Abstract
In the past decade, apoptosis pathway has gained a serious consideration being a critical cellular process in determining the cancer progression. Inverse relationship between cancer progression and apoptosis rate has been well established in the literature. It causes apoptosis proteins under the investigative scanner for developing anticancer therapies, which certainly got a success in the case of few apoptosis proteins as drug targets. In the present study, we have developed a dedicated database of 82 apoptosis proteins called ApoCanD. This database comprises of crucial information of apoptosis proteins in the context of cancer. Genomic status of proteins in the form of mutation, copy number variation and expression in thousands of tumour samples and cancer cell lines are the major bricks of this database. In analysis, we have found that TP53 and MYD88 are the two most frequently mutated proteins in cancer. Availability of other information e.g. gene essentiality data, tertiary structure, sequence alignments, sequences profiles, post-translational modifications makes it even more useful for the researchers. A user-friendly web interface is provided to ameliorate the use of ApoCanD. We anticipate that, this database will facilitate the research community working in the field of apoptosis and cancer. The database can be accessed at: http://crdd.osdd.net/raghava/apocand.
Similar content being viewed by others
Introduction
Apoptosis is a crucial process in deciding the cell fate and received enormous attention among the biologists in the past decade1,2. Dysregulation of apoptosis network can lead to the pathological conditions and cancer is one of them3,4. Many studies have been done in the past to show the relation between apoptosis and cancer progression5,6,7. Not only cancer establishment and progression, apoptosis dysregulation also imparts drug resistance to almost all types of cancer8,9,10. Cancer molds expression levels of crucial apoptosis proteins in such a way that apoptosis suppressed, which helps in prolongation of cancer survival11,12. Expression level of anti-apoptotic proteins e.g. IAP proteins13,14,15 is increased and expression level of apoptotic initiator is decreased in cancer e.g. Caspases16,17,18 and SMAC/DIABLO19,20,21. Because of this many apoptosis proteins are considered as target of anticancer therapies to ameliorate the cancer treatment22,23,24,25. Moreover, few molecules targeting apoptosis proteins have entered into the clinical trials also26,27,28. To make anticancer therapy against apoptosis proteins more pragmatic, we need a database that comprises of substantial background information about these proteins in cancer e.g. genomic information, gene essentiality data, structure etc. A database of apoptosis proteins was developed by Diez et al. in 2010 called DeathBase29. This database covers many aspects of apoptosis proteins like sequence and structure information, evolutionary information, ontology and direct links to the other databases. On the other side, DeathBase lacks the genomic information e.g. mutations, copy number variation and expression, gene essentiality, which is a preliminary requirement to understand these proteins in the context of cancer. It also does not provide any information regarding tertiary structure of these proteins. So, DeathBase is implausible for studying the apoptosis proteins in the context of cancer. For the better understanding of apoptosis proteins in cancer, still there is great demand for dedicated database of apoptosis proteins, which compiles information regarding their genomic status in large number of tumour samples and cancer cell lines along with the tertiary structure and gene essentiality data at one platform. With this tenet in mind, we have developed a dedicated database called ApoCanD, which comprises of compelling information about mutations, copy number variation, expression, gene essentiality, modeled tertiary structure, sequence alignment, sequences profiles from position-specific scoring matrix (PSSM) and hidden Markov models (HMM), structure domains, post-translational modification (PTM) and direct links to essential databases. Information in ApoCanD will complement the research community in designing anticancer therapies against apoptosis proteins.
Database Aim
ApoCanD is dedicated to understanding the role of apoptosis proteins in the cancer progression and drug resistance development. To address this issue, we have compiled three types of genomic information of apoptosis proteins e.g. mutation status, copy number variation and gene expression levels in tumour samples and cell lines. Along with this, normal variations from 1000 genome project30 are also compiled, which allow comparative benchmarking of normal and cancer samples. Gene expression and copy number variation data of these proteins is also included to understand their role in cancer. All the proteins are not essential for the survival of a particular cancer, so we have included the gene essentiality data in ApoCanD. Other basic information like, their tertiary structure, sequence alignment and profiles, post-translation modification etc. on a user-friendly web platform to draw fundamental conclusions about cancer. This web platform will provide plenty of opportunities for the researchers in exploring the role of apoptosis in cancer.
Backend Information
ApoCanD is built on Apache HTTP server, which is platform independent and available as open-source software. MySQL database is used to store the information in backend. Front end is developed by PHP, HTML, CSS and Java integration. Perl is also used as programming language to process the data in web presenting form.
Content of Database
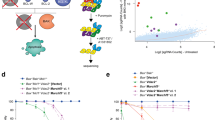
ApoCanD contains various types of information about each apoptosis protein, which is collected from different resources. Figure 1 shows the procedure of curation of ApoCanD.
Apoptosis Genes
We have followed the same criteria adopted in the DeathBase29 for the selection of apoptosis proteins e.g. they should be the part of the core machinery of apoptosis, they should regulate the core machinery of apoptosis, proteins having characteristic apoptotic domains or they are homologous to the central proteins of apoptosis. Following these criteria, we have selected 82 proteins and done our study on these proteins for the development of ApoCanD. Figure 2 represents the distribution of these proteins by various categories e.g. cellular location, pathway, chromosomal location and protein family.
Mutation Data
Mutation data of each apoptosis protein is collected from two primary resources namely, Cancer Cell Line Encyclopedia (CCLE), released on 24-Oct-2012 (CCLE_hybrid_capture1650_hg19_NoCommonSNPs_NoNeutralVariants_CDS_2012.05.07.maf)31 and Catalogue of Somatic Mutation in Cancer (COSMIC), version 67 (CosmicMutantExport_v67_241013.tsv) for tumour samples and cell line mutation data (CosmicCLP_MutantExport.tsv)32. CCLE contains the mutation status of more than 1600 proteins in 947 cancer cell lines, from this data we have filtered out the total 1368 mutations of apoptosis proteins. Similarly, we have filtered out 32157 mutations of apoptosis proteins available in cosmic data, which contains mutation data from both tumour samples and cancer cell lines. Out of the total mutations of 33525, major type of mutation is substitution mutation (30957) as shown in Figure 3.
Gene Expression Data and Copy Number Variation
Copy number variation and gene expression data of apoptosis proteins were collected from the CCLE. In CCLE, expression data was obtained from Affymetrix U133 plus array and further normalized by RMA technique using quantile normalization. Similarly, copy number variation data was obtained from Affymetrix SNP 6.0 arrays.
Gene Essentiality Data
Every gene is not required for the survival, so it is essential to check the importance of each gene in survival of cancer cells. So, we have compiled the shRNA dropout profiles of apoptosis genes from COLT-cancer database33. In COLT-cancer database, gene essential profiles of approximately 16000 genes are present, which is identified in 72 breast, ovarian and pancreatic cancer cell lines. Gene essentiality data is given in the form of two parameters. First GARP score, which quantifies the shRNA dropout rate, lower GARP score (more negative) represents the high essentiality of that gene. Second parameter is P-value, which describes the significance of GARP score. Figure 4 represents the gene essentiality data of XIAP in 72 cancer cell lines, where it shows the essentiality of this gene for HPDE and OVCA1369 cell lines.
Tertiary Structure
Structural information is essential for the designing of targeted therapy against any protein target. So, we found out the crystal structure of apoptosis proteins in PDB and out of 82 apoptosis proteins, structures of 61 proteins were available in PDB. Although for most of the proteins, structures were available in PDB, still we modeled the structure of all the proteins using HH-suite 2.0.1634 and Modeller 9.1335. Modeled structure of each protein can be visualized in Jmol applet and their respective PDB files can be downloaded by clicking on download button. We have also provided the PDB IDs of known structures and directly linked them to the PDB website.
Structure Domains
Many of the apoptosis proteins contain characteristic domains e.g. BIR domains in IAPs proteins. We have mapped the Pfam36 and Superfamily37 domains in all the apoptosis proteins. PFAM domains were searched by querying at Pfam website using the fasta sequences and E-value were kept at default value of 1. For Superfamily domains, we used standalone version of Superfamily and ran it with default parameters. We got 233 Pfam domains and 139 Superfamily domains in 82 apoptosis proteins.
Sequence Alignment
In ApoCanD, we have generated four types of sequence alignments of apoptosis proteins. First, alignment with variants obtained from 1000 genome project. In 1000 genome project, VCF file of more than 1000 genomes are given. We have converted the VCF file format to ANNOVAR input and then extracted the respective variations for each protein using ANNOVAR software38. These variations were mapped on to the wild-type protein sequences and these variants were aligned with wild-type sequences. Second, alignment with CCLE mutants, here mutants available in cancer cell lines were obtained from the CCLE and aligned with wild-type sequences. Third, alignments were generated with cancer mutants available in COSMIC. Fourth, we have obtained the homologous proteins of each human apoptosis proteins in other species from the NCBI and aligned them with human apoptosis proteins. In the case of homologous proteins, we have also generated the evolutionary tree. For the sequence alignment, we have used ClustalW39 and for the better visualization of sequence alignments and evolutionary tree, Jalview40 was used.
Sequence Profiles
Two types of sequence profiles were generated and included in ApoCanD e.g. HMM and PSSM, which tells about the conservation score at each position of the protein. HMM profiles were made by using ‘jackhmmer’ and ‘hmmbuild’ modules of HMMER software41 and PSSM profiles were generated by the ‘blastpgp’ and ‘makemat’ module of BLAST software42. We have created these alignments with three types of sequence databases e.g. Uniprot database, mutated sequences database (CCLE and COSMIC) and normal variant database (1000 Genomes). HMM profiles with mutants were proved to be excellent in predicting the functional impact of a mutation43, so we generated them for each apoptosis protein.
Post-Translational Modifications
Post-translational modifications (PTMs) play a crucial role in the functioning of proteins44,45,46. So, we included the PTM information in ApoCanD, which were compiled from the dbPTM47. In ApoCanD, position of the modification, amino acid (where modification occurred) and type of modification is given for each apoptosis protein. Major types of PTMs are phosphorylation, acetylation and ubiquitylation.
Querying the Database
For the maximal use of ApoCanD data with ease, we have provided three methods to query or access the database.
Tools
In tools section, we have provided five modules to access the data. (i) Simple search: Here user can query the data by entering a simple keyword e.g. protein name, cellular location, pathway, domain name, family etc. Simple search returns an aesthetic table containing all the major information available about the query keyword. (ii) Advanced search: It allows searching on the basis of three fields e.g. cellular location, pathway and family. Advance search uses logical operators (AND/OR) in returning the final result. ‘All’ option is given in all the three fields, which represents the OR function. Selecting ‘All’ option in all the three fields returns all the information about 82 proteins available in ApoCanD. (iii) BLAST: Here user can do the BLAST of a query protein with the apoptosis proteins available in ApoCanD to see the similarity of query protein with apoptosis proteins. (iv) Alignment: Here user can do the alignment of their query sequence with four different types of protein sequences, (a) With wild type apoptosis protein sequences, (b) With CCLE mutants, (c) With COSMIC mutants and (d) With 1000 Genome variants. Alignment can be visualized on Jalview applet, which gives three types of information i.e. conservation, quality and consensus along with the protein sequence alignment. This module also provides the option to view the dendrogram tree of query sequence with the respective protein type selected. (v) Cancer Sensitivity: This tool helps users to predict the nature of a protein sequence change, whether it would be a normal variation or a cancer sensitive mutation. This tool is based on the similarity score of the query sequence with the HMM profile of normal variants from 1000 Genome project and the cancer mutants from CCLE and COSMIC. Altered query sequence will be declared as cancer sensitive, if the similarity score is higher with cancer mutants HMM profile and vice versa. This tool uses “hmmsearch” module of HMMER suite (version 3.1b1)48 for calculating the similarity with respective HMM profiles.
Browse
Browse section has four inbuilt modules, which will help users to browse the ApoCanD database to draw the substantial information regarding apoptosis pathway proteins. (i) Protein: This module enlists all the 82 apoptosis proteins in a single aesthetic table and each protein is linked to its respective summary page. This table consists of five other fields apart from the gene name, Uniprot link, Deathbase link, homologues proteins link, PDB link and PubChem/ChEMBL bioassay link. Clicking on any of them opens up the respective information page on the host website. (ii) Chromosomes: In this module, apoptosis proteins have been divided on the basis of their gene location on their respective chromosomes. Here, user can browse the apoptosis proteins on the basis of the chromosomes distribution of their genes. Chromosome distribution of apoptosis genes is also depicted by the Circos plot, which also shows the gene expression and copy number variations (CNV) of apoptosis genes. (iii) Mutation Source: Mutational information of apoptosis genes was taken from CCLE and COSMIC and normal variants were taken from 1000 Genome project. Then, we identified the frequencies of each protein for cancer mutation and normal variation. Further, we calculated the ratio of the cancer mutation frequency and normal variant frequency for each protein, which built the list of highly mutated proteins in cancer and could be a target for anticancer therapy. TP53 has the highest frequency of mutation in cancer, which is approximately 1100 times more mutated in cancer as compared to the normal (Table 1). Second most mutated protein (100 times) is MYD88, which is an adapter protein involved in Toll-like receptor signaling pathway49. This protein is a well-established target for cancer therapy, whose mutation is responsible for the proliferation of cancer cells50. This module will help users to identify such apoptosis proteins, which are more prevalent for mutations in cancer and could be a target for anticancer therapy. (iv) Genomic Features: Here, we allow the users to select the apoptosis proteins on the basis of their genomic features i.e. mutation frequency, gene expression and copy number variation (CNV). User can select a particular range available in the option to fish out the apoptosis proteins.
Pathway
On this page, diagram of apoptosis pathway (intrinsic and extrinsic) is provided, which is designed by the CellDesigner software51 and adopted from the KEGG pathways52 as shown in Figure 5. To keep it simple, we have linked this pathway diagram with respective summary page of each protein. User can just click on a protein to open its summary page. At the end of the page, legends of pathway diagram are given to understand the pathway diagram.
Information
This section is dedicated to the relevant information about ApoCanD, which provides the following four type of information, (i) Statistics: This page summarizes the statistics about the distribution of ApoCanD proteins i.e. family wise, pathway wise, cellular location wise, frequency of mutation types and chromosome wise. (ii) Publication: This page provides the list of most recent papers about the apoptosis available in PubMed. (iii) Related Links: Here, user can find the relevant links of databases related to apoptosis. (iv) Acknowledgment: This page acknowledges the authors of databases and software used in the construction of ApoCanD.
Downloads
To complement and growth of this research field, we have provided all the data of ApoCanD for download. A dedicated download page is built, which gives access to the user to download the data without any condition. Data available for download includes mutation, copy number variation, expression data; modeled tertiary structures and domains; sequence alignment and profiles (PSSM and HMM); post-translational modifications for all the apoptosis proteins available in ApoCanD.
Discussion and Future Direction
Apoptosis pathway has a crucial role in cancer survival and development of drug resistance against a number of anticancer drugs. It is an urgent need of the present time to understand the role and mechanism of apoptosis proteins in cancer. Database of apoptosis proteins developed earlier just compiled the information about these proteins, which is not enough to understand their role in cancer. So, we developed a dedicated database of apoptosis proteins in the present study, which is in the context of cancer. ApoCanD is a critical step in this direction, which compiles fundamental information required for apoptosis proteins to gain deeper insight into their role in cancer. Genomics data available in ApoCanD helps researchers to identify the mutation, copy number variation and expression level status of apoptosis proteins in thousand of tumour samples and cancer cell lines. For example, TP53 protein is mutated in most of the cancer cell lines; X-linked inhibitor of apoptosis proteins (XIAP) is over-expressed in most of the cancer cell lines and inhibits caspases, which helps cancer cells to inhibit apoptosis and grow without any regulatory check. On the other hand SMAC/DIABLO is under-expressed, which further suppresses apoptosis. There are many such relevant information can be drawn from the ApoCanD data. Further, gene essentiality data makes it convenient to find out, which gene is essential for the survival of a particular cancer cell line and could act as therapeutic target for drug development. Structure and sequence alignment information in ApoCanD makes it easier to look for a possible strategy to target a protein for the therapeutic point of view. Figure 6 illustrates the applications of ApoCanD. Mutation and expression data available in ApoCanD can be used to develop predictive features of drug response by applying machine learning algorithm like naïve bayes, elastic net, support vector machine etc31,53,54,55. Moreover, mutation and copy number variation data can be explored to predict cancer subtype-specific drug response55,56,57,58. Apoptosis is a critical pathway in cancer and found to be deregulated in cancer5,6,7 and explored for fishing cancer therapeutic targets10,11,23,59.Genomic data available in ApoCanD can be explored for designing map of apoptosis pathway, which can be used for assigning cancer associated apoptosis genes as biomarkers60. Hudson et al.61 shown a comparative study of mutation data from CCLE and COSMIC, where they found discrepancies in these two large mutation datasets. These discrepancies were attributed to differences in computational protocols e.g. dbSNP filtering, acquisition/loss of mutation, passaging of cell lines etc. Unfortunately, these factors cannot be avoided in such large genomic studies and sometime limit down the impact of such studies. But from somewhere at some point, we have to start our efforts to address the critical disease problems. In future, we will try to develop quantitative structure-activity relationship (QSAR) models for most of the apoptosis proteins to develop nifty chemical molecules against them. Integration of such models on ApoCanD platform will make it more useful for the researchers, where they can choose their drug target and design active molecules against them at one platform. We keep on increasing the quantity and quality of the data in ApoCanD to makes it a cutting edge tool for the researchers in the apoptosis field.
Additional Information
How to cite this article: Kumar, R. and Raghava, G. P. S. ApoCanD: Database of human apoptotic proteins in the context of cancer. Sci. Rep. 6, 20797; doi: 10.1038/srep20797 (2016).
References
Zimmermann, K. C. & Green, D. R. How cells die: apoptosis pathways. J. Allergy Clin. Immunol. 108, S99–103 (2001).
Elmore, S. Apoptosis: a review of programmed cell death. Toxicol. Pathol. 35, 495–516 (2007).
Testa, U. & Riccioni, R. Deregulation of apoptosis in acute myeloid leukemia. Haematologica 92, 81–94 (2007).
Pettigrew, C. A. & Cotter, T. G. Deregulation of cell death (apoptosis): implications for tumor development. Discov. Med. 8, 61–3 (2009).
Wong, R. S. Y. Apoptosis in cancer: from pathogenesis to treatment. J. Exp. Clin. Cancer Res. 30, 87 (2011).
Lowe, S. W. Apoptosis in cancer. Carcinogenesis 21, 485–495 (2000).
Johnstone, R. W., Ruefli, A. A. & Lowe, S. W. Apoptosis. Cell 108, 153–164 (2002).
Indran, I. R., Tufo, G., Pervaiz, S. & Brenner, C. Recent advances in apoptosis, mitochondria and drug resistance in cancer cells. Biochim. Biophys. Acta 1807, 735–45 (2011).
Pommier, Y., Sordet, O., Antony, S., Hayward, R. L. & Kohn, K. W. Apoptosis defects and chemotherapy resistance: molecular interaction maps and networks. Oncogene 23, 2934–49 (2004).
Giménez-Bonafé, P., Tortosa, A. & Pérez-Tomás, R. Overcoming drug resistance by enhancing apoptosis of tumor cells. Curr. Cancer Drug Targets 9, 320–40 (2009).
Yip, K. W. & Reed, J. C. Bcl-2 family proteins and cancer. Oncogene 27, 6398–406 (2008).
Backus, H. H. J. et al. Differential expression of cell cycle and apoptosis related proteins in colorectal mucosa, primary colon tumours and liver metastases. J. Clin. Pathol. 55, 206–11 (2002).
Li, S. et al. XIAP expression is associated with pancreatic carcinoma outcome. Mol. Clin. Oncol. 1, 305–308 (2013).
Krepela, E. et al. Increased expression of inhibitor of apoptosis proteins, survivin and XIAP, in non-small cell lung carcinoma. Int. J. Oncol. 35, 1449–62 (2009).
Bilim, V., Kasahara, T., Hara, N., Takahashi, K. & Tomita, Y. Role of XIAP in the malignant phenotype of transitional cell cancer (TCC) and therapeutic activity of XIAP antisense oligonucleotides against multidrug-resistant TCC in vitro. Int. J. Cancer 103, 29–37 (2003).
Terry, M. R. et al. Caspase-2 impacts lung tumorigenesis and chemotherapy response in vivo. Cell Death Differ. (2014).
Devarajan, E. et al. Down-regulation of caspase 3 in breast cancer: a possible mechanism for chemoresistance. Oncogene 21, 8843–51 (2002).
Winter, R. N., Kramer, A., Borkowski, A. & Kyprianou, N. Loss of Caspase-1 and Caspase-3 Protein Expression in Human Prostate Cancer. Cancer Res. 61, 1227–1232 (2001).
Endo, K. et al. Clinical significance of Smac/DIABLO expression in colorectal cancer. Oncol. Rep. 21, 351–5 (2009).
Martinez-Ruiz, G., Maldonado, V., Ceballos-Cancino, G., Grajeda, J. P. R. & Melendez-Zajgla, J. Role of Smac/DIABLO in cancer progression. J. Exp. Clin. Cancer Res. 27, 48 (2008).
Mizutani, Y. et al. Downregulation of Smac/DIABLO expression in renal cell carcinoma and its prognostic significance. J. Clin. Oncol. 23, 448–54 (2005).
Gillissen, B. et al. Targeted therapy of the XIAP/proteasome pathway overcomes TRAIL-resistance in carcinoma by switching apoptosis signaling to a Bax/Bak-independent ‘type I’ mode. Cell Death Dis. 4, e643 (2013).
Wang, S. Design of small-molecule Smac mimetics as IAP antagonists. Curr. Top. Microbiol. Immunol. 348, 89–113 (2011).
Chen, D. J. & Huerta, S. Smac mimetics as new cancer therapeutics. Anticancer. Drugs 20, 646–58 (2009).
Lessene, G., Czabotar, P. E. & Colman, P. M. BCL-2 family antagonists for cancer therapy. Nat. Rev. Drug Discov. 7, 989–1000 (2008).
Souers, A. J. et al. ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat. Med. 19, 202–8 (2013).
Oltersdorf, T. et al. An inhibitor of Bcl-2 family proteins induces regression of solid tumours. Nature 435, 677–81 (2005).
Benetatos, C. A. et al. Birinapant (TL32711), a bivalent SMAC mimetic, targets TRAF2-associated cIAPs, abrogates TNF-induced NF-κB activation and is active in patient-derived xenograft models. Mol. Cancer Ther. 13, 867–79 (2014).
Díez, J., Walter, D., Muñoz-Pinedo, C. & Gabaldón, T. DeathBase: a database on structure, evolution and function of proteins involved in apoptosis and other forms of cell death. Cell Death Differ. 17, 735–6 (2010).
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–7 (2012).
Forbes, S. A. et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39, D945–50 (2011).
Koh, J. L. Y. et al. COLT-Cancer: functional genetic screening resource for essential genes in human cancer cell lines. Nucleic Acids Res. 40, D957–63 (2012).
Remmert, M., Biegert, A., Hauser, A. & Söding, J. HHblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat. Methods 9, 173–5 (2012).
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci. Chapter 2, Unit 2.9 (2007).
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–30 (2014).
Wilson, D. et al. SUPERFAMILY--sophisticated comparative genomics, data mining, visualization and phylogeny. Nucleic Acids Res. 37, D380–6 (2009).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Chenna, R. et al. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 31, 3497–500 (2003).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–91 (2009).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–37 (2011).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–10 (1990).
Shihab, H. A. et al. Predicting the functional, molecular and phenotypic consequences of amino acid substitutions using hidden Markov models. Hum. Mutat. 34, 57–65 (2013).
Grillari, J., Grillari-Voglauer, R. & Jansen-Dürr, P. Post-Translational Modification of Cellular Proteins by Ubiquitin and Ubiquitin-Like Molecules: Role in Cellular Senescence and Ageing. (2000).
Park, E. S. et al. Integrative analysis of proteomic signatures, mutations and drug responsiveness in the NCI 60 cancer cell line set. Mol. Cancer Ther. 9, 257–67 (2010).
Karve, T. M. & Cheema, A. K. Small changes huge impact: the role of protein posttranslational modifications in cellular homeostasis and disease. J. Amino Acids 2011, 207691 (2011).
Lu, C.-T. et al. DbPTM 3.0: an informative resource for investigating substrate site specificity and functional association of protein post-translational modifications. Nucleic Acids Res. 41, D295–305 (2013).
Eddy, S. R. Accelerated Profile HMM Searches. PLoS Comput. Biol. 7, e1002195 (2011).
Kawai, T. et al. Interferon-alpha induction through Toll-like receptors involves a direct interaction of IRF7 with MyD88 and TRAF6. Nat. Immunol. 5, 1061–8 (2004).
Wang, J. Q., Jeelall, Y. S., Ferguson, L. L. & Horikawa, K. Toll-Like Receptors and Cancer: MYD88 Mutation and Inflammation. Front. Immunol. 5, 367 (2014).
Funahashi, A., Morohashi, M., Kitano, H. & Tanimura, N. CellDesigner: a process diagram editor for gene-regulatory and biochemical networks. BIOSILICO 1, 159–162 (2003).
Kanehisa, M. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Garnett, M. J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–5 (2012).
Wang, E. et al. Predictive genomics: a cancer hallmark network framework for predicting tumor clinical phenotypes using genome sequencing data. Semin. Cancer Biol. 30, 4–12 (2015).
Zaman, N. et al. Signaling network assessment of mutations and copy number variations predict breast cancer subtype-specific drug targets. Cell Rep. 5, 216–23 (2013).
Niepel, M. et al. Profiles of Basal and stimulated receptor signaling networks predict drug response in breast cancer lines. Sci. Signal. 6, ra84 (2013).
Sasidharan Nair, P. & Vihinen, M. VariBench: a benchmark database for variations. Hum. Mutat. 34, 42–9 (2013).
Costello, J. C. et al. A community effort to assess and improve drug sensitivity prediction algorithms. Nat. Biotechnol. 32, 1202–1212 (2014).
Brunckhorst, M. K., Lerner, D., Wang, S. & Yu, Q. AT-406, an orally active antagonist of multiple inhibitor of apoptosis proteins, inhibits progression of human ovarian cancer. Cancer Biol. Ther. 13, 804–11 (2012).
Cui, Q. et al. A map of human cancer signaling. Mol. Syst. Biol. 3, 152 (2007).
Hudson, A. M. et al. Discrepancies in cancer genomic sequencing highlight opportunities for driver mutation discovery. Cancer Res. 74, 6390–6 (2014).
Acknowledgements
We are thankful to the funding agencies CSIR (Projects: Open Source Drug Discovery and GENESIS BSC0121) and Department of Biotechnology (project BTISNET), Govt. of India.
Author information
Authors and Affiliations
Contributions
R.K. developed the datasets and web interface. R.K. and G.P.S.R. wrote the manuscript. R.K. and G.P.S.R. conceived the idea.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kumar, R., Raghava, G. ApoCanD: Database of human apoptotic proteins in the context of cancer. Sci Rep 6, 20797 (2016). https://doi.org/10.1038/srep20797
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep20797
This article is cited by
-
Derivation of a novel antimicrobial peptide from the Red Sea Brine Pools modified to enhance its anticancer activity against U2OS cells
BMC Biotechnology (2024)
-
Drug efficacy and toxicity prediction: an innovative application of transcriptomic data
Cell Biology and Toxicology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.