculation in those plants33. We found that no increase in StRDR1 transcript levels was detectable in either inoculated or non-inoculated leaves of potato plants in response to infection with TMV, indicating that RDR1 transcript accumulation in potato is less sensitive to wounding, consistent with results indicating that StRDR1 was not responsive to chewing by the Colorado potato beetle49. Although HR induction is associated with increased SA biosynthesis there was no evidence found for increased StRDR1 transcript accumulation following elicitation of necrotic HR lesions by either PVX or PVYO.
We successfully down-regulated the levels of the transcripts for the potato RDR1 genes (StRDR1a and StRDR1b) by expression of either of two hairpin constructs in transgenic plants. However, this did not cause any consistent or reproducible increase in susceptibility to TMV or breakdown of resistance to PVYO or PVX in the inoculated leaves. These results were surprising in light of multiple previous reports that have demonstrated that RDR1 is involved in defence against viruses: TMV and tobacco rattle virus in Arabidopsis23,28; TMV, PVX and PVY in tobacco22,36, and against TMV, as well as two other tobamoviruses (turnip vein-clearing virus and sunn-hemp mosaic virus) in N. benthamiana25,50. These results suggest either that RDR1 is not as critical for antiviral defence in potato as it appears to be in other species, or (in the case of resistance to PVX and PVY on this host) that RDR1 is not critical for virus restriction during the HR, and in the case of TMV, its inability to escape from the inoculated leaves may be due to factors other than RDR1.
One explanation for these results might be that there is redundancy between potato RDRs. For example, StRDR6a and b might compensate for decreased levels of StRDR1a and b. Alternatively, StRDR1a and StRDR1b may be involved in processes other than antiviral defence, such as the regulation of endogenous mRNA, which appears to be one of the roles for RDR1 in N. attenuata and tobacco32,36. Given that there are eight RDR genes in potato, there may have been more scope during the evolution of this plant for RDR diversification and specialization than in other species. Thus, we speculate that RDRs 1a and 1b may have lost their roles in antiviral resistance (while retaining other roles) without any loss in plant fitness since the expression of RDRs 6a and 6b may be adequate for the antiviral defence needs of potato.
Methods
Bioinformatics and Phylogenetic analysis of plant RDR genes
Preliminary identification of putative RDR protein or nucleic acid sequences in potato, Arabidopsis, tobacco, tomato and other plant species was done by searching TAIR (http://www.arabidopsis.org/; date of access 17/09/2015), Swissprot (http://web.expasy.org/docs/swiss-prot_guideline.html; date of access 17/09/2015), Potato Genome Sequencing Consortium (PGSC)51 (http://www.potatogenome.net; date of access 17/09/2015), the Tomato Genome Sequencing Consortium52 (https://solgenomics.net/organism/Solanum_lycopersicum/genome; date of access 17/09/2015), and GoMapMan53 (http://www.gomapman.org/; date of access 17/09/2015). The PGSC sequence data used were obtained from a doubled monoploid (DM) potato clone from Solanum tuberosum, group phureja51. To identify all relevant potato gene sequences a non-stringent basic local alignment search tool (BLAST: ref. 54) search was run against the PGSC double monoploid gene/loci (DMg) data, by querying with Arabidopsis RDR sequences. Putative RDR sequences were downloaded in FASTA format. Transcript and protein sequences were identified for each of the potato genes by running another BLAST search against DM transcript (DMt), DM coding sequence (DMc), and DM protein (DMp) data accessible through the potato genome browser. These sequences were downloaded for each of the potato RDR genes for use in alignments and the DMt, DMc and DMp identifiers were entered into a spreadsheet next to their corresponding RDR1 gene. Genes were classified into ‘gene families’ according to sequence similarity by checking E-values obtained from BLAST results when one DMg retrieves the other DMg sequences. Alignments of the DMc, non-redundant DMp, and Arabidopsis RDR sequences of interest were done using CLC Sequence Viewer (Qiagen, Hilden, Germany).
Putative RDR gene sequences were also obtained using BLAST search of Phytozome55 (http://www.phytozome.net; date of access 17/09/2015) against known orthogroups. The search string employed was the Arabidopsis RDR1 sequence. Amino acid alignments were done with MAFFT56 (http://mafft.cbrc.jp/alignment/software/; date of access 17/09/2015) using an E-INS-I alignment strategy. Alignments were refined with Se-Al57 (http://tree.bio.ed.ac.uk/software/seal/; date of access 17/09/2015) and converted to corresponding aligned DNA sequences with RevTrans58 (http://www.cbs.dtu.dk/services/RevTrans/; date of access 17/09/2015). All phylogenetic analyses were conducted on nucleotide alignments. For maximum likelihood analyses the program GARLI (Genetic Algorithm for Rapid Likelihood Inference; version 0.942) was employed. GARLI conducts maximum likelihood heuristic phylogenetic searches under the GTR (generalized time reversible) model of nucleotide substitution, in addition to using models that incorporate among-site rate variation, either assuming a gamma distribution (Γ) or a proportion of invariable sites (I), or both. Analyses were run with default options, except that the “significanttopochange” parameter was reduced to 0.01 to make searches more stringent. A strict consensus of five replicate GARLI analyses was done, selecting the topology with the highest likelihood score.
Potato plant growth conditions and treatments
Lines of non-transformed and transformed potato (Solanum tuberosum L.) plants of cultivar Pentland Dell were maintained by micro-propagation in tissue culture under sterile conditions. Potato plants were maintained on MS30 agar (Murashige and Skoog basal salts supplemented with 3% (w/v) sucrose and 1% (w/v) agar, adjusted to pH5.8). Micro-propagated lines were passaged using 1 cm stem pieces that included a petiole. Tissue cultured plant material was maintained at 22 °C and 16 hours of light.
Plants were also grown in Levington M3 compost (Scotts, Ipswich UK) and medium vermiculite (Scotts) at a ratio of 5:1, respectively with 0.002% (w/v) ‘Intercept 5GR’ (Scotts) either in a glasshouse (with supplementary lighting in winter), or in a growth chamber (Conviron Ltd., Winnipeg, Manitoba, Canada) under 16 hours of light, 200 μE.m−2.s−1, 22 °C, 60% humidity). To maintain the lines, cuttings were taken every four weeks by removing the apical 10 cm of the plant (cut below a node), dipped in rooting powder and planted in fresh compost mix.
For treatment with SA, plants were sprayed with 1 mM SA33. Potato plants were infected with PVY strain O (PVYO) by mechanical inoculation of two lower leaves with Carborundum as an abrasive using sap extracted from two systemically infected leaves of PVY-infected Nicotiana benthamiana plants in 0.8 ml of 0.1 M potassium phosphate buffer pH 7.259. Mechanical inoculation with PVX strain UK360 and TMV strain U1 used purified virions diluted to 50 μg/ml in potassium phosphate buffer61. Mock-inoculation used phosphate buffer only. Four plants were used for each treatment and experiments were carried out at least three times.
Transformation constructs
The silencing construct design was based on that used previously in tobacco for silencing NtRDR1 transcripts by Rakhshandehroo and colleagues36 and two target sequences (Supplementary Fig. S2) were chosen by lining up known RDR1 gene sequences. DNA for the silencing constructs was amplified from potato DNA by PCR using Phusion DNA polymerase (Invitrogen, Waltham MA). Primers used for amplifying the first and second RNAi constructs are listed in Supplementary Table S2 (primer numbers 1–4). The Gateway cloning entry vector pDONR 207 (Invitrogen) was used to generate entry clones for each of the two RNAi constructs, following the manufacturers instructions. For RNAi sequences inserted into pDONR 207 plasmid, the Gateway LR reaction (Invitrogen) was used to recombine the construct sequences into the destination vector pK7GWIWG2(II)0 used for potato plant transformation62.
Transformation
Micro-propagated Pentland Dell plants were used to obtain leaf and stem pieces for transformation. Stem sections (c. 5 mm in length) excised from inter-nodal regions were transferred to liquid Murashige and Skoog basal salts pH 5.8 containing 2% sucrose for co-cultivation with Agrobacterium tumefaciens LBA4404. A. tumefaciens cells harboring either Construct 1 or 2 were incubated in liquid LB medium58 at 28 °C for 48 hours prior to use in plant transformation. Co-cultivation was carried out at room temperature with gentle agitation at 25 rpm for 20 minutes. Stem pieces were transferred to half-strength MS30 medium amended with 0.2 mg/l naphthalene acetic acid, 2.5 mg/l zeatin riboside and 2 mg/l gibberellic acid for 48 hours, then transferred to the same medium with the addition of 250 μg/ml cefotaxime and 40 μg/ml kanamycin. Plant pieces were transferred onto fresh medium every 7 days for 4–5 weeks until callus had developed. Calli were transferred onto MS30 agar amended with 0.02 mg/l naphthalene acetic acid, 2 mg/l zeatin riboside, 2 mg/l gibberellic acid, 250 μg/ml cefotaxime and 40 μg/ml kanamycin, with weekly transfers. Shoots that developed from calli were transferred to MS30 agar amended 250 μg/ml cefotaxime and 40 μg/ml kanamycin to promote root formation.
Genomic DNA was extracted using the method of Doyle and Doyle63 from young leaves of transformed and non-transformed plants and PCR with primers designed to amplify regions within the RNAi constructs used to verify transgene insertion into the plant genome (primers 5–7 in Supplementary Table S2). Identification of potato lines transformed with the empty destination vector was achieved through PCR of genomic DNA from the empty vector potato line. The PCR primers used were situated either side of one of the insert regions on the destination vector, one in the 35S promoter (primer 5) and the other in the intron (primer 8).
Twelve independent transformed lines were chosen for RTqPCR analysis to assess down-regulation of StRDR1 transcript expression. One plant from each independent line was sampled. Reverse transcription PCR and first-strand cDNA synthesis was done according to the manufacturers instructions. RTqPCR using Sybr Green (Sigma) was used to assess the degree of silencing in transformed and non-transformed lines (non-transformed lines were used as the reference samples). Cyclophilin was used as the reference transcript (primers 9 and 10 in Supplementary Table S2) as its expression was identified as being stable under the experimental conditions and virus-inoculated non-transformed plants were used as the reference samples during RTqPCR analysis. RTqPCR procedures and equipment used have been described previously33. StRDR1-specific RTqPCR primers were used to amplify the region of the StRDR1 gene that lies between the regions of the RDR1 gene that are included in two RNAi constructs (primers 11 and 12 listed in Supplementary Table S2). Primers were designed with sequences matching both StRDR1a and StRDR1b, so that RDR1 expression analysis accounted for both of the potato RDR1 genes.
Analysis of viral infection of RDR1-silenced and non-transformed potato plants
During sampling, leaves were first photographed and then frozen in liquid nitrogen. Harvested leaves of the same potato line/treatment were pooled and stored at −80 °C prior to RNA extraction and RTqPCR analysis to assess relative virus accumulation. Both inoculated leaf tissue and upper, younger leaves were harvested for RNA extraction. Upper, non-inoculated leaves were sampled to assess whether silencing of RDR1 might allow systemic viral movement. RTqPCR primers were designed to amplify the coat proteins of each of the viruses used, PVX, PVYO and TMV (primers 13–18 in Supplementary Table S2). RTqPCR was performed and results analysed as previously described33.
Additional Information
How to cite this article: Hunter, L. J. R. et al. RNA-dependent RNA polymerase 1 in potato (Solanum tuberosum) and its relationship to other plant RNA-dependent RNA polymerases. Sci. Rep. 6, 23082; doi: 10.1038/srep23082 (2016).
References
Csorba, T., Pantaleo, V. & Burgyan, J. RNA silencing: an antiviral mechanism. Adv. Virus Res. 75, 35–71 (2009).
Fagard, M., Boutet, S., Morel, J.-B., Bellini, C. & Vaucheret, H. AGO1, QDE-2 and RDE-1 are related proteins required for post-transcriptional gene silencing in plants, quelling in fungi, and RNA interference in animals. Proc. Nat. Acad. Sci. USA 97, 11650–11654 (2000).
Vaucheret, H., Vazquez, F., Crété, P. & Bertel, D. P. The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Genes Dev. 18, 1187–1197 (2004).
Li, J., Yang, Z., Yu, B., Liu, J. & Chen, X. Methylation protects miRNAs and siRNAs from a 3′-end uridylation activity in Arabidopsis . Curr. Biol. 15, 1501–1507 (2005).
Montgomery, T. A. et al. Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 133, 128–141 (2008).
Montgomery, T. A. et al. AGO1-miR173 complex initiates phased siRNA formation in plants. Proc. Nat. Acad. Sci. USA 105, 20055–20062 (2008).
Qu, F., Ye, X. & Morris, T. J. 2008. Arabidopsis DRB4, AGO1, AGO7 and RDR6 participate in a DCL4-initiated antiviral RNA silencing pathway negatively regulated by DCL1. Proc. Nat. Acad. Sci. USA 105, 14732–14737 (2008).
Harvey, J. J. W. et al. An antiviral defense role of AGO2 in plants. PloS One 6, e14639 (2011).
Qi, Y., He, X., Wang, X., Kohany, O., Jurka, J. & Hannon, G. J. Distinct catalytic and non-catalytic roles of ARGONAUTE4 in RNA-directed DNA methylation. Nature 443, 1008–1012 (2006).
Havecker, E. R. et al. The Arabidopsis RNA-directed DNA methylation argonautes functionally diverge based on their expression and interaction with target loci. Plant Cell 22, 321–334 (2010).
Hammond, S. M., Boettcher, S., Caudy, A. A., Kobayashi, R. & Hannon, G. J. Argonaute2, a link between genetic and biochemical analyses of RNAi. Science 293, 1146–1150 (2001).
Mourrain, P. et al. Arabidopsis SGS2 and SGS3 genes are required for posttranscriptional gene silencing and natural virus resistance. Cell 101, 533–542 (2000).
Cogoni, C. & Macino, G. Gene silencing in Neurospora crassa requires a protein homologous to RNA-dependent RNA polymerase. Nature 399, 166–169 (1999).
Smardon, A. et al. EGO-1 is related to RNA-directed RNA polymerase and functions in germ-line development and RNA interference in C. elegans . Current Biology 10, 169–178 (2000).
Vaistij, F. E., Jones, L. & Baulcombe, D. C. Spreading of RNA targeting and DNA methylation in RNA silencing requires transcription of the target gene and a putative RNA-dependent RNA polymerase. Plant Cell 14, 857–867 (2002).
Schiebel, W., Haas, B., Marinkovic, S., Klanner, A. & Sänger, H. L. RNA-directed RNA polymerase from tomato leaves. II Catalytic in vitro properties. J. Biol. Chem. 268, 11858–11867 (1993).
Qi, X., Bao, F. S. & Xie, Z. Small RNA deep sequencing reveals role for Arabidopsis thaliana RNA-dependent RNA polymerases in viral siRNA biogenesis. PLoS One 4, e4971 (2009).
Molnár, A. et al. Plant virus-derived small interfering RNAs originate predominantly from highly structured single-stranded viral RNAs. J. Virol. 79, 7812–7818 (2005).
Wassenegger, M. & Krczal, G. Nomenclature and functions of RNA-directed RNA polymerases. Trends Plant Sci. 11, 142–151 (2006).
Zong, J., Yao, X., Yin, J., Zhang, D. & Ma, H. Evolution of the RNA-dependent RNA polymerase (RdRP) genes: duplications and possible losses before and after the divergence of major eukaryotic groups. Gene 447, 29–39 (2009).
Butterbach, P. et al. Tomato yellow leaf curl virus resistance by Ty-1 involves increased cytosine methylation of viral genomes and is compromised by cucumber mosaic virus infection. Proc. Nat. Acad. Sci. USA 111, 12942–12947 (2014).
Xie, Z., Fan, B., Chen, C. & Chen, Z. An important role of an inducible RNA-dependent RNA polymerase in plant antiviral defense. Proc. Nat. Acad. Sci. USA 98, 6516–6521 (2001).
Yu, D., Fan, B., MacFarlane, S. A. & Chen, Z. Analysis of the involvement of an inducible Arabidopsis RNA-dependent RNA polymerase in antiviral defense. Molec. Plant-Microbe Interact. 16, 206–216 (2003).
Muangsan, N., Beclin, C., Vaucheret, H. & Robertson, D. Geminivirus VIGS of endogenous genes requires SGS2/SDE1 and SGS3 and defines a new branch in the genetic pathway for silencing in plants. Plant J. 38, 1004–1014 (2004).
Yang, S.-J., Carter, S. A., Cole, A. B., Cheng, N.-H. & Nelson, R. S. A natural variant of a host RNA-dependent RNA polymerase is associated with increased susceptibility to viruses by Nicotiana benthamiana . Proc. Nat. Acad. Sci. USA 101, 6297–6302 (2004).
Schwach, F., Vaistij, F. E., Jones, L. & Baulcombe, D. C. An RNA-dependent RNA polymerase prevents meristem invasion by potato virus X and is required for the activity but not the production of a systemic silencing signal. Plant Physiol. 138, 1842–1852 (2005).
Diaz-Pendon, J. A., Li, F., Li, W. & Ding, S. Suppression of antiviral silencing by cucumber mosaic virus 2b protein in Arabidopsis is associated with drastically reduced accumulation of three classes of viral small interfering RNAs. Plant Cell 19, 2053–2063 (2007).
Donaire, L. et al. Structural and genetic requirements for the biogenesis of Tobacco rattle virus-derived small interfering RNAs. J. Virol. 82, 5167–5177 (2008).
Garcia-Ruiz, H. et al. Arabidopsis RNA-dependent RNA polymerases and dicer-like proteins in antiviral defense and small interfering RNA biogenesis during Turnip mosaic virus infection. Plant Cell 22, 481–496 (2010).
He, J., Dong, Z., Jia, Z., Wang, J. & Wang, G. Isolation, expression and functional analysis of a putative RNA-dependent RNA polymerase gene from maize (Zea mays L.). Molec. Biol. Rep. 37, 865–874 (2010).
Yang, H., Wang, M., Gao, Z., Zhu, C. & Guo, X. Isolation of a novel RNA-dependent RNA polymerase 6 from Nicotiana glutinosa, NgRDR6, and analysis of its response to biotic and abiotic stresses. Molec. Biol. Rep. 38, 929–937 (2011).
Pandey, S. P. & Baldwin, I. T. RNA-directed RNA polymerase 1 (RdR1) mediates the resistance of Nicotiana attenuata to herbivore attack in nature. Plant J. 50, 40–53 (2007).
Hunter, L. J. R. et al. Regulation of RNA-dependent RNA polymerase 1 and isochorismate synthase gene expression in Arabidopsis. PLoS One 8, e66530 (2013).
Liao, Y. W. K. et al. The role of hydrogen peroxide and nitric oxide in the induction of plant-encoded RNA-dependent RNA polymerase 1 in the basal defense against tobacco mosaic virus. PLoS One 8, e76090 (2013).
Xu, T. et al. Expressional and regulatory characterization of Arabidopsis RNA-dependent RNA polymerase 1. Planta 237, 1561–1569 (2013).
Rakhshandehroo, F., Takeshita, M., Squires, J. & Palukaitis, P. The influence of RNA-dependent RNA polymerase 1 on Potato virus Y infection and on other antiviral response genes. Molec. Plant-Microbe Interact. 22, 1312–1318 (2009).
International Potato Center. Potato Facts and Figures (2016) (Date of access: 22/01/2016). Available: http://cipotato.org/potato/facts/.
Palukaitis, P. Resistance to viruses of potato and their vectors. Plant Pathol. J. 28, 248–258 (2012).
Barker, H. & Dale, M. F. B. Resistance to viruses in potato. In Natural Resistance Mechanisms of Plants to Viruses ( G. Loebenstein & J. P. Carr Eds) pp 341–367 Springer (2006).
Liu, Y., Gao, Q., Wu, B., Ai, T. & Guo, X. NgRDR1, an RNA-dependent RNA polymerase isolated from Nicotiana glutinosa, was involved in biotic and abiotic stresses. Plant Physiol. Biochem. & 47, 359–368 (2009).
Wylie, S. J. et al. Differential responses to virus challenge of laboratory and wild accessions of Australian species of Nicotiana, and comparative analysis of RDR1 gene sequences. PLoS One 10, e0121787 (2015).
Madsen, C. T. et al. Identification and characterization of barley RNA-directed RNA polymerases. Biochimica et Biophysica Acta 1789, 375–385 (2009).
Jung, S., Cho, I., Sosinski, B., Abbott, A. & Main, D. Comparative genomic sequence analysis of strawberry and other rosids reveals significant microsynteny. BMC Res . Notes 3, 168 (2010).
Di Serio, F. et al. Deep sequencing of the small RNAs derived from two symptomatic variants of a chloroplastic viroid: implications for their genesis and for pathogenesis. PloS One 4, e7539 (2009).
Wang, M. et al. Characterization and functional analysis of GhRDR6, a novel RDR6 gene from cotton (Gossypium hirsutum L.). Biosci. Rep. 32, 139–151 (2012).
Jones, R. A. C. Breakdown of potato virus X resistance gene Nx: selection of a group four strain from strain group three. Plant Pathol. 31, 325–331 (1982).
Malamy, J., Carr, J. P., Klessig, D. F. & Raskin, I. Salicylic acid- A likely endogenous signal in the resistance response of tobacco to viral infection. Science 250, 1002–1004 (1990).
Huang, W. E. et al. Quantitative in situ assay of salicylic acid in tobacco leaves using a genetically modified biosensor strain of Acinetobacter sp. ADP1. Plant J. 46, 1073–1083 (2006).
Petek, M. et al. Potato virus Y infection hinders potato defence response and renders plants more vulnerable to Colorado potato beetle attack. Molec. Ecol. 23, 5378–5391 (2014).
Lee, W. S. et al. Salicylic acid treatment and expression of an RNA-dependent RNA polymerase 1 transgene inhibits lethal symptoms and meristem invasion during tobacco mosaic virus infection in Nicotiana benthamiana . BMC Plant Biol. 16, 15 (2016).
Potato Genome Sequencing Consortium. Genome sequence and analysis of the tuber crop potato. Nature 475, 189–195 (2011).
Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485, 635–641 (2012).
Ramšak, Ž. et al. GoMapMan: Integration, consolidation and visualization of plant gene annotations within the MapMan ontology. Nucleic Acids Res. 42(D1), D1167–D1175 (2014).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–10 (1990).
Goodstein, D. M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40(D1), D1178–D1186 (2012).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007).
Wernersson, R. & Pedersen, A. G. RevTrans - Constructing alignments of coding DNA from aligned amino acid sequences. Nucleic Acids Res. 31, 3537–3539 (2003).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press. N.Y. (1989).
Köhm, B. A., Goulden, M. G., Gilbert, J. E., Kavanaugh, T. A. & Baulcombe D. C. A potato virus X resistance gene mediates an induced, non-specific resistance in protoplasts. Plant Cell 5, 913–920 (1993).
Lee, W. S., Fu, S. F., Verchot-Lubicz, J. & Carr, J. P. Genetic modification of alternative respiration in Nicotiana benthamiana affects basal and salicylic acid-induced resistance to potato virus X. BMC Plant Biol. 11, 41 (2011).
Karimi, M., Inzé, D. & Depicker, A. Gateway vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 7, 193–195 (2002).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Acknowledgements
We thank Drs. David E. Hanke, Ruth LeFevre and Krys Kelly for useful discussions. LJRH was supported by a studentship co-funded by the James Hutton Institute (formerly Scottish Crop Research Institute) and the UK Biotechnological and Biological Sciences Research Council (BBSRC). Work in the JPC lab is funded by The Leverhulme Trust (RPG-2012-667), BBSRC (BB/D014376/1, BB/J011762/1) and the Cambridge University Newton Trust. SFB was funded by Leverhulme grant F/09-741/G to Professor Beverley Glover. KG was funded by an EMBO Short Term Fellowship. Work in the PP lab is funded by grant number NRF-2013R1A2A2A01016282 from the Korean National Research Foundation.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: L.J.R.H., S.A.M., P.P., S.F.B. and J.P.C. Performed the experiments: L.J.R.H., S.F.B., A.M.M. and A.E.P. Analysed the data: L.J.R.H., S.F.B., K.G. and J.P.C. Wrote the paper: L.J.R.H., S.F.B., S.A.M., P.P. and J.P.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hunter, L., Brockington, S., Murphy, A. et al. RNA-dependent RNA polymerase 1 in potato (Solanum tuberosum) and its relationship to other plant RNA-dependent RNA polymerases. Sci Rep 6, 23082 (2016). https://doi.org/10.1038/srep23082
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep23082
This article is cited by
-
Atypical DNA methylation, sRNA-size distribution, and female gametogenesis in Utricularia gibba
Scientific Reports (2021)
-
Genome-wide identification and characterization of DCL, AGO and RDR gene families in Saccharum spontaneum
Scientific Reports (2020)
-
Dicer-like and RNA-dependent RNA polymerase gene family identification and annotation in the cultivated Solanum tuberosum and its wild relative S. commersonii
Planta (2018)
-
Quantitative trait loci affecting intensity of violet flower colour in potato
Euphytica (2017)
-
Resistance to tomato leaf curl New Delhi virus in melon is controlled by a major QTL located in chromosome 11
Plant Cell Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.