Abstract
For a network, the accomplishment of its functions despite perturbations is called robustness. Although this property has been extensively studied, in most cases, the network is modified by removing nodes. In our approach, it is no longer perturbed by site percolation, but evolves after site invasion. The process transforming resident/healthy nodes into invader/mutant/diseased nodes is described by the Moran model. We explore the sources of robustness (or its counterpart, the propensity to spread favourable innovations) of the US high-voltage power grid network, the Internet2 academic network, and the C. elegans connectome. We compare them to three modular and non-modular benchmark networks, and samples of one thousand random networks with the same degree distribution. It is found that, contrary to what happens with networks of small order, fixation probability and robustness are poorly correlated with most of standard statistics, but they depend strongly on the degree distribution. While community detection techniques are able to detect the existence of a central core in Internet2, they are not effective in detecting hierarchical structures whose topological complexity arises from the repetition of a few rules. Box counting dimension and Rent’s rule are applied to show a subtle trade-off between topological and wiring complexity.
Similar content being viewed by others
Introduction
A biological or technological system is robust if it continues to function despite perturbations. Different names have been used for this concept depending on the nature of the system, the particular feature to be considered robust, and the kind of change to be applied. Scale-free networks1 have been proposed as models of complex networks enjoying an unexpected degree of robustness or error tolerance when a fraction of nodes is removed at random. The Word-Wide-Web, the Internet and several social networks have been regarded as interesting examples by many authors. However, these real networks seem extremely vulnerable to targeted attacks on the highly connected nodes. Similar conclusions were stated for living systems like metabolic networks2 and PPI networks3 after computational removal of randomly chosen nodes. These results suggest an evolutionary selection of the topological structure of biological networks in both senses, global generating mechanisms that give rise to power laws, as local correlating degree connectivity and influence in the network.
To understand the paradoxical “robust yet fragile” nature of all these networks, other scenarios have been also explored. Highly Optimised Tolerance (HOT) models4 were proposed from the double perspective of the existence of technological and economical constraints limiting the network topology and the accomplishment of their functional objectives. Abilene, the backbone for the Internet2 academic network, was used to illustrate the performance of this kind of models after node removal4.
Our aim was to explore the topological sources of another kind of robustness of biological and technological networks. In our approach, such a network is no longer perturbed by site percolation, but it evolves after the attack of pathogens like viruses or prions. The evolutionary process transforming healthy (or resident) nodes into diseased (or invader) nodes is described by the Moran model on networks introduced by Lieberman et al.5. Morbidity depends on the relative fitness of the pathogen with respect to the immune answer of healthy nodes. More precisely, at the beginning, all nodes are healthy. Then, one node chosen at random becomes infected by the pathogen. At successive steps, a diseased or healthy node is chosen at random with probability proportional to the relative fitness  or
or  . Next, a randomly chosen neighbour of the node is infected or cured. The (average) fixation probability is the probability that all the nodes of the network become infected. For a homogeneous or well-mixed networks this invasion process coincides with the classical Moran process6. In fact, isothermal networks has the same fixation probability that the homogeneous networks5. However, there are networks structures acting like evolutionary amplifiers favouring the disease spread across the network5,7,8,9.
. Next, a randomly chosen neighbour of the node is infected or cured. The (average) fixation probability is the probability that all the nodes of the network become infected. For a homogeneous or well-mixed networks this invasion process coincides with the classical Moran process6. In fact, isothermal networks has the same fixation probability that the homogeneous networks5. However, there are networks structures acting like evolutionary amplifiers favouring the disease spread across the network5,7,8,9.
In this context, we called robustness against invasion to a measure of the proximity of the network to the isothermal equilibrium. Here, we attempted to explore the topological sources of this kind of pathogen tolerance or its counterpart, the propensity to spread favourable innovations10. Therefore, motivated by the aforementioned works about usual robustness, we studied the robustness against invasion of the US Power Grid (PG)11 and Internet2 (I2) technological networks, and the neuronal network of the hermaphrodite nematode Caenorhabditis elegans (CE)12,13, see Data.
On the other hand, many researchers are interested in modularity and hierarchical modularity of technological and biological networks14,15,16,17,18,19,20, searching for adaptive, spatial, or economical constraints on their evolution4,21,22,23. According to them, modular architecture allows a faster adaptation to environmental changes, and their robustness is a major advantage when networks evolve under natural selection. Thus, we added a hierarchical modular network (HR)20 and a random toy model (TW)21 to contrast modularity and to test spatial aggregation propensity. Modular networks have the property of small-worldness24 characterised by a relative short average path length and a high clustering coefficient, favouring a low wiring cost. However, there are small-world networks that are not modular, so we also added the scale-free Barabási-Albert model (BA)25 to our study.
We computed the fixation probability using the Monte Carlo method on the associated Embedded Markov Chain, called the EMC method7. This allowed us to estimate the robustness against invasion of these networks. In an effort to understanding the structural causes of this robustness, we compared the fixation probability of the four deterministic networks PG, I2, CE and HR to that of samples of  random networks having the same degree distribution.
random networks having the same degree distribution.
Results
Mathematical model
Consider an undirected connected network  with node set
with node set  with no loops or multiple edges. Denote by
with no loops or multiple edges. Denote by  the degree of the node
the degree of the node  . The Moran process on
. The Moran process on  with fitness
with fitness  is the following Markov chain: start with a population
is the following Markov chain: start with a population  of
of  healthy nodes. Afterward, one single node
healthy nodes. Afterward, one single node  is chosen, with probability
is chosen, with probability  , to become a diseased node. At successive steps one node
, to become a diseased node. At successive steps one node  is selected at random with probability
is selected at random with probability  , where
, where  is
is  or
or  depending on whether
depending on whether  is healthy or diseased, and
is healthy or diseased, and  is the number of diseased nodes in that moment. Next, a neighbour of
is the number of diseased nodes in that moment. Next, a neighbour of  , randomly chosen with probability
, randomly chosen with probability  , becomes healthy or diseased depending on whether
, becomes healthy or diseased depending on whether  is healthy or diseased. The fixation probability, denoted
is healthy or diseased. The fixation probability, denoted  , is the probability that the whole population becomes diseased at some time step. For more details, see Supplementary Information.
, is the probability that the whole population becomes diseased at some time step. For more details, see Supplementary Information.
Fixation function and robustness
The computation of the fixation probability leads to a system of  equations, see Supplementary Information. Hence, except where global or partial symmetries reduce the number of equations 5,7, analytically solving the system is infeasible if the order of the network is not a very small. Thus, we computed the fixation probability of the three real networks PG, I2 and CE and the deterministic hierarchical one HR applying the EMC method7 and using
equations, see Supplementary Information. Hence, except where global or partial symmetries reduce the number of equations 5,7, analytically solving the system is infeasible if the order of the network is not a very small. Thus, we computed the fixation probability of the three real networks PG, I2 and CE and the deterministic hierarchical one HR applying the EMC method7 and using  trials and fitness values
trials and fitness values  varying from
varying from  to
to  with step size of
with step size of  . Each of these fixation functions is compared with the exact and asymptotic fixation functions
. Each of these fixation functions is compared with the exact and asymptotic fixation functions  and
and  for equivalent sized complete and complete bipartite networks, which are given in Methods, Equations (2) and (3) respectively. Each of these fixation functions was also compared to the fixation function on average from a benchmark sample of
for equivalent sized complete and complete bipartite networks, which are given in Methods, Equations (2) and (3) respectively. Each of these fixation functions was also compared to the fixation function on average from a benchmark sample of  networks having the same degree distribution. Graphical representations of these fixation functions in Fig. 1 show a strong dependence on the degree distribution, which indeed increases for the most robust networks CE and HR. We repeated the computation for the random networks BA and TW, which also are robust networks.
networks having the same degree distribution. Graphical representations of these fixation functions in Fig. 1 show a strong dependence on the degree distribution, which indeed increases for the most robust networks CE and HR. We repeated the computation for the random networks BA and TW, which also are robust networks.
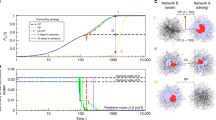
(a–d) Fixation probability functions for the US Power Grid, Internet2, C. elegans neuronal and hierarchical HR networks and asymptotically uniform samples of  networks with the same degree distributions. We used
networks with the same degree distributions. We used  and
and  trials respectively for every fitness value
trials respectively for every fitness value  varying from
varying from  to
to  with step size of
with step size of  . (e–f) Fixation probability functions for the Toy Worm and Barabási-Albert random models with samples of
. (e–f) Fixation probability functions for the Toy Worm and Barabási-Albert random models with samples of  networks and
networks and  trials for the same values of
trials for the same values of  .
.
To give a more accurate measure of the robustness of these networks, we introduce the robustness against invasion

For details see Methods. We estimated the value of  approximating
approximating  and
and  by the supremum of the differences
by the supremum of the differences  and
and  where
where  varies from
varies from  to
to  with steps of size
with steps of size  . The data is shown in Table 1, which includes both average and standard deviation of
. The data is shown in Table 1, which includes both average and standard deviation of  for random or randomised networks.
for random or randomised networks.
Heterogeneity
The notions of heterogeneity26,27 and heat heterogeneity10 (i.e. the variance of degree and temperature respectively) were proposed as strong indicators of the fixation probability for some small networks. We compared both heterogeneities with the fixation probability in the neutral case  and in the non-neutral case
and in the non-neutral case  . The most interesting cases
. The most interesting cases  and
and  are shown in Fig. 2. The rest of the cases are similar to
are shown in Fig. 2. The rest of the cases are similar to  , see Supplementary Information.
, see Supplementary Information.
Comparing fixation probability in the neutral and non-neutral case with (a,b) variance of the degree distribution and, (c,d) heat heterogeneity10.
For the networks considered here, both heterogeneity and heat heterogeneity show a poor correlation with  , see Table 1 and Fig. 2. In fact, Kendall’s rank correlation coefficient values
, see Table 1 and Fig. 2. In fact, Kendall’s rank correlation coefficient values  for
for  and
and  for
for  . Figs 1 and 2 support the idea that robustness and fixation probability depend strongly on the degree distribution. So it is natural to study the correlation of the fixation probability in both neutral and non-neutral cases with other statistics directly related with the degree distribution such as mean degree (
. Figs 1 and 2 support the idea that robustness and fixation probability depend strongly on the degree distribution. So it is natural to study the correlation of the fixation probability in both neutral and non-neutral cases with other statistics directly related with the degree distribution such as mean degree ( ), degree median (
), degree median ( ), degree skewness and degree kurtosis, the network’s global scale such as size (
), degree skewness and degree kurtosis, the network’s global scale such as size ( ), edge number, diameter and temperature entropy (
), edge number, diameter and temperature entropy ( ), or the network’s topology such clustering coefficient (
), or the network’s topology such clustering coefficient ( ), transitivity ratio, average path length (
), transitivity ratio, average path length ( ), power law exponent,
), power law exponent,  -modularity,
-modularity,  -modularity and fractal dimension (
-modularity and fractal dimension ( ). The values of these statistics are shown in Tables 1 and 2.
). The values of these statistics are shown in Tables 1 and 2.
We concluded that  and
and  are the best correlated quantities with the fixation probability in the neutral case
are the best correlated quantities with the fixation probability in the neutral case  , with
, with  (having mutual Kendall’s
(having mutual Kendall’s  ). In the non-neutral case
). In the non-neutral case  the most correlated statistics are the median degree with
the most correlated statistics are the median degree with  , the mean degree with
, the mean degree with  , and the average path length
, and the average path length  with
with  , together with
, together with  and
and  having
having  , see Table 3. Nevertheless, at least for the networks considered here, robustness is moderately well correlated with the median degree with
, see Table 3. Nevertheless, at least for the networks considered here, robustness is moderately well correlated with the median degree with  , the mean degree
, the mean degree  with
with  and the ratio
and the ratio  with
with  , but it is rather poorly correlated with the other basic statistics. But it is reasonable to think that the correlations with the median and mean degree are biased by the nature of the considered networks, see Discussion.
, but it is rather poorly correlated with the other basic statistics. But it is reasonable to think that the correlations with the median and mean degree are biased by the nature of the considered networks, see Discussion.
Modularity and hierarchical modularity
For many years, researchers have been interested in modularity and hierarchical modularity, searching for dynamic and evolutionary constraints that justify why biological and technological networks have a modular architecture. We were initially interested in two types of modularity. In the first one, the network splits into densely connected modules which are sparsely interconnected. The second one uses the idea that modular architecture reduces the information flow between modules.
Thus, we computed the  -modularity28, using Louvain method proposed by Blondel et al.29. The results can be seen in Table 2, where
-modularity28, using Louvain method proposed by Blondel et al.29. The results can be seen in Table 2, where  is compared with the robustness
is compared with the robustness  and the fixation probabilities
and the fixation probabilities  and
and  , see also Fig. 3. For the networks considered here, the
, see also Fig. 3. For the networks considered here, the  -modularity has positive Kendall’s
-modularity has positive Kendall’s  with respect to the fixation probability in the non-neutral case
with respect to the fixation probability in the non-neutral case  and negative
and negative  with respect to the robustness against invasion. The CE neuronal network shows a poor
with respect to the robustness against invasion. The CE neuronal network shows a poor  -modularity,
-modularity,  , but significantly higher than that of the corresponding randomised network,
, but significantly higher than that of the corresponding randomised network,  . In general,
. In general,  -modularity is sensitive to the randomization process. The
-modularity is sensitive to the randomization process. The  -modularity of the hierarchical HR network is moderate,
-modularity of the hierarchical HR network is moderate,  , contrary to what happens with the technological networks PG and I2 with a high
, contrary to what happens with the technological networks PG and I2 with a high  -modularity and a central core or “hub subcomplex” in their structure.
-modularity and a central core or “hub subcomplex” in their structure.
We also adopted the point of view by Rosvall et al.30,31 replacing  by the Infomap code or minimum description length
by the Infomap code or minimum description length  , see Table 2 and Fig. 3. For the networks considered here the correlations of
, see Table 2 and Fig. 3. For the networks considered here the correlations of  with the robustness
with the robustness  and the fixation probability
and the fixation probability  are a little worse than those of
are a little worse than those of  . Both modularities has a mutual (negative) moderate correlation with
. Both modularities has a mutual (negative) moderate correlation with  , but they seem to have different nature according to the clustering in Fig. 4.
, but they seem to have different nature according to the clustering in Fig. 4.
Community structure
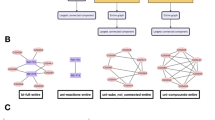
The idea that high  -modularity is correlated with the existence of a central core or “hub subcomplex” was explicitly tested on the Internet2 network and the hierarchical HR networks of level
-modularity is correlated with the existence of a central core or “hub subcomplex” was explicitly tested on the Internet2 network and the hierarchical HR networks of level  . We used Louvain algorithm29 to detect the community structure of these networks. For I2 network, it produces a similar but not identical community structure to the actual one, suggesting a more suitable placement of some connectors, see Supplementary Information. As counterpoint to the robustness, the spreading of favourable innovations can be enhanced by this kind of structures10 that incorporate trade-offs between performance and available resources4. Thus, community detection techniques could allow us to optimise the accomplishment of the functional objectives of a network with the same resources. However, these techniques are not equally effective in detecting other hierarchical modular structures, like the HR network, more topologically complex but constructed by repeating a simple rule, see Supplementary Information for details. Consequently, we say that a network is repetitive if it is obtained from the repetition of a reduced number of deterministic or random rules that encode its topological and dynamical complexity.
. We used Louvain algorithm29 to detect the community structure of these networks. For I2 network, it produces a similar but not identical community structure to the actual one, suggesting a more suitable placement of some connectors, see Supplementary Information. As counterpoint to the robustness, the spreading of favourable innovations can be enhanced by this kind of structures10 that incorporate trade-offs between performance and available resources4. Thus, community detection techniques could allow us to optimise the accomplishment of the functional objectives of a network with the same resources. However, these techniques are not equally effective in detecting other hierarchical modular structures, like the HR network, more topologically complex but constructed by repeating a simple rule, see Supplementary Information for details. Consequently, we say that a network is repetitive if it is obtained from the repetition of a reduced number of deterministic or random rules that encode its topological and dynamical complexity.
Topological complexity
Song et al. investigated the role of the fractal modular architecture in the robustness of some biological networks32,33. But the renormalisation mechanism that characterises this architecture continues to operate in the non-fractal case: there are models and examples of real networks showing a fractal behaviour in small scales, although they behave globally as small-worlds. We have seen that the BA model, and the CE and HR networks are robust small-worlds, while the PG and I2 networks are modular networks with central cores. From the point of view of both small-worldness (measured by the ratio  ) and modularity (measured by
) and modularity (measured by  ), the robust TW random model appears to be at an intermediate position. It has no central core because of the construction itself.
), the robust TW random model appears to be at an intermediate position. It has no central core because of the construction itself.
Our computations of the fractal dimension  are consistent with the global picture sketched above, except for the random TW model which has the lowest fractal dimension. See Table 2. Like the over-representation of some motifs in both C. elegans neuronal18 and random TW21 networks, the robustness of both networks could be favoured by some kind of spatial aggregation. By its construction, TW has similar properties to the planar lattice, including its fractal dimension. There also might be spatial reasons for the properties of the C. elegans connectome, but obviously these cannot be the same as the given above.
are consistent with the global picture sketched above, except for the random TW model which has the lowest fractal dimension. See Table 2. Like the over-representation of some motifs in both C. elegans neuronal18 and random TW21 networks, the robustness of both networks could be favoured by some kind of spatial aggregation. By its construction, TW has similar properties to the planar lattice, including its fractal dimension. There also might be spatial reasons for the properties of the C. elegans connectome, but obviously these cannot be the same as the given above.
Nevertheless, similarly to what happens with Rent’s rule34,35,36 to analyse internal communications in integrated circuits, fractality measures could be perturbed by the existence of two regions in the log-log plots. Namely, a Region I where the linear scaling show a fractal topology, and a Region II where the scaling is not linear but exponential. By restriction to Region I, we obtained new estimates, but the new data do not alter the picture above. See Supplementary Material for these results.
Wiring complexity
In fact, Rent’s rule has been applied to some biological networks including C. elegans connectome22,36,37. Our estimates of Rent’s exponents somewhat differ from those obtained by these authors. This is due to the different ways in which boxes decompositions are constructed, using box counting instead of min-cut partition algorithms. Values for the random TW ( ), the PG (
), the PG ( ) and the I2 (
) and the I2 ( ) networks belong to the range of values that requires a cost-efficient wiring architecture for VLSI circuits. But the HR network has also a similar Rent’s exponent (
) networks belong to the range of values that requires a cost-efficient wiring architecture for VLSI circuits. But the HR network has also a similar Rent’s exponent ( ) showing that a high topological complexity is compatible with a moderate wiring complexity. On the contrary, the BA model and the CE connectome exhibit high interconnection complexity with Rent’s exponents
) showing that a high topological complexity is compatible with a moderate wiring complexity. On the contrary, the BA model and the CE connectome exhibit high interconnection complexity with Rent’s exponents  and
and  respectively. In all cases, randomisation process increases Rent’s exponents as well as the fractal dimension. Now, a combination of repetitiveness with some randomness, which is missing in the HR network, could justify a gain of topological and wiring complexity without affecting the robustness of network like the CE connectome.
respectively. In all cases, randomisation process increases Rent’s exponents as well as the fractal dimension. Now, a combination of repetitiveness with some randomness, which is missing in the HR network, could justify a gain of topological and wiring complexity without affecting the robustness of network like the CE connectome.
Finally, Fig. 4 shows a global picture of the correlations. The hierarchical clustering of different statistics (with respect to the taxicab metric) is illustrated with a dendrogram. Most notable is the clustering of the modularity measures, in especial  and
and  , around the robustness and the fixation probability in the non-neutral case
, around the robustness and the fixation probability in the non-neutral case  .
.
Discussion
We computed the fixation functions for US Power Grid, Internet2 and C. elegans neuronal networks and asymptotically uniform samples of  randomly constructed networks with the same degree distributions. We completed these calculations with the fixation probability functions of four other benchmark networks: the hierarchical network constructed by Ravasz et al.20, the corresponding randomised sample of
randomly constructed networks with the same degree distributions. We completed these calculations with the fixation probability functions of four other benchmark networks: the hierarchical network constructed by Ravasz et al.20, the corresponding randomised sample of  networks with the same degree distribution, the random toy model constructed by Artzy-Randrup et al.21 and the Barabási-Albert model25, both with the same sample size as above. Moreover, we introduced the robustness against invasion to measure the proximity of a network to the isothermal equilibrium, that is, the equivalence of the network to a homogeneous population from the point of view of the Moran process. This quantity is interpreted as pathogens tolerance.
networks with the same degree distribution, the random toy model constructed by Artzy-Randrup et al.21 and the Barabási-Albert model25, both with the same sample size as above. Moreover, we introduced the robustness against invasion to measure the proximity of a network to the isothermal equilibrium, that is, the equivalence of the network to a homogeneous population from the point of view of the Moran process. This quantity is interpreted as pathogens tolerance.
We distinguished two different groups: a first group of robust small-worlds formed by the BA model and the CE and HR networks with the corresponding randomised networks, and a second group of modular networks with central cores formed by PG and I2 with their corresponding randomised benchmarks. Regarding small-worldness and modularity, the TW model appears to be in an intermediate position. In the paper, we attempted to explore the topological sources of the robustness against invasion.
Initially, heterogeneity26,27 and heat heterogeneity10, defined as the variances of the degree and temperature distributions, were proposed as statistics well correlated with the fixation probability on some networks of small order. But neither heterogeneity, nor heat heterogeneity seem to have a determining effect on the fixation probability of these technological and biological large networks. In fact, we have shown that there is a strong dependence on the whole degree distribution. Consequently, it is not surprising that specific statistical correlations are not too high.
In the neutral case  , we proved that the size and the temperature entropy8 are equally very well (negatively) correlated with the fixation probability
, we proved that the size and the temperature entropy8 are equally very well (negatively) correlated with the fixation probability  . In the non-neutral case
. In the non-neutral case  , the median degree and the mean degree are also (negatively) well correlated to the fixation probability
, the median degree and the mean degree are also (negatively) well correlated to the fixation probability  , which now gives a measure of the propensity to spread favourable innovations10. With respect to the robustness
, which now gives a measure of the propensity to spread favourable innovations10. With respect to the robustness  , the ratio
, the ratio  between the average path length
between the average path length  and the clustering coefficient
and the clustering coefficient  is also moderately correlated. But as evidenced by our own numerical experiments (to be published elsewhere), correlations with respect the median and mean degree are much lower on networks of small order.
is also moderately correlated. But as evidenced by our own numerical experiments (to be published elsewhere), correlations with respect the median and mean degree are much lower on networks of small order.
Small-worldness of the C. elegans neuronal network was evidenced by other authors. In some cases, they used the same model as we did to describe the CE connectome15. In other cases, the model is similar, but not exactly the same12,22,23, even if the data have been obtained at the same source13. Our estimations for  and
and  coincide with those given by Kim et al.15, except that we rounded up to
coincide with those given by Kim et al.15, except that we rounded up to  decimal places.
decimal places.
To precise the global picture of the biological and technological networks considered here, we also estimated the  -modularity, the
-modularity, the  -modularity and the fractal dimension of these networks. As explained in Methods, the
-modularity and the fractal dimension of these networks. As explained in Methods, the  -modularity was calculated by using the Louvain method proposed by Blondel et al.29. In fact, the same method was already used by Basset et al.22 to estimate
-modularity was calculated by using the Louvain method proposed by Blondel et al.29. In fact, the same method was already used by Basset et al.22 to estimate  for the CE connectome obtaining the same low value
for the CE connectome obtaining the same low value  . On the other hand, using an older algorithm38, Pan et al.23 gave a slightly lower estimate
. On the other hand, using an older algorithm38, Pan et al.23 gave a slightly lower estimate  where all the numbers are still rounded up to
where all the numbers are still rounded up to  decimal places. In fact, this authors make another estimate assuming that communities in CE connectome correspond to ganglia where
decimal places. In fact, this authors make another estimate assuming that communities in CE connectome correspond to ganglia where  . The same approach (but using a different algorithm) was applied by Kim et al.15 obtaining a value
. The same approach (but using a different algorithm) was applied by Kim et al.15 obtaining a value  . Note however that these results could be biased by the edge/link swapping algorithm to randomise CE connectome. Regarding fractality, the use of the box counting method for networks with low diameter represents the main limitation, because these networks are rapidly covered by a single box. This applies to the BA model and the CE and HR networks, as well as their randomised benchmarks. In the particular case of the CE connectome, Bassett et al.22 used a standard partitioning algorithm to estimate what they call topological fractal dimension
. Note however that these results could be biased by the edge/link swapping algorithm to randomise CE connectome. Regarding fractality, the use of the box counting method for networks with low diameter represents the main limitation, because these networks are rapidly covered by a single box. This applies to the BA model and the CE and HR networks, as well as their randomised benchmarks. In the particular case of the CE connectome, Bassett et al.22 used a standard partitioning algorithm to estimate what they call topological fractal dimension  . Explicitly, they obtained the value
. Explicitly, they obtained the value  which is higher than the box-counting dimension
which is higher than the box-counting dimension  and the box-counting dimension
and the box-counting dimension  in restriction to the Region I. Nevertheless, their results using the box-counting method coincide with ours, both included in the corresponding Supplementary Informations.
in restriction to the Region I. Nevertheless, their results using the box-counting method coincide with ours, both included in the corresponding Supplementary Informations.
Our results are also consistent with previous observations correlating  -modularity with the existence of a central core or “hub subcomplex” in the network, although one must be careful about interpreting the
-modularity with the existence of a central core or “hub subcomplex” in the network, although one must be careful about interpreting the  -modularity values obtained here: high for the technological networks PG and I2, both modular with central core, medium for HR and TW, and low for BA and CE. The idea that a high
-modularity values obtained here: high for the technological networks PG and I2, both modular with central core, medium for HR and TW, and low for BA and CE. The idea that a high  -modularity is correlated with the existence of this kind of core has been tested by describing explicitly the community structure of the Internet2 network and the HR networks of level
-modularity is correlated with the existence of this kind of core has been tested by describing explicitly the community structure of the Internet2 network and the HR networks of level  . While community detection techniques are able to identify the central core of Internet2, and even suggest improvements in its efficiency, they are not equally effective in detecting other hierarchical modular structures where topological complexity comes from the repetition of a single or finite set of rules. The BA model and the HR network are examples of such repetitive networks. BA shows low
. While community detection techniques are able to identify the central core of Internet2, and even suggest improvements in its efficiency, they are not equally effective in detecting other hierarchical modular structures where topological complexity comes from the repetition of a single or finite set of rules. The BA model and the HR network are examples of such repetitive networks. BA shows low  -modularity and high fractal dimension whereas the
-modularity and high fractal dimension whereas the  -modularity and the fractal dimension of HR are moderate and high respectively. Moreover, as we proved, the randomisation processes on these examples always increases the fractal dimension, which measures their topological complexity.
-modularity and the fractal dimension of HR are moderate and high respectively. Moreover, as we proved, the randomisation processes on these examples always increases the fractal dimension, which measures their topological complexity.
When networks are decomposed into boxes in order to compute their fractal dimension, it is natural to ask for the relationship between the number of interconnections and the box size. We applied Rent’s rule to estimate the interconnection or wiring complexity of the network. Although this power law was initially formulated for VSLI circuits34, a number of authors used Rent’s rule to analyse other biological and technological networks22,36,37. Using the box counting method, we privileged topological aspects against geometrical ones, even if those are certainly interesting. Based on our own method to determine Region II (and also Region III introduced by Stroobandt39), we obtained estimates of Rent’s exponents that slightly differ from those obtained by these authors, but allowed us to draw a reasonable schema: the BA model and the CE connectome have a high wiring complexity, while the other networks have moderate values corresponding to cost-efficient wiring architectures. Moreover, randomisation processes increases Rent’s exponents, a property which was also observed by Reda36 for other biological and technological networks. The HR network shows that a high topological complexity and a moderate wiring complexity are compatible for a network.
Values for Kendall’s coefficient correlating robustness and fixation probability in the non-neutral case  with
with  -modularity,
-modularity,  -modularity and fractal dimension, see Table 3, suggest a certain correlation with a suitable combination of topological and wiring complexity. However, as we have seen, HR and TW are robust networks with a similar wiring complexity, but a very different topological complexity. The reason for the low fractal dimension of TW resides in its own geometrical construction favouring a sort of spatial aggregation, which is similar to that of a planar lattice. On the contrary, a combination of repetitiveness with some randomness, not present in HR, could justify a gain of topological and wiring complexity without affecting the robustness against invasion of a network like the CE connectome.
-modularity and fractal dimension, see Table 3, suggest a certain correlation with a suitable combination of topological and wiring complexity. However, as we have seen, HR and TW are robust networks with a similar wiring complexity, but a very different topological complexity. The reason for the low fractal dimension of TW resides in its own geometrical construction favouring a sort of spatial aggregation, which is similar to that of a planar lattice. On the contrary, a combination of repetitiveness with some randomness, not present in HR, could justify a gain of topological and wiring complexity without affecting the robustness against invasion of a network like the CE connectome.
Summing up, from the comparison of the C. elegans neuronal network with the technological networks PG and I2 and the benchmark models TW and BA emerges the idea of a subtle trade-off between high complexity and low cost. In fact, according to a number of authors14,16,17,19,22 and based on their observations, this phenomenon could have been favoured by the evolution. Now, we have a triple challenge: firstly, analyse in more detail the influence of the degree distribution on the robustness, secondly, give a measure of the repetitiveness for networks of any order, and thirdly, quantify its effect on the robustness.
Methods
Robustness
For a homogeneous network  of size
of size  , the Moran process on
, the Moran process on  reduces to the classical Moran process where all nodes have the same fixation probability
reduces to the classical Moran process where all nodes have the same fixation probability

Let  be a weight-balanced network, that is, the weights of entering edges
be a weight-balanced network, that is, the weights of entering edges  and leaving edges
and leaving edges  are equal for any node
are equal for any node  . Then, according to the Circulation Theorem5, the number of elements of each state performs a biased random walk on the integer interval
. Then, according to the Circulation Theorem5, the number of elements of each state performs a biased random walk on the integer interval  with forward bias
with forward bias  and absorbing states
and absorbing states  and
and  . Therefore, the fixation probability is the same as that of the homogeneous network of size
. Therefore, the fixation probability is the same as that of the homogeneous network of size  . In fact, if
. In fact, if  is undirected, the weight of entering edges of a node
is undirected, the weight of entering edges of a node  can be interpreted as the temperature
can be interpreted as the temperature  , where
, where  means that
means that  is a neighbour of
is a neighbour of  and
and  is the number of neighbours of
is the number of neighbours of  . Hence, an undirected weight-balanced network is isothermal because
. Hence, an undirected weight-balanced network is isothermal because  for all nodes
for all nodes  .
.
On the other hand, the fixation probability of complete bipartite networks  converges to the same limit that the fixation probability
converges to the same limit that the fixation probability

of the Moran process with fitness  as
as  7. In other words, these structures are evolutionary amplifiers favouring advantageous invaders.
7. In other words, these structures are evolutionary amplifiers favouring advantageous invaders.
Let  be the fixation function of a network of size
be the fixation function of a network of size  . In order to measure the distance between
. In order to measure the distance between  and
and  , we used the norm
, we used the norm  and the ratio
and the ratio  The robustness against invasion of a network is the quotient
The robustness against invasion of a network is the quotient

Hence, any isothermal network has robustness against invasion  .
.
Networks data
The US Power Grid (PG) network is the high-voltage power grid in the Western States in the USA11. The nodes are generators, transformers, or substations and the edges are high-voltage transmission lines. Originally used by Watts and Strogatz24, this undirected network has  nodes and
nodes and  edges.
edges.
The Internet2 (I2) network assembles data from Internet2 community, available now through the Global Research Network Operations Center (GlobalNOC) at Indiana University40, which were collected in April 2013. First, we considered the list of active Internet2 Connectors at October 2012. It is an essential part of the Internet2 Combined Infrastructure Topology as described at September 2010. Secondly, we added the list of active Internet2 Primary Participants at April 2013 according to the GlobalNOC website40. For more details on the lists of connectors and primary participants, see Supplementary Information. Thus, we obtain a network with  nodes and
nodes and  edges. The initial Abilene network (which was replaced by Internet2 in October 2007) was previously analised by Doyle et al.4.
edges. The initial Abilene network (which was replaced by Internet2 in October 2007) was previously analised by Doyle et al.4.
The C. elegans neuronal (CE) network used here incorporates original data from41 and updates based upon later work42,12. This version of C. elegans connectome has  somatic neurones,
somatic neurones,  chemical synapses,
chemical synapses,  gap junctions, and
gap junctions, and  neuromuscular junctions13. According to our aim, we do not distinguish directionality of connections in this network that combines undirected gap junctions with directed chemical synapses, ignoring neuromuscular junctions and synaptic multiplicities. Thus, all the unidirectional connections between two different neurones will be replaced by bidirectional ones leading to a total of
neuromuscular junctions13. According to our aim, we do not distinguish directionality of connections in this network that combines undirected gap junctions with directed chemical synapses, ignoring neuromuscular junctions and synaptic multiplicities. Thus, all the unidirectional connections between two different neurones will be replaced by bidirectional ones leading to a total of  edges connecting
edges connecting  nodes. Since neurones RIBL/R and VA08 have auto-connections, we restrict our attention to
nodes. Since neurones RIBL/R and VA08 have auto-connections, we restrict our attention to  connections, cf. refs 12,15,22,23. There were efforts on the description of prionic diffusion within the brain43 and the construction of replication models44, which are consistent with the Moran process. An analysis of the human brain cluster structure detected by Gallos et al.45 using fMRI techniques (where modular clusters belong to a region of
connections, cf. refs 12,15,22,23. There were efforts on the description of prionic diffusion within the brain43 and the construction of replication models44, which are consistent with the Moran process. An analysis of the human brain cluster structure detected by Gallos et al.45 using fMRI techniques (where modular clusters belong to a region of  activated voxels) would require additional computational effort.
activated voxels) would require additional computational effort.
The Hierarchical (HR) network was constructed by Ravasz et al.20 as an example of hierarchical modular organisation, see Fig. 5. The one used here has four different levels leading to a total of  nodes and
nodes and  edges.
edges.
Hierarchical network of level  constructed by Ravasz et al.20.
constructed by Ravasz et al.20.
The Toy Worm (TW) network21 is a random network sampled as follows: it consists of  networks of order
networks of order  which were obtained from a square of
which were obtained from a square of  points in the integer lattice. Let
points in the integer lattice. Let  be the density function of the standard normal distribution. Two different nodes
be the density function of the standard normal distribution. Two different nodes  and
and  in the square are connected by an edge with probability
in the square are connected by an edge with probability  depending on the Euclidian distance
depending on the Euclidian distance  . The distribution of the number of connected components and the order frequencies of the maximal ones are showed in Supplemetary Information. The mean order is
. The distribution of the number of connected components and the order frequencies of the maximal ones are showed in Supplemetary Information. The mean order is  with a standard deviation of 1.26.
with a standard deviation of 1.26.
Finally, the Barabási-Albert (BA) model is another random network constructed using the preferential attachment process25. Here, we sampled  networks of order
networks of order  starting from
starting from  initial nodes and attaching each new node
initial nodes and attaching each new node  to
to  nodes with probability
nodes with probability  or
or  for
for  , depending on whether
, depending on whether  or
or  . We needed
. We needed  iterations to complete each network having
iterations to complete each network having  edges.
edges.
Random networks with prescribed degree distribution and fixation probability computation
The problem of generating random networks with a prescribed degree distribution was discussed by many authors46,47,48,49. In our case, random network generation was done in NetworkX50 using the Markov chain scheme proposed by Gkantsidis et al.47.
We compared the robustness of the three real networks PG, I2 and CE and the hierarchical deterministic one HR to that of their respective benchmark families of random networks having the same degree distribution. Each family consists of an asymptotically uniform sample of  networks generated by using the ‘swap’ algorithm47, where
networks generated by using the ‘swap’ algorithm47, where  true double-edge swaps were done on each real or deterministic network.
true double-edge swaps were done on each real or deterministic network.
On the other hand, to estimate the average fixation probability for each element of the sample we used the EMC method7. We computed the fixation probability function  using sequences of
using sequences of  trials for each fitness value
trials for each fitness value  varying from
varying from  to
to  with step size of
with step size of  for each of the
for each of the  networks in the sample. For the other random networks
networks in the sample. For the other random networks  and
and  , we did the same simulation from the initial sample of
, we did the same simulation from the initial sample of  networks. The fixation probability function of the original networks PG, I2, CE and HR was computed using
networks. The fixation probability function of the original networks PG, I2, CE and HR was computed using  trials. Finally, we estimated its robustness against invasion (4).
trials. Finally, we estimated its robustness against invasion (4).
Statistics
We analysed several statistical properties of the four networks PG, I2, CE and HR and their respective randomised benchmarks, as well as those of the two random networks BA and TW. Firstly, for each real or deterministic network, we considered some standard measures such as mean degree, degree median, degree variance, degree skewness and degree kurtosis related with the degree distribution. We also considered the heat heterogeneity10 defined as the variance of the temperature distribution  , where
, where  is the temperature of node
is the temperature of node  and
and  is the mean temperature. As above, for the other random networks, we computed these measures on average from the initial sample of
is the mean temperature. As above, for the other random networks, we computed these measures on average from the initial sample of  networks. Secondly, for any undirected connected network
networks. Secondly, for any undirected connected network  , in addition to the size
, in addition to the size  and the edge number
and the edge number  , we consider other global measures such as the diameter
, we consider other global measures such as the diameter  , where
, where  is the length of the shortest path between the nodes i and j, and the temperature entropy8
is the length of the shortest path between the nodes i and j, and the temperature entropy8 . Finally, we studied small-word properties of
. Finally, we studied small-word properties of  starting by the clustering coefficient
starting by the clustering coefficient  , where
, where  is the density of the induced sub-network
is the density of the induced sub-network  consisting of the set
consisting of the set  of neighbours of
of neighbours of  and the set
and the set  of edges between neighbours of
of edges between neighbours of  . The transitivity ratio is a measure of the network transitivity defined as the ratio between the number of nodes in triangles and the triplets of nodes connected by two edges. The average path length of
. The transitivity ratio is a measure of the network transitivity defined as the ratio between the number of nodes in triangles and the triplets of nodes connected by two edges. The average path length of  is given by
is given by  . Power law exponents were calculated using the power law Python package51 to adequately fit the tail of each degree distribution by a power law with respect to a optimal minimal value
. Power law exponents were calculated using the power law Python package51 to adequately fit the tail of each degree distribution by a power law with respect to a optimal minimal value  .
.
Rank correlation measures
A rank correlation coefficient is a measure of monotone dependence between two numerical random variables when ranked according to their values. Giving a set  consisting of
consisting of  individuals, two quantitive properties of these individuals are represented by two vectors
individuals, two quantitive properties of these individuals are represented by two vectors  and
and  . To each pair of individuals
. To each pair of individuals  and
and  , we associate antisymmetric
, we associate antisymmetric  -score
-score  and
and  -score
-score  . A rank correlation coefficient52 is defined by
. A rank correlation coefficient52 is defined by

When  is the difference between the ranks of
is the difference between the ranks of  and
and  , Spearman’s rank correlation coefficient is obtained. Here, we considered Kendall’s
, Spearman’s rank correlation coefficient is obtained. Here, we considered Kendall’s  derived from (5) by choosing
derived from (5) by choosing

In our case, each deterministic, random or randomised network is treated as a single network in order to state monotone dependence between two statistics (on average for random or randomised networks). For ordered sets with  elements, the upper critical value at a significance level of
elements, the upper critical value at a significance level of  is
is  . Statistical computations and graphics were done using the Python scientific stack53,54,55,56.
. Statistical computations and graphics were done using the Python scientific stack53,54,55,56.
Modularity and hierarchical modularity
A quantity called  -modularity28,38 was introduced by Girvan and Newman to measure the decomposability of a network into modules. Given a partition
-modularity28,38 was introduced by Girvan and Newman to measure the decomposability of a network into modules. Given a partition  of the vertex set
of the vertex set  of an undirected connected network
of an undirected connected network  , the modularity of
, the modularity of  is equal to
is equal to  , where
, where  is the entry of the adjacency matrix corresponding to two nodes
is the entry of the adjacency matrix corresponding to two nodes  and
and  in some module
in some module  . Several algorithms were proposed to detected the modular structure of the network by finding partitions with the largest value of
. Several algorithms were proposed to detected the modular structure of the network by finding partitions with the largest value of  . See a comparative analysis in ref. 38. Recently, methods to study hierarchical modularity were introduced starting from the idea of decomposing modules into submodules, which in turn are decomposed into sub-submodules, and so on. Here, we used the Louvain method29, which takes advantage of the hierarchical structure of the network to accelerate the optimisation of
. See a comparative analysis in ref. 38. Recently, methods to study hierarchical modularity were introduced starting from the idea of decomposing modules into submodules, which in turn are decomposed into sub-submodules, and so on. Here, we used the Louvain method29, which takes advantage of the hierarchical structure of the network to accelerate the optimisation of  .
.
However, as discussed in the comparative analysis by Lancichinetti et al.57, there are other approaches to identify community structures. We also used the Infomap method by Rosvall et al.30,31. The modular structure of the network is now achieved by optimising a function, called the minimum description length  , which gives account of the balance between information flow and data compression30,31.
, which gives account of the balance between information flow and data compression30,31.
Fractal dimension
An alternative definition of modular network was introduced by Song et al.32,33 representing modules by boxes of different length scales. Any network  admits a decomposition into disjoint boxes of size
admits a decomposition into disjoint boxes of size  , i.e. finite sets of nodes of diameter
, i.e. finite sets of nodes of diameter  . The fractality of
. The fractality of  can be formulated as an invariance property by the renormalisation flow, which replaces each box with a “renormalised” node. The fractal dimension
can be formulated as an invariance property by the renormalisation flow, which replaces each box with a “renormalised” node. The fractal dimension  of
of  can be estimated by two equivalent ways: as the exponent of a power law for the number of boxes of size
can be estimated by two equivalent ways: as the exponent of a power law for the number of boxes of size 

or as the exponent of a power law for the average number of nodes  in a box of size
in a box of size  when
when  varies from
varies from  to the first integer
to the first integer  such that
such that  .
.
In general, for scale-free networks, it is convenient to use the first method based on power law (7) to find the fractal scaling, although several algorithms can be used to calculate the fractal dimensions, each of them with its pros and cons58,59. Here, we implemented the greedy colouring algorithm proposed by Song et al.58 according to the description given by Locci et al.59. The fractal dimension of PG, I2, CE and HR networks was estimated using the ordinary least squares regression on the data gathered from the above algorithm. For each random and randomised network, each of the  sample elements was partitioned and then
sample elements was partitioned and then  was estimated both computing fractal dimension on average (with the corresponding standard deviations) and fitting a single line to the whole data set. In all cases, standard errors in the fit of the power law (on average in random ones) were also computed and included in Table 2. Naturally, errors decrease as the diameter of network increase. But there is another source of possible error in the estimation of
was estimated both computing fractal dimension on average (with the corresponding standard deviations) and fitting a single line to the whole data set. In all cases, standard errors in the fit of the power law (on average in random ones) were also computed and included in Table 2. Naturally, errors decrease as the diameter of network increase. But there is another source of possible error in the estimation of  related with random choices in the construction of the partitions. However, other authors found low values of the standard deviation60 which have been corroborate by our own numerical experiments. So we have chosen at random a single partition for each of
related with random choices in the construction of the partitions. However, other authors found low values of the standard deviation60 which have been corroborate by our own numerical experiments. So we have chosen at random a single partition for each of  sample elements.
sample elements.
Additional Information
How to cite this article: Alcalde Cuesta, F. et al. Exploring the topological sources of robustness against invasion in biological and technological networks. Sci. Rep. 6, 20666; doi: 10.1038/srep20666 (2016).
References
Barabási, A.-L., Jeong, H. & Albert, R. Error and attack tolerance of complex networks. Nature 406, 378–382 (2000).
Jeong, H., Tombor, B., Albert, R., Oltvai, Z. N. & Barabasi, A.-L. The large-scale organization of metabolic networks. Nature 407, 651–654 (2000).
Jeong, H., Mason, S. P., Barabasi, A.-L. & Oltvai, Z. N. Lethality and centrality in protein networks. Nature 411, 41–42 (2001).
Doyle, J. C. et al. The “robust yet fragile” nature of the Internet. PNAS 102, 14497–14502 (2005).
Lieberman, E., Hauert, C. & Nowak, M. A. Evolutionary dynamics on graphs. Nature 433, 312–316 (2005).
Moran, P. A. P. Random processes in genetics. Proc. Cambridge Philos. Soc. 54, 60–71 (1958).
Alcalde Cuesta, F., González Sequeiros, P. & Lozano Rojo, A. Fast and asymptotic computation of the fixation probability for Moran processes on graphs. Biosystems 129, 25–35 (2015).
Voorhees, B. & Murray, A. Fixation probabilities for simple digraphs. Philos. T. R. Soc. A 469 (2013).
Daz, J. et al. On the fixation probability of superstars. Philos. T. R. Soc. A 469 (2013).
Tan, S. & Lu, J. Characterizing the effect of population heterogeneity on evolutionary dynamics on complex networks. Sci. Rep. 4, 5034 (2014).
Koblenz Network Collection. U.S. Power Grid Network Dataset. (2015) Available at: http://konect.uni-koblenz.de. Accessed: 15/4/2015.
Varshney, L. R., Chen, B. L., Paniagua, E., Hall, D. H. & Chklovskii, D. B. Structural properties of the Caenorhabditis elegans neuronal network . PLoS Comput. Biol. 7, e1001066 (2011).
Varshney, L. R., Chen, B. L., Paniagua, E., Hall, D. H. & Chklovskii, D. B. Neuronal connectivity II. (2015) Available at: http://www.wormatlas.org/neuronalwiring.html. Accessed: 15/4/2015.
Bullmore, E. & Sporns, O. Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198 (2009).
Kim, J. S. & Kaiser, M. From Caenorhabditis elegans to the human connectome: a specific modular organization increases metabolic, functional and developmental efficiency. Philos. T. R. Soc. B 369, doi: 10.1098/rstb.2013.0529 (2014).
Meunier, D., Lambiotte, R. & Bullmore, E. T. Modular and hierarchically modular organization of brain networks. Front. Neurosci. 4, doi: 10.3389/fnins.2010.00200 (2010).
Meunier, D., Lambiotte, R., Fornito, A., Ersche, K. & Bullmore, E. T. Hierarchical modularity in human brain functional networks. Front. Neuroinform. 3, doi: 10.3389/neuro.11.037 (2009).
Milo, R. et al. Network motifs: Simple building blocks of complex networks. Science 298, 824–827 (2002).
Sporns, O. Structure and function of complex brain networks. Dialogues Clin. Neurosci. 15, 247–262 (2013).
Ravasz, E., Somera, A. L., Mongru, D. A., Oltvai, Z. N. & Barabási, A.-L. Hierarchical organization of modularity in metabolic networks. Science 297, 1551–1555 (2002).
Artzy-Randrup, Y., Fleishman, S. J., Ben-Tal, N. & Stone, L. Comment on “Network motifs: Simple building blocks of complex networks” and “Superfamilies of evolved and designed networks”. Science 305, 1107 (2004).
Bassett, D. S. et al. Efficient physical embedding of topologically complex information processing networks in brains and computer circuits. PLoS Comput. Biol. 6, e1000748 (2010).
Pan, R. K., Chatterjee, N. & Sinha, S. Mesoscopic organization reveals the constraints governing Caenorhabditis elegans nervous system. PLoS ONE 5, e9240 (2010).
Watts, D. J. & Strogatz, S. H. Collective dynamics of ‘small-world’ networks. Nature 393, 440–442 (1998).
Barabási, A.-L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Broom, M., Rychtář, J. & Stadler, B. T. Evolutionary dynamics on small-order graphs. J. Intesdiscip. Math. 12, 129–140 (2009).
Broom, M., Rychtář, J. & Stadler, B. T. Evolutionary dynamics on graphs–the effect of graph structure and initial placement on mutant spread. J. Stat. Theory Pract. 5, 369–381 (2011).
Newman, M. E. J. & Girvan, M. Finding and evaluating community structure in networks. Phys. Rev. E 69, 026113 (2004).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech.-Theory E. 2008, P10008 (2008).
Rosvall, M. & Bergstrom, C. T. Maps of random walks on complex networks reveal community structure. PNAS 105, 1118–1123 (2008).
Rosvall, M. & Bergstrom, C. T. Multilevel compression of random walks on networks reveals hierarchical organization in large integrated systems. PLoS ONE 6, e18209 (2011).
Song, C., Havlin, S. & Makse, H. A. Self-similarity of complex networks. Nature 433, 392–395 (2005).
Song, C., Havlin, S. & Makse, H. A. Origins of fractality in the growth of complex networks. Nat. Phys. 2, 275–281 (2006).
Landman, B. & Russo, R. L. On a pin versus block relationship for partitions of logic graphs. IEEE T. Comput. C- 20, 1469–1479 (1971).
Christie, P. & Stroobandt, D. The interpretation and application of Rent’s rule. IEEE T. VLSI Syst. 8, 639–648 (2000).
Reda, S. Using circuit structural analysis techniques for networks in systems biology. In Proceedings of the 11th International Workshop on System Level Interconnect Prediction, SLIP ‘09, 37–44 (ACM, 2009).
Partzsch, J. & Schüffny, R. On the routing complexity of neural network models - Rent’s rule revisited. In ESANN (2009).
Newman, M. E. J. Modularity and community structure in networks. PNAS 103, 8577–8582 (2006).
Stroobandt, D. On an efficient method for estimating the interconnection complexity of designs and on the existence of region III in rent’s rule. In Proceedings of the Ninth Great Lakes Symposium on VLSI, 330–331 (IEEE, 1999).
GlobalNOC. Internet2 Maps & Documentation. (2015) Available at: http://noc.net.internet2.edu/i2network/maps-documentation.html. Accessed:3/4/2015.
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans . Philos. T. R. Soc. B 314, 1–340 (1986).
Chen, B. L., Hall, D. H. & Chklovskii, D. B. Wiring optimization can relate neuronal structure and function. PNAS 103, 4723–4728 (2006).
Hecker, R. et al. Replication of distinct scrapie prion isolates is region specific in brains of transgenic mice and hamsters. Genes. Dev. 6, 1213–1228 (1992).
Masel, J., Jansen, V. A. & Nowak, M. A. Quantifying the kinetic parameters of prion replication. Biophys. Chem. 77, 139–152 (1999).
Gallos, L. K., Sigman, M. & Makse, H. A. The conundrum of functional brain networks: Small-world efficiency or fractal modularity. Frontiers in Physiology 3, 123 (2012).
McKay, B. D. & Wormald, N. C. Uniform generation of random regular graphs of moderate degree. J. Algorithm. 11, 52–67 (1990).
Gkantsidis, C., Mihail, M. & Zegura, E. The Markov chain simulation method for generating connected power law random graphs. In In Proc. 5th Workshop on Algorithm Engineering and Experiments (ALENEX03), Baltimore, USA. SIAM (2003, January 11).
Viger, F. & Latapy, M. Efficient and simple generation of random simple connected graphs with prescribed degree sequence. In Computing and Combinatorics, (ed. Wang, L. ) vol. 3595 of Lecture Notes in Computer Science, 440–449 (Springer, 2005).
Bayati, M., Kim, J. & Saberi, A. A sequential algorithm for generating random graphs. Algorithmica 58, 860–910 (2010).
Hagberg, A. A., Schult, D. A. & Swart, P. J. Exploring network structure, dynamics, and function using NetworkX. In Proceedings of the 7th Python in Science Conference (SciPy2008), Pasadena, USA (eds Varoquaux, G. et al.), 11–15 (2008, August 19). URL http://conference.scipy.org/proceedings/SciPy2008/paper_2/. Accessed: 1/5/2013.
Alstott, J., Bullmore, E. & Plenz, D. Powerlaw: A Python package for analysis of heavy-tailed distributions. PLoS ONE 9, e85777 (2014).
Kendall, M. Rank correlation methods (Charles Griffin & Co. Ltd., 1975), fourth edn.
Walt, S. v. d., Colbert, S. C. & Varoquaux, G. The numpy array: A structure for efficient numerical computation. Computing in Science & Engineering 13, 22–30 (2011).
McKinney, W. Data structures for statistical computing in python. In Proceedings of the 9th Python in Science Conference, Austin, USA (eds van der Walt, S. et al.), 51–56 (2010, June 28). URL http://conference.scipy.org/proceedings/scipy2010/mckinney.html Accessed: 10/3/2014.
Statsmodels developers. Statsmodels 0.6.1. URL http://statsmodels.sourceforge.net. Accessed: 1/5/2013.
Waskom, M. et al. Seaborn: statistical data visualization, Stanford, California. doi: 10.5281/zenodo.19108 (2015).
Lancichinetti, A. & Fortunato, S. Community detection algorithms: A comparative analysis. Phys. Rev. E 80, 056117 (2009).
Song, C., Gallos, L. K., Havlin, S. & Makse, H. A. How to calculate the fractal dimension of a complex network: the box covering algorithm. J. Stat. Mech.-Theory E. 2007, P03006 (2007).
Locci, M., Concas, G., Tonelli, R. & Turnu, I. Three algorithms for analyzing fractal software networks. WSEAS Trans. Info. Sci. and App. 7, 371–380 (20101
Concas, G., Locci, M. F., Marchesi, M., Pinna, S. & Turnu, I. Fractal dimension in software networks. EPL 76, 1221 (2006).
Acknowledgements
This work was supported by the Spanish Excellence Grant MTM2013-46337-C2-2-P, Galician Grant GPC2015/006 and the European Regional Development Fund. Third author was also supported by the European Social Fund and Government of Aragón (Grant E15), and University of Zaragoza/CUD Grant UZCUD2015-CIE-05.
Author information
Authors and Affiliations
Contributions
F.A., P.G. and Á.L. designed and performed the research as well as wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Alcalde Cuesta, F., González Sequeiros, P. & Lozano Rojo, Á. Exploring the topological sources of robustness against invasion in biological and technological networks. Sci Rep 6, 20666 (2016). https://doi.org/10.1038/srep20666
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep20666
This article is cited by
-
Link Prediction Model Based on the Topological Feature Learning for Complex Networks
Arabian Journal for Science and Engineering (2020)
-
A method for validating Rent’s rule for technological and biological networks
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.





 -modularity, (c,d)
-modularity, (c,d)  -modularity and, (e,f) fractal dimension.
-modularity and, (e,f) fractal dimension.
